Project
Epithelial-Mesenchymal Transition, human (Scheel, 2011)
Navigation
Downloads
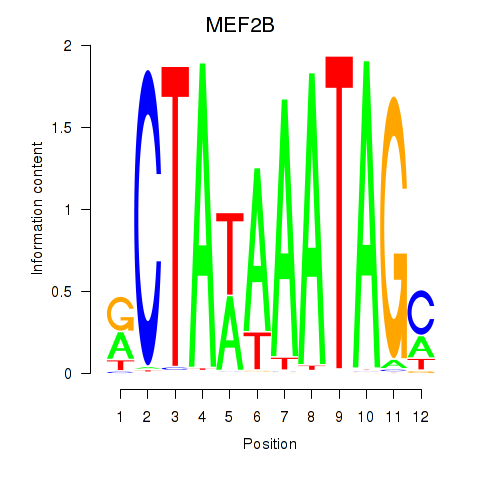
Results for MEF2B
Z-value: 0.56
Transcription factors associated with MEF2B
| Gene Symbol | Gene ID | Gene Info |
|---|---|---|
|
MEF2B
|
ENSG00000213999.11 | MEF2B |
Activity-expression correlation:
| Gene | Promoter | Pearson corr. coef. | P-value | Plot |
|---|---|---|---|---|
| MEF2B | hg19_v2_chr19_-_19302931_19302974 | -0.84 | 9.7e-03 | Click! |
Activity profile of MEF2B motif
Sorted Z-values of MEF2B motif
Network of associatons between targets according to the STRING database.
First level regulatory network of MEF2B
| Promoter | Score | Transcript | Gene | Gene Info |
|---|---|---|---|---|
| chr20_+_58251716 | 0.96 |
ENST00000355648.4 |
PHACTR3 |
phosphatase and actin regulator 3 |
| chr1_+_77333117 | 0.81 |
ENST00000477717.1 |
ST6GALNAC5 |
ST6 (alpha-N-acetyl-neuraminyl-2,3-beta-galactosyl-1,3)-N-acetylgalactosaminide alpha-2,6-sialyltransferase 5 |
| chr15_+_40531243 | 0.71 |
ENST00000558055.1 ENST00000455577.2 |
PAK6 |
p21 protein (Cdc42/Rac)-activated kinase 6 |
| chr11_+_1860200 | 0.70 |
ENST00000381911.1 |
TNNI2 |
troponin I type 2 (skeletal, fast) |
| chr15_+_40531621 | 0.68 |
ENST00000560346.1 |
PAK6 |
p21 protein (Cdc42/Rac)-activated kinase 6 |
| chr11_+_1860682 | 0.63 |
ENST00000381906.1 |
TNNI2 |
troponin I type 2 (skeletal, fast) |
| chr11_+_1860832 | 0.62 |
ENST00000252898.7 |
TNNI2 |
troponin I type 2 (skeletal, fast) |
| chr12_+_44229846 | 0.59 |
ENST00000551577.1 ENST00000266534.3 |
TMEM117 |
transmembrane protein 117 |
| chr2_-_165424973 | 0.55 |
ENST00000543549.1 |
GRB14 |
growth factor receptor-bound protein 14 |
| chr7_-_16872932 | 0.53 |
ENST00000419572.2 ENST00000412973.1 |
AGR2 |
anterior gradient 2 |
| chr21_-_42219065 | 0.52 |
ENST00000400454.1 |
DSCAM |
Down syndrome cell adhesion molecule |
| chr5_-_16509101 | 0.49 |
ENST00000399793.2 |
FAM134B |
family with sequence similarity 134, member B |
| chr3_+_186739636 | 0.44 |
ENST00000440338.1 ENST00000448044.1 |
ST6GAL1 |
ST6 beta-galactosamide alpha-2,6-sialyltranferase 1 |
| chr4_-_143227088 | 0.42 |
ENST00000511838.1 |
INPP4B |
inositol polyphosphate-4-phosphatase, type II, 105kDa |
| chr4_-_143226979 | 0.39 |
ENST00000514525.1 |
INPP4B |
inositol polyphosphate-4-phosphatase, type II, 105kDa |
| chr1_-_116311402 | 0.39 |
ENST00000261448.5 |
CASQ2 |
calsequestrin 2 (cardiac muscle) |
| chr4_-_25865159 | 0.38 |
ENST00000502949.1 ENST00000264868.5 ENST00000513691.1 ENST00000514872.1 |
SEL1L3 |
sel-1 suppressor of lin-12-like 3 (C. elegans) |
| chr2_-_208030647 | 0.35 |
ENST00000309446.6 |
KLF7 |
Kruppel-like factor 7 (ubiquitous) |
| chr1_+_212782012 | 0.34 |
ENST00000341491.4 ENST00000366985.1 |
ATF3 |
activating transcription factor 3 |
| chr14_+_76776957 | 0.34 |
ENST00000512784.1 |
ESRRB |
estrogen-related receptor beta |
| chr1_-_204329013 | 0.32 |
ENST00000272203.3 ENST00000414478.1 |
PLEKHA6 |
pleckstrin homology domain containing, family A member 6 |
