Project
Epithelial-Mesenchymal Transition, human (Scheel, 2011)
Navigation
Downloads
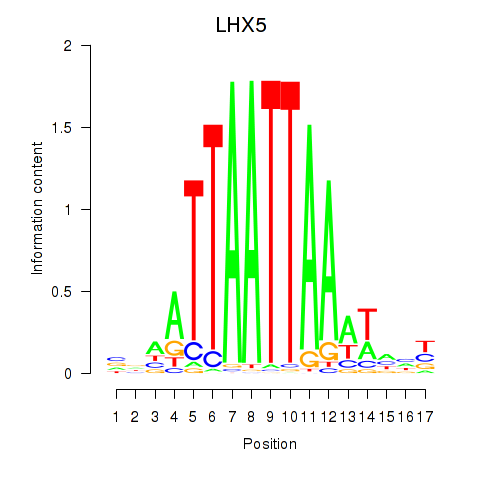
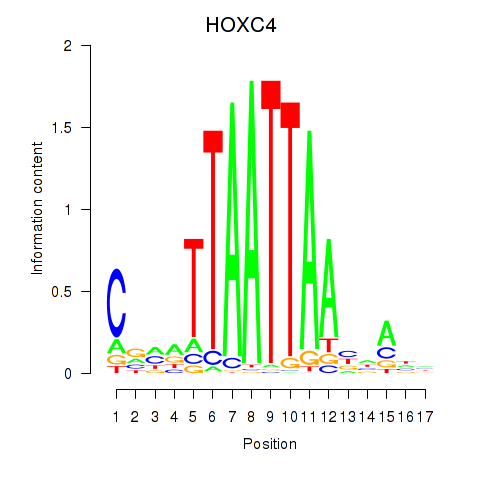
Results for OTP_PHOX2B_LHX1_LMX1A_LHX5_HOXC4
Z-value: 0.84
Transcription factors associated with OTP_PHOX2B_LHX1_LMX1A_LHX5_HOXC4
| Gene Symbol | Gene ID | Gene Info |
|---|---|---|
|
OTP
|
ENSG00000171540.6 | OTP |
|
PHOX2B
|
ENSG00000109132.5 | PHOX2B |
|
LHX1
|
ENSG00000132130.7 | LHX1 |
|
LMX1A
|
ENSG00000162761.10 | LMX1A |
|
LHX5
|
ENSG00000089116.3 | LHX5 |
|
HOXC4
|
ENSG00000198353.6 | HOXC4 |
|
HOXC4
|
ENSG00000273266.1 | HOXC4 |
Activity-expression correlation:
| Gene | Promoter | Pearson corr. coef. | P-value | Plot |
|---|---|---|---|---|
| OTP | hg19_v2_chr5_-_76935513_76935513 | -0.82 | 1.3e-02 | Click! |
| LMX1A | hg19_v2_chr1_-_165324983_165325012 | -0.78 | 2.2e-02 | Click! |
| LHX1 | hg19_v2_chr17_+_35294075_35294102 | -0.76 | 3.0e-02 | Click! |
| HOXC4 | hg19_v2_chr12_+_54447637_54447661 | -0.38 | 3.6e-01 | Click! |
| LHX5 | hg19_v2_chr12_-_113909877_113909877 | -0.29 | 4.9e-01 | Click! |
| PHOX2B | hg19_v2_chr4_-_41750922_41750987 | -0.20 | 6.4e-01 | Click! |
Activity profile of OTP_PHOX2B_LHX1_LMX1A_LHX5_HOXC4 motif
Sorted Z-values of OTP_PHOX2B_LHX1_LMX1A_LHX5_HOXC4 motif
Network of associatons between targets according to the STRING database.
First level regulatory network of OTP_PHOX2B_LHX1_LMX1A_LHX5_HOXC4
| Promoter | Score | Transcript | Gene | Gene Info |
|---|---|---|---|---|
| chr19_+_50016610 | 1.76 |
ENST00000596975.1 |
FCGRT |
Fc fragment of IgG, receptor, transporter, alpha |
| chr5_+_95066823 | 1.75 |
ENST00000506817.1 ENST00000379982.3 |
RHOBTB3 |
Rho-related BTB domain containing 3 |
| chr5_-_20575959 | 1.59 |
ENST00000507958.1 |
CDH18 |
cadherin 18, type 2 |
| chr19_+_50016411 | 1.49 |
ENST00000426395.3 ENST00000600273.1 ENST00000599988.1 |
FCGRT |
Fc fragment of IgG, receptor, transporter, alpha |
| chr17_+_1674982 | 1.40 |
ENST00000572048.1 ENST00000573763.1 |
SERPINF1 |
serpin peptidase inhibitor, clade F (alpha-2 antiplasmin, pigment epithelium derived factor), member 1 |
| chr3_+_157154578 | 1.26 |
ENST00000295927.3 |
PTX3 |
pentraxin 3, long |
| chr11_-_63376013 | 1.21 |
ENST00000540943.1 |
PLA2G16 |
phospholipase A2, group XVI |
| chr2_+_152214098 | 1.14 |
ENST00000243347.3 |
TNFAIP6 |
tumor necrosis factor, alpha-induced protein 6 |
| chr6_+_114178512 | 0.89 |
ENST00000368635.4 |
MARCKS |
myristoylated alanine-rich protein kinase C substrate |
| chr20_-_33735070 | 0.89 |
ENST00000374491.3 ENST00000542871.1 ENST00000374492.3 |
EDEM2 |
ER degradation enhancer, mannosidase alpha-like 2 |
| chr3_+_45927994 | 0.85 |
ENST00000357632.2 ENST00000395963.2 |
CCR9 |
chemokine (C-C motif) receptor 9 |
| chr9_-_99540328 | 0.81 |
ENST00000223428.4 ENST00000375231.1 ENST00000374641.3 |
ZNF510 |
zinc finger protein 510 |
| chr3_-_191000172 | 0.76 |
ENST00000427544.2 |
UTS2B |
urotensin 2B |
| chr1_-_92371839 | 0.66 |
ENST00000370399.2 |
TGFBR3 |
transforming growth factor, beta receptor III |
| chr7_+_134528635 | 0.63 |
ENST00000445569.2 |
CALD1 |
caldesmon 1 |
| chr19_-_14064114 | 0.56 |
ENST00000585607.1 ENST00000538517.2 ENST00000587458.1 ENST00000538371.2 |
PODNL1 |
podocan-like 1 |
| chr3_-_164796269 | 0.55 |
ENST00000264382.3 |
SI |
sucrase-isomaltase (alpha-glucosidase) |
| chr10_+_94451574 | 0.53 |
ENST00000492654.2 |
HHEX |
hematopoietically expressed homeobox |
| chr11_-_121986923 | 0.52 |
ENST00000560104.1 |
BLID |
BH3-like motif containing, cell death inducer |
| chr10_+_90660832 | 0.51 |
ENST00000371924.1 |
STAMBPL1 |
STAM binding protein-like 1 |
| chr11_+_5710919 | 0.51 |
ENST00000379965.3 ENST00000425490.1 |
TRIM22 |
tripartite motif containing 22 |
| chr3_+_12329397 | 0.50 |
ENST00000397015.2 |
PPARG |
peroxisome proliferator-activated receptor gamma |
| chr1_+_78769549 | 0.48 |
ENST00000370758.1 |
PTGFR |
prostaglandin F receptor (FP) |
| chr16_-_66583701 | 0.48 |
ENST00000527800.1 ENST00000525974.1 ENST00000563369.2 |
TK2 |
thymidine kinase 2, mitochondrial |
| chrX_+_22050546 | 0.48 |
ENST00000379374.4 |
PHEX |
phosphate regulating endopeptidase homolog, X-linked |
| chr20_-_18477862 | 0.46 |
ENST00000337227.4 |
RBBP9 |
retinoblastoma binding protein 9 |
| chr1_-_57045228 | 0.45 |
ENST00000371250.3 |
PPAP2B |
phosphatidic acid phosphatase type 2B |
| chr2_-_40680578 | 0.42 |
ENST00000455476.1 |
SLC8A1 |
