Project
Epithelial-Mesenchymal Transition, human (Scheel, 2011)
Navigation
Downloads
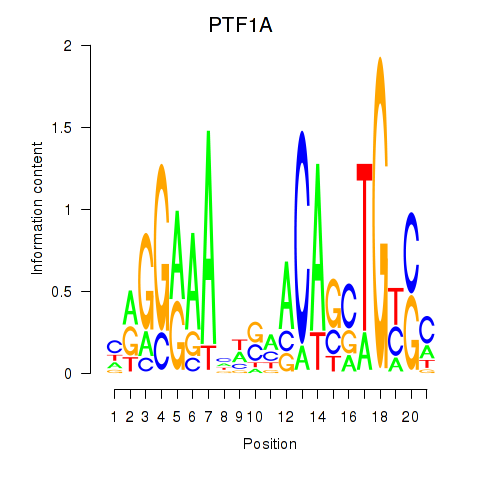
Results for PTF1A
Z-value: 0.51
Transcription factors associated with PTF1A
| Gene Symbol | Gene ID | Gene Info |
|---|---|---|
|
PTF1A
|
ENSG00000168267.5 | PTF1A |
Activity profile of PTF1A motif
Sorted Z-values of PTF1A motif
Network of associatons between targets according to the STRING database.
First level regulatory network of PTF1A
| Promoter | Score | Transcript | Gene | Gene Info |
|---|---|---|---|---|
| chr20_+_44637526 | 0.39 |
ENST00000372330.3 |
MMP9 |
matrix metallopeptidase 9 (gelatinase B, 92kDa gelatinase, 92kDa type IV collagenase) |
| chr19_+_42259329 | 0.37 |
ENST00000199764.6 |
CEACAM6 |
carcinoembryonic antigen-related cell adhesion molecule 6 (non-specific cross reacting antigen) |
| chr10_+_120116527 | 0.34 |
ENST00000445161.1 |
LINC00867 |
long intergenic non-protein coding RNA 867 |
| chr13_+_113633620 | 0.31 |
ENST00000421756.1 ENST00000375601.3 |
MCF2L |
MCF.2 cell line derived transforming sequence-like |
| chr8_+_32405785 | 0.30 |
ENST00000287842.3 |
NRG1 |
neuregulin 1 |
| chr8_+_32405728 | 0.29 |
ENST00000523079.1 ENST00000338921.4 ENST00000356819.4 ENST00000287845.5 ENST00000341377.5 |
NRG1 |
neuregulin 1 |
| chr1_-_9129598 | 0.29 |
ENST00000535586.1 |
SLC2A5 |
solute carrier family 2 (facilitated glucose/fructose transporter), member 5 |
| chr2_-_27435125 | 0.28 |
ENST00000414408.1 ENST00000310574.3 |
SLC5A6 |
solute carrier family 5 (sodium/multivitamin and iodide cotransporter), member 6 |
| chr18_+_33877654 | 0.25 |
ENST00000257209.4 ENST00000445677.1 ENST00000590592.1 ENST00000359247.4 |
FHOD3 |
formin homology 2 domain containing 3 |
| chr4_+_40198527 | 0.23 |
ENST00000381799.5 |
RHOH |
ras homolog family member H |
| chr16_+_5121814 | 0.20 |
ENST00000262374.5 ENST00000586840.1 |
ALG1 |
ALG1, chitobiosyldiphosphodolichol beta-mannosyltransferase |
| chr2_-_237412331 | 0.19 |
ENST00000418802.1 |
IQCA1 |
IQ motif containing with AAA domain 1 |
| chr7_-_72993033 | 0.19 |
ENST00000305632.5 |
TBL2 |
transducin (beta)-like 2 |
| chr1_-_231114542 | 0.18 |
ENST00000522821.1 ENST00000366661.4 ENST00000366662.4 ENST00000414259.1 ENST00000522399.1 |
TTC13 |
tetratricopeptide repeat domain 13 |
| chr2_-_160761179 | 0.18 |
ENST00000554112.1 ENST00000553424.1 ENST00000263636.4 ENST00000504764.1 ENST00000505052.1 |
LY75 LY75-CD302 |
lymphocyte antigen 75 LY75-CD302 readthrough |
| chr2_+_97426631 | 0.18 |
ENST00000377075.2 |
CNNM4 |
cyclin M4 |
| chr1_+_23037323 | 0.18 |
ENST00000544305.1 ENST00000374630.3 ENST00000400191.3 ENST00000374632.3 |
EPHB2 |
EPH receptor B2 |
| chr10_-_101841588 | 0.17 |
ENST00000370418.3 |
CPN1 |
carboxypeptidase N, polypeptide 1 |
| chr1_-_205649580 | 0.16 |
ENST00000367145.3 |
SLC45A3 |
solute carrier family 45, member 3 |
| chrX_-_48216101 | 0.16 |
ENST00000298396.2 ENST00000376893.3 |
SSX3 |
synovial sarcoma, X breakpoint 3 |
| chr3_-_172241250 | 0.16 |
ENST00000420541.2 ENST00000241261.2 |
TNFSF10 |
tumor necrosis factor (ligand) superfamily, member 10 |
| chr17_+_53342311 | 0.15 |
ENST00000226067.5 |
HLF |
hepatic leukemia factor |
| chr7_-_152373216 | 0.15 |
ENST00000359321.1 |
XRCC2 |
X-ray repair complementing defective repair in Chinese hamster cells 2 |
| chr22_-_50050986 | 0.15 |
ENST00000405854.1 |
C22orf34 |
chromosome 22 open reading frame 34 |
| chr22_-_50051151 | 0.14 |
ENST00000400023.1 ENST00000444628.1 |
C22orf34 |
chromosome 22 open reading frame 34 |
