Project
Epithelial-Mesenchymal Transition, human (Scheel, 2011)
Navigation
Downloads
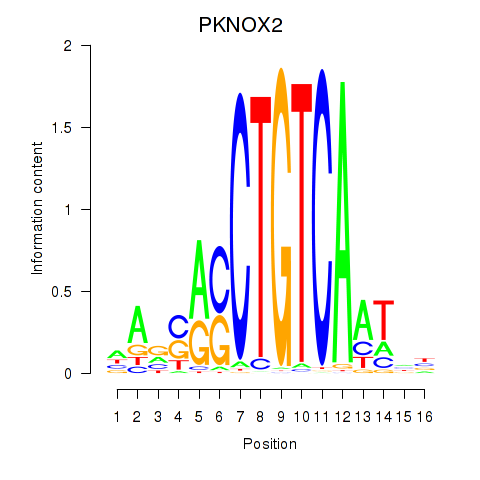
Results for PKNOX2
Z-value: 0.36
Transcription factors associated with PKNOX2
| Gene Symbol | Gene ID | Gene Info |
|---|---|---|
|
PKNOX2
|
ENSG00000165495.11 | PKNOX2 |
Activity-expression correlation:
| Gene | Promoter | Pearson corr. coef. | P-value | Plot |
|---|---|---|---|---|
| PKNOX2 | hg19_v2_chr11_+_125034586_125034604 | -0.80 | 1.8e-02 | Click! |
Activity profile of PKNOX2 motif
Sorted Z-values of PKNOX2 motif
Network of associatons between targets according to the STRING database.
First level regulatory network of PKNOX2
| Promoter | Score | Transcript | Gene | Gene Info |
|---|---|---|---|---|
| chr15_+_33010175 | 0.48 |
ENST00000300177.4 ENST00000560677.1 ENST00000560830.1 |
GREM1 |
gremlin 1, DAN family BMP antagonist |
| chr16_+_2880157 | 0.30 |
ENST00000382280.3 |
ZG16B |
zymogen granule protein 16B |
| chr9_-_79307096 | 0.27 |
ENST00000376717.2 ENST00000223609.6 ENST00000443509.2 |
PRUNE2 |
prune homolog 2 (Drosophila) |
| chr16_+_2880369 | 0.27 |
ENST00000572863.1 |
ZG16B |
zymogen granule protein 16B |
| chr4_-_157892167 | 0.26 |
ENST00000541126.1 |
PDGFC |
platelet derived growth factor C |
| chr4_-_157892055 | 0.25 |
ENST00000422544.2 |
PDGFC |
platelet derived growth factor C |
| chr11_+_6411670 | 0.24 |
ENST00000530395.1 ENST00000527275.1 |
SMPD1 |
sphingomyelin phosphodiesterase 1, acid lysosomal |
| chr16_+_2880254 | 0.23 |
ENST00000570670.1 |
ZG16B |
zymogen granule protein 16B |
| chr11_+_6411636 | 0.22 |
ENST00000299397.3 ENST00000356761.2 ENST00000342245.4 |
SMPD1 |
sphingomyelin phosphodiesterase 1, acid lysosomal |
| chr16_+_2880296 | 0.22 |
ENST00000571723.1 |
ZG16B |
zymogen granule protein 16B |
| chr4_-_157892498 | 0.21 |
ENST00000502773.1 |
PDGFC |
platelet derived growth factor C |
| chr5_+_118407053 | 0.16 |
ENST00000311085.8 ENST00000539542.1 |
DMXL1 |
Dmx-like 1 |
| chr6_-_87804815 | 0.16 |
ENST00000369582.2 |
CGA |
glycoprotein hormones, alpha polypeptide |
| chr9_-_73477826 | 0.15 |
ENST00000396285.1 ENST00000396292.4 ENST00000396280.5 ENST00000358082.3 ENST00000408909.2 ENST00000377097.3 |
TRPM3 |
transient receptor potential cation channel, subfamily M, member 3 |
| chr3_-_52719810 | 0.14 |
ENST00000424867.1 ENST00000394830.3 ENST00000431678.1 ENST00000450271.1 |
PBRM1 |
polybromo 1 |
| chr2_+_202316392 | 0.14 |
ENST00000194530.3 ENST00000392249.2 |
STRADB |
STE20-related kinase adaptor beta |
| chr18_-_54305658 | 0.14 |
ENST00000586262.1 ENST00000217515.6 |
TXNL1 |
thioredoxin-like 1 |
| chrX_-_39923656 | 0.13 |
