Project
Epithelial-Mesenchymal Transition, human (Scheel, 2011)
Navigation
Downloads
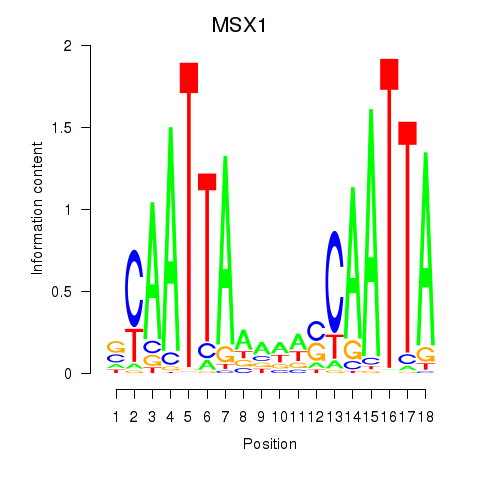
Results for MSX1
Z-value: 0.82
Transcription factors associated with MSX1
| Gene Symbol | Gene ID | Gene Info |
|---|---|---|
|
MSX1
|
ENSG00000163132.6 | MSX1 |
Activity-expression correlation:
| Gene | Promoter | Pearson corr. coef. | P-value | Plot |
|---|---|---|---|---|
| MSX1 | hg19_v2_chr4_+_4861385_4861398 | 0.71 | 4.7e-02 | Click! |
Activity profile of MSX1 motif
Sorted Z-values of MSX1 motif
Network of associatons between targets according to the STRING database.
First level regulatory network of MSX1
| Promoter | Score | Transcript | Gene | Gene Info |
|---|---|---|---|---|
| chr13_-_38172863 | 3.53 |
ENST00000541481.1 ENST00000379743.4 ENST00000379742.4 ENST00000379749.4 ENST00000541179.1 ENST00000379747.4 |
POSTN |
periostin, osteoblast specific factor |
| chr5_-_75919253 | 1.13 |
ENST00000296641.4 |
F2RL2 |
coagulation factor II (thrombin) receptor-like 2 |
| chr2_-_145278475 | 1.08 |
ENST00000558170.2 |
ZEB2 |
zinc finger E-box binding homeobox 2 |
| chr8_+_97597148 | 0.98 |
ENST00000521590.1 |
SDC2 |
syndecan 2 |
| chr2_-_158345462 | 0.79 |
ENST00000439355.1 ENST00000540637.1 |
CYTIP |
cytohesin 1 interacting protein |
| chr9_+_79074068 | 0.76 |
ENST00000444201.2 ENST00000376730.4 |
GCNT1 |
glucosaminyl (N-acetyl) transferase 1, core 2 |
| chr1_-_72566613 | 0.75 |
ENST00000306821.3 |
NEGR1 |
neuronal growth regulator 1 |
| chr6_+_28249332 | 0.71 |
ENST00000259883.3 |
PGBD1 |
piggyBac transposable element derived 1 |
| chr6_+_28249299 | 0.71 |
ENST00000405948.2 |
PGBD1 |
piggyBac transposable element derived 1 |
| chr9_+_90112117 | 0.71 |
ENST00000358077.5 |
DAPK1 |
death-associated protein kinase 1 |
| chr5_+_82767583 | 0.68 |
ENST00000512590.2 ENST00000513960.1 ENST00000513984.1 ENST00000502527.2 |
VCAN |
versican |
| chr5_+_82767487 | 0.65 |
ENST00000343200.5 ENST00000342785.4 |
VCAN |
versican |
| chr5_+_82767284 | 0.64 |
ENST00000265077.3 |
VCAN |
versican |
| chr14_+_52327109 | 0.64 |
ENST00000335281.4 |
GNG2 |
guanine nucleotide binding protein (G protein), gamma 2 |
| chr19_-_53662257 | 0.62 |
ENST00000599096.1 ENST00000334197.7 ENST00000597183.1 ENST00000601804.1 ENST00000601469.2 ENST00000452676.2 |
ZNF347 |
zinc finger protein 347 |
| chr5_-_75919217 | 0.58 |
ENST00000504899.1 |
F2RL2 |
coagulation factor II (thrombin) receptor-like 2 |
| chr18_-_70532906 | 0.48 |
ENST00000299430.2 ENST00000397929.1 |
NETO1 |
neuropilin (NRP) and tolloid (TLL)-like 1 |
| chr5_+_67586465 | 0.48 |
ENST00000336483.5 |
PIK3R1 |
phosphoinositide-3-kinase, regulatory subunit 1 (alpha) |
| chr4_-_57547454 | 0.46 |
ENST00000556376.2 |
HOPX |
HOP homeobox |
| chr11_+_12766583 | 0.34 |
ENST00000361985.2 |
TEAD1 |
TEA domain family member 1 (SV40 transcriptional enhancer factor) |
| chr1_+_210501589 | 0.31 |
ENST00000413764.2 ENST00000541565.1 |
HHAT |
hedgehog acyltransferase |
| chr11_+_12132117 | 0.30 |
ENST00000256194.4 |
MICAL2 |
microtubule associated monooxygenase, calponin and LIM domain containing 2 |
| chr16_-_66583701 | 0.29 |
ENST00000527800.1 ENST00000525974.1 ENST00000563369.2 |
TK2 |
thymidine kinase 2, mitochondrial |
| chr6_+_28092338 | 0.29 |
ENST00000340487.4 |
ZSCAN16 |
zinc finger and SCAN domain containing 16 |