| chr13_-_31736027 | 0.14 |
ENST00000380406.5 ENST00000320027.5 ENST00000380405.4 |
HSPH1 |
heat shock 105kDa/110kDa protein 1 |
| chr16_+_2802623 | 0.13 |
ENST00000576924.1 ENST00000575009.1 ENST00000576415.1 ENST00000571378.1 |
SRRM2 |
serine/arginine repetitive matrix 2 |
| chr16_+_71392616 | 0.13 |
ENST00000349553.5 ENST00000302628.4 ENST00000562305.1 |
CALB2 |
calbindin 2 |
| chr13_-_31736132 | 0.13 |
ENST00000429785.2 |
HSPH1 |
heat shock 105kDa/110kDa protein 1 |
| chr10_+_49892904 | 0.13 |
ENST00000360890.2 |
WDFY4 |
WDFY family member 4 |
| chr11_-_1629693 | 0.13 |
ENST00000399685.1 |
KRTAP5-3 |
keratin associated protein 5-3 |
| chr1_+_26856236 | 0.12 |
ENST00000374168.2 ENST00000374166.4 |
RPS6KA1 |
ribosomal protein S6 kinase, 90kDa, polypeptide 1 |
| chr13_+_113812956 | 0.12 |
ENST00000375547.2 ENST00000342783.4 |
PROZ |
protein Z, vitamin K-dependent plasma glycoprotein |
| chr1_+_231114795 | 0.12 |
ENST00000310256.2 ENST00000366658.2 ENST00000450711.1 ENST00000435927.1 |
ARV1 |
ARV1 homolog (S. cerevisiae) |
| chr20_-_44259893 | 0.12 |
ENST00000326000.1 |
WFDC9 |
WAP four-disulfide core domain 9 |
| chr2_-_237416071 | 0.12 |
ENST00000309507.5 ENST00000431676.2 |
IQCA1 |
IQ motif containing with AAA domain 1 |
| chr1_+_150480576 | 0.11 |
ENST00000346569.6 |
ECM1 |
extracellular matrix protein 1 |
| chr1_-_20446020 | 0.11 |
ENST00000375105.3 |
PLA2G2D |
phospholipase A2, group IID |
| chr19_+_6464243 | 0.11 |
ENST00000600229.1 ENST00000356762.3 |
CRB3 |
crumbs homolog 3 (Drosophila) |
| chr4_-_83719983 | 0.11 |
ENST00000319540.4 |
SCD5 |
stearoyl-CoA desaturase 5 |
| chr12_+_52281744 | 0.11 |
ENST00000301190.6 ENST00000538991.1 |
ANKRD33 |
ankyrin repeat domain 33 |
| chr1_+_32084641 | 0.11 |
ENST00000373706.5 |
HCRTR1 |
hypocretin (orexin) receptor 1 |
| chr2_+_85922491 | 0.11 |
ENST00000526018.1 |
GNLY |
granulysin |
| chr8_+_11351494 | 0.11 |
ENST00000259089.4 |
BLK |
B lymphoid tyrosine kinase |
| chr2_-_237416181 | 0.11 |
ENST00000409907.3 |
IQCA1 |
IQ motif containing with AAA domain 1 |
| chr1_+_100503643 | 0.11 |
ENST00000370152.3 |
HIAT1 |
hippocampus abundant transcript 1 |
| chr7_-_99097863 | 0.11 |
ENST00000426306.2 ENST00000337673.6 |
ZNF394 |
zinc finger protein 394 |
| chr1_+_150480551 | 0.11 |
ENST00000369049.4 ENST00000369047.4 |
ECM1 |
extracellular matrix protein 1 |
| chr11_+_7597639 | 0.10 |
ENST00000533792.1 |
PPFIBP2 |
PTPRF interacting protein, binding protein 2 (liprin beta 2) |
| chr2_-_196933536 | 0.10 |
ENST00000312428.6 ENST00000410072.1 |
DNAH7 |
dynein, axonemal, heavy chain 7 |
| chr11_+_71238313 | 0.10 |
ENST00000398536.4 |
KRTAP5-7 |
keratin associated protein 5-7 |
| chrX_-_134305322 | 0.10 |
ENST00000276241.6 ENST00000344129.2 |
CXorf48 |
cancer/testis antigen 55 |
| chr8_+_11351876 | 0.10 |
ENST00000529894.1 |
BLK |
B lymphoid tyrosine kinase |
| chr3_-_155011483 | 0.10 |
ENST00000489090.1 |
RP11-451G4.2 |
RP11-451G4.2 |
| chr9_+_125281420 | 0.09 |
ENST00000340750.1 |
OR1J4 |
olfactory receptor, family 1, subfamily J, member 4 |
| chr1_+_171454639 | 0.09 |
ENST00000392078.3 ENST00000426496.2 |
PRRC2C |
proline-rich coiled-coil 2C |
| chr19_+_54495542 | 0.09 |
ENST00000252729.2 ENST00000352529.1 |
CACNG6 |
calcium channel, voltage-dependent, gamma subunit 6 |
| chr20_+_29956369 | 0.09 |
ENST00000253381.2 |
DEFB118 |
defensin, beta 118 |
| chr1_-_114447620 | 0.09 |
ENST00000369569.1 ENST00000432415.1 ENST00000369571.2 |
AP4B1 |
adaptor-related protein complex 4, beta 1 subunit |
| chr1_+_171454659 | 0.09 |
ENST00000367742.3 ENST00000338920.4 |