solute carrier family 8 (sodium/calcium exchanger), member 1 |
| chr20_+_15177480 | 0.40 |
ENST00000402914.1 |
MACROD2 |
MACRO domain containing 2 |
| chr10_-_28571015 | 0.39 |
ENST00000375719.3 ENST00000375732.1 |
MPP7 |
membrane protein, palmitoylated 7 (MAGUK p55 subfamily member 7) |
| chr12_+_8309630 | 0.38 |
ENST00000396570.3 |
ZNF705A |
zinc finger protein 705A |
| chrX_+_43515467 | 0.37 |
ENST00000338702.3 ENST00000542639.1 |
MAOA |
monoamine oxidase A |
| chr5_+_140762268 | 0.37 |
ENST00000518325.1 |
PCDHGA7 |
protocadherin gamma subfamily A, 7 |
| chr13_-_38172863 | 0.36 |
ENST00000541481.1 ENST00000379743.4 ENST00000379742.4 ENST00000379749.4 ENST00000541179.1 ENST00000379747.4 |
POSTN |
periostin, osteoblast specific factor |
| chr12_+_59989918 | 0.35 |
ENST00000547379.1 ENST00000549465.1 |
SLC16A7 |
solute carrier family 16 (monocarboxylate transporter), member 7 |
| chr7_+_138145076 | 0.35 |
ENST00000343526.4 |
TRIM24 |
tripartite motif containing 24 |
| chr17_-_53800217 | 0.35 |
ENST00000424486.2 |
TMEM100 |
transmembrane protein 100 |
| chr4_+_169418195 | 0.34 |
ENST00000261509.6 ENST00000335742.7 |
PALLD |
palladin, cytoskeletal associated protein |
| chr14_-_51027838 | 0.33 |
ENST00000555216.1 |
MAP4K5 |
mitogen-activated protein kinase kinase kinase kinase 5 |
| chr5_+_81575281 | 0.32 |
ENST00000380167.4 |
ATP6AP1L |
ATPase, H+ transporting, lysosomal accessory protein 1-like |
| chr16_-_66584059 | 0.32 |
ENST00000417693.3 ENST00000544898.1 ENST00000569718.1 ENST00000527284.1 ENST00000299697.7 ENST00000451102.2 |
TK2 |
thymidine kinase 2, mitochondrial |
| chr15_-_20193370 | 0.31 |
ENST00000558565.2 |
IGHV3OR15-7 |
immunoglobulin heavy variable 3/OR15-7 (pseudogene) |
| chr16_-_28937027 | 0.31 |
ENST00000358201.4 |
RABEP2 |
rabaptin, RAB GTPase binding effector protein 2 |
| chr3_+_12329358 | 0.30 |
ENST00000309576.6 |
PPARG |
peroxisome proliferator-activated receptor gamma |
| chr19_-_44388116 | 0.30 |
ENST00000587539.1 |
ZNF404 |
zinc finger protein 404 |
| chr3_-_9994021 | 0.26 |
ENST00000411976.2 ENST00000412055.1 |
PRRT3 |
proline-rich transmembrane protein 3 |
| chr8_-_42623747 | 0.26 |
ENST00000534622.1 |
CHRNA6 |
cholinergic receptor, nicotinic, alpha 6 (neuronal) |
| chr11_-_327537 | 0.25 |
ENST00000602735.1 |
IFITM3 |
interferon induced transmembrane protein 3 |
| chr9_+_90112767 | 0.25 |
ENST00000408954.3 |
DAPK1 |
death-associated protein kinase 1 |
| chr13_+_24144796 | 0.24 |
ENST00000403372.2 |
TNFRSF19 |
tumor necrosis factor receptor superfamily, member 19 |
| chr9_+_90112590 | 0.24 |
ENST00000472284.1 |
DAPK1 |
death-associated protein kinase 1 |
| chr9_+_125133315 | 0.24 |
ENST00000223423.4 ENST00000362012.2 |
PTGS1 |
prostaglandin-endoperoxide synthase 1 (prostaglandin G/H synthase and cyclooxygenase) |
| chr7_+_141811539 | 0.24 |
ENST00000550469.2 ENST00000477922.3 |
RP11-1220K2.2 |
Putative inactive maltase-glucoamylase-like protein LOC93432 |
| chrX_+_51486481 | 0.23 |
ENST00000340438.4 |
GSPT2 |
G1 to S phase transition 2 |
| chr11_+_114168085 | 0.23 |
ENST00000541754.1 |
NNMT |
nicotinamide N-methyltransferase |
| chr7_-_144435985 | 0.23 |
ENST00000549981.1 |
TPK1 |
thiamin pyrophosphokinase 1 |
| chr10_-_27529486 | 0.22 |
ENST00000375888.1 |
ACBD5 |
acyl-CoA binding domain containing 5 |
| chr8_+_104384616 | 0.22 |
ENST00000520337.1 |
CTHRC1 |
collagen triple helix repeat containing 1 |
| chr4_+_113568207 | 0.21 |
ENST00000511529.1 |
LARP7 |
La ribonucleoprotein domain family, member 7 |
| chr15_+_93443419 | 0.20 |
ENST00000557381.1 ENST00000420239.2 |
CHD2 |
chromodomain helicase DNA binding protein 2 |
| chr3_+_20081515 | 0.20 |
ENST00000263754.4 |
KAT2B |
K(lysine) acetyltransferase 2B |
| chr3_-_157221380 | 0.20 |
ENST00000468233.1 |
VEPH1 |
ventricular zone expressed PH domain-containing 1 |
| chr12_-_54653313 | 0.20 |
ENST00000550411.1 ENST00000439541.2 |
CBX5 |
chromobox homolog 5 |
| chr4_+_169418255 | 0.19 |
ENST00000505667.1 ENST00000511948.1 |
PALLD |
palladin, cytoskeletal associated protein |
| chr1_+_201617450 | 0.19 |
ENST00000295624.6 ENST00000367297.4 ENST00000367300.3 |
NAV1 |
neuron navigator 1 |
| chr21_+_33671160 | 0.18 |
ENST00000303645.5 |
MRAP |
melanocortin 2 receptor accessory protein |
| chr1_+_78383813 | 0.18 |
ENST00000342754.5 |
NEXN |
nexilin (F actin binding protein) |
| chr4_+_71108300 | 0.18 |
ENST00000304954.3 |
CSN3 |
casein kappa |
