Project
Epithelial-Mesenchymal Transition, human (Scheel, 2011)
Navigation
Downloads
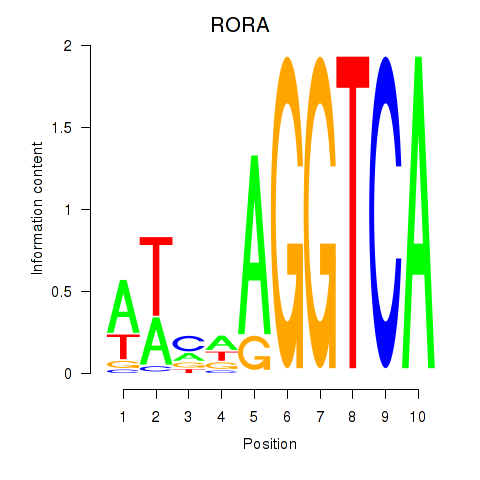
Results for RORA
Z-value: 0.34
Transcription factors associated with RORA
| Gene Symbol | Gene ID | Gene Info |
|---|---|---|
|
RORA
|
ENSG00000069667.11 | RORA |
Activity-expression correlation:
| Gene | Promoter | Pearson corr. coef. | P-value | Plot |
|---|---|---|---|---|
| RORA | hg19_v2_chr15_-_60884706_60884743 | 0.58 | 1.3e-01 | Click! |
Activity profile of RORA motif
Sorted Z-values of RORA motif
Network of associatons between targets according to the STRING database.
First level regulatory network of RORA
| Promoter | Score | Transcript | Gene | Gene Info |
|---|---|---|---|---|
| chr18_-_56296182 | 0.10 |
ENST00000361673.3 |
ALPK2 |
alpha-kinase 2 |
| chr3_+_141105235 | 0.10 |
ENST00000503809.1 |
ZBTB38 |
zinc finger and BTB domain containing 38 |
| chr19_+_1248547 | 0.09 |
ENST00000586757.1 ENST00000300952.2 |
MIDN |
midnolin |
| chrX_+_10124977 | 0.07 |
ENST00000380833.4 |
CLCN4 |
chloride channel, voltage-sensitive 4 |
| chr20_-_13971255 | 0.07 |
ENST00000284951.5 ENST00000378072.5 |
SEL1L2 |
sel-1 suppressor of lin-12-like 2 (C. elegans) |
| chr11_-_7985193 | 0.07 |
ENST00000328600.2 |
NLRP10 |
NLR family, pyrin domain containing 10 |
| chr17_-_33390667 | 0.07 |
ENST00000378516.2 ENST00000268850.7 ENST00000394597.2 |
RFFL |
ring finger and FYVE-like domain containing E3 ubiquitin protein ligase |
| chr12_+_65672423 | 0.07 |
ENST00000355192.3 ENST00000308259.5 ENST00000540804.1 ENST00000535664.1 ENST00000541189.1 |
MSRB3 |
methionine sulfoxide reductase B3 |
| chr11_+_66624527 | 0.07 |
ENST00000393952.3 |
LRFN4 |
leucine rich repeat and fibronectin type III domain containing 4 |
| chr11_-_790060 | 0.06 |
ENST00000330106.4 |
CEND1 |
cell cycle exit and neuronal differentiation 1 |
| chr6_-_9933500 | 0.06 |
ENST00000492169.1 |
OFCC1 |
orofacial cleft 1 candidate 1 |
| chr16_+_31366536 | 0.06 |
ENST00000562522.1 |
ITGAX |
integrin, alpha X (complement component 3 receptor 4 subunit) |
| chrX_-_21776281 | 0.05 |
ENST00000379494.3 |
SMPX |
small muscle protein, X-linked |
| chr12_+_65672702 | 0.05 |
ENST00000538045.1 ENST00000535239.1 |
MSRB3 |
methionine sulfoxide reductase B3 |
| chr10_-_65028817 | 0.05 |
ENST00000542921.1 |
JMJD1C |
jumonji domain containing 1C |
| chr11_+_64879317 | 0.05 |
ENST00000526809.1 ENST00000279263.7 ENST00000524986.1 ENST00000534371.1 ENST00000540748.1 ENST00000525385.1 ENST00000345348.5 ENST00000531321.1 ENST00000529414.1 ENST00000526085.1 ENST00000530750.1 |
TM7SF2 |
transmembrane 7 superfamily member 2 |
| chr16_+_31366455 | 0.05 |
ENST00000268296.4 |
ITGAX |
integrin, alpha X (complement component 3 receptor 4 subunit) |
| chr10_+_77056134 | 0.05 |
ENST00000528121.1 ENST00000416398.1 |
ZNF503-AS1 |
ZNF503 antisense RNA 1 |
| chr9_-_130966497 | 0.05 |
ENST00000393608.1 ENST00000372948.3 |
CIZ1 |