PRRC2C |
proline-rich coiled-coil 2C |
| chr1_-_114447683 | 0.09 |
ENST00000256658.4 ENST00000369564.1 |
AP4B1 |
adaptor-related protein complex 4, beta 1 subunit |
| chr3_-_52868931 | 0.09 |
ENST00000486659.1 |
MUSTN1 |
musculoskeletal, embryonic nuclear protein 1 |
| chr12_+_112451120 | 0.08 |
ENST00000261735.3 ENST00000455836.1 |
ERP29 |
endoplasmic reticulum protein 29 |
| chr1_+_151682909 | 0.08 |
ENST00000326413.3 ENST00000442233.2 |
RIIAD1 AL589765.1 |
regulatory subunit of type II PKA R-subunit (RIIa) domain containing 1 Uncharacterized protein; cDNA FLJ36032 fis, clone TESTI2017069 |
| chr4_+_81951957 | 0.08 |
ENST00000282701.2 |
BMP3 |
bone morphogenetic protein 3 |
| chr9_-_104145795 | 0.08 |
ENST00000259407.2 |
BAAT |
bile acid CoA: amino acid N-acyltransferase (glycine N-choloyltransferase) |
| chr5_-_16916624 | 0.08 |
ENST00000513882.1 |
MYO10 |
myosin X |
| chr3_+_101546827 | 0.08 |
ENST00000461724.1 ENST00000483180.1 ENST00000394054.2 |
NFKBIZ |
nuclear factor of kappa light polypeptide gene enhancer in B-cells inhibitor, zeta |
| chr19_+_11670245 | 0.08 |
ENST00000585493.1 |
ZNF627 |
zinc finger protein 627 |
| chr10_+_43633914 | 0.08 |
ENST00000374466.3 ENST00000374464.1 |
CSGALNACT2 |
chondroitin sulfate N-acetylgalactosaminyltransferase 2 |
| chr1_-_204654826 | 0.08 |
ENST00000367177.3 |
LRRN2 |
leucine rich repeat neuronal 2 |
| chr6_-_41715128 | 0.07 |
ENST00000356667.4 ENST00000373025.3 ENST00000425343.2 |
PGC |
progastricsin (pepsinogen C) |
| chr13_-_79979952 | 0.07 |
ENST00000438724.1 |
RBM26 |
RNA binding motif protein 26 |
| chr20_+_42544782 | 0.07 |
ENST00000423191.2 ENST00000372999.1 |
TOX2 |
TOX high mobility group box family member 2 |
| chr12_+_12966250 | 0.07 |
ENST00000352940.4 ENST00000358007.3 ENST00000544400.1 |
DDX47 |
DEAD (Asp-Glu-Ala-Asp) box polypeptide 47 |
| chr3_-_52869205 | 0.07 |
ENST00000446157.2 |
MUSTN1 |
musculoskeletal, embryonic nuclear protein 1 |
| chr13_-_79979919 | 0.07 |
ENST00000267229.7 |
RBM26 |
RNA binding motif protein 26 |
| chr12_+_75784850 | 0.07 |
ENST00000550916.1 ENST00000435775.1 ENST00000378689.2 ENST00000378692.3 ENST00000320460.4 ENST00000547164.1 |
GLIPR1L2 |
GLI pathogenesis-related 1 like 2 |
| chr8_-_99306611 | 0.07 |
ENST00000341166.3 |
NIPAL2 |
NIPA-like domain containing 2 |
| chr17_+_31318886 | 0.07 |
ENST00000269053.3 ENST00000394638.1 |
SPACA3 |
sperm acrosome associated 3 |
| chr19_+_6464502 | 0.07 |
ENST00000308243.7 |
CRB3 |
crumbs homolog 3 (Drosophila) |
| chr12_-_90102594 | 0.07 |
ENST00000428670.3 |
ATP2B1 |
ATPase, Ca++ transporting, plasma membrane 1 |
| chr17_-_3375033 | 0.07 |
ENST00000268981.5 ENST00000397168.3 ENST00000572969.1 ENST00000355380.4 ENST00000571553.1 ENST00000574797.1 ENST00000575375.1 |
SPATA22 |
spermatogenesis associated 22 |
| chr16_-_2004683 | 0.06 |
ENST00000268661.7 |
RPL3L |
ribosomal protein L3-like |
| chr2_+_202122703 | 0.06 |
ENST00000447616.1 ENST00000358485.4 |
CASP8 |
caspase 8, apoptosis-related cysteine peptidase |
| chr9_-_97090926 | 0.06 |
ENST00000335456.7 ENST00000253262.4 ENST00000341207.4 |
NUTM2F |
NUT family member 2F |
| chr2_+_120187465 | 0.06 |
ENST00000409826.1 ENST00000417645.1 |
TMEM37 |
transmembrane protein 37 |
| chr1_-_9129895 | 0.06 |
ENST00000473209.1 |
SLC2A5 |
solute carrier family 2 (facilitated glucose/fructose transporter), member 5 |
| chr17_+_40996590 | 0.06 |
ENST00000253799.3 ENST00000452774.2 |
AOC2 |
amine oxidase, copper containing 2 (retina-specific) |
| chr1_-_78149041 | 0.06 |
