Project
Epithelial-Mesenchymal Transition, human (Scheel, 2011)
Navigation
Downloads
Results for RFX5
Z-value: 0.36
Transcription factors associated with RFX5
| Gene Symbol | Gene ID | Gene Info |
|---|---|---|
|
RFX5
|
ENSG00000143390.13 | RFX5 |
Activity-expression correlation:
| Gene | Promoter | Pearson corr. coef. | P-value | Plot |
|---|---|---|---|---|
| RFX5 | hg19_v2_chr1_-_151319283_151319314 | 0.74 | 3.5e-02 | Click! |
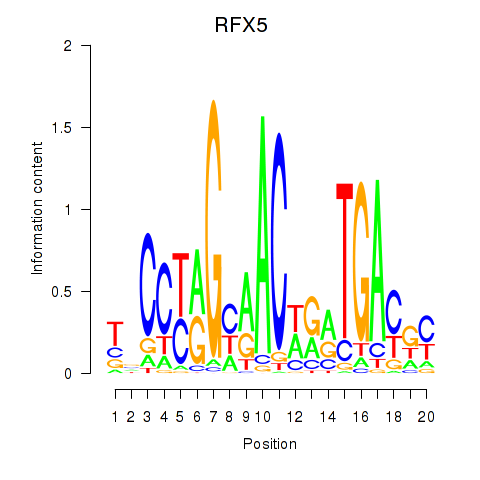
Activity profile of RFX5 motif
Sorted Z-values of RFX5 motif
Network of associatons between targets according to the STRING database.
First level regulatory network of RFX5
| Promoter | Score | Transcript | Gene | Gene Info |
|---|---|---|---|---|
| chr6_+_33043703 | 0.34 |
ENST00000418931.2 ENST00000535465.1 |
HLA-DPB1 |
major histocompatibility complex, class II, DP beta 1 |
| chr19_+_45418067 | 0.22 |
ENST00000589078.1 ENST00000586638.1 |
APOC1 |
apolipoprotein C-I |
| chr19_+_45417921 | 0.21 |
ENST00000252491.4 ENST00000592885.1 ENST00000589781.1 |
APOC1 |
apolipoprotein C-I |
| chr6_+_32821924 | 0.19 |
ENST00000374859.2 ENST00000453265.2 |
PSMB9 |
proteasome (prosome, macropain) subunit, beta type, 9 |
| chr6_+_32812568 | 0.15 |
ENST00000414474.1 |
PSMB9 |
proteasome (prosome, macropain) subunit, beta type, 9 |
| chr6_-_32557610 | 0.11 |
ENST00000360004.5 |
HLA-DRB1 |
major histocompatibility complex, class II, DR beta 1 |
| chr6_-_32498046 | 0.10 |
ENST00000374975.3 |
HLA-DRB5 |
major histocompatibility complex, class II, DR beta 5 |
| chr6_+_26440700 | 0.09 |
ENST00000494393.1 ENST00000482451.1 ENST00000244519.2 ENST00000339789.4 ENST00000471353.1 ENST00000361232.3 ENST00000487627.1 ENST00000496719.1 ENST00000490254.1 ENST00000487272.1 |
BTN3A3 |
butyrophilin, subfamily 3, member A3 |
| chr6_-_32812420 | 0.09 |
ENST00000374881.2 |
PSMB8 |
proteasome (prosome, macropain) subunit, beta type, 8 |
| chr3_-_169587621 | 0.07 |
ENST00000523069.1 ENST00000316428.5 ENST00000264676.5 |
LRRC31 |
leucine rich repeat containing 31 |
| chr1_+_181003067 | 0.06 |
ENST00000434571.2 ENST00000367579.3 ENST00000282990.6 ENST00000367580.5 |
MR1 |
major histocompatibility complex, class I-related |
| chr6_-_32784687 | 0.06 |
ENST00000447394.1 ENST00000438763.2 |
HLA-DOB |
major histocompatibility complex, class II, DO beta |
| chr6_+_26365443 | 0.06 |
ENST00000527422.1 ENST00000356386.2 ENST00000396934.3 ENST00000377708.2 ENST00000396948.1 ENST00000508906.2 |
BTN3A2 |
butyrophilin, subfamily 3, member A2 |
| chr1_+_85527987 | 0.06 |
ENST00000326813.8 ENST00000294664.6 ENST00000528899.1 |
WDR63 |
WD repeat domain 63 |
| chr11_+_45868957 | 0.06 |
ENST00000443527.2 |
CRY2 |
cryptochrome 2 (photolyase-like) |
| chr10_+_82116529 | 0.05 |
ENST00000411538.1 ENST00000256039.2 |
DYDC2 |
DPY30 domain containing 2 |
| chr3_+_66271410 | 0.05 |
ENST00000336733.6 |
SLC25A26 |
solute carrier family 25 (S-adenosylmethionine carrier), member 26 |
| chr11_-_72433346 | 0.05 |
ENST00000334211.8 |
ARAP1 |
ArfGAP with RhoGAP domain, ankyrin repeat and PH domain 1 |
| chr11_+_8008867 | 0.05 |
ENST00000309828.4 ENST00000449102.2 |
EIF3F |
eukaryotic translation initiation factor 3, subunit F |
| chr17_-_19266045 | 0.05 |
ENST00000395616.3 |
B9D1 |
B9 protein domain 1 |
| chr3_+_66271489 | 0.04 |
ENST00000536651.1 |
SLC25A26 |
solute carrier family 25 (S-adenosylmethionine carrier), member 26 |
| chr9_+_135937365 | 0.04 |