| chr14_-_22005343 | 0.32 |
ENST00000327430.3 |
SALL2 |
spalt-like transcription factor 2 |
| chr16_+_84328429 | 0.31 |
ENST00000568638.1 |
WFDC1 |
WAP four-disulfide core domain 1 |
| chr16_+_84328252 | 0.31 |
ENST00000219454.5 |
WFDC1 |
WAP four-disulfide core domain 1 |
| chr14_-_21493884 | 0.29 |
ENST00000556974.1 ENST00000554419.1 ENST00000298687.5 ENST00000397858.1 ENST00000360463.3 ENST00000350792.3 ENST00000397847.2 |
NDRG2 |
NDRG family member 2 |
| chr17_-_9694614 | 0.29 |
ENST00000330255.5 ENST00000571134.1 |
DHRS7C |
dehydrogenase/reductase (SDR family) member 7C |
| chr3_+_127634069 | 0.28 |
ENST00000405109.1 |
KBTBD12 |
kelch repeat and BTB (POZ) domain containing 12 |
| chr3_+_127634312 | 0.27 |
ENST00000407609.3 |
KBTBD12 |
kelch repeat and BTB (POZ) domain containing 12 |
| chr12_+_52445191 | 0.26 |
ENST00000243050.1 ENST00000394825.1 ENST00000550763.1 ENST00000394824.2 ENST00000548232.1 ENST00000562373.1 |
NR4A1 |
nuclear receptor subfamily 4, group A, member 1 |
| chr4_-_143767428 | 0.25 |
ENST00000513000.1 ENST00000509777.1 ENST00000503927.1 |
INPP4B |
inositol polyphosphate-4-phosphatase, type II, 105kDa |
| chr8_+_1993173 | 0.24 |
ENST00000523438.1 |
MYOM2 |
myomesin 2 |
| chr7_-_105332084 | 0.23 |
ENST00000472195.1 |
ATXN7L1 |
ataxin 7-like 1 |
| chr11_+_7597639 | 0.23 |
ENST00000533792.1 |
PPFIBP2 |
PTPRF interacting protein, binding protein 2 (liprin beta 2) |
| chr1_+_84630367 | 0.23 |
ENST00000370680.1 |
PRKACB |
protein kinase, cAMP-dependent, catalytic, beta |
| chr8_+_1993152 | 0.22 |
ENST00000262113.4 |
MYOM2 |
myomesin 2 |
| chr10_+_71561649 | 0.22 |
ENST00000398978.3 ENST00000354547.3 ENST00000357811.3 |
COL13A1 |
collagen, type XIII, alpha 1 |
| chr6_+_44194762 | 0.22 |
ENST00000371708.1 |
SLC29A1 |
solute carrier family 29 (equilibrative nucleoside transporter), member 1 |
| chr1_+_92632542 | 0.21 |
ENST00000409154.4 ENST00000370378.4 |
KIAA1107 |
KIAA1107 |
| chr14_-_21493123 | 0.21 |
ENST00000556147.1 ENST00000554489.1 ENST00000555657.1 ENST00000557274.1 ENST00000555158.1 ENST00000554833.1 ENST00000555384.1 ENST00000556420.1 ENST00000554893.1 ENST00000553503.1 ENST00000555733.1 ENST00000553867.1 ENST00000397856.3 ENST00000397855.3 ENST00000556008.1 ENST00000557182.1 ENST00000554483.1 ENST00000556688.1 ENST00000397853.3 ENST00000556329.2 ENST00000554143.1 ENST00000397851.2 ENST00000555142.1 ENST00000557676.1 ENST00000556924.1 |
NDRG2 |
NDRG family member 2 |
| chr10_+_71561630 | 0.21 |
ENST00000398974.3 ENST00000398971.3 ENST00000398968.3 ENST00000398966.3 ENST00000398964.3 ENST00000398969.3 ENST00000356340.3 ENST00000398972.3 ENST00000398973.3 |
COL13A1 |
collagen, type XIII, alpha 1 |
| chr2_-_208031542 | 0.21 |
ENST00000423015.1 |
KLF7 |
Kruppel-like factor 7 (ubiquitous) |
| chr16_-_88717482 | 0.18 |
ENST00000261623.3 |
CYBA |
cytochrome b-245, alpha polypeptide |
| chr14_-_21493649 | 0.17 |
ENST00000553442.1 ENST00000555869.1 ENST00000556457.1 ENST00000397844.2 ENST00000554415.1 |
NDRG2 |