ENST00000414381.1 ENST00000370798.1 |
ZZZ3 |
zinc finger, ZZ-type containing 3 |
| chr8_-_38126675 | 0.06 |
ENST00000531823.1 ENST00000534339.1 ENST00000524616.1 ENST00000422581.2 ENST00000424479.2 ENST00000419686.2 |
PPAPDC1B |
phosphatidic acid phosphatase type 2 domain containing 1B |
| chr19_+_42300548 | 0.06 |
ENST00000344550.4 |
CEACAM3 |
carcinoembryonic antigen-related cell adhesion molecule 3 |
| chr12_+_120884222 | 0.06 |
ENST00000551765.1 ENST00000229384.5 |
GATC |
glutamyl-tRNA(Gln) amidotransferase, subunit C |
| chr2_-_228497888 | 0.06 |
ENST00000264387.4 ENST00000409066.1 |
C2orf83 |
chromosome 2 open reading frame 83 |
| chr19_-_48389651 | 0.06 |
ENST00000222002.3 |
SULT2A1 |
sulfotransferase family, cytosolic, 2A, dehydroepiandrosterone (DHEA)-preferring, member 1 |
| chr8_-_99306564 | 0.06 |
ENST00000430223.2 |
NIPAL2 |
NIPA-like domain containing 2 |
| chr17_+_27895609 | 0.06 |
ENST00000581411.2 ENST00000301057.7 |
TP53I13 |
tumor protein p53 inducible protein 13 |
| chr1_-_9129735 | 0.06 |
ENST00000377424.4 |
SLC2A5 |
solute carrier family 2 (facilitated glucose/fructose transporter), member 5 |
| chr17_+_79859985 | 0.06 |
ENST00000333383.7 |
NPB |
neuropeptide B |
| chr19_+_19779686 | 0.05 |
ENST00000415784.2 |
ZNF101 |
zinc finger protein 101 |
| chr7_-_142207004 | 0.05 |
ENST00000426318.2 |
TRBV10-2 |
T cell receptor beta variable 10-2 |
| chr1_-_114447412 | 0.05 |
ENST00000369567.1 ENST00000369566.3 |
AP4B1 |
adaptor-related protein complex 4, beta 1 subunit |
| chr15_+_68924327 | 0.05 |
ENST00000543950.1 |
CORO2B |
coronin, actin binding protein, 2B |
| chr8_-_22089845 | 0.05 |
ENST00000454243.2 |
PHYHIP |
phytanoyl-CoA 2-hydroxylase interacting protein |
| chr19_-_42947121 | 0.05 |
ENST00000601181.1 |
CXCL17 |
chemokine (C-X-C motif) ligand 17 |
| chr2_+_173600565 | 0.05 |
ENST00000397081.3 |
RAPGEF4 |
Rap guanine nucleotide exchange factor (GEF) 4 |
| chr2_-_239140011 | 0.05 |
ENST00000409376.1 ENST00000409070.1 ENST00000409942.1 |
AC016757.3 |
Protein LOC151174 |
| chr8_-_125577940 | 0.05 |
ENST00000519168.1 ENST00000395508.2 |
MTSS1 |
metastasis suppressor 1 |
| chr16_-_46723066 | 0.04 |
ENST00000299138.7 |
VPS35 |
vacuolar protein sorting 35 homolog (S. cerevisiae) |
| chr11_+_71249071 | 0.04 |
ENST00000398534.3 |
KRTAP5-8 |
keratin associated protein 5-8 |
| chr2_+_27435179 | 0.04 |
ENST00000606999.1 ENST00000405489.3 |
ATRAID |
all-trans retinoic acid-induced differentiation factor |
| chr1_+_12185949 | 0.04 |
ENST00000413146.2 |
TNFRSF8 |
tumor necrosis factor receptor superfamily, member 8 |
| chr4_+_183370146 | 0.04 |
ENST00000510504.1 |
TENM3 |
teneurin transmembrane protein 3 |
| chr6_-_159466136 | 0.04 |
ENST00000367066.3 ENST00000326965.6 |
TAGAP |
T-cell activation RhoGTPase activating protein |
| chr11_-_102576537 | 0.04 |
ENST00000260229.4 |
MMP27 |
matrix metallopeptidase 27 |
| chr2_-_1748214 | 0.04 |
ENST00000433670.1 ENST00000425171.1 ENST00000252804.4 |
PXDN |
peroxidasin homolog (Drosophila) |
| chr4_-_14889791 | 0.04 |
ENST00000509654.1 ENST00000515031.1 ENST00000505089.2 |
LINC00504 |
long intergenic non-protein coding RNA 504 |
| chr19_-_11039261 | 0.04 |
ENST00000590329.1 ENST00000587943.1 ENST00000585858.1 ENST00000586748.1 ENST00000586575.1 ENST00000253031.2 |
YIPF2 |
Yip1 domain family, member 2 |
| chr19_-_39694894 | 0.04 |
ENST00000318438.6 |
SYCN |
syncollin |
| chr2_+_173600514 | 0.04 |
ENST00000264111.6 |
RAPGEF4 |
Rap guanine nucleotide exchange factor (GEF) 4 |
| chr4_+_156587979 | 0.04 |
ENST00000511507.1 |