| chr19_-_53757941 | 0.28 |
ENST00000594517.1 ENST00000601413.1 ENST00000594681.1 |
ZNF677 |
zinc finger protein 677 |
| chr18_-_5396271 | 0.28 |
ENST00000579951.1 |
EPB41L3 |
erythrocyte membrane protein band 4.1-like 3 |
| chr5_+_140762268 | 0.26 |
ENST00000518325.1 |
PCDHGA7 |
protocadherin gamma subfamily A, 7 |
| chr2_+_158114051 | 0.25 |
ENST00000259056.4 |
GALNT5 |
UDP-N-acetyl-alpha-D-galactosamine:polypeptide N-acetylgalactosaminyltransferase 5 (GalNAc-T5) |
| chr7_+_148287657 | 0.25 |
ENST00000307003.2 |
C7orf33 |
chromosome 7 open reading frame 33 |
| chr11_+_101785727 | 0.25 |
ENST00000263468.8 |
KIAA1377 |
KIAA1377 |
| chr16_-_66584059 | 0.24 |
ENST00000417693.3 ENST00000544898.1 ENST00000569718.1 ENST00000527284.1 ENST00000299697.7 ENST00000451102.2 |
TK2 |
thymidine kinase 2, mitochondrial |
| chr17_-_5321549 | 0.24 |
ENST00000572809.1 |
NUP88 |
nucleoporin 88kDa |
| chr20_+_56964169 | 0.23 |
ENST00000475243.1 |
VAPB |
VAMP (vesicle-associated membrane protein)-associated protein B and C |
| chr8_-_25281747 | 0.21 |
ENST00000421054.2 |
GNRH1 |
gonadotropin-releasing hormone 1 (luteinizing-releasing hormone) |
| chr6_+_136172820 | 0.21 |
ENST00000308191.6 |
PDE7B |
phosphodiesterase 7B |
| chr1_+_224370873 | 0.21 |
ENST00000323699.4 ENST00000391877.3 |
DEGS1 |
delta(4)-desaturase, sphingolipid 1 |
| chr6_-_109777128 | 0.21 |
ENST00000358807.3 ENST00000358577.3 |
MICAL1 |
microtubule associated monooxygenase, calponin and LIM domain containing 1 |
| chr6_+_26240561 | 0.20 |
ENST00000377745.2 |
HIST1H4F |
histone cluster 1, H4f |
| chr19_-_53758094 | 0.20 |
ENST00000601828.1 ENST00000598513.1 ENST00000599012.1 ENST00000333952.4 ENST00000598806.1 |
ZNF677 |
zinc finger protein 677 |
| chr17_-_8113542 | 0.20 |
ENST00000578549.1 ENST00000535053.1 ENST00000582368.1 |
AURKB |
aurora kinase B |
| chr10_+_17270214 | 0.20 |
ENST00000544301.1 |
VIM |
vimentin |
| chr16_+_21689835 | 0.20 |
ENST00000286149.4 ENST00000388958.3 |
OTOA |
otoancorin |
| chr13_+_50570019 | 0.20 |
ENST00000442421.1 |
TRIM13 |
tripartite motif containing 13 |
| chr12_+_54422142 | 0.19 |
ENST00000243108.4 |
HOXC6 |
homeobox C6 |
| chr13_+_49551020 | 0.18 |
ENST00000541916.1 |
FNDC3A |
fibronectin type III domain containing 3A |
| chr11_-_102401469 | 0.18 |
ENST00000260227.4 |
MMP7 |
matrix metallopeptidase 7 (matrilysin, uterine) |
| chr12_-_10978957 | 0.17 |
ENST00000240619.2 |
TAS2R10 |
taste receptor, type 2, member 10 |
| chr1_-_54405773 | 0.17 |
ENST00000371376.1 |
HSPB11 |
heat shock protein family B (small), member 11 |
| chr1_+_84630645 | 0.17 |
ENST00000394839.2 |
PRKACB |
protein kinase, cAMP-dependent, catalytic, beta |
| chr10_+_13142225 | 0.16 |
ENST00000378747.3 |
OPTN |
optineurin |
| chr11_+_61197508 | 0.16 |
ENST00000541135.1 ENST00000301761.2 |
RP11-286N22.8 SDHAF2 |
Uncharacterized protein succinate dehydrogenase complex assembly factor 2 |
| chr6_+_131894284 | 0.15 |
ENST00000368087.3 ENST00000356962.2 |
ARG1 |
arginase 1 |
| chr6_+_6588316 | 0.15 |
ENST00000379953.2 |
LY86 |
lymphocyte antigen 86 |
| chr4_-_186696425 | 0.15 |
ENST00000430503.1 ENST00000319454.6 ENST00000450341.1 |
SORBS2 |
sorbin and SH3 domain containing 2 |
| chr16_-_71518504 | 0.15 |
ENST00000565100.2 |
ZNF19 |
zinc finger protein 19 |
| chr1_+_158985457 | 0.15 |
ENST00000567661.1 ENST00000474473.1 |