| chr1_+_225600404 | 0.17 |
ENST00000366845.2 |
AC092811.1 |
AC092811.1 |
| chr1_+_201617264 | 0.17 |
ENST00000367296.4 |
NAV1 |
neuron navigator 1 |
| chr12_+_131438443 | 0.17 |
ENST00000261654.5 |
GPR133 |
G protein-coupled receptor 133 |
| chr3_+_138340049 | 0.16 |
ENST00000464668.1 |
FAIM |
Fas apoptotic inhibitory molecule |
| chr5_-_88119580 | 0.16 |
ENST00000539796.1 |
MEF2C |
myocyte enhancer factor 2C |
| chr2_-_74618964 | 0.16 |
ENST00000417090.1 ENST00000409868.1 |
DCTN1 |
dynactin 1 |
| chr1_+_117544366 | 0.16 |
ENST00000256652.4 ENST00000369470.1 |
CD101 |
CD101 molecule |
| chr7_-_22862406 | 0.16 |
ENST00000372879.4 |
TOMM7 |
translocase of outer mitochondrial membrane 7 homolog (yeast) |
| chr7_-_140482926 | 0.15 |
ENST00000496384.2 |
BRAF |
v-raf murine sarcoma viral oncogene homolog B |
| chr6_+_26402517 | 0.15 |
ENST00000414912.2 |
BTN3A1 |
butyrophilin, subfamily 3, member A1 |
| chrX_-_100872911 | 0.15 |
ENST00000361910.4 ENST00000539247.1 ENST00000538627.1 |
ARMCX6 |
armadillo repeat containing, X-linked 6 |
| chr3_+_69812701 | 0.15 |
ENST00000472437.1 |
MITF |
microphthalmia-associated transcription factor |
| chr12_+_64798095 | 0.15 |
ENST00000332707.5 |
XPOT |
exportin, tRNA |
| chr15_-_37393406 | 0.15 |
ENST00000338564.5 ENST00000558313.1 ENST00000340545.5 |
MEIS2 |
Meis homeobox 2 |
| chr3_+_69812877 | 0.15 |
ENST00000457080.1 ENST00000328528.6 |
MITF |
microphthalmia-associated transcription factor |
| chr2_+_109237717 | 0.14 |
ENST00000409441.1 |
LIMS1 |
LIM and senescent cell antigen-like domains 1 |
| chr3_-_157221357 | 0.14 |
ENST00000494677.1 |
VEPH1 |
ventricular zone expressed PH domain-containing 1 |
| chr1_-_153518270 | 0.14 |
ENST00000354332.4 ENST00000368716.4 |
S100A4 |
S100 calcium binding protein A4 |
| chr9_-_215744 | 0.13 |
ENST00000382387.2 |
C9orf66 |
chromosome 9 open reading frame 66 |
| chr3_+_138340067 | 0.13 |
ENST00000479848.1 |
FAIM |
Fas apoptotic inhibitory molecule |
| chr10_-_4285923 | 0.13 |
ENST00000418372.1 ENST00000608792.1 |
LINC00702 |
long intergenic non-protein coding RNA 702 |
| chr1_+_212965170 | 0.13 |
ENST00000532324.1 ENST00000366974.4 ENST00000530441.1 ENST00000526641.1 ENST00000531963.1 ENST00000366973.4 ENST00000526997.1 ENST00000488246.2 |
TATDN3 |
TatD DNase domain containing 3 |
| chr7_+_129932974 | 0.13 |
ENST00000445470.2 ENST00000222482.4 ENST00000492072.1 ENST00000473956.1 ENST00000493259.1 ENST00000486598.1 |
CPA4 |
carboxypeptidase A4 |
| chr20_-_56265680 | 0.13 |
ENST00000414037.1 |
PMEPA1 |
prostate transmembrane protein, androgen induced 1 |
| chr19_-_51920952 | 0.12 |
ENST00000356298.5 ENST00000339313.5 ENST00000529627.1 ENST00000439889.2 ENST00000353836.5 ENST00000432469.2 |
SIGLEC10 |
sialic acid binding Ig-like lectin 10 |
| chr18_-_32870148 | 0.12 |
ENST00000589178.1 ENST00000333206.5 ENST00000592278.1 ENST00000592211.1 ENST00000420878.3 ENST00000383091.2 ENST00000586922.2 |
ZSCAN30 RP11-158H5.7 |
zinc finger and SCAN domain containing 30 RP11-158H5.7 |
| chr14_+_55493920 | 0.12 |
ENST00000395472.2 ENST00000555846.1 |
SOCS4 |
suppressor of cytokine signaling 4 |
| chrX_+_36254051 | 0.12 |
ENST00000378657.4 |
CXorf30 |
chromosome X open reading frame 30 |
| chr7_+_138915102 | 0.12 |
ENST00000486663.1 |
UBN2 |
ubinuclein 2 |
| chr1_-_212965104 | 0.12 |
ENST00000422588.2 ENST00000366975.6 ENST00000366977.3 ENST00000366976.1 |
NSL1 |
NSL1, MIS12 kinetochore complex component |
| chr2_-_37068530 | 0.11 |
ENST00000593798.1 |
AC007382.1 |
Uncharacterized protein |
| chr3_-_114477962 | 0.11 |
ENST00000471418.1 |
ZBTB20 |
zinc finger and BTB domain containing 20 |
| chr3_-_64211112 | 0.11 |
ENST00000295902.6 |
PRICKLE2 |
prickle homolog 2 (Drosophila) |
| chr3_-_114477787 | 0.11 |
ENST00000464560.1 |
ZBTB20 |
zinc finger and BTB domain containing 20 |
| chr2_-_231860596 | 0.11 |
ENST00000441063.1 ENST00000434094.1 ENST00000418330.1 ENST00000457803.1 ENST00000414876.1 ENST00000446741.1 ENST00000426904.1 |
AC105344.2 |
SPATA3 antisense RNA 1 (head to head) |
| chr12_+_26348582 | 0.11 |
ENST00000535504.1 |
SSPN |
sarcospan |
| chrX_+_55744166 | 0.11 |
ENST00000374941.4 ENST00000414239.1 |
RRAGB |
Ras-related GTP binding B |
| chr20_+_22034809 | 0.10 |
ENST00000449427.1 |
RP11-125P18.1 |
RP11-125P18.1 |
| chr11_-_102576537 | 0.10 |
ENST00000260229.4 |
MMP27 |
matrix metallopeptidase 27 |
| chr15_+_64680003 | 0.10 |
ENST00000261884.3 |
TRIP4 |