GUCY1A3 |
guanylate cyclase 1, soluble, alpha 3 |
| chr11_+_193065 | 0.04 |
ENST00000342878.2 |
SCGB1C1 |
secretoglobin, family 1C, member 1 |
| chr14_+_70346125 | 0.04 |
ENST00000361956.3 ENST00000381280.4 |
SMOC1 |
SPARC related modular calcium binding 1 |
| chr19_+_29493456 | 0.03 |
ENST00000591143.1 ENST00000592653.1 |
CTD-2081K17.2 |
CTD-2081K17.2 |
| chr3_+_167453493 | 0.03 |
ENST00000295777.5 ENST00000472747.2 |
SERPINI1 |
serpin peptidase inhibitor, clade I (neuroserpin), member 1 |
| chr1_-_76076759 | 0.03 |
ENST00000370855.5 |
SLC44A5 |
solute carrier family 44, member 5 |
| chr18_-_53177984 | 0.03 |
ENST00000543082.1 |
TCF4 |
transcription factor 4 |
| chr12_+_1929421 | 0.03 |
ENST00000543818.1 |
LRTM2 |
leucine-rich repeats and transmembrane domains 2 |
| chr10_+_76871454 | 0.03 |
ENST00000372687.4 |
SAMD8 |
sterile alpha motif domain containing 8 |
| chr3_-_195938256 | 0.03 |
ENST00000296326.3 |
ZDHHC19 |
zinc finger, DHHC-type containing 19 |
| chr12_+_1929644 | 0.03 |
ENST00000299194.1 |
LRTM2 |
leucine-rich repeats and transmembrane domains 2 |
| chr5_-_75013193 | 0.03 |
ENST00000514838.2 ENST00000506164.1 ENST00000502826.1 ENST00000503835.1 ENST00000428202.2 ENST00000380475.2 |
POC5 |
POC5 centriolar protein |
| chr2_-_89417335 | 0.03 |
ENST00000490686.1 |
IGKV1-17 |
immunoglobulin kappa variable 1-17 |
| chr7_-_142099977 | 0.03 |
ENST00000390359.3 |
TRBV7-8 |
T cell receptor beta variable 7-8 |
| chr11_-_1643368 | 0.03 |
ENST00000399682.1 |
KRTAP5-4 |
keratin associated protein 5-4 |
| chr8_-_38126635 | 0.03 |
ENST00000529359.1 |
PPAPDC1B |
phosphatidic acid phosphatase type 2 domain containing 1B |
| chr11_+_96123158 | 0.03 |
ENST00000332349.4 ENST00000458427.1 |
JRKL |
jerky homolog-like (mouse) |
| chr5_+_56205878 | 0.03 |
ENST00000423328.1 |
SETD9 |
SET domain containing 9 |
| chr6_-_46620522 | 0.02 |
ENST00000275016.2 |
CYP39A1 |
cytochrome P450, family 39, subfamily A, polypeptide 1 |
| chr1_+_114447763 | 0.02 |
ENST00000369563.3 |
DCLRE1B |
DNA cross-link repair 1B |
| chr11_+_207477 | 0.02 |
ENST00000526104.1 |
RIC8A |
RIC8 guanine nucleotide exchange factor A |
| chrX_+_48242863 | 0.02 |
ENST00000376886.2 ENST00000375517.3 |
SSX4 |
synovial sarcoma, X breakpoint 4 |
| chr17_-_34313685 | 0.02 |
ENST00000435911.2 ENST00000586216.1 ENST00000394509.4 |
CCL14 |
chemokine (C-C motif) ligand 14 |
| chr19_+_54496132 | 0.02 |
ENST00000346968.2 |
CACNG6 |
calcium channel, voltage-dependent, gamma subunit 6 |
| chr8_-_22089533 | 0.02 |
ENST00000321613.3 |
PHYHIP |
phytanoyl-CoA 2-hydroxylase interacting protein |
| chr11_-_130298888 | 0.02 |
ENST00000257359.6 |
ADAMTS8 |
ADAM metallopeptidase with thrombospondin type 1 motif, 8 |
| chr20_+_48892848 | 0.02 |
ENST00000422459.1 |
RP11-290F20.3 |
RP11-290F20.3 |
| chr16_+_2802316 | 0.02 |
ENST00000301740.8 |
SRRM2 |
serine/arginine repetitive matrix 2 |
| chr9_+_99691286 | 0.02 |
ENST00000372322.3 |
NUTM2G |
NUT family member 2G |
| chr14_+_77292715 | 0.02 |
ENST00000393774.3 ENST00000555189.1 ENST00000450042.2 |
C14orf166B |
chromosome 14 open reading frame 166B |
| chrX_+_154299753 | 0.02 |
ENST00000369459.2 ENST00000369462.1 ENST00000411985.1 ENST00000399042.1 |
BRCC3 |
BRCA1/BRCA2-containing complex, subunit 3 |
| chr19_-_17014408 | 0.02 |
ENST00000594249.1 |
CPAMD8 |
C3 and PZP-like, alpha-2-macroglobulin domain containing 8 |
| chr12_-_57644952 | 0.02 |
ENST00000554578.1 ENST00000546246.2 ENST00000553489.1 ENST00000332782.2 |
STAC3 |
SH3 and cysteine rich domain 3 |
| chrX_+_154299690 | 0.02 |