NDRG family member 2 |
| chr16_-_88717423 | 0.17 |
ENST00000568278.1 ENST00000569359.1 ENST00000567174.1 |
CYBA |
cytochrome b-245, alpha polypeptide |
| chr6_+_12007897 | 0.17 |
ENST00000437559.1 |
RP11-456H18.2 |
RP11-456H18.2 |
| chr10_-_69455873 | 0.15 |
ENST00000433211.2 |
CTNNA3 |
catenin (cadherin-associated protein), alpha 3 |
| chr1_+_145413268 | 0.15 |
ENST00000421822.2 ENST00000336751.5 ENST00000497365.1 ENST00000475797.1 |
HFE2 |
hemochromatosis type 2 (juvenile) |
| chrX_-_63450480 | 0.14 |
ENST00000362002.2 |
ASB12 |
ankyrin repeat and SOCS box containing 12 |
| chr6_-_137314371 | 0.14 |
ENST00000432330.1 ENST00000418699.1 |
RP11-55K22.5 |
RP11-55K22.5 |
| chr8_+_107282368 | 0.13 |
ENST00000521369.2 |
RP11-395G23.3 |
RP11-395G23.3 |
| chr1_-_156460391 | 0.13 |
ENST00000360595.3 |
MEF2D |
myocyte enhancer factor 2D |
| chr1_-_26394114 | 0.13 |
ENST00000374272.3 |
TRIM63 |
tripartite motif containing 63, E3 ubiquitin protein ligase |
| chr6_+_12007963 | 0.13 |
ENST00000607445.1 |
RP11-456H18.2 |
RP11-456H18.2 |
| chr3_-_52486841 | 0.12 |
ENST00000496590.1 |
TNNC1 |
troponin C type 1 (slow) |
| chr9_-_74980113 | 0.12 |
ENST00000376962.5 ENST00000376960.4 ENST00000237937.3 |
ZFAND5 |
zinc finger, AN1-type domain 5 |
| chr7_-_37488834 | 0.12 |
ENST00000310758.4 |
ELMO1 |
engulfment and cell motility 1 |
| chr19_-_16008880 | 0.12 |
ENST00000011989.7 ENST00000221700.6 |
CYP4F2 |
cytochrome P450, family 4, subfamily F, polypeptide 2 |
| chr14_+_56046914 | 0.11 |
ENST00000413890.2 ENST00000395309.3 ENST00000554567.1 ENST00000555498.1 |
KTN1 |
kinectin 1 (kinesin receptor) |
| chr16_+_12058961 | 0.11 |
ENST00000053243.1 |
TNFRSF17 |
tumor necrosis factor receptor superfamily, member 17 |
| chr17_-_48785216 | 0.11 |
ENST00000285243.6 |
ANKRD40 |
ankyrin repeat domain 40 |
| chr4_+_8321882 | 0.10 |
ENST00000509453.1 ENST00000503186.1 |
RP11-774O3.2 RP11-774O3.1 |
RP11-774O3.2 RP11-774O3.1 |
| chr10_+_71561704 | 0.10 |
ENST00000520267.1 |
COL13A1 |
collagen, type XIII, alpha 1 |
| chr8_-_27115931 | 0.10 |
ENST00000523048.1 |
STMN4 |
stathmin-like 4 |
| chr10_+_135340859 | 0.10 |
ENST00000252945.3 ENST00000421586.1 ENST00000418356.1 |
CYP2E1 |
cytochrome P450, family 2, subfamily E, polypeptide 1 |
| chr12_+_25205446 | 0.10 |
ENST00000557489.1 ENST00000354454.3 ENST00000536173.1 |
LRMP |
lymphoid-restricted membrane protein |
| chr3_+_35683651 | 0.10 |
ENST00000187397.4 |
ARPP21 |
cAMP-regulated phosphoprotein, 21kDa |
| chrX_-_11284095 | 0.10 |
ENST00000303025.6 ENST00000534860.1 |
ARHGAP6 |
Rho GTPase activating protein 6 |
| chr11_-_84634447 | 0.10 |
ENST00000532653.1 |
DLG2 |
discs, large homolog 2 (Drosophila) |
| chr12_-_99548270 | 0.09 |
ENST00000546568.1 ENST00000332712.7 ENST00000546960.1 |
ANKS1B |
ankyrin repeat and sterile alpha motif domain containing 1B |
| chr5_+_132009675 | 0.09 |
ENST00000231449.2 ENST00000350025.2 |
IL4 |
interleukin 4 |
| chr16_+_12059050 | 0.09 |
ENST00000396495.3 |
TNFRSF17 |
tumor necrosis factor receptor superfamily, member 17 |