ENST00000413905.1 |
BCOR |
BCL6 corepressor |
| chr2_-_202316260 | 0.13 |
ENST00000332624.3 |
TRAK2 |
trafficking protein, kinesin binding 2 |
| chr3_-_127541679 | 0.11 |
ENST00000265052.5 |
MGLL |
monoglyceride lipase |
| chr3_-_101395936 | 0.11 |
ENST00000461821.1 |
ZBTB11 |
zinc finger and BTB domain containing 11 |
| chrX_-_153775426 | 0.10 |
ENST00000393562.2 |
G6PD |
glucose-6-phosphate dehydrogenase |
| chr12_+_103981044 | 0.10 |
ENST00000388887.2 |
STAB2 |
stabilin 2 |
| chr2_-_73460334 | 0.10 |
ENST00000258083.2 |
PRADC1 |
protease-associated domain containing 1 |
| chr22_-_31885514 | 0.09 |
ENST00000397525.1 |
EIF4ENIF1 |
eukaryotic translation initiation factor 4E nuclear import factor 1 |
| chr5_-_81046904 | 0.09 |
ENST00000515395.1 |
SSBP2 |
single-stranded DNA binding protein 2 |
| chr5_-_81046841 | 0.09 |
ENST00000509013.2 ENST00000505980.1 ENST00000509053.1 |
SSBP2 |
single-stranded DNA binding protein 2 |
| chr6_+_110299501 | 0.09 |
ENST00000414000.2 |
GPR6 |
G protein-coupled receptor 6 |
| chr20_-_39946237 | 0.09 |
ENST00000441102.2 ENST00000559234.1 |
ZHX3 |
zinc fingers and homeoboxes 3 |
| chr5_-_81046922 | 0.09 |
ENST00000514493.1 ENST00000320672.4 |
SSBP2 |
single-stranded DNA binding protein 2 |
| chr6_+_108881012 | 0.09 |
ENST00000343882.6 |
FOXO3 |
forkhead box O3 |
| chr22_-_50946113 | 0.08 |
ENST00000216080.5 ENST00000474879.2 ENST00000380796.3 |
LMF2 |
lipase maturation factor 2 |
| chr10_+_94608245 | 0.08 |
ENST00000443748.2 ENST00000260762.6 |
EXOC6 |
exocyst complex component 6 |
| chr6_-_116575226 | 0.08 |
ENST00000420283.1 |
TSPYL4 |
TSPY-like 4 |
| chr7_-_148580563 | 0.08 |
ENST00000476773.1 |
EZH2 |
enhancer of zeste homolog 2 (Drosophila) |
| chr3_-_52001448 | 0.08 |
ENST00000461554.1 ENST00000395013.3 ENST00000428823.2 ENST00000483411.1 ENST00000461544.1 ENST00000355852.2 |
PCBP4 |
poly(rC) binding protein 4 |
| chr11_+_12132117 | 0.08 |
ENST00000256194.4 |
MICAL2 |
microtubule associated monooxygenase, calponin and LIM domain containing 2 |
| chr6_-_11382478 | 0.07 |
ENST00000397378.3 ENST00000513989.1 ENST00000508546.1 ENST00000504387.1 |
NEDD9 |
neural precursor cell expressed, developmentally down-regulated 9 |
| chr3_-_127541194 | 0.07 |
ENST00000453507.2 |
MGLL |
monoglyceride lipase |
| chr15_-_37392086 | 0.07 |
ENST00000561208.1 |
MEIS2 |
Meis homeobox 2 |
| chr4_+_166300084 | 0.07 |
ENST00000402744.4 |
CPE |
carboxypeptidase E |
| chr5_-_132073111 | 0.06 |
ENST00000403231.1 |
KIF3A |
kinesin family member 3A |
| chr7_-_137028498 | 0.06 |
ENST00000393083.2 |
PTN |
pleiotrophin |
| chr18_+_3449330 | 0.06 |
ENST00000549253.1 |