ENST00000340647.4 ENST00000330045.7 |
BRCC3 |
BRCA1/BRCA2-containing complex, subunit 3 |
| chr21_+_44073860 | 0.02 |
ENST00000335512.4 ENST00000539837.1 ENST00000291539.6 ENST00000380328.2 ENST00000398232.3 ENST00000398234.3 ENST00000398236.3 ENST00000328862.6 ENST00000335440.6 ENST00000398225.3 ENST00000398229.3 ENST00000398227.3 |
PDE9A |
phosphodiesterase 9A |
| chr21_+_44073916 | 0.01 |
ENST00000349112.3 ENST00000398224.3 |
PDE9A |
phosphodiesterase 9A |
| chr16_-_75569068 | 0.01 |
ENST00000336257.3 ENST00000565039.1 |
CHST5 |
carbohydrate (N-acetylglucosamine 6-O) sulfotransferase 5 |
| chr9_-_19033185 | 0.01 |
ENST00000380534.4 |
FAM154A |
family with sequence similarity 154, member A |
| chr9_-_19033237 | 0.01 |
ENST00000380530.1 ENST00000542071.1 |
FAM154A |
family with sequence similarity 154, member A |
| chr2_-_86422523 | 0.01 |
ENST00000442664.2 ENST00000409051.2 ENST00000449247.2 |
IMMT |
inner membrane protein, mitochondrial |
| chr19_-_46526304 | 0.01 |
ENST00000008938.4 |
PGLYRP1 |
peptidoglycan recognition protein 1 |
| chr19_-_10491234 | 0.01 |
ENST00000524462.1 ENST00000531836.1 ENST00000525621.1 |
TYK2 |
tyrosine kinase 2 |
| chr20_+_30697298 | 0.01 |
ENST00000398022.2 |
TM9SF4 |
transmembrane 9 superfamily protein member 4 |
| chr10_-_96829246 | 0.01 |
ENST00000371270.3 ENST00000535898.1 ENST00000539050.1 |
CYP2C8 |
cytochrome P450, family 2, subfamily C, polypeptide 8 |
| chr12_+_31079652 | 0.01 |
ENST00000546076.1 ENST00000535215.1 ENST00000544427.1 ENST00000261177.9 |
TSPAN11 |
tetraspanin 11 |
| chr13_+_49280951 | 0.01 |
ENST00000282018.3 |
CYSLTR2 |
cysteinyl leukotriene receptor 2 |
| chr11_-_96123022 | 0.01 |
ENST00000542662.1 |
CCDC82 |
coiled-coil domain containing 82 |
| chr1_-_9129631 | 0.01 |
ENST00000377414.3 |
SLC2A5 |
solute carrier family 2 (facilitated glucose/fructose transporter), member 5 |
| chrX_+_15808569 | 0.01 |
ENST00000380308.3 ENST00000307771.7 |
ZRSR2 |
zinc finger (CCCH type), RNA-binding motif and serine/arginine rich 2 |
| chr11_+_64052692 | 0.01 |
ENST00000377702.4 |
GPR137 |
G protein-coupled receptor 137 |
| chr14_-_50154921 | 0.00 |
ENST00000553805.2 ENST00000554396.1 ENST00000216367.5 ENST00000539565.2 |
POLE2 |
polymerase (DNA directed), epsilon 2, accessory subunit |
| chr3_-_193272874 | 0.00 |
ENST00000342695.4 |
ATP13A4 |
ATPase type 13A4 |
| chr6_-_159466042 | 0.00 |
ENST00000338313.5 |
TAGAP |
T-cell activation RhoGTPase activating protein |
| chr3_-_88108212 | 0.00 |
ENST00000482016.1 |
CGGBP1 |
CGG triplet repeat binding protein 1 |
| chr6_-_62996066 | 0.00 |
ENST00000281156.4 |
KHDRBS2 |
KH domain containing, RNA binding, signal transduction associated 2 |
| chr7_+_150688166 | 0.00 |
ENST00000461406.1 |
NOS3 |
nitric oxide synthase 3 (endothelial cell) |
| chr3_+_51863433 | 0.00 |
ENST00000444293.1 |
IQCF3 |
IQ motif containing F3 |
| chrX_-_44402231 | 0.00 |
ENST00000378045.4 |
FUNDC1 |
FUN14 domain containing 1 |
| chr4_-_83719884 | 0.00 |
ENST00000282709.4 ENST00000273908.4 |
SCD5 |
stearoyl-CoA desaturase 5 |
| chr4_-_175750364 | 0.00 |
ENST00000340217.5 ENST00000274093.3 |
GLRA3 |
glycine receptor, alpha 3 |
| chr3_+_40518599 | 0.00 |
ENST00000314686.5 ENST00000447116.2 ENST00000429348.2 ENST00000456778.1 |
ZNF619 |
zinc finger protein 619 |
| chr14_-_58893832 | 0.00 |
ENST00000556007.2 |
TIMM9 |
translocase of inner mitochondrial membrane 9 homolog (yeast) |
| chr22_-_17640110 | 0.00 |
ENST00000399852.3 ENST00000336737.4 |
CECR5 |
cat eye syndrome chromosome region, candidate 5 |