ENST00000372080.4 ENST00000351304.7 |
CEL |
carboxyl ester lipase |
| chr6_+_26383318 | 0.04 |
ENST00000469230.1 ENST00000490025.1 ENST00000356709.4 ENST00000352867.2 ENST00000493275.1 ENST00000472507.1 ENST00000482536.1 ENST00000432533.2 ENST00000482842.1 |
BTN2A2 |
butyrophilin, subfamily 2, member A2 |
| chr6_-_32634425 | 0.04 |
ENST00000399082.3 ENST00000399079.3 ENST00000374943.4 ENST00000434651.2 |
HLA-DQB1 |
major histocompatibility complex, class II, DQ beta 1 |
| chr16_+_67381263 | 0.03 |
ENST00000541146.1 ENST00000563189.1 ENST00000290940.7 |
LRRC36 |
leucine rich repeat containing 36 |
| chr1_+_150254936 | 0.03 |
ENST00000447007.1 ENST00000369095.1 ENST00000369094.1 |
C1orf51 |
chromosome 1 open reading frame 51 |
| chr12_-_126467906 | 0.03 |
ENST00000507313.1 ENST00000545784.1 |
LINC00939 |
long intergenic non-protein coding RNA 939 |
| chr2_-_105030466 | 0.03 |
ENST00000449772.1 |
AC068535.3 |
AC068535.3 |
| chr11_-_85430088 | 0.03 |
ENST00000533057.1 ENST00000533892.1 |
SYTL2 |
synaptotagmin-like 2 |
| chr17_-_19265982 | 0.03 |
ENST00000268841.6 ENST00000261499.4 ENST00000575478.1 |
B9D1 |
B9 protein domain 1 |
| chr16_+_67381289 | 0.03 |
ENST00000435835.3 |
LRRC36 |
leucine rich repeat containing 36 |
| chr11_-_6640585 | 0.02 |
ENST00000533371.1 ENST00000528657.1 ENST00000436873.2 ENST00000299427.6 |
TPP1 |
tripeptidyl peptidase I |
| chr6_+_26383404 | 0.02 |
ENST00000416795.2 ENST00000494184.1 |
BTN2A2 |
butyrophilin, subfamily 2, member A2 |
| chr17_-_19265855 | 0.02 |
ENST00000440841.1 ENST00000395615.1 ENST00000461069.2 |
B9D1 |
B9 protein domain 1 |
| chr8_-_98290087 | 0.02 |
ENST00000322128.3 |
TSPYL5 |
TSPY-like 5 |
| chr8_-_99306611 | 0.02 |
ENST00000341166.3 |
NIPAL2 |
NIPA-like domain containing 2 |
| chr1_+_249132462 | 0.02 |
ENST00000306562.3 |
ZNF672 |
zinc finger protein 672 |
| chr20_+_306177 | 0.02 |
ENST00000544632.1 |
SOX12 |
SRY (sex determining region Y)-box 12 |
| chr11_-_118436606 | 0.02 |
ENST00000530872.1 |
IFT46 |
intraflagellar transport 46 homolog (Chlamydomonas) |
| chr1_-_53608289 | 0.02 |
ENST00000371491.4 |
SLC1A7 |
solute carrier family 1 (glutamate transporter), member 7 |
| chr10_-_82116467 | 0.02 |
ENST00000454362.1 |
DYDC1 |
DPY30 domain containing 1 |
| chrX_-_40594755 | 0.01 |
ENST00000324817.1 |
MED14 |
mediator complex subunit 14 |
| chr8_-_99306564 | 0.01 |
ENST00000430223.2 |
NIPAL2 |
NIPA-like domain containing 2 |
| chr13_-_30424821 | 0.01 |
ENST00000380680.4 |
UBL3 |
ubiquitin-like 3 |
| chr19_-_51014345 | 0.01 |
ENST00000391815.3 ENST00000594350.1 ENST00000601423.1 |
JOSD2 |
Josephin domain containing 2 |
| chr1_-_27339317 | 0.01 |
ENST00000289166.5 |
FAM46B |
family with sequence similarity 46, member B |
| chr11_-_118436707 | 0.01 |
ENST00000264020.2 ENST00000264021.3 |
IFT46 |
intraflagellar transport 46 homolog (Chlamydomonas) |
| chr3_-_27410847 | 0.01 |
ENST00000429845.2 ENST00000341435.5 ENST00000435750.1 |
NEK10 |
NIMA-related kinase 10 |
| chr20_+_306221 | 0.01 |
ENST00000342665.2 |
SOX12 |
SRY (sex determining region Y)-box 12 |
| chr6_-_32731299 | 0.01 |
ENST00000435145.2 ENST00000437316.2 |
HLA-DQB2 |
major histocompatibility complex, class II, DQ beta 2 |
| chr19_+_19496728 | 0.01 |
ENST00000537887.1 ENST00000417582.2 |
GATAD2A |
GATA zinc finger domain containing 2A |
| chr22_+_31518938 | 0.01 |
ENST00000412985.1 ENST00000331075.5 ENST00000412277.2 ENST00000420017.1 ENST00000400294.2 ENST00000405300.1 ENST00000404390.3 |