thyroid hormone receptor interactor 4 |
| chr20_+_31870927 | 0.10 |
ENST00000253354.1 |
BPIFB1 |
BPI fold containing family B, member 1 |
| chr13_+_46276441 | 0.10 |
ENST00000310521.1 ENST00000533564.1 |
SPERT |
spermatid associated |
| chr14_+_55494323 | 0.10 |
ENST00000339298.2 |
SOCS4 |
suppressor of cytokine signaling 4 |
| chr17_-_8198636 | 0.10 |
ENST00000577745.1 ENST00000579192.1 ENST00000396278.1 |
SLC25A35 |
solute carrier family 25, member 35 |
| chr20_-_29978286 | 0.10 |
ENST00000376315.2 |
DEFB119 |
defensin, beta 119 |
| chr6_+_42749759 | 0.10 |
ENST00000314073.5 |
GLTSCR1L |
GLTSCR1-like |
| chr3_-_149293990 | 0.10 |
ENST00000472417.1 |
WWTR1 |
WW domain containing transcription regulator 1 |
| chr1_-_190446759 | 0.10 |
ENST00000367462.3 |
BRINP3 |
bone morphogenetic protein/retinoic acid inducible neural-specific 3 |
| chr12_-_46121554 | 0.10 |
ENST00000609803.1 |
LINC00938 |
long intergenic non-protein coding RNA 938 |
| chr7_-_22862448 | 0.10 |
ENST00000358435.4 |
TOMM7 |
translocase of outer mitochondrial membrane 7 homolog (yeast) |
| chr13_+_103459704 | 0.10 |
ENST00000602836.1 |
BIVM-ERCC5 |
BIVM-ERCC5 readthrough |
| chr17_+_74463650 | 0.10 |
ENST00000392492.3 |
AANAT |
aralkylamine N-acetyltransferase |
| chrX_+_55744228 | 0.10 |
ENST00000262850.7 |
RRAGB |
Ras-related GTP binding B |
| chr10_-_50396407 | 0.10 |
ENST00000374153.2 ENST00000374151.3 |
C10orf128 |
chromosome 10 open reading frame 128 |
| chr3_+_186353756 | 0.09 |
ENST00000431018.1 ENST00000450521.1 ENST00000539949.1 |
FETUB |
fetuin B |
| chr2_-_74619152 | 0.09 |
ENST00000440727.1 ENST00000409240.1 |
DCTN1 |
dynactin 1 |
| chr6_+_26440700 | 0.09 |
ENST00000494393.1 ENST00000482451.1 ENST00000244519.2 ENST00000339789.4 ENST00000471353.1 ENST00000361232.3 ENST00000487627.1 ENST00000496719.1 ENST00000490254.1 ENST00000487272.1 |
BTN3A3 |
butyrophilin, subfamily 3, member A3 |
| chr12_+_8666126 | 0.09 |
ENST00000299665.2 |
CLEC4D |
C-type lectin domain family 4, member D |
| chrX_+_77166172 | 0.09 |
ENST00000343533.5 ENST00000350425.4 ENST00000341514.6 |
ATP7A |
ATPase, Cu++ transporting, alpha polypeptide |
| chr2_-_191115229 | 0.09 |
ENST00000409820.2 ENST00000410045.1 |
HIBCH |
3-hydroxyisobutyryl-CoA hydrolase |
| chr7_-_102283238 | 0.09 |
ENST00000340457.8 |
UPK3BL |
uroplakin 3B-like |
| chr3_-_141747950 | 0.09 |
ENST00000497579.1 |
TFDP2 |
transcription factor Dp-2 (E2F dimerization partner 2) |
| chr5_-_125930929 | 0.08 |
ENST00000553117.1 ENST00000447989.2 ENST00000409134.3 |
ALDH7A1 |
aldehyde dehydrogenase 7 family, member A1 |
| chr14_+_50234309 | 0.08 |
ENST00000298307.5 |
KLHDC2 |
kelch domain containing 2 |
| chrX_-_106243451 | 0.08 |
ENST00000355610.4 ENST00000535534.1 |
MORC4 |
MORC family CW-type zinc finger 4 |
| chr7_-_143599207 | 0.08 |
ENST00000355951.2 ENST00000479870.1 ENST00000478172.1 |
FAM115A |
family with sequence similarity 115, member A |
| chr12_+_8662057 | 0.08 |
ENST00000382064.2 |
CLEC4D |
C-type lectin domain family 4, member D |
| chr17_+_18086392 | 0.08 |
ENST00000541285.1 |
ALKBH5 |
alkB, alkylation repair homolog 5 (E. coli) |
| chr10_+_18549645 | 0.08 |
ENST00000396576.2 |
CACNB2 |
calcium channel, voltage-dependent, beta 2 subunit |
| chr6_+_47749718 | 0.08 |
ENST00000489301.2 ENST00000371211.2 ENST00000393699.2 |
OPN5 |
opsin 5 |
| chr2_-_241497374 | 0.08 |
ENST00000373318.2 ENST00000406958.1 ENST00000391987.1 ENST00000373320.4 |
ANKMY1 |
ankyrin repeat and MYND domain containing 1 |
| chr7_+_133812052 | 0.08 |
ENST00000285928.2 |
LRGUK |
leucine-rich repeats and guanylate kinase domain containing |
| chr4_-_138453559 | 0.08 |
ENST00000511115.1 |
PCDH18 |
protocadherin 18 |
| chr13_-_61989655 | 0.07 |
ENST00000409204.4 |
PCDH20 |
protocadherin 20 |
| chr13_+_53602894 | 0.07 |
ENST00000219022.2 |
OLFM4 |
olfactomedin 4 |
| chr15_-_98417780 | 0.07 |
ENST00000503874.3 |
LINC00923 |
long intergenic non-protein coding RNA 923 |
| chr3_-_99569821 | 0.07 |
ENST00000487087.1 |
FILIP1L |
filamin A interacting protein 1-like |
| chr3_-_157221128 | 0.07 |
ENST00000392833.2 ENST00000362010.2 |
VEPH1 |
ventricular zone expressed PH domain-containing 1 |
| chr4_+_71091786 | 0.07 |
ENST00000317987.5 |
FDCSP |
follicular dendritic cell secreted protein |
| chr12_+_26348246 | 0.07 |
ENST00000422622.2 |
SSPN |
sarcospan |
| chr14_+_22465771 | 0.07 |
ENST00000390445.2 |
TRAV17 |
T cell receptor alpha variable 17 |