| chr14_+_56046990 | 0.09 |
ENST00000438792.2 ENST00000395314.3 ENST00000395308.1 |
KTN1 |
kinectin 1 (kinesin receptor) |
| chr11_-_47400062 | 0.09 |
ENST00000533030.1 |
SPI1 |
spleen focus forming virus (SFFV) proviral integration oncogene |
| chr6_+_152128371 | 0.09 |
ENST00000443427.1 |
ESR1 |
estrogen receptor 1 |
| chr18_-_35145728 | 0.08 |
ENST00000361795.5 ENST00000603232.1 |
CELF4 |
CUGBP, Elav-like family member 4 |
| chr7_-_121944491 | 0.08 |
ENST00000331178.4 ENST00000427185.2 ENST00000442488.2 |
FEZF1 |
FEZ family zinc finger 1 |
| chr8_+_107282389 | 0.08 |
ENST00000577661.1 ENST00000445937.1 |
RP11-395G23.3 OXR1 |
RP11-395G23.3 oxidation resistance 1 |
| chr17_-_42200958 | 0.08 |
ENST00000336057.5 |
HDAC5 |
histone deacetylase 5 |
| chr3_+_188664988 | 0.07 |
ENST00000433971.1 |
TPRG1 |
tumor protein p63 regulated 1 |
| chr12_-_131200810 | 0.07 |
ENST00000536002.1 ENST00000544034.1 |
RIMBP2 RP11-662M24.2 |
RIMS binding protein 2 RP11-662M24.2 |
| chr4_+_41362796 | 0.07 |
ENST00000508501.1 ENST00000512946.1 ENST00000313860.7 ENST00000512632.1 ENST00000512820.1 |
LIMCH1 |
LIM and calponin homology domains 1 |
| chr3_+_46924829 | 0.07 |
ENST00000313049.5 |
PTH1R |
parathyroid hormone 1 receptor |
| chr11_-_84634217 | 0.07 |
ENST00000524982.1 |
DLG2 |
discs, large homolog 2 (Drosophila) |
| chr17_+_20059302 | 0.07 |
ENST00000395530.2 |
SPECC1 |
sperm antigen with calponin homology and coiled-coil domains 1 |
| chr3_-_11610255 | 0.07 |
ENST00000424529.2 |
VGLL4 |
vestigial like 4 (Drosophila) |
| chr17_-_34417479 | 0.06 |
ENST00000225245.5 |
CCL3 |
chemokine (C-C motif) ligand 3 |
| chr12_-_7899935 | 0.06 |
ENST00000543765.1 |
CLEC4C |
C-type lectin domain family 4, member C |
| chr1_+_22778337 | 0.06 |
ENST00000404138.1 ENST00000400239.2 ENST00000375647.4 ENST00000374651.4 |
ZBTB40 |
zinc finger and BTB domain containing 40 |
| chr19_+_16607122 | 0.06 |
ENST00000221671.3 ENST00000594035.1 ENST00000599550.1 ENST00000594813.1 |
C19orf44 |
chromosome 19 open reading frame 44 |
| chr16_-_31439735 | 0.06 |
ENST00000287490.4 |
COX6A2 |
cytochrome c oxidase subunit VIa polypeptide 2 |
| chr17_-_42200996 | 0.06 |
ENST00000587135.1 ENST00000225983.6 ENST00000393622.2 ENST00000588703.1 |
HDAC5 |
histone deacetylase 5 |
| chr20_+_44035847 | 0.06 |
ENST00000372712.2 |
DBNDD2 |
dysbindin (dystrobrevin binding protein 1) domain containing 2 |
| chr10_-_96829246 | 0.05 |
ENST00000371270.3 ENST00000535898.1 ENST00000539050.1 |
CYP2C8 |
cytochrome P450, family 2, subfamily C, polypeptide 8 |
| chr6_-_123957942 | 0.05 |
ENST00000398178.3 |
TRDN |
triadin |
| chr6_+_50061315 | 0.05 |
ENST00000415106.1 |
RP11-397G17.1 |
RP11-397G17.1 |
| chr4_+_44680429 | 0.05 |
ENST00000281543.5 |
GUF1 |
GUF1 GTPase homolog (S. cerevisiae) |
| chr11_-_19223523 | 0.05 |
ENST00000265968.3 |
CSRP3 |
cysteine and glycine-rich protein 3 (cardiac LIM protein) |
| chr12_-_111358372 | 0.05 |
ENST00000548438.1 ENST00000228841.8 |
MYL2 |