Gene Ontology Analysis
Gene overrepresentation in biological_process category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.1 | 0.6 | GO:0021840 | directional guidance of interneurons involved in migration from the subpallium to the cortex(GO:0021840) chemorepulsion involved in interneuron migration from the subpallium to the cortex(GO:0021842) |
| 0.1 | 0.4 | GO:0097319 | fructose transport(GO:0015755) fructose import(GO:0032445) carbohydrate import into cell(GO:0097319) carbohydrate import across plasma membrane(GO:0098704) fructose import across plasma membrane(GO:1990539) |
| 0.1 | 0.2 | GO:0015772 | disaccharide transport(GO:0015766) sucrose transport(GO:0015770) oligosaccharide transport(GO:0015772) |
| 0.0 | 0.3 | GO:1903750 | regulation of intrinsic apoptotic signaling pathway in response to hydrogen peroxide(GO:1903750) negative regulation of intrinsic apoptotic signaling pathway in response to hydrogen peroxide(GO:1903751) |
| 0.0 | 0.0 | GO:1903181 | regulation of dopamine biosynthetic process(GO:1903179) positive regulation of dopamine biosynthetic process(GO:1903181) |
| 0.0 | 0.4 | GO:2001268 | positive regulation of keratinocyte migration(GO:0051549) negative regulation of cysteine-type endopeptidase activity involved in apoptotic signaling pathway(GO:2001268) |
| 0.0 | 0.2 | GO:0030070 | insulin processing(GO:0030070) |
| 0.0 | 0.1 | GO:1903964 | monounsaturated fatty acid metabolic process(GO:1903964) monounsaturated fatty acid biosynthetic process(GO:1903966) |
| 0.0 | 0.1 | GO:0002818 | intracellular defense response(GO:0002818) |
| 0.0 | 0.3 | GO:0051182 | coenzyme transport(GO:0051182) |
| 0.0 | 0.1 | GO:0002361 | CD4-positive, CD25-positive, alpha-beta regulatory T cell differentiation(GO:0002361) |
| 0.0 | 0.1 | GO:0043335 | protein unfolding(GO:0043335) |
| 0.0 | 0.1 | GO:0032383 | regulation of intracellular lipid transport(GO:0032377) regulation of intracellular sterol transport(GO:0032380) regulation of intracellular cholesterol transport(GO:0032383) |
| 0.0 | 0.2 | GO:1904783 | positive regulation of NMDA glutamate receptor activity(GO:1904783) |
| 0.0 | 0.1 | GO:0050653 | chondroitin sulfate proteoglycan biosynthetic process, polysaccharide chain biosynthetic process(GO:0050653) |
| 0.0 | 0.1 | GO:0070681 | glutaminyl-tRNAGln biosynthesis via transamidation(GO:0070681) |
| 0.0 | 0.1 | GO:2000491 | positive regulation of hepatic stellate cell activation(GO:2000491) |
| 0.0 | 0.1 | GO:0000707 | meiotic DNA recombinase assembly(GO:0000707) |
| 0.0 | 0.1 | GO:0000711 | meiotic DNA repair synthesis(GO:0000711) |
| 0.0 | 0.1 | GO:0052056 | modulation by virus of host molecular function(GO:0039506) suppression by virus of host molecular function(GO:0039507) suppression by virus of host catalytic activity(GO:0039513) modulation by virus of host catalytic activity(GO:0039516) suppression by virus of host cysteine-type endopeptidase activity involved in apoptotic process(GO:0039650) negative regulation by symbiont of host catalytic activity(GO:0052053) negative regulation by symbiont of host molecular function(GO:0052056) modulation by symbiont of host catalytic activity(GO:0052148) |
| 0.0 | 0.2 | GO:0051639 | actin filament network formation(GO:0051639) |
| 0.0 | 0.3 | GO:1903830 | magnesium ion transmembrane transport(GO:1903830) |
| 0.0 | 0.0 | GO:0032679 | TRAIL production(GO:0032639) regulation of TRAIL production(GO:0032679) positive regulation of TRAIL production(GO:0032759) |
| 0.0 | 0.1 | GO:0071934 | thiamine transmembrane transport(GO:0071934) |