| chr5_+_140593509 | 0.07 |
ENST00000341948.4 |
PCDHB13 |
protocadherin beta 13 |
| chr2_+_166095898 | 0.07 |
ENST00000424833.1 ENST00000375437.2 ENST00000357398.3 |
SCN2A |
sodium channel, voltage-gated, type II, alpha subunit |
| chrX_+_108779004 | 0.07 |
ENST00000218004.1 |
NXT2 |
nuclear transport factor 2-like export factor 2 |
| chr17_+_66511540 | 0.07 |
ENST00000588188.2 |
PRKAR1A |
protein kinase, cAMP-dependent, regulatory, type I, alpha |
| chr12_+_26164645 | 0.07 |
ENST00000542004.1 |
RASSF8 |
Ras association (RalGDS/AF-6) domain family (N-terminal) member 8 |
| chr13_+_36050881 | 0.07 |
ENST00000537702.1 |
NBEA |
neurobeachin |
| chr9_-_116065551 | 0.07 |
ENST00000297894.5 |
RNF183 |
ring finger protein 183 |
| chr1_+_205682497 | 0.06 |
ENST00000598338.1 |
AC119673.1 |
AC119673.1 |
| chr4_-_119759795 | 0.06 |
ENST00000419654.2 |
SEC24D |
SEC24 family member D |
| chr12_+_4130143 | 0.06 |
ENST00000543206.1 |
RP11-320N7.2 |
RP11-320N7.2 |
| chr8_-_116504448 | 0.06 |
ENST00000518018.1 |
TRPS1 |
trichorhinophalangeal syndrome I |
| chr16_-_71842941 | 0.06 |
ENST00000423132.2 ENST00000433195.2 ENST00000569748.1 ENST00000570017.1 |
AP1G1 |
adaptor-related protein complex 1, gamma 1 subunit |
| chr15_+_94899183 | 0.06 |
ENST00000557742.1 |
MCTP2 |
multiple C2 domains, transmembrane 2 |
| chr3_-_122283424 | 0.06 |
ENST00000477522.2 ENST00000360356.2 |
PARP9 |
poly (ADP-ribose) polymerase family, member 9 |
| chr9_+_34652164 | 0.06 |
ENST00000441545.2 ENST00000553620.1 |
IL11RA |
interleukin 11 receptor, alpha |
| chr4_+_186990298 | 0.06 |
ENST00000296795.3 ENST00000513189.1 |
TLR3 |
toll-like receptor 3 |
| chr5_-_130500922 | 0.06 |
ENST00000513012.1 ENST00000508488.1 ENST00000506908.1 ENST00000304043.5 |
HINT1 |
histidine triad nucleotide binding protein 1 |
| chr8_-_90996459 | 0.06 |
ENST00000517337.1 ENST00000409330.1 |
NBN |
nibrin |
| chr5_-_126409159 | 0.06 |
ENST00000607731.1 ENST00000535381.1 ENST00000296662.5 ENST00000509733.3 |
C5orf63 |
chromosome 5 open reading frame 63 |
| chr6_-_41039567 | 0.06 |
ENST00000468811.1 |
OARD1 |
O-acyl-ADP-ribose deacylase 1 |
| chr1_-_231005310 | 0.06 |
ENST00000470540.1 |
C1orf198 |
chromosome 1 open reading frame 198 |
| chr4_+_158493642 | 0.05 |
ENST00000507108.1 ENST00000455598.1 ENST00000509450.1 |
RP11-364P22.1 |
RP11-364P22.1 |
| chr14_-_39639523 | 0.05 |
ENST00000330149.5 ENST00000554018.1 ENST00000347691.5 |
TRAPPC6B |
trafficking protein particle complex 6B |
| chr2_-_21022818 | 0.05 |
ENST00000440866.2 ENST00000541941.1 ENST00000402479.2 ENST00000435420.2 ENST00000432947.1 ENST00000403006.2 ENST00000419825.2 ENST00000381090.3 ENST00000237822.3 ENST00000412261.1 |
C2orf43 |
chromosome 2 open reading frame 43 |
| chr9_-_26947453 | 0.05 |
ENST00000397292.3 |
PLAA |
phospholipase A2-activating protein |
| chr1_+_168250194 | 0.05 |
ENST00000367821.3 |
TBX19 |
T-box 19 |
| chr7_+_13141097 | 0.05 |
ENST00000411542.1 |
AC011288.2 |
AC011288.2 |
| chr3_-_157217328 | 0.05 |
ENST00000392832.2 ENST00000543418.1 |
VEPH1 |
ventricular zone expressed PH domain-containing 1 |
| chr7_+_150020329 | 0.05 |
ENST00000323078.7 |
LRRC61 |
leucine rich repeat containing 61 |
| chrX_+_79675965 | 0.05 |
ENST00000308293.5 |
FAM46D |
family with sequence similarity 46, member D |
| chr8_-_57123815 | 0.05 |
ENST00000316981.3 ENST00000423799.2 ENST00000429357.2 |
PLAG1 |
pleiomorphic adenoma gene 1 |
| chr7_+_150020363 | 0.05 |
ENST00000359623.4 ENST00000493307.1 |
LRRC61 |
leucine rich repeat containing 61 |
| chr4_-_152149033 | 0.05 |
ENST00000514152.1 |
SH3D19 |
SH3 domain containing 19 |
| chr3_+_141457030 | 0.05 |
ENST00000273480.3 |
RNF7 |
ring finger protein 7 |
| chr19_-_37663332 | 0.04 |
ENST00000392157.2 |
ZNF585A |
zinc finger protein 585A |
| chr1_-_101360331 | 0.04 |
ENST00000416479.1 ENST00000370113.3 |
EXTL2 |
exostosin-like glycosyltransferase 2 |
| chr8_-_7673238 | 0.04 |
ENST00000335021.2 |
DEFB107A |
defensin, beta 107A |
| chr7_-_107642348 | 0.04 |
ENST00000393561.1 |
LAMB1 |
laminin, beta 1 |
| chr15_-_75748115 | 0.04 |
ENST00000360439.4 |
SIN3A |
SIN3 transcription regulator family member A |
| chr3_-_186262166 | 0.04 |
ENST00000307944.5 |
CRYGS |
crystallin, gamma S |
| chr6_+_29079668 | 0.04 |
ENST00000377169.1 |
OR2J3 |
olfactory receptor, family 2, subfamily J, member 3 |
| chr13_+_77522632 | 0.04 |
ENST00000377462.1 |
IRG1 |
immunoresponsive 1 homolog (mouse) |