myosin, light chain 2, regulatory, cardiac, slow |
| chr6_+_42984723 | 0.05 |
ENST00000332245.8 |
KLHDC3 |
kelch domain containing 3 |
| chr5_-_131347306 | 0.05 |
ENST00000296869.4 ENST00000379249.3 ENST00000379272.2 ENST00000379264.2 |
ACSL6 |
acyl-CoA synthetase long-chain family member 6 |
| chr9_+_116327326 | 0.05 |
ENST00000342620.5 |
RGS3 |
regulator of G-protein signaling 3 |
| chr2_+_88367299 | 0.05 |
ENST00000419482.2 ENST00000444564.2 |
SMYD1 |
SET and MYND domain containing 1 |
| chr2_+_168043793 | 0.05 |
ENST00000409273.1 ENST00000409605.1 |
XIRP2 |
xin actin-binding repeat containing 2 |
| chr8_-_27115903 | 0.05 |
ENST00000350889.4 ENST00000519997.1 ENST00000519614.1 ENST00000522908.1 ENST00000265770.7 |
STMN4 |
stathmin-like 4 |
| chr8_+_76452097 | 0.04 |
ENST00000396423.2 |
HNF4G |
hepatocyte nuclear factor 4, gamma |
| chr20_+_61147651 | 0.04 |
ENST00000370527.3 ENST00000370524.2 |
C20orf166 |
chromosome 20 open reading frame 166 |
| chr6_+_152128686 | 0.04 |
ENST00000206249.3 |
ESR1 |
estrogen receptor 1 |
| chr12_+_25205666 | 0.04 |
ENST00000547044.1 |
LRMP |
lymphoid-restricted membrane protein |
| chr18_-_35145981 | 0.04 |
ENST00000420428.2 ENST00000412753.1 |
CELF4 |
CUGBP, Elav-like family member 4 |
| chr14_+_39735411 | 0.04 |
ENST00000603904.1 |
RP11-407N17.3 |
cTAGE family member 5 isoform 4 |
| chr12_-_371994 | 0.03 |
ENST00000343164.4 ENST00000436453.1 ENST00000445055.2 ENST00000546319.1 |
SLC6A13 |
solute carrier family 6 (neurotransmitter transporter), member 13 |
| chr8_+_95565947 | 0.03 |
ENST00000523011.1 |
RP11-267M23.4 |
RP11-267M23.4 |
| chr15_+_40226304 | 0.03 |
ENST00000559624.1 ENST00000382727.2 ENST00000263791.5 ENST00000560648.1 |
EIF2AK4 |
eukaryotic translation initiation factor 2 alpha kinase 4 |
| chr3_-_113897545 | 0.03 |
ENST00000467632.1 |
DRD3 |
dopamine receptor D3 |
| chr15_+_42651691 | 0.03 |
ENST00000357568.3 ENST00000349748.3 ENST00000318023.7 ENST00000397163.3 |
CAPN3 |
calpain 3, (p94) |
| chr2_-_157198860 | 0.03 |
ENST00000409572.1 |
NR4A2 |
nuclear receptor subfamily 4, group A, member 2 |
| chr8_-_95565673 | 0.03 |
ENST00000437199.1 ENST00000297591.5 ENST00000421249.2 |
KIAA1429 |
KIAA1429 |
| chr21_-_33984456 | 0.03 |
ENST00000431216.1 ENST00000553001.1 ENST00000440966.1 |
AP000275.65 C21orf59 |
Uncharacterized protein chromosome 21 open reading frame 59 |
| chr3_+_49057876 | 0.03 |
ENST00000326912.4 |
NDUFAF3 |
NADH dehydrogenase (ubiquinone) complex I, assembly factor 3 |
| chr20_+_44657845 | 0.03 |
ENST00000243964.3 |
SLC12A5 |
solute carrier family 12 (potassium/chloride transporter), member 5 |
| chr12_+_25205568 | 0.03 |
ENST00000548766.1 ENST00000556887.1 |
LRMP |
lymphoid-restricted membrane protein |
| chr12_-_7899958 | 0.03 |
ENST00000360345.3 |
CLEC4C |
C-type lectin domain family 4, member C |
| chr18_+_3252206 | 0.02 |
ENST00000578562.2 |
MYL12A |
myosin, light chain 12A, regulatory, non-sarcomeric |
| chr1_+_26348259 | 0.02 |
ENST00000374280.3 |
EXTL1 |
exostosin-like glycosyltransferase 1 |