TGIF1 |
TGFB-induced factor homeobox 1 |
| chr11_-_111741994 | 0.05 |
ENST00000398006.2 |
ALG9 |
ALG9, alpha-1,2-mannosyltransferase |
| chr17_-_72358001 | 0.05 |
ENST00000375366.3 |
BTBD17 |
BTB (POZ) domain containing 17 |
| chr5_-_127873896 | 0.05 |
ENST00000502468.1 |
FBN2 |
fibrillin 2 |
| chr7_-_137028534 | 0.05 |
ENST00000348225.2 |
PTN |
pleiotrophin |
| chr15_-_61521495 | 0.05 |
ENST00000335670.6 |
RORA |
RAR-related orphan receptor A |
| chr8_+_11666649 | 0.05 |
ENST00000528643.1 ENST00000525777.1 |
FDFT1 |
farnesyl-diphosphate farnesyltransferase 1 |
| chrX_-_71792477 | 0.05 |
ENST00000421523.1 ENST00000415409.1 ENST00000373559.4 ENST00000373556.3 ENST00000373560.2 ENST00000373583.1 ENST00000429103.2 ENST00000373571.1 ENST00000373554.1 |
HDAC8 |
histone deacetylase 8 |
| chr18_+_3449695 | 0.05 |
ENST00000343820.5 |
TGIF1 |
TGFB-induced factor homeobox 1 |
| chr16_-_1275257 | 0.05 |
ENST00000234798.4 |
TPSG1 |
tryptase gamma 1 |
| chr15_+_57668695 | 0.05 |
ENST00000281282.5 |
CGNL1 |
cingulin-like 1 |
| chr3_+_52828805 | 0.04 |
ENST00000416872.2 ENST00000449956.2 |
ITIH3 |
inter-alpha-trypsin inhibitor heavy chain 3 |
| chr4_-_175750364 | 0.04 |
ENST00000340217.5 ENST00000274093.3 |
GLRA3 |
glycine receptor, alpha 3 |
| chr9_-_132805430 | 0.04 |
ENST00000446176.2 ENST00000355681.3 ENST00000420781.1 |
FNBP1 |
formin binding protein 1 |
| chr2_+_128848881 | 0.04 |
ENST00000259253.6 |
UGGT1 |
UDP-glucose glycoprotein glucosyltransferase 1 |
| chr11_-_842509 | 0.04 |
ENST00000322028.4 |
POLR2L |
polymerase (RNA) II (DNA directed) polypeptide L, 7.6kDa |
| chr9_-_123639600 | 0.04 |
ENST00000373896.3 |
PHF19 |
PHD finger protein 19 |
| chr9_-_123639304 | 0.04 |
ENST00000436309.1 |
PHF19 |
PHD finger protein 19 |
| chr20_+_43514320 | 0.04 |
ENST00000372839.3 ENST00000428262.1 ENST00000445830.1 |
YWHAB |
tyrosine 3-monooxygenase/tryptophan 5-monooxygenase activation protein, beta |
| chr12_-_121734489 | 0.04 |
ENST00000412367.2 ENST00000402834.4 ENST00000404169.3 |
CAMKK2 |
calcium/calmodulin-dependent protein kinase kinase 2, beta |
| chr5_-_58882219 | 0.04 |
ENST00000505453.1 ENST00000360047.5 |
PDE4D |
phosphodiesterase 4D, cAMP-specific |
| chr19_+_39759154 | 0.04 |
ENST00000331982.5 |
IFNL2 |
interferon, lambda 2 |
| chr17_-_26220366 | 0.03 |
ENST00000460380.2 ENST00000508862.1 ENST00000379102.3 ENST00000582441.1 |
LYRM9 RP1-66C13.4 |
LYR motif containing 9 Uncharacterized protein |
| chr18_+_54318566 | 0.03 |
ENST00000589935.1 ENST00000357574.3 |
WDR7 |
WD repeat domain 7 |
| chr6_+_41021027 | 0.03 |
ENST00000244669.2 |
APOBEC2 |