| chr3_+_140396881 | 0.04 |
ENST00000286349.3 |
TRIM42 |
tripartite motif containing 42 |
| chr20_-_29978383 | 0.04 |
ENST00000339144.3 ENST00000376321.3 |
DEFB119 |
defensin, beta 119 |
| chr14_-_45252031 | 0.04 |
ENST00000556405.1 |
RP11-398E10.1 |
RP11-398E10.1 |
| chr3_+_141106643 | 0.04 |
ENST00000514251.1 |
ZBTB38 |
zinc finger and BTB domain containing 38 |
| chr8_+_121137333 | 0.04 |
ENST00000309791.4 ENST00000297848.3 ENST00000247781.3 |
COL14A1 |
collagen, type XIV, alpha 1 |
| chr9_-_21202204 | 0.04 |
ENST00000239347.3 |
IFNA7 |
interferon, alpha 7 |
| chr19_-_14945933 | 0.04 |
ENST00000322301.3 |
OR7A5 |
olfactory receptor, family 7, subfamily A, member 5 |
| chr12_+_26348429 | 0.04 |
ENST00000242729.2 |
SSPN |
sarcospan |
| chr9_+_111624577 | 0.04 |
ENST00000333999.3 |
ACTL7A |
actin-like 7A |
Gene Ontology Analysis
Gene overrepresentation in biological_process category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.4 | 3.2 | GO:0002414 | immunoglobulin transcytosis in epithelial cells(GO:0002414) |
| 0.3 | 1.5 | GO:0071279 | cellular response to cobalt ion(GO:0071279) |
| 0.3 | 0.9 | GO:0036512 | trimming of terminal mannose on B branch(GO:0036509) trimming of first mannose on A branch(GO:0036511) trimming of second mannose on A branch(GO:0036512) |
| 0.2 | 1.3 | GO:0052199 | negative regulation of catalytic activity in other organism involved in symbiotic interaction(GO:0052199) |
| 0.2 | 0.8 | GO:2000230 | negative regulation of pancreatic stellate cell proliferation(GO:2000230) |
| 0.2 | 0.8 | GO:0046104 | thymidine metabolic process(GO:0046104) pyrimidine deoxyribonucleoside metabolic process(GO:0046125) |
| 0.2 | 0.5 | GO:0010621 | negative regulation of transcription by transcription factor localization(GO:0010621) gall bladder development(GO:0061010) |
| 0.2 | 0.7 | GO:0034699 | transforming growth factor beta receptor complex assembly(GO:0007181) response to luteinizing hormone(GO:0034699) |
| 0.1 | 1.2 | GO:0046485 | ether lipid metabolic process(GO:0046485) |
| 0.1 | 0.5 | GO:1904383 | response to sodium phosphate(GO:1904383) |
| 0.1 | 0.6 | GO:0044245 | polysaccharide digestion(GO:0044245) |
| 0.1 | 0.2 | GO:0002184 | cytoplasmic translational termination(GO:0002184) |
| 0.1 | 0.8 | GO:0042492 | gamma-delta T cell differentiation(GO:0042492) |
| 0.1 | 0.4 | GO:0032571 | response to vitamin K(GO:0032571) bone regeneration(GO:1990523) |
| 0.1 | 0.2 | GO:0042357 | thiamine diphosphate metabolic process(GO:0042357) |
| 0.1 | 0.4 | GO:0051725 | protein de-ADP-ribosylation(GO:0051725) |
| 0.1 | 0.5 | GO:0071799 | response to prostaglandin D(GO:0071798) cellular response to prostaglandin D stimulus(GO:0071799) |
| 0.0 | 0.2 | GO:0018076 | N-terminal peptidyl-lysine acetylation(GO:0018076) |
| 0.0 | 0.3 | GO:0070562 | regulation of vitamin D receptor signaling pathway(GO:0070562) |
| 0.0 | 0.4 | GO:0071896 | protein localization to adherens junction(GO:0071896) |
| 0.0 | 0.4 | GO:0006848 | pyruvate transport(GO:0006848) pyruvate transmembrane transport(GO:1901475) |
| 0.0 | 0.3 | GO:0060842 | arterial endothelial cell differentiation(GO:0060842) |
| 0.0 | 0.4 | GO:0098735 | positive regulation of the force of heart contraction(GO:0098735) |
| 0.0 | 0.2 | GO:0003172 | primary heart field specification(GO:0003138) sinoatrial valve development(GO:0003172) sinoatrial valve morphogenesis(GO:0003185) |
| 0.0 | 0.9 | GO:0051764 | actin crosslink formation(GO:0051764) |
| 0.0 | 0.1 | GO:0030186 | melatonin metabolic process(GO:0030186) melatonin biosynthetic process(GO:0030187) |
| 0.0 | 0.2 | GO:0030242 | pexophagy(GO:0030242) |
| 0.0 | 0.3 | GO:0090063 | positive regulation of microtubule nucleation(GO:0090063) |
| 0.0 | 1.0 | GO:0030728 | ovulation(GO:0030728) |
| 0.0 | 0.2 | GO:0043932 | ossification involved in bone remodeling(GO:0043932) |
| 0.0 | 0.2 | GO:2000301 | negative regulation of synaptic vesicle exocytosis(GO:2000301) |
| 0.0 | 0.1 | GO:0019285 | glycine betaine biosynthetic process from choline(GO:0019285) glycine betaine metabolic process(GO:0031455) glycine betaine biosynthetic process(GO:0031456) |
| 0.0 | 0.3 | GO:0045040 | protein import into mitochondrial outer membrane(GO:0045040) |
| 0.0 | 0.2 | GO:1990253 | cellular response to leucine starvation(GO:1990253) |
| 0.0 | 0.5 | GO:0003334 | keratinocyte development(GO:0003334) |
| 0.0 | 0.4 | GO:0042420 | dopamine catabolic process(GO:0042420) |