| chr18_+_12407895 | 0.02 |
ENST00000590956.1 ENST00000336990.4 ENST00000440960.1 ENST00000588729.1 |
SLMO1 |
slowmo homolog 1 (Drosophila) |
| chr17_-_5322786 | 0.02 |
ENST00000225696.4 |
NUP88 |
nucleoporin 88kDa |
| chr11_-_47399942 | 0.02 |
ENST00000227163.4 |
SPI1 |
spleen focus forming virus (SFFV) proviral integration oncogene |
| chr20_+_10199468 | 0.02 |
ENST00000254976.2 ENST00000304886.2 |
SNAP25 |
synaptosomal-associated protein, 25kDa |
| chr18_-_47792851 | 0.02 |
ENST00000398545.4 |
CCDC11 |
coiled-coil domain containing 11 |
| chr17_+_7184986 | 0.02 |
ENST00000317370.8 ENST00000571308.1 |
SLC2A4 |
solute carrier family 2 (facilitated glucose transporter), member 4 |
| chr10_-_97200772 | 0.02 |
ENST00000371241.1 ENST00000354106.3 ENST00000371239.1 ENST00000361941.3 ENST00000277982.5 ENST00000371245.3 |
SORBS1 |
sorbin and SH3 domain containing 1 |
| chr7_-_113559104 | 0.01 |
ENST00000284601.3 |
PPP1R3A |
protein phosphatase 1, regulatory subunit 3A |
| chr10_-_75415825 | 0.01 |
ENST00000394810.2 |
SYNPO2L |
synaptopodin 2-like |
| chr20_-_54580523 | 0.01 |
ENST00000064571.2 |
CBLN4 |
cerebellin 4 precursor |
| chr13_+_39261224 | 0.01 |
ENST00000280481.7 |
FREM2 |
FRAS1 related extracellular matrix protein 2 |
| chr11_+_75110530 | 0.01 |
ENST00000531188.1 ENST00000530164.1 ENST00000422465.2 ENST00000278572.6 ENST00000534440.1 ENST00000527446.1 ENST00000526608.1 ENST00000527273.1 ENST00000524851.1 |
RPS3 |
ribosomal protein S3 |
| chr12_-_49393092 | 0.01 |
ENST00000421952.2 |
DDN |
dendrin |
| chr2_+_220299547 | 0.00 |
ENST00000312358.7 |
SPEG |
SPEG complex locus |
| chr19_-_51220176 | 0.00 |
ENST00000359082.3 ENST00000293441.1 |
SHANK1 |
SH3 and multiple ankyrin repeat domains 1 |
| chr7_+_143013198 | 0.00 |
ENST00000343257.2 |
CLCN1 |
chloride channel, voltage-sensitive 1 |
| chr3_-_57326704 | 0.00 |
ENST00000487349.1 ENST00000389601.3 |
ASB14 |
ankyrin repeat and SOCS box containing 14 |
Gene Ontology Analysis
Gene overrepresentation in biological_process category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.1 | 0.3 | GO:0061394 | regulation of transcription from RNA polymerase II promoter in response to arsenic-containing substance(GO:0061394) |
| 0.1 | 0.3 | GO:0000117 | regulation of transcription involved in G2/M transition of mitotic cell cycle(GO:0000117) |
| 0.1 | 0.5 | GO:1903899 | lung goblet cell differentiation(GO:0060480) positive regulation of PERK-mediated unfolded protein response(GO:1903899) |
| 0.1 | 0.4 | GO:1904844 | response to L-glutamine(GO:1904844) cellular response to L-glutamine(GO:1904845) |
| 0.1 | 0.4 | GO:0002074 | extraocular skeletal muscle development(GO:0002074) |
| 0.0 | 0.2 | GO:0015862 | uridine transport(GO:0015862) |
| 0.0 | 0.7 | GO:0090361 | platelet-derived growth factor production(GO:0090360) regulation of platelet-derived growth factor production(GO:0090361) |
| 0.0 | 0.5 | GO:0048842 | positive regulation of axon extension involved in axon guidance(GO:0048842) |