IFI16 |
interferon, gamma-inducible protein 16 |
| chr5_+_175490540 | 0.14 |
ENST00000515817.1 |
FAM153B |
family with sequence similarity 153, member B |
| chr4_+_113568207 | 0.14 |
ENST00000511529.1 |
LARP7 |
La ribonucleoprotein domain family, member 7 |
| chr15_-_95870310 | 0.14 |
ENST00000508732.2 |
CTD-2536I1.1 |
CTD-2536I1.1 |
| chr2_-_169746878 | 0.13 |
ENST00000282074.2 |
SPC25 |
SPC25, NDC80 kinetochore complex component |
| chr14_+_52327350 | 0.13 |
ENST00000555472.1 ENST00000556766.1 |
GNG2 |
guanine nucleotide binding protein (G protein), gamma 2 |
| chr11_+_62037622 | 0.12 |
ENST00000227918.2 ENST00000525380.1 |
SCGB2A2 |
secretoglobin, family 2A, member 2 |
| chr13_+_103459704 | 0.12 |
ENST00000602836.1 |
BIVM-ERCC5 |
BIVM-ERCC5 readthrough |
| chr11_+_61197572 | 0.11 |
ENST00000542074.1 ENST00000534878.1 ENST00000537782.1 ENST00000543265.1 |
SDHAF2 |
succinate dehydrogenase complex assembly factor 2 |
| chr9_+_133539981 | 0.11 |
ENST00000253008.2 |
PRDM12 |
PR domain containing 12 |
| chr8_-_121824374 | 0.11 |
ENST00000517992.1 |
SNTB1 |
syntrophin, beta 1 (dystrophin-associated protein A1, 59kDa, basic component 1) |
| chr15_-_77712477 | 0.11 |
ENST00000560626.2 |
PEAK1 |
pseudopodium-enriched atypical kinase 1 |
| chr21_+_33671160 | 0.11 |
ENST00000303645.5 |
MRAP |
melanocortin 2 receptor accessory protein |
| chr19_+_16059818 | 0.11 |
ENST00000322107.1 |
OR10H4 |
olfactory receptor, family 10, subfamily H, member 4 |
| chr10_+_135050908 | 0.11 |
ENST00000325980.9 |
VENTX |
VENT homeobox |
| chr9_-_15510287 | 0.10 |
ENST00000397519.2 |
PSIP1 |
PC4 and SFRS1 interacting protein 1 |
| chr5_-_177207634 | 0.10 |
ENST00000513554.1 ENST00000440605.3 |
FAM153A |
family with sequence similarity 153, member A |
| chr14_+_55494323 | 0.10 |
ENST00000339298.2 |
SOCS4 |
suppressor of cytokine signaling 4 |
| chrX_-_103499602 | 0.09 |
ENST00000372588.4 |
ESX1 |
ESX homeobox 1 |
| chr10_+_13141585 | 0.09 |
ENST00000378764.2 |
OPTN |
optineurin |
| chr10_-_32667660 | 0.09 |
ENST00000375110.2 |
EPC1 |
enhancer of polycomb homolog 1 (Drosophila) |
| chr3_-_47555167 | 0.09 |
ENST00000296149.4 |
ELP6 |
elongator acetyltransferase complex subunit 6 |
| chr14_+_55595762 | 0.09 |
ENST00000254301.9 |
LGALS3 |
lectin, galactoside-binding, soluble, 3 |
| chr11_+_94245617 | 0.09 |
ENST00000542198.1 |
RP11-867G2.2 |
long intergenic non-protein coding RNA 1171 |
| chr7_+_7811992 | 0.09 |
ENST00000406829.1 |
RPA3-AS1 |
RPA3 antisense RNA 1 |
| chr3_+_114012819 | 0.09 |
ENST00000383671.3 |
TIGIT |
T cell immunoreceptor with Ig and ITIM domains |
| chr15_+_79166065 | 0.09 |
ENST00000559690.1 ENST00000559158.1 |
MORF4L1 |
mortality factor 4 like 1 |
| chr14_+_55493920 | 0.09 |
ENST00000395472.2 ENST00000555846.1 |
SOCS4 |
suppressor of cytokine signaling 4 |
| chr6_+_118869452 | 0.08 |
ENST00000357525.5 |
PLN |
phospholamban |
| chr3_+_108308513 | 0.08 |
ENST00000361582.3 |
DZIP3 |
DAZ interacting zinc finger protein 3 |
| chr14_-_75389925 | 0.08 |
ENST00000556776.1 |
RPS6KL1 |
ribosomal protein S6 kinase-like 1 |
| chr20_+_5987890 | 0.08 |
ENST00000378868.4 |
CRLS1 |
cardiolipin synthase 1 |
| chr11_-_66964638 | 0.08 |
ENST00000444002.2 |
AP001885.1 |
AP001885.1 |
| chr15_-_42500351 | 0.08 |
ENST00000348544.4 ENST00000318006.5 |
VPS39 |
vacuolar protein sorting 39 homolog (S. cerevisiae) |