| 0.0 | 0.1 | GO:0002225 | positive regulation of antimicrobial peptide production(GO:0002225) positive regulation of antimicrobial humoral response(GO:0002760) positive regulation of antibacterial peptide production(GO:0002803) |
| 0.0 | 0.1 | GO:0051673 | membrane disruption in other organism(GO:0051673) |
Gene overrepresentation in cellular_component category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.0 | 0.1 | GO:0036156 | inner dynein arm(GO:0036156) |
| 0.0 | 0.1 | GO:0030956 | glutamyl-tRNA(Gln) amidotransferase complex(GO:0030956) |
| 0.0 | 0.1 | GO:0033063 | Rad51B-Rad51C-Rad51D-XRCC2 complex(GO:0033063) |
| 0.0 | 0.1 | GO:0097013 | phagocytic vesicle lumen(GO:0097013) |
| 0.0 | 0.1 | GO:0032541 | cortical endoplasmic reticulum(GO:0032541) |
| 0.0 | 0.6 | GO:0030673 | axolemma(GO:0030673) |
| 0.0 | 0.3 | GO:0071682 | endocytic vesicle lumen(GO:0071682) |
| 0.0 | 0.2 | GO:0031089 | platelet dense granule lumen(GO:0031089) |
Gene overrepresentation in molecular_function category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.1 | 0.3 | GO:0008523 | sodium-dependent multivitamin transmembrane transporter activity(GO:0008523) |
| 0.1 | 0.4 | GO:0005353 | fructose transmembrane transporter activity(GO:0005353) |
| 0.1 | 0.6 | GO:0005176 | ErbB-2 class receptor binding(GO:0005176) |
| 0.1 | 0.2 | GO:0008515 | sucrose:proton symporter activity(GO:0008506) sucrose transmembrane transporter activity(GO:0008515) disaccharide transmembrane transporter activity(GO:0015154) oligosaccharide transmembrane transporter activity(GO:0015157) |
| 0.0 | 0.2 | GO:0005134 | interleukin-2 receptor binding(GO:0005134) |
| 0.0 | 0.1 | GO:0016499 | orexin receptor activity(GO:0016499) |
| 0.0 | 0.1 | GO:0016215 | stearoyl-CoA 9-desaturase activity(GO:0004768) acyl-CoA desaturase activity(GO:0016215) |
| 0.0 | 0.2 | GO:0005005 | transmembrane-ephrin receptor activity(GO:0005005) |
| 0.0 | 0.1 | GO:0052594 | tryptamine:oxygen oxidoreductase (deaminating) activity(GO:0052593) aminoacetone:oxygen oxidoreductase(deaminating) activity(GO:0052594) aliphatic-amine oxidase activity(GO:0052595) phenethylamine:oxygen oxidoreductase (deaminating) activity(GO:0052596) |
| 0.0 | 0.3 | GO:0000774 | adenyl-nucleotide exchange factor activity(GO:0000774) |
| 0.0 | 0.1 | GO:0050567 | glutaminyl-tRNA synthase (glutamine-hydrolyzing) activity(GO:0050567) |
| 0.0 | 0.1 | GO:0015403 | thiamine uptake transmembrane transporter activity(GO:0015403) |
| 0.0 | 0.1 | GO:0047237 | glucuronylgalactosylproteoglycan 4-beta-N-acetylgalactosaminyltransferase activity(GO:0047237) |
| 0.0 | 0.3 | GO:0015095 | magnesium ion transmembrane transporter activity(GO:0015095) |
| 0.0 | 0.1 | GO:0000150 | recombinase activity(GO:0000150) |
| 0.0 | 0.1 | GO:0097199 | cysteine-type endopeptidase activity involved in apoptotic signaling pathway(GO:0097199) |
Gene overrepresentation in C2:CP category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.0 | 0.6 | PID ERBB NETWORK PATHWAY | ErbB receptor signaling network |
Gene overrepresentation in C2:CP:REACTOME category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.0 | 0.6 | REACTOME DOWNREGULATION OF ERBB2 ERBB3 SIGNALING | Genes involved in Downregulation of ERBB2:ERBB3 signaling |
| 0.0 | 0.4 | REACTOME FACILITATIVE NA INDEPENDENT GLUCOSE TRANSPORTERS | Genes involved in Facilitative Na+-independent glucose transporters |
| 0.0 | 0.3 | REACTOME VITAMIN B5 PANTOTHENATE METABOLISM | Genes involved in Vitamin B5 (pantothenate) metabolism |