| 0.0 | 0.1 | GO:0032304 | negative regulation of icosanoid secretion(GO:0032304) |
| 0.0 | 1.3 | GO:0097503 | sialylation(GO:0097503) |
| 0.0 | 0.4 | GO:0086029 | Purkinje myocyte to ventricular cardiac muscle cell signaling(GO:0086029) Purkinje myocyte to ventricular cardiac muscle cell communication(GO:0086068) |
| 0.0 | 0.3 | GO:0043435 | response to corticotropin-releasing hormone(GO:0043435) cellular response to corticotropin-releasing hormone stimulus(GO:0071376) |
| 0.0 | 0.1 | GO:0014878 | response to electrical stimulus involved in regulation of muscle adaptation(GO:0014878) |
| 0.0 | 0.1 | GO:0032972 | diaphragm contraction(GO:0002086) regulation of muscle filament sliding speed(GO:0032972) |
| 0.0 | 0.1 | GO:1903660 | negative regulation of complement-dependent cytotoxicity(GO:1903660) |
| 0.0 | 1.8 | GO:0003009 | skeletal muscle contraction(GO:0003009) |
| 0.0 | 0.1 | GO:0044027 | hypermethylation of CpG island(GO:0044027) |
| 0.0 | 0.1 | GO:1900383 | regulation of synaptic plasticity by receptor localization to synapse(GO:1900383) |
| 0.0 | 0.2 | GO:0086073 | bundle of His cell-Purkinje myocyte adhesion involved in cell communication(GO:0086073) |
| 0.0 | 0.1 | GO:0010193 | response to ozone(GO:0010193) |
| 0.0 | 0.1 | GO:0060523 | prostate epithelial cord elongation(GO:0060523) |
| 0.0 | 0.1 | GO:0097338 | response to clozapine(GO:0097338) |
| 0.0 | 0.1 | GO:0021797 | forebrain anterior/posterior pattern specification(GO:0021797) |
| 0.0 | 0.1 | GO:1902866 | regulation of retina development in camera-type eye(GO:1902866) |
| 0.0 | 0.1 | GO:1903781 | endoplasmic reticulum membrane organization(GO:0090158) positive regulation of cardiac conduction(GO:1903781) |
| 0.0 | 1.1 | GO:0036092 | phosphatidylinositol-3-phosphate biosynthetic process(GO:0036092) |
| 0.0 | 0.7 | GO:0001958 | endochondral ossification(GO:0001958) replacement ossification(GO:0036075) |
| 0.0 | 0.0 | GO:0035995 | detection of muscle stretch(GO:0035995) |
| 0.0 | 0.0 | GO:0098735 | positive regulation of the force of heart contraction(GO:0098735) |
Gene overrepresentation in cellular_component category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.1 | 0.5 | GO:0005600 | collagen type XIII trimer(GO:0005600) transmembrane collagen trimer(GO:0030936) |
| 0.1 | 0.3 | GO:1990622 | CHOP-ATF3 complex(GO:1990622) |
| 0.1 | 2.1 | GO:0005861 | troponin complex(GO:0005861) |
| 0.0 | 0.4 | GO:0030314 | junctional membrane complex(GO:0030314) |
| 0.0 | 0.3 | GO:0032039 | integrator complex(GO:0032039) |
| 0.0 | 0.4 | GO:0043020 | NADPH oxidase complex(GO:0043020) |
| 0.0 | 0.4 | GO:0032982 | myosin filament(GO:0032982) |
| 0.0 | 0.1 | GO:0097550 | transcriptional preinitiation complex(GO:0097550) |
| 0.0 | 0.1 | GO:1990712 | HFE-transferrin receptor complex(GO:1990712) |
| 0.0 | 0.2 | GO:0005916 | fascia adherens(GO:0005916) |
Gene overrepresentation in molecular_function category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.2 | 1.1 | GO:0052828 | phosphatidylinositol-3,4-bisphosphate 4-phosphatase activity(GO:0016316) inositol-1,3,4-trisphosphate 4-phosphatase activity(GO:0017161) inositol-3,4-bisphosphate 4-phosphatase activity(GO:0052828) |