| chr14_+_55595960 | 0.08 |
ENST00000554715.1 |
LGALS3 |
lectin, galactoside-binding, soluble, 3 |
| chr8_+_28748765 | 0.07 |
ENST00000355231.5 |
HMBOX1 |
homeobox containing 1 |
| chr14_+_22964877 | 0.07 |
ENST00000390494.1 |
TRAJ43 |
T cell receptor alpha joining 43 |
| chr5_-_11588907 | 0.07 |
ENST00000513598.1 ENST00000503622.1 |
CTNND2 |
catenin (cadherin-associated protein), delta 2 |
| chr12_-_11150474 | 0.07 |
ENST00000538986.1 |
TAS2R20 |
taste receptor, type 2, member 20 |
| chr11_-_114430570 | 0.07 |
ENST00000251921.2 |
NXPE1 |
neurexophilin and PC-esterase domain family, member 1 |
| chr2_+_196313239 | 0.07 |
ENST00000413290.1 |
AC064834.1 |
AC064834.1 |
| chr7_+_23338819 | 0.07 |
ENST00000466681.1 |
MALSU1 |
mitochondrial assembly of ribosomal large subunit 1 |
| chr16_+_69345243 | 0.07 |
ENST00000254950.11 |
VPS4A |
vacuolar protein sorting 4 homolog A (S. cerevisiae) |
| chr20_+_34129770 | 0.07 |
ENST00000348547.2 ENST00000357394.4 ENST00000447986.1 ENST00000279052.6 ENST00000416206.1 ENST00000411577.1 ENST00000413587.1 |
ERGIC3 |
ERGIC and golgi 3 |
| chr15_+_51669444 | 0.07 |
ENST00000396399.2 |
GLDN |
gliomedin |
| chr1_+_202091980 | 0.06 |
ENST00000367282.5 |
GPR37L1 |
G protein-coupled receptor 37 like 1 |
| chr4_-_100140331 | 0.06 |
ENST00000407820.2 ENST00000394897.1 ENST00000508558.1 ENST00000394899.2 |
ADH6 |
alcohol dehydrogenase 6 (class V) |
| chr13_-_81801115 | 0.06 |
ENST00000567258.1 |
LINC00564 |
long intergenic non-protein coding RNA 564 |
| chr17_-_37934466 | 0.06 |
ENST00000583368.1 |
IKZF3 |
IKAROS family zinc finger 3 (Aiolos) |
| chr19_-_7812446 | 0.06 |
ENST00000394173.4 ENST00000301357.8 |
CD209 |
CD209 molecule |
| chr20_+_5731027 | 0.06 |
ENST00000378979.4 ENST00000303142.6 |
C20orf196 |
chromosome 20 open reading frame 196 |
| chr8_+_24241969 | 0.05 |
ENST00000522298.1 |
ADAMDEC1 |
ADAM-like, decysin 1 |
| chr2_-_109605663 | 0.05 |
ENST00000409271.1 ENST00000258443.2 ENST00000376651.1 |
EDAR |
ectodysplasin A receptor |
| chr3_-_21792838 | 0.05 |
ENST00000281523.2 |
ZNF385D |
zinc finger protein 385D |
| chr10_+_135204338 | 0.05 |
ENST00000468317.2 |
RP11-108K14.8 |
Mitochondrial GTPase 1 |
| chr6_-_31830655 | 0.05 |
ENST00000375631.4 |
NEU1 |
sialidase 1 (lysosomal sialidase) |
| chr6_-_76072719 | 0.05 |
ENST00000370020.1 |
FILIP1 |
filamin A interacting protein 1 |
| chr1_+_160160283 | 0.05 |
ENST00000368079.3 |
CASQ1 |
calsequestrin 1 (fast-twitch, skeletal muscle) |
| chr9_+_75229616 | 0.05 |
ENST00000340019.3 |
TMC1 |
transmembrane channel-like 1 |
| chr8_+_24241789 | 0.05 |
ENST00000256412.4 ENST00000538205.1 |
ADAMDEC1 |
ADAM-like, decysin 1 |
| chr1_-_167487808 | 0.05 |
ENST00000392122.3 |
CD247 |
CD247 molecule |
| chr14_-_101295407 | 0.05 |
ENST00000596284.1 |
AL117190.2 |
AL117190.2 |
| chr21_+_17792672 | 0.05 |
ENST00000602620.1 |
LINC00478 |
long intergenic non-protein coding RNA 478 |
| chr14_+_77582905 | 0.05 |
ENST00000557408.1 |
TMEM63C |
transmembrane protein 63C |
| chr1_+_8021954 | 0.05 |
ENST00000377491.1 ENST00000377488.1 |
PARK7 |
parkinson protein 7 |
| chr3_-_93781750 | 0.04 |
ENST00000314636.2 |
DHFRL1 |
dihydrofolate reductase-like 1 |
| chr21_+_33671264 | 0.04 |
ENST00000339944.4 |
MRAP |
melanocortin 2 receptor accessory protein |
| chr7_+_107220899 | 0.04 |