INPP5J |
inositol polyphosphate-5-phosphatase J |
| chr6_-_24667180 | 0.01 |
ENST00000545995.1 |
TDP2 |
tyrosyl-DNA phosphodiesterase 2 |
| chr5_-_55412774 | 0.01 |
ENST00000434982.2 |
ANKRD55 |
ankyrin repeat domain 55 |
| chr6_-_24666819 | 0.01 |
ENST00000341060.3 |
TDP2 |
tyrosyl-DNA phosphodiesterase 2 |
| chr6_-_32731243 | 0.00 |
ENST00000427449.1 ENST00000411527.1 |
HLA-DQB2 |
major histocompatibility complex, class II, DQ beta 2 |
| chr16_-_74808710 | 0.00 |
ENST00000219368.3 ENST00000544337.1 |
FA2H |
fatty acid 2-hydroxylase |
| chr6_-_133079022 | 0.00 |
ENST00000525289.1 ENST00000326499.6 |
VNN2 |
vanin 2 |
| chr19_+_3933579 | 0.00 |
ENST00000593949.1 |
NMRK2 |
nicotinamide riboside kinase 2 |
| chr1_+_46049706 | 0.00 |
ENST00000527470.1 ENST00000525515.1 ENST00000537798.1 ENST00000402363.3 ENST00000528238.1 ENST00000350030.3 ENST00000470768.1 ENST00000372052.4 ENST00000351223.3 |
NASP |
nuclear autoantigenic sperm protein (histone-binding) |
| chr10_-_82116505 | 0.00 |
ENST00000372202.1 ENST00000421924.2 ENST00000453477.1 |
DYDC1 |
DPY30 domain containing 1 |
| chr2_+_242088963 | 0.00 |
ENST00000438799.1 ENST00000402734.1 ENST00000423280.1 |
PPP1R7 |
protein phosphatase 1, regulatory subunit 7 |
| chr6_-_167571817 | 0.00 |
ENST00000366834.1 |
GPR31 |
G protein-coupled receptor 31 |
| chr4_-_143227088 | 0.00 |
ENST00000511838.1 |
INPP4B |
inositol polyphosphate-4-phosphatase, type II, 105kDa |
| chr13_-_47012325 | 0.00 |
ENST00000409879.2 |
KIAA0226L |
KIAA0226-like |
| chr4_-_143226979 | 0.00 |
ENST00000514525.1 |
INPP4B |
inositol polyphosphate-4-phosphatase, type II, 105kDa |
Gene Ontology Analysis
Gene overrepresentation in biological_process category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.1 | 0.4 | GO:0010900 | negative regulation of phosphatidylcholine catabolic process(GO:0010900) |
| 0.0 | 0.1 | GO:0002581 | negative regulation of antigen processing and presentation of peptide or polysaccharide antigen via MHC class II(GO:0002581) |
| 0.0 | 0.1 | GO:2000118 | regulation of sodium-dependent phosphate transport(GO:2000118) |
| 0.0 | 0.0 | GO:0030037 | actin filament reorganization involved in cell cycle(GO:0030037) |
| 0.0 | 0.1 | GO:0051182 | coenzyme transport(GO:0051182) |
Gene overrepresentation in cellular_component category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.0 | 0.4 | GO:1990111 | spermatoproteasome complex(GO:1990111) |
| 0.0 | 0.4 | GO:0042613 | MHC class II protein complex(GO:0042613) |
| 0.0 | 0.4 | GO:0042627 | chylomicron(GO:0042627) |
Gene overrepresentation in molecular_function category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.0 | 0.4 | GO:0060228 | phosphatidylcholine-sterol O-acyltransferase activator activity(GO:0060228) |
| 0.0 | 0.1 | GO:0051185 | coenzyme transporter activity(GO:0051185) |
| 0.0 | 0.6 | GO:0042605 | peptide antigen binding(GO:0042605) |
| 0.0 | 0.4 | GO:0070003 | threonine-type endopeptidase activity(GO:0004298) threonine-type peptidase activity(GO:0070003) |
| 0.0 | 0.1 | GO:0032395 | MHC class II receptor activity(GO:0032395) |
| 0.0 | 0.1 | GO:0008158 | hedgehog receptor activity(GO:0008158) |
Gene overrepresentation in C2:CP:REACTOME category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.0 | 0.5 | REACTOME TRANSLOCATION OF ZAP 70 TO IMMUNOLOGICAL SYNAPSE | Genes involved in Translocation of ZAP-70 to Immunological synapse |