| 0.1 | 2.1 | GO:0031014 | troponin T binding(GO:0031014) |
| 0.1 | 0.8 | GO:0001665 | alpha-N-acetylgalactosaminide alpha-2,6-sialyltransferase activity(GO:0001665) |
| 0.1 | 0.4 | GO:0003835 | beta-galactoside alpha-2,6-sialyltransferase activity(GO:0003835) |
| 0.0 | 0.1 | GO:0097258 | tocopherol omega-hydroxylase activity(GO:0052870) alpha-tocopherol omega-hydroxylase activity(GO:0052871) 20-hydroxy-leukotriene B4 omega oxidase activity(GO:0097258) 20-aldehyde-leukotriene B4 20-monooxygenase activity(GO:0097259) |
| 0.0 | 0.1 | GO:0005136 | interleukin-4 receptor binding(GO:0005136) |
| 0.0 | 0.1 | GO:0001025 | RNA polymerase III transcription factor binding(GO:0001025) |
| 0.0 | 1.0 | GO:0008157 | protein phosphatase 1 binding(GO:0008157) |
| 0.0 | 0.1 | GO:0098821 | BMP receptor activity(GO:0098821) |
| 0.0 | 0.5 | GO:0002162 | dystroglycan binding(GO:0002162) |
| 0.0 | 0.1 | GO:0042835 | BRE binding(GO:0042835) |
| 0.0 | 0.4 | GO:0016175 | superoxide-generating NADPH oxidase activity(GO:0016175) |
| 0.0 | 0.1 | GO:0004991 | parathyroid hormone receptor activity(GO:0004991) |
| 0.0 | 0.2 | GO:0086083 | cell adhesive protein binding involved in bundle of His cell-Purkinje myocyte communication(GO:0086083) |
| 0.0 | 1.4 | GO:0048365 | Rac GTPase binding(GO:0048365) |
| 0.0 | 0.1 | GO:0038052 | RNA polymerase II transcription factor activity, estrogen-activated sequence-specific DNA binding(GO:0038052) |
| 0.0 | 0.4 | GO:0097493 | structural molecule activity conferring elasticity(GO:0097493) |
| 0.0 | 0.1 | GO:0004698 | calcium-dependent protein kinase C activity(GO:0004698) |
| 0.0 | 0.1 | GO:0034875 | oxidoreductase activity, acting on CH or CH2 groups, quinone or similar compound as acceptor(GO:0033695) caffeine oxidase activity(GO:0034875) |
| 0.0 | 0.3 | GO:0004745 | retinol dehydrogenase activity(GO:0004745) |
Gene overrepresentation in C2:CP category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.0 | 1.4 | SIG REGULATION OF THE ACTIN CYTOSKELETON BY RHO GTPASES | Genes related to regulation of the actin cytoskeleton |
Gene overrepresentation in C2:CP:REACTOME category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.1 | 1.4 | REACTOME ACTIVATION OF RAC | Genes involved in Activation of Rac |
| 0.1 | 1.1 | REACTOME SYNTHESIS OF PIPS AT THE EARLY ENDOSOME MEMBRANE | Genes involved in Synthesis of PIPs at the early endosome membrane |
| 0.0 | 0.5 | REACTOME DSCAM INTERACTIONS | Genes involved in DSCAM interactions |
| 0.0 | 1.8 | REACTOME STRIATED MUSCLE CONTRACTION | Genes involved in Striated Muscle Contraction |
| 0.0 | 0.4 | REACTOME N GLYCAN ANTENNAE ELONGATION | Genes involved in N-Glycan antennae elongation |
| 0.0 | 0.5 | REACTOME TIE2 SIGNALING | Genes involved in Tie2 Signaling |
| 0.0 | 0.5 | REACTOME COLLAGEN FORMATION | Genes involved in Collagen formation |