ENST00000379117.2 ENST00000473124.1 |
BCAP29 |
B-cell receptor-associated protein 29 |
| chr1_+_160160346 | 0.04 |
ENST00000368078.3 |
CASQ1 |
calsequestrin 1 (fast-twitch, skeletal muscle) |
| chr5_-_147162078 | 0.04 |
ENST00000507386.1 |
JAKMIP2 |
janus kinase and microtubule interacting protein 2 |
| chr7_+_99425633 | 0.04 |
ENST00000354829.2 ENST00000421837.2 ENST00000417625.1 ENST00000342499.4 ENST00000444905.1 ENST00000415413.1 ENST00000312017.5 ENST00000222382.5 |
CYP3A43 |
cytochrome P450, family 3, subfamily A, polypeptide 43 |
| chr1_-_167487758 | 0.04 |
ENST00000362089.5 |
CD247 |
CD247 molecule |
| chr18_+_61637159 | 0.04 |
ENST00000397985.2 ENST00000353706.2 ENST00000542677.1 ENST00000397988.3 ENST00000448851.1 |
SERPINB8 |
serpin peptidase inhibitor, clade B (ovalbumin), member 8 |
| chr9_-_70465758 | 0.03 |
ENST00000489273.1 |
CBWD5 |
COBW domain containing 5 |
| chr11_-_92931098 | 0.03 |
ENST00000326402.4 |
SLC36A4 |
solute carrier family 36 (proton/amino acid symporter), member 4 |
| chr6_-_32908765 | 0.03 |
ENST00000416244.2 |
HLA-DMB |
major histocompatibility complex, class II, DM beta |
| chr22_+_41697520 | 0.03 |
ENST00000352645.4 |
ZC3H7B |
zinc finger CCCH-type containing 7B |
| chrX_+_11311533 | 0.03 |
ENST00000380714.3 ENST00000380712.3 ENST00000348912.4 |
AMELX |
amelogenin, X-linked |
| chr1_-_149908217 | 0.03 |
ENST00000369140.3 |
MTMR11 |
myotubularin related protein 11 |
| chr10_-_29923893 | 0.03 |
ENST00000355867.4 |
SVIL |
supervillin |
| chr2_-_136288113 | 0.03 |
ENST00000401392.1 |
ZRANB3 |
zinc finger, RAN-binding domain containing 3 |
| chr7_+_138818490 | 0.03 |
ENST00000430935.1 ENST00000495038.1 ENST00000474035.2 ENST00000478836.2 ENST00000464848.1 ENST00000343187.4 |
TTC26 |
tetratricopeptide repeat domain 26 |
| chr19_-_20150014 | 0.03 |
ENST00000358523.5 ENST00000397162.1 ENST00000601100.1 |
ZNF682 |
zinc finger protein 682 |
| chr9_+_44868935 | 0.03 |
ENST00000448436.2 |
RP11-160N1.10 |
RP11-160N1.10 |
| chr19_+_55043977 | 0.02 |
ENST00000335056.3 |
KIR3DX1 |
killer cell immunoglobulin-like receptor, three domains, X1 |
| chr9_-_86593238 | 0.02 |
ENST00000351839.3 |
HNRNPK |
heterogeneous nuclear ribonucleoprotein K |
| chr1_+_76540386 | 0.02 |
ENST00000328299.3 |
ST6GALNAC3 |
ST6 (alpha-N-acetyl-neuraminyl-2,3-beta-galactosyl-1,3)-N-acetylgalactosaminide alpha-2,6-sialyltransferase 3 |
| chrX_-_122756660 | 0.02 |
ENST00000441692.1 |
THOC2 |
THO complex 2 |
| chr6_+_108616243 | 0.02 |
ENST00000421954.1 |
LACE1 |
lactation elevated 1 |
| chr3_+_93781728 | 0.02 |
ENST00000314622.4 |
NSUN3 |
NOP2/Sun domain family, member 3 |
| chr11_-_5345582 | 0.01 |
ENST00000328813.2 |
OR51B2 |
olfactory receptor, family 51, subfamily B, member 2 |
| chr19_-_7812397 | 0.01 |
ENST00000593660.1 ENST00000354397.6 ENST00000593821.1 ENST00000602261.1 ENST00000315591.8 ENST00000394161.5 ENST00000204801.8 ENST00000601256.1 ENST00000601951.1 ENST00000315599.7 |
CD209 |
CD209 molecule |
| chr11_+_60681346 | 0.01 |
ENST00000227525.3 |
TMEM109 |
transmembrane protein 109 |
| chr13_-_52733980 | 0.01 |
ENST00000339406.3 |
NEK3 |
NIMA-related kinase 3 |
| chr4_+_159442878 | 0.01 |
ENST00000307765.5 ENST00000423548.1 |
RXFP1 |
relaxin/insulin-like family peptide receptor 1 |
| chr8_-_72274095 | 0.00 |
ENST00000303824.7 |
EYA1 |
eyes absent homolog 1 (Drosophila) |