apolipoprotein B mRNA editing enzyme, catalytic polypeptide-like 2 |
| chr14_-_21492113 | 0.03 |
ENST00000554094.1 |
NDRG2 |
NDRG family member 2 |
| chr10_-_101945771 | 0.03 |
ENST00000370408.2 ENST00000407654.3 |
ERLIN1 |
ER lipid raft associated 1 |
| chr9_-_123639445 | 0.03 |
ENST00000312189.6 |
PHF19 |
PHD finger protein 19 |
| chr17_-_35969409 | 0.03 |
ENST00000394378.2 ENST00000585472.1 ENST00000591288.1 ENST00000502449.2 ENST00000345615.4 ENST00000346661.4 ENST00000585689.1 ENST00000339208.6 |
SYNRG |
synergin, gamma |
| chr1_-_179112173 | 0.03 |
ENST00000408940.3 ENST00000504405.1 |
ABL2 |
c-abl oncogene 2, non-receptor tyrosine kinase |
| chr3_-_3221358 | 0.03 |
ENST00000424814.1 ENST00000450014.1 ENST00000231948.4 ENST00000432408.2 |
CRBN |
cereblon |
| chr1_-_179112189 | 0.03 |
ENST00000512653.1 ENST00000344730.3 |
ABL2 |
c-abl oncogene 2, non-receptor tyrosine kinase |
| chr20_-_3065362 | 0.03 |
ENST00000380293.3 |
AVP |
arginine vasopressin |
| chr8_+_57124245 | 0.03 |
ENST00000521831.1 ENST00000355315.3 ENST00000303759.3 ENST00000517636.1 ENST00000517933.1 ENST00000518801.1 ENST00000523975.1 ENST00000396723.5 ENST00000523061.1 ENST00000521524.1 |
CHCHD7 |
coiled-coil-helix-coiled-coil-helix domain containing 7 |
| chr6_-_84418841 | 0.03 |
ENST00000369694.2 ENST00000195649.6 |
SNAP91 |
synaptosomal-associated protein, 91kDa |
| chr3_+_185080908 | 0.03 |
ENST00000265026.3 |
MAP3K13 |
mitogen-activated protein kinase kinase kinase 13 |
| chr2_+_128848740 | 0.03 |
ENST00000375990.3 |
UGGT1 |
UDP-glucose glycoprotein glucosyltransferase 1 |
| chr4_+_169418255 | 0.03 |
ENST00000505667.1 ENST00000511948.1 |
PALLD |
palladin, cytoskeletal associated protein |
| chr11_-_19262486 | 0.03 |
ENST00000250024.4 |
E2F8 |
E2F transcription factor 8 |
| chr6_-_84418860 | 0.03 |
ENST00000521743.1 |
SNAP91 |
synaptosomal-associated protein, 91kDa |
| chr1_-_35658736 | 0.03 |
ENST00000357214.5 |
SFPQ |
splicing factor proline/glutamine-rich |
| chr20_-_61847586 | 0.02 |
ENST00000370339.3 |
YTHDF1 |
YTH domain family, member 1 |
| chr2_-_211179883 | 0.02 |
ENST00000352451.3 |
MYL1 |
myosin, light chain 1, alkali; skeletal, fast |
| chr6_-_32140886 | 0.02 |
ENST00000395496.1 |
AGPAT1 |
1-acylglycerol-3-phosphate O-acyltransferase 1 |
| chr4_+_169418195 | 0.02 |
ENST00000261509.6 ENST00000335742.7 |
PALLD |
palladin, cytoskeletal associated protein |
| chr12_+_113495492 | 0.02 |
ENST00000257600.3 |
DTX1 |
deltex homolog 1 (Drosophila) |
| chr11_+_120207787 | 0.02 |
ENST00000397843.2 ENST00000356641.3 |
ARHGEF12 |
Rho guanine nucleotide exchange factor (GEF) 12 |
| chr6_-_37225391 | 0.02 |
ENST00000356757.2 |