| 0.0 | 0.1 | GO:0031860 | telomeric 3' overhang formation(GO:0031860) |
| 0.0 | 0.1 | GO:0035553 | oxidative RNA demethylation(GO:0035513) oxidative single-stranded RNA demethylation(GO:0035553) |
| 0.0 | 0.1 | GO:0043321 | regulation of natural killer cell degranulation(GO:0043321) positive regulation of natural killer cell degranulation(GO:0043323) |
| 0.0 | 0.4 | GO:0071447 | cellular response to hydroperoxide(GO:0071447) |
| 0.0 | 1.7 | GO:0042147 | retrograde transport, endosome to Golgi(GO:0042147) |
| 0.0 | 0.3 | GO:0034356 | NAD biosynthesis via nicotinamide riboside salvage pathway(GO:0034356) |
| 0.0 | 0.1 | GO:0006574 | valine catabolic process(GO:0006574) |
| 0.0 | 0.1 | GO:0034344 | microglial cell activation involved in immune response(GO:0002282) type III interferon production(GO:0034343) regulation of type III interferon production(GO:0034344) positive regulation of interferon-alpha biosynthetic process(GO:0045356) |
| 0.0 | 0.5 | GO:0070536 | protein K63-linked deubiquitination(GO:0070536) |
| 0.0 | 0.2 | GO:0019371 | cyclooxygenase pathway(GO:0019371) |
| 0.0 | 0.1 | GO:0034334 | adherens junction maintenance(GO:0034334) |
| 0.0 | 0.1 | GO:1904879 | positive regulation of calcium ion transmembrane transport via high voltage-gated calcium channel(GO:1904879) |
| 0.0 | 0.2 | GO:0007175 | negative regulation of epidermal growth factor-activated receptor activity(GO:0007175) |
| 0.0 | 0.0 | GO:0021812 | neuronal-glial interaction involved in cerebral cortex radial glia guided migration(GO:0021812) |
| 0.0 | 0.0 | GO:1901675 | negative regulation of histone H3-K27 acetylation(GO:1901675) |
| 0.0 | 0.1 | GO:2000389 | regulation of neutrophil extravasation(GO:2000389) |
| 0.0 | 0.0 | GO:0051097 | negative regulation of helicase activity(GO:0051097) |
Gene overrepresentation in cellular_component category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.3 | 0.9 | GO:0042585 | germinal vesicle(GO:0042585) |
| 0.2 | 0.7 | GO:0034673 | inhibin-betaglycan-ActRII complex(GO:0034673) |
| 0.1 | 1.4 | GO:0043203 | axon hillock(GO:0043203) |
| 0.1 | 0.2 | GO:0018444 | translation release factor complex(GO:0018444) |
| 0.1 | 0.3 | GO:0005726 | perichromatin fibrils(GO:0005726) |
| 0.0 | 0.6 | GO:0030478 | actin cap(GO:0030478) |
| 0.0 | 0.2 | GO:1990131 | Gtr1-Gtr2 GTPase complex(GO:1990131) |
| 0.0 | 0.3 | GO:0005742 | mitochondrial outer membrane translocase complex(GO:0005742) |
| 0.0 | 0.4 | GO:0097025 | MPP7-DLG1-LIN7 complex(GO:0097025) |
| 0.0 | 0.1 | GO:0043224 | nuclear SCF ubiquitin ligase complex(GO:0043224) |
| 0.0 | 2.5 | GO:1904724 | tertiary granule lumen(GO:1904724) |
| 0.0 | 0.1 | GO:0000444 | MIS12/MIND type complex(GO:0000444) |
| 0.0 | 0.9 | GO:0044322 | endoplasmic reticulum quality control compartment(GO:0044322) |
| 0.0 | 0.2 | GO:0000125 | PCAF complex(GO:0000125) |
| 0.0 | 0.3 | GO:0033180 | proton-transporting V-type ATPase, V1 domain(GO:0033180) |
| 0.0 | 0.4 | GO:0043194 | axon initial segment(GO:0043194) |
| 0.0 | 0.3 | GO:0005869 | dynactin complex(GO:0005869) |
| 0.0 | 0.1 | GO:0002177 | manchette(GO:0002177) |
| 0.0 | 0.1 | GO:0042406 | extrinsic component of endoplasmic reticulum membrane(GO:0042406) |
| 0.0 | 0.0 | GO:0005607 | laminin-2 complex(GO:0005607) |
| 0.0 | 1.8 | GO:0032588 | trans-Golgi network membrane(GO:0032588) |
| 0.0 | 1.2 | GO:0031903 | peroxisomal membrane(GO:0005778) microbody membrane(GO:0031903) |
| 0.0 | 0.2 | GO:0031618 | nuclear pericentric heterochromatin(GO:0031618) |
| 0.0 | 0.0 | GO:0030934 | anchoring collagen complex(GO:0030934) |
| 0.0 | 0.0 | GO:0005873 | plus-end kinesin complex(GO:0005873) |
Gene overrepresentation in molecular_function category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.8 | 3.2 | GO:0019770 | IgG receptor activity(GO:0019770) |
| 0.3 | 0.8 | GO:0004137 | deoxycytidine kinase activity(GO:0004137) |
| 0.2 | 1.2 | GO:0052740 | 1-acyl-2-lysophosphatidylserine acylhydrolase activity(GO:0052740) |
| 0.2 | 0.6 | GO:0004574 | oligo-1,6-glucosidase activity(GO:0004574) |
| 0.2 | 0.5 | GO:0004958 | prostaglandin F receptor activity(GO:0004958) |
| 0.1 | 1.3 | GO:0001849 | complement component C1q binding(GO:0001849) |
| 0.1 | 0.8 | GO:0050692 | DBD domain binding(GO:0050692) |
| 0.1 | 0.4 | GO:0005477 | pyruvate secondary active transmembrane transporter activity(GO:0005477) |
| 0.1 | 0.9 | GO:0004571 | mannosyl-oligosaccharide 1,2-alpha-mannosidase activity(GO:0004571) |