| chr11_+_92085262 | 0.00 |
ENST00000298047.6 ENST00000409404.2 ENST00000541502.1 |
FAT3 |
FAT atypical cadherin 3 |
| chr7_+_2636522 | 0.00 |
ENST00000423196.1 |
IQCE |
IQ motif containing E |
| chr4_-_164534657 | 0.00 |
ENST00000339875.5 |
MARCH1 |
membrane-associated ring finger (C3HC4) 1, E3 ubiquitin protein ligase |
| chr12_-_82752565 | 0.00 |
ENST00000256151.7 |
CCDC59 |
coiled-coil domain containing 59 |
Gene Ontology Analysis
Gene overrepresentation in biological_process category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.7 | 3.5 | GO:0032571 | response to vitamin K(GO:0032571) bone regeneration(GO:1990523) |
| 0.2 | 1.1 | GO:1902748 | positive regulation of lens fiber cell differentiation(GO:1902748) |
| 0.2 | 0.5 | GO:0019417 | sulfur oxidation(GO:0019417) |
| 0.2 | 1.0 | GO:0008218 | bioluminescence(GO:0008218) |
| 0.1 | 0.5 | GO:0046104 | thymidine metabolic process(GO:0046104) pyrimidine deoxyribonucleoside metabolic process(GO:0046125) |
| 0.1 | 0.5 | GO:2000312 | regulation of kainate selective glutamate receptor activity(GO:2000312) |
| 0.1 | 0.3 | GO:0034553 | respiratory chain complex II assembly(GO:0034552) mitochondrial respiratory chain complex II assembly(GO:0034553) mitochondrial respiratory chain complex II biogenesis(GO:0097032) |
| 0.1 | 1.7 | GO:0070493 | thrombin receptor signaling pathway(GO:0070493) |
| 0.1 | 2.0 | GO:0030208 | dermatan sulfate biosynthetic process(GO:0030208) |
| 0.1 | 0.8 | GO:0060352 | cell adhesion molecule production(GO:0060352) |
| 0.1 | 0.2 | GO:1990637 | response to prolactin(GO:1990637) |
| 0.0 | 0.3 | GO:0044878 | mitotic cytokinesis checkpoint(GO:0044878) |
| 0.0 | 0.1 | GO:1900111 | positive regulation of histone H3-K9 dimethylation(GO:1900111) |
| 0.0 | 0.3 | GO:0002175 | protein localization to paranode region of axon(GO:0002175) |
| 0.0 | 0.2 | GO:2000521 | negative regulation of immunological synapse formation(GO:2000521) |
| 0.0 | 0.8 | GO:0007191 | adenylate cyclase-activating dopamine receptor signaling pathway(GO:0007191) |
| 0.0 | 0.2 | GO:0090467 | L-arginine import(GO:0043091) arginine import(GO:0090467) |
| 0.0 | 0.7 | GO:0071447 | cellular response to hydroperoxide(GO:0071447) |
| 0.0 | 0.3 | GO:0001920 | negative regulation of receptor recycling(GO:0001920) |
| 0.0 | 0.1 | GO:0086092 | regulation of the force of heart contraction by cardiac conduction(GO:0086092) regulation of calcium ion binding(GO:1901876) negative regulation of calcium ion binding(GO:1901877) regulation of calcium ion import into sarcoplasmic reticulum(GO:1902080) negative regulation of calcium ion import into sarcoplasmic reticulum(GO:1902081) |
| 0.0 | 0.2 | GO:0000055 | ribosomal large subunit export from nucleus(GO:0000055) ribosomal small subunit export from nucleus(GO:0000056) |
| 0.0 | 0.2 | GO:0097338 | response to clozapine(GO:0097338) |
| 0.0 | 0.5 | GO:0051531 | NFAT protein import into nucleus(GO:0051531) |
| 0.0 | 0.1 | GO:0090071 | negative regulation of ribosome biogenesis(GO:0090071) |
| 0.0 | 0.2 | GO:0031666 | positive regulation of lipopolysaccharide-mediated signaling pathway(GO:0031666) |
| 0.0 | 0.2 | GO:0060020 | Bergmann glial cell differentiation(GO:0060020) |
| 0.0 | 0.5 | GO:0001829 | trophectodermal cell differentiation(GO:0001829) |