TMEM217 |
transmembrane protein 217 |
| chr13_+_33160553 | 0.02 |
ENST00000315596.10 |
PDS5B |
PDS5, regulator of cohesion maintenance, homolog B (S. cerevisiae) |
| chr9_+_137772652 | 0.02 |
ENST00000350339.2 ENST00000291744.6 |
FCN2 |
ficolin (collagen/fibrinogen domain containing lectin) 2 |
| chr1_+_161736072 | 0.02 |
ENST00000367942.3 |
ATF6 |
activating transcription factor 6 |
| chr2_+_191208196 | 0.02 |
ENST00000392329.2 ENST00000322522.4 ENST00000430311.1 ENST00000541441.1 |
INPP1 |
inositol polyphosphate-1-phosphatase |
| chr8_-_57123815 | 0.02 |
ENST00000316981.3 ENST00000423799.2 ENST00000429357.2 |
PLAG1 |
pleiomorphic adenoma gene 1 |
| chr11_-_71810258 | 0.01 |
ENST00000544594.1 |
LAMTOR1 |
late endosomal/lysosomal adaptor, MAPK and MTOR activator 1 |
| chr15_+_68346501 | 0.01 |
ENST00000249636.6 |
PIAS1 |
protein inhibitor of activated STAT, 1 |
| chr17_+_39868577 | 0.01 |
ENST00000329402.3 |
GAST |
gastrin |
| chr12_+_49621658 | 0.01 |
ENST00000541364.1 |
TUBA1C |
tubulin, alpha 1c |
| chr22_+_17082732 | 0.01 |
ENST00000558085.2 ENST00000592918.1 ENST00000400593.2 ENST00000592107.1 ENST00000426585.1 ENST00000591299.1 |
TPTEP1 |
transmembrane phosphatase with tensin homology pseudogene 1 |
| chr2_+_219524473 | 0.01 |
ENST00000439945.1 ENST00000431802.1 |
BCS1L |
BC1 (ubiquinol-cytochrome c reductase) synthesis-like |
| chr16_-_30457048 | 0.01 |
ENST00000500504.2 ENST00000542752.1 |
SEPHS2 |
selenophosphate synthetase 2 |
| chrX_-_77225135 | 0.01 |
ENST00000458128.1 |
PGAM4 |
phosphoglycerate mutase family member 4 |
| chr4_-_149365827 | 0.01 |
ENST00000344721.4 |
NR3C2 |
nuclear receptor subfamily 3, group C, member 2 |
| chr19_+_7733929 | 0.01 |
ENST00000221515.2 |
RETN |
resistin |
| chr22_+_50946645 | 0.01 |
ENST00000420993.2 ENST00000395698.3 ENST00000395701.3 ENST00000523045.1 ENST00000299821.11 |
NCAPH2 |
non-SMC condensin II complex, subunit H2 |
| chr1_+_74701062 | 0.01 |
ENST00000326637.3 |
TNNI3K |
TNNI3 interacting kinase |
| chr5_+_125758865 | 0.01 |
ENST00000542322.1 ENST00000544396.1 |
GRAMD3 |
GRAM domain containing 3 |
| chr11_+_74660278 | 0.01 |
ENST00000263672.6 ENST00000530257.1 ENST00000526361.1 ENST00000532972.1 |
SPCS2 |
signal peptidase complex subunit 2 homolog (S. cerevisiae) |
| chr16_+_50776021 | 0.01 |
ENST00000566679.2 ENST00000564634.1 ENST00000398568.2 |
CYLD |
cylindromatosis (turban tumor syndrome) |
| chr1_+_154966058 | 0.01 |
ENST00000392487.1 |
LENEP |
lens epithelial protein |
| chr3_+_46742823 | 0.01 |
ENST00000326431.3 |
TMIE |
transmembrane inner ear |
| chr11_-_46867780 | 0.01 |
ENST00000529230.1 ENST00000415402.1 ENST00000312055.5 |