| 0.1 | 0.2 | GO:0031783 | corticotropin hormone receptor binding(GO:0031780) type 5 melanocortin receptor binding(GO:0031783) |
| 0.1 | 0.7 | GO:0005114 | type II transforming growth factor beta receptor binding(GO:0005114) |
| 0.1 | 0.4 | GO:0099580 | ion antiporter activity involved in regulation of postsynaptic membrane potential(GO:0099580) |
| 0.1 | 0.2 | GO:0008112 | nicotinamide N-methyltransferase activity(GO:0008112) pyridine N-methyltransferase activity(GO:0030760) |
| 0.1 | 0.2 | GO:0004666 | prostaglandin-endoperoxide synthase activity(GO:0004666) |
| 0.0 | 0.8 | GO:0016493 | C-C chemokine receptor activity(GO:0016493) |
| 0.0 | 0.4 | GO:0008131 | primary amine oxidase activity(GO:0008131) |
| 0.0 | 1.1 | GO:0005540 | hyaluronic acid binding(GO:0005540) |
| 0.0 | 0.5 | GO:0061578 | Lys63-specific deubiquitinase activity(GO:0061578) |
| 0.0 | 0.3 | GO:0034056 | estrogen response element binding(GO:0034056) |
| 0.0 | 0.1 | GO:0032767 | copper-dependent protein binding(GO:0032767) |
| 0.0 | 0.2 | GO:0003747 | translation release factor activity(GO:0003747) translation termination factor activity(GO:0008079) |
| 0.0 | 0.7 | GO:0071837 | HMG box domain binding(GO:0071837) |
| 0.0 | 0.2 | GO:0016778 | diphosphotransferase activity(GO:0016778) |
| 0.0 | 0.2 | GO:0004468 | lysine N-acetyltransferase activity, acting on acetyl phosphate as donor(GO:0004468) |
| 0.0 | 0.1 | GO:0004060 | arylamine N-acetyltransferase activity(GO:0004060) |
| 0.0 | 0.2 | GO:0016812 | hydrolase activity, acting on carbon-nitrogen (but not peptide) bonds, in cyclic amides(GO:0016812) |
| 0.0 | 0.3 | GO:0015266 | protein channel activity(GO:0015266) |
| 0.0 | 0.1 | GO:0035515 | oxidative RNA demethylase activity(GO:0035515) |
| 0.0 | 0.6 | GO:0005523 | tropomyosin binding(GO:0005523) |
| 0.0 | 0.5 | GO:0051371 | muscle alpha-actinin binding(GO:0051371) |
| 0.0 | 0.9 | GO:0005080 | protein kinase C binding(GO:0005080) |
| 0.0 | 0.5 | GO:0017075 | syntaxin-1 binding(GO:0017075) |
| 0.0 | 0.3 | GO:0008349 | MAP kinase kinase kinase kinase activity(GO:0008349) |
| 0.0 | 0.3 | GO:0046933 | proton-transporting ATP synthase activity, rotational mechanism(GO:0046933) |
| 0.0 | 0.1 | GO:0005502 | 11-cis retinal binding(GO:0005502) |
| 0.0 | 0.3 | GO:0022848 | acetylcholine-gated cation channel activity(GO:0022848) |
Gene overrepresentation in C2:CP category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.0 | 1.3 | ST MYOCYTE AD PATHWAY | Myocyte Adrenergic Pathway is a specific case of the generalized Adrenergic Pathway. |
| 0.0 | 1.0 | PID WNT NONCANONICAL PATHWAY | Noncanonical Wnt signaling pathway |
Gene overrepresentation in C2:CP:REACTOME category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.1 | 0.9 | REACTOME REGULATION OF INSULIN SECRETION BY ACETYLCHOLINE | Genes involved in Regulation of Insulin Secretion by Acetylcholine |
| 0.1 | 1.2 | REACTOME ACYL CHAIN REMODELLING OF PS | Genes involved in Acyl chain remodelling of PS |
| 0.1 | 0.6 | REACTOME DIGESTION OF DIETARY CARBOHYDRATE | Genes involved in Digestion of dietary carbohydrate |
| 0.0 | 0.9 | REACTOME CALNEXIN CALRETICULIN CYCLE | Genes involved in Calnexin/calreticulin cycle |
| 0.0 | 1.6 | REACTOME ADHERENS JUNCTIONS INTERACTIONS | Genes involved in Adherens junctions interactions |
| 0.0 | 0.5 | REACTOME PROSTANOID LIGAND RECEPTORS | Genes involved in Prostanoid ligand receptors |
| 0.0 | 0.5 | REACTOME ROLE OF DCC IN REGULATING APOPTOSIS | Genes involved in Role of DCC in regulating apoptosis |
| 0.0 | 0.8 | REACTOME G1 S SPECIFIC TRANSCRIPTION | Genes involved in G1/S-Specific Transcription |
| 0.0 | 0.4 | REACTOME NOREPINEPHRINE NEUROTRANSMITTER RELEASE CYCLE | Genes involved in Norepinephrine Neurotransmitter Release Cycle |
| 0.0 | 0.3 | REACTOME BILE SALT AND ORGANIC ANION SLC TRANSPORTERS | Genes involved in Bile salt and organic anion SLC transporters |
| 0.0 | 0.3 | REACTOME PRESYNAPTIC NICOTINIC ACETYLCHOLINE RECEPTORS | Genes involved in Presynaptic nicotinic acetylcholine receptors |
| 0.0 | 0.2 | REACTOME NOTCH HLH TRANSCRIPTION PATHWAY | Genes involved in Notch-HLH transcription pathway |
| 0.0 | 0.7 | REACTOME CHEMOKINE RECEPTORS BIND CHEMOKINES | Genes involved in Chemokine receptors bind chemokines |
| 0.0 | 0.3 | REACTOME PLATELET CALCIUM HOMEOSTASIS | Genes involved in Platelet calcium homeostasis |