| 0.0 | 0.2 | GO:0006987 | activation of signaling protein activity involved in unfolded protein response(GO:0006987) |
| 0.0 | 0.0 | GO:1903384 | enzyme active site formation(GO:0018307) neuron intrinsic apoptotic signaling pathway in response to hydrogen peroxide(GO:0036482) detoxification of mercury ion(GO:0050787) positive regulation of mitochondrial electron transport, NADH to ubiquinone(GO:1902958) regulation of protein K48-linked deubiquitination(GO:1903093) negative regulation of protein K48-linked deubiquitination(GO:1903094) regulation of dopamine biosynthetic process(GO:1903179) positive regulation of dopamine biosynthetic process(GO:1903181) regulation of hydrogen peroxide-induced neuron intrinsic apoptotic signaling pathway(GO:1903383) negative regulation of hydrogen peroxide-induced neuron intrinsic apoptotic signaling pathway(GO:1903384) negative regulation of ubiquitin-specific protease activity(GO:2000157) |
| 0.0 | 0.0 | GO:0036292 | DNA rewinding(GO:0036292) |
| 0.0 | 0.1 | GO:0046968 | peptide antigen transport(GO:0046968) |
| 0.0 | 0.1 | GO:1902774 | late endosome to lysosome transport(GO:1902774) |
| 0.0 | 0.1 | GO:0000395 | mRNA 5'-splice site recognition(GO:0000395) |
| 0.0 | 0.1 | GO:1904386 | response to thyroxine(GO:0097068) response to L-phenylalanine derivative(GO:1904386) |
| 0.0 | 0.0 | GO:2001190 | positive regulation of T cell activation via T cell receptor contact with antigen bound to MHC molecule on antigen presenting cell(GO:2001190) |
Gene overrepresentation in cellular_component category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.1 | 0.5 | GO:1990578 | perinuclear endoplasmic reticulum membrane(GO:1990578) |
| 0.1 | 0.3 | GO:0071148 | TEAD-1-YAP complex(GO:0071148) |
| 0.0 | 0.5 | GO:0098839 | postsynaptic density membrane(GO:0098839) |
| 0.0 | 3.5 | GO:0031594 | neuromuscular junction(GO:0031594) |
| 0.0 | 0.2 | GO:0032133 | chromosome passenger complex(GO:0032133) |
| 0.0 | 0.2 | GO:0098554 | cytoplasmic side of endoplasmic reticulum membrane(GO:0098554) |
| 0.0 | 0.1 | GO:0031262 | Ndc80 complex(GO:0031262) |
| 0.0 | 3.0 | GO:0005796 | Golgi lumen(GO:0005796) |
| 0.0 | 0.1 | GO:0032777 | Piccolo NuA4 histone acetyltransferase complex(GO:0032777) |
| 0.0 | 0.8 | GO:0005834 | heterotrimeric G-protein complex(GO:0005834) |
| 0.0 | 0.3 | GO:0033270 | paranode region of axon(GO:0033270) |
| 0.0 | 0.2 | GO:0005952 | cAMP-dependent protein kinase complex(GO:0005952) |
| 0.0 | 0.1 | GO:0014802 | terminal cisterna(GO:0014802) |
| 0.0 | 0.1 | GO:0000836 | Hrd1p ubiquitin ligase complex(GO:0000836) |
Gene overrepresentation in molecular_function category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.3 | 0.8 | GO:0003829 | beta-1,3-galactosyl-O-glycosyl-glycoprotein beta-1,6-N-acetylglucosaminyltransferase activity(GO:0003829) |
| 0.2 | 1.7 | GO:0015057 | thrombin receptor activity(GO:0015057) |
| 0.2 | 0.5 | GO:0004137 | deoxycytidine kinase activity(GO:0004137) thymidine kinase activity(GO:0004797) |
| 0.1 | 0.2 | GO:0033149 | FFAT motif binding(GO:0033149) |
| 0.1 | 0.8 | GO:0031681 | G-protein beta-subunit binding(GO:0031681) |
| 0.1 | 2.0 | GO:0005540 | hyaluronic acid binding(GO:0005540) |
| 0.1 | 0.5 | GO:0043559 | insulin binding(GO:0043559) |
| 0.0 | 0.1 | GO:0031783 | corticotropin hormone receptor binding(GO:0031780) type 5 melanocortin receptor binding(GO:0031783) |