CKAP5 |
cytoskeleton associated protein 5 |
| chr2_-_179672142 | 0.00 |
ENST00000342992.6 ENST00000360870.5 ENST00000460472.2 ENST00000589042.1 ENST00000591111.1 ENST00000342175.6 ENST00000359218.5 |
TTN |
titin |
| chr10_+_28822636 | 0.00 |
ENST00000442148.1 ENST00000448193.1 |
WAC |
WW domain containing adaptor with coiled-coil |
| chr3_+_32147997 | 0.00 |
ENST00000282541.5 |
GPD1L |
glycerol-3-phosphate dehydrogenase 1-like |
| chr10_+_70661014 | 0.00 |
ENST00000373585.3 |
DDX50 |
DEAD (Asp-Glu-Ala-Asp) box polypeptide 50 |
| chr2_-_219524193 | 0.00 |
ENST00000450560.1 ENST00000449707.1 ENST00000432460.1 ENST00000411696.2 |
ZNF142 |
zinc finger protein 142 |
| chr6_+_31674639 | 0.00 |
ENST00000556581.1 ENST00000375832.4 ENST00000503322.1 |
LY6G6F MEGT1 |
lymphocyte antigen 6 complex, locus G6F HCG43720, isoform CRA_a; Lymphocyte antigen 6 complex locus protein G6f; Megakaryocyte-enhanced gene transcript 1 protein; Uncharacterized protein |
| chr11_-_64013288 | 0.00 |
ENST00000542235.1 |
PPP1R14B |
protein phosphatase 1, regulatory (inhibitor) subunit 14B |
| chr5_+_125758813 | 0.00 |
ENST00000285689.3 ENST00000515200.1 |
GRAMD3 |
GRAM domain containing 3 |
| chr6_-_114664180 | 0.00 |
ENST00000312719.5 |
HS3ST5 |
heparan sulfate (glucosamine) 3-O-sulfotransferase 5 |
| chr16_-_57481278 | 0.00 |
ENST00000567751.1 ENST00000568940.1 ENST00000563341.1 ENST00000565961.1 ENST00000569370.1 ENST00000567518.1 ENST00000565786.1 ENST00000394391.4 |
CIAPIN1 |
cytokine induced apoptosis inhibitor 1 |
| chr11_+_842928 | 0.00 |
ENST00000397408.1 |
TSPAN4 |
tetraspanin 4 |
| chr1_+_181057638 | 0.00 |
ENST00000367577.4 |
IER5 |
immediate early response 5 |
| chr6_+_42123141 | 0.00 |
ENST00000418175.1 ENST00000541991.1 ENST00000053469.4 ENST00000394237.1 ENST00000372963.1 |
GUCA1A RP1-139D8.6 |
guanylate cyclase activator 1A (retina) RP1-139D8.6 |
| chr1_+_66258846 | 0.00 |
ENST00000341517.4 |
PDE4B |
phosphodiesterase 4B, cAMP-specific |
Gene Ontology Analysis
Gene overrepresentation in biological_process category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.2 | 0.5 | GO:1900158 | negative regulation of bone mineralization involved in bone maturation(GO:1900158) |
| 0.1 | 0.5 | GO:0023021 | termination of signal transduction(GO:0023021) |
| 0.0 | 0.1 | GO:0000415 | negative regulation of histone H3-K36 methylation(GO:0000415) |
| 0.0 | 0.1 | GO:0098935 | dendritic transport(GO:0098935) anterograde dendritic transport(GO:0098937) |
| 0.0 | 0.1 | GO:1904395 | positive regulation of skeletal muscle acetylcholine-gated channel clustering(GO:1904395) negative regulation of neuromuscular junction development(GO:1904397) |
| 0.0 | 0.1 | GO:0009051 | pentose-phosphate shunt, oxidative branch(GO:0009051) |
| 0.0 | 0.1 | GO:0001544 | initiation of primordial ovarian follicle growth(GO:0001544) |
| 0.0 | 0.2 | GO:2000124 | regulation of endocannabinoid signaling pathway(GO:2000124) |
| 0.0 | 0.1 | GO:0036333 | hepatocyte homeostasis(GO:0036333) response to tetrachloromethane(GO:1904772) |
| 0.0 | 0.1 | GO:0019417 | sulfur oxidation(GO:0019417) |
| 0.0 | 0.6 | GO:0007171 | activation of transmembrane receptor protein tyrosine kinase activity(GO:0007171) |
| 0.0 | 0.1 | GO:0097359 | UDP-glucosylation(GO:0097359) |
| 0.0 | 0.1 | GO:0030070 | insulin processing(GO:0030070) |
| 0.0 | 0.0 | GO:0071922 | establishment of sister chromatid cohesion(GO:0034085) cohesin loading(GO:0071921) regulation of cohesin loading(GO:0071922) |
| 0.0 | 0.2 | GO:0070072 | proton-transporting V-type ATPase complex assembly(GO:0070070) vacuolar proton-transporting V-type ATPase complex assembly(GO:0070072) |
| 0.0 | 1.0 | GO:0001895 | retina homeostasis(GO:0001895) |
Gene overrepresentation in cellular_component category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.1 | 0.2 | GO:0043291 | RAVE complex(GO:0043291) |
| 0.0 | 0.5 | GO:0042599 | lamellar body(GO:0042599) |
| 0.0 | 0.1 | GO:0016939 | kinesin II complex(GO:0016939) |
| 0.0 | 0.0 | GO:0016935 | glycine-gated chloride channel complex(GO:0016935) |
Gene overrepresentation in molecular_function category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.0 | 0.5 | GO:0016015 | morphogen activity(GO:0016015) |
| 0.0 | 0.5 | GO:0004767 | sphingomyelin phosphodiesterase activity(GO:0004767) |
| 0.0 | 0.1 | GO:0004345 | glucose-6-phosphate dehydrogenase activity(GO:0004345) |
| 0.0 | 0.1 | GO:0035373 | chondroitin sulfate proteoglycan binding(GO:0035373) |
| 0.0 | 0.7 | GO:0005161 | platelet-derived growth factor receptor binding(GO:0005161) |
| 0.0 | 0.1 | GO:0003980 | UDP-glucose:glycoprotein glucosyltransferase activity(GO:0003980) |
| 0.0 | 0.1 | GO:0000026 | alpha-1,2-mannosyltransferase activity(GO:0000026) |
| 0.0 | 0.0 | GO:0004310 | farnesyl-diphosphate farnesyltransferase activity(GO:0004310) squalene synthase activity(GO:0051996) |
| 0.0 | 0.2 | GO:0047372 | acylglycerol lipase activity(GO:0047372) |
Gene overrepresentation in C2:CP:REACTOME category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.0 | 0.7 | REACTOME VEGF LIGAND RECEPTOR INTERACTIONS | Genes involved in VEGF ligand-receptor interactions |
| 0.0 | 0.2 | REACTOME GLYCOPROTEIN HORMONES | Genes involved in Glycoprotein hormones |