| 0.0 | 0.2 | GO:1990254 | keratin filament binding(GO:1990254) |
| 0.0 | 0.1 | GO:0030572 | cardiolipin synthase activity(GO:0008808) phosphatidyltransferase activity(GO:0030572) CDP-diacylglycerol-phosphatidylglycerol phosphatidyltransferase activity(GO:0043337) |
| 0.0 | 0.2 | GO:0035174 | histone serine kinase activity(GO:0035174) |
| 0.0 | 0.2 | GO:0019863 | IgE binding(GO:0019863) |
| 0.0 | 0.7 | GO:0017075 | syntaxin-1 binding(GO:0017075) |
| 0.0 | 1.1 | GO:0070412 | R-SMAD binding(GO:0070412) |
| 0.0 | 1.4 | GO:0005044 | scavenger receptor activity(GO:0005044) |
| 0.0 | 0.5 | GO:0071949 | FAD binding(GO:0071949) |
| 0.0 | 0.3 | GO:1904264 | ubiquitin protein ligase activity involved in ERAD pathway(GO:1904264) |
| 0.0 | 3.7 | GO:0008201 | heparin binding(GO:0008201) |
| 0.0 | 0.3 | GO:0001134 | transcription factor activity, transcription factor recruiting(GO:0001134) |
| 0.0 | 0.2 | GO:0033038 | bitter taste receptor activity(GO:0033038) |
| 0.0 | 0.1 | GO:0042030 | ATPase inhibitor activity(GO:0042030) |
| 0.0 | 0.1 | GO:1990226 | histone methyltransferase binding(GO:1990226) |
| 0.0 | 0.0 | GO:0016532 | superoxide dismutase copper chaperone activity(GO:0016532) |
| 0.0 | 0.0 | GO:0015173 | aromatic amino acid transmembrane transporter activity(GO:0015173) |
| 0.0 | 0.2 | GO:0004691 | cAMP-dependent protein kinase activity(GO:0004691) |
| 0.0 | 0.1 | GO:0052795 | exo-alpha-(2->3)-sialidase activity(GO:0052794) exo-alpha-(2->6)-sialidase activity(GO:0052795) exo-alpha-(2->8)-sialidase activity(GO:0052796) |
| 0.0 | 0.3 | GO:0070530 | K63-linked polyubiquitin binding(GO:0070530) |
| 0.0 | 0.0 | GO:0046848 | hydroxyapatite binding(GO:0046848) |
Gene overrepresentation in C2:CP category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.1 | 2.5 | PID THROMBIN PAR4 PATHWAY | PAR4-mediated thrombin signaling events |
| 0.1 | 2.0 | PID TOLL ENDOGENOUS PATHWAY | Endogenous TLR signaling |
| 0.0 | 0.8 | PID HEDGEHOG 2PATHWAY | Signaling events mediated by the Hedgehog family |
| 0.0 | 1.0 | PID SYNDECAN 2 PATHWAY | Syndecan-2-mediated signaling events |
| 0.0 | 0.7 | PID IFNG PATHWAY | IFN-gamma pathway |
| 0.0 | 3.5 | NABA ECM GLYCOPROTEINS | Genes encoding structural ECM glycoproteins |
Gene overrepresentation in C2:CP:REACTOME category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.1 | 2.0 | REACTOME CS DS DEGRADATION | Genes involved in CS/DS degradation |
| 0.0 | 0.7 | REACTOME ROLE OF DCC IN REGULATING APOPTOSIS | Genes involved in Role of DCC in regulating apoptosis |
| 0.0 | 2.5 | REACTOME THROMBIN SIGNALLING THROUGH PROTEINASE ACTIVATED RECEPTORS PARS | Genes involved in Thrombin signalling through proteinase activated receptors (PARs) |
| 0.0 | 1.0 | REACTOME A TETRASACCHARIDE LINKER SEQUENCE IS REQUIRED FOR GAG SYNTHESIS | Genes involved in A tetrasaccharide linker sequence is required for GAG synthesis |
| 0.0 | 0.5 | REACTOME IL 7 SIGNALING | Genes involved in Interleukin-7 signaling |
| 0.0 | 0.5 | REACTOME G1 S SPECIFIC TRANSCRIPTION | Genes involved in G1/S-Specific Transcription |
| 0.0 | 0.2 | REACTOME HORMONE LIGAND BINDING RECEPTORS | Genes involved in Hormone ligand-binding receptors |
| 0.0 | 0.1 | REACTOME APOBEC3G MEDIATED RESISTANCE TO HIV1 INFECTION | Genes involved in APOBEC3G mediated resistance to HIV-1 infection |