CDKN1A interacting zinc finger protein 1 |
| chr5_+_73109339 | 0.05 |
ENST00000296799.4 |
ARHGEF28 |
Rho guanine nucleotide exchange factor (GEF) 28 |
| chr14_-_107114267 | 0.05 |
ENST00000454421.2 |
IGHV3-64 |
immunoglobulin heavy variable 3-64 |
| chr8_-_134072593 | 0.05 |
ENST00000427060.2 |
SLA |
Src-like-adaptor |
| chr7_+_128399002 | 0.05 |
ENST00000493278.1 |
CALU |
calumenin |
| chr1_-_241799232 | 0.05 |
ENST00000366553.1 |
CHML |
choroideremia-like (Rab escort protein 2) |
| chr1_+_36690011 | 0.05 |
ENST00000354618.5 ENST00000469141.2 ENST00000478853.1 |
THRAP3 |
thyroid hormone receptor associated protein 3 |
| chr10_-_103578162 | 0.04 |
ENST00000361464.3 ENST00000357797.5 ENST00000370094.3 |
MGEA5 |
meningioma expressed antigen 5 (hyaluronidase) |
| chr3_+_10068095 | 0.04 |
ENST00000287647.3 ENST00000383807.1 ENST00000383806.1 ENST00000419585.1 |
FANCD2 |
Fanconi anemia, complementation group D2 |
| chr7_+_112063192 | 0.04 |
ENST00000005558.4 |
IFRD1 |
interferon-related developmental regulator 1 |
| chr16_+_2076869 | 0.04 |
ENST00000424542.2 ENST00000432365.2 |
SLC9A3R2 |
solute carrier family 9, subfamily A (NHE3, cation proton antiporter 3), member 3 regulator 2 |
| chr17_+_74075263 | 0.04 |
ENST00000334586.5 ENST00000392503.2 |
ZACN |
zinc activated ligand-gated ion channel |
| chr15_+_43809797 | 0.04 |
ENST00000399453.1 ENST00000300231.5 |
MAP1A |
microtubule-associated protein 1A |
| chr6_+_149068464 | 0.04 |
ENST00000367463.4 |
UST |
uronyl-2-sulfotransferase |
| chr20_+_3776936 | 0.04 |
ENST00000439880.2 |
CDC25B |
cell division cycle 25B |
| chr2_-_207024233 | 0.04 |
ENST00000423725.1 ENST00000233190.6 |
NDUFS1 |
NADH dehydrogenase (ubiquinone) Fe-S protein 1, 75kDa (NADH-coenzyme Q reductase) |
| chr7_-_105319536 | 0.04 |
ENST00000477775.1 |
ATXN7L1 |
ataxin 7-like 1 |
| chr11_-_104480019 | 0.04 |
ENST00000536529.1 ENST00000545630.1 ENST00000538641.1 |
RP11-886D15.1 |
RP11-886D15.1 |
| chr11_-_64527425 | 0.04 |
ENST00000377432.3 |
PYGM |
phosphorylase, glycogen, muscle |
| chr12_+_69139886 | 0.04 |
ENST00000398004.2 |
SLC35E3 |
solute carrier family 35, member E3 |
| chr1_-_150207017 | 0.04 |
ENST00000369119.3 |
ANP32E |
acidic (leucine-rich) nuclear phosphoprotein 32 family, member E |
| chr20_+_37590942 | 0.04 |
ENST00000373325.2 ENST00000252011.3 ENST00000373323.4 |
DHX35 |
DEAH (Asp-Glu-Ala-His) box polypeptide 35 |
| chr2_+_109237717 | 0.04 |
ENST00000409441.1 |
LIMS1 |
LIM and senescent cell antigen-like domains 1 |
| chr7_-_107642348 | 0.04 |
ENST00000393561.1 |
LAMB1 |
laminin, beta 1 |
| chr3_+_45067659 | 0.04 |
ENST00000296130.4 |
CLEC3B |
C-type lectin domain family 3, member B |
| chr2_+_198365095 | 0.04 |
ENST00000409468.1 |
HSPE1 |
heat shock 10kDa protein 1 |
| chr3_+_46449049 | 0.04 |
ENST00000357392.4 ENST00000400880.3 ENST00000433848.1 |
CCRL2 |
chemokine (C-C motif) receptor-like 2 |
| chr16_+_29991673 | 0.03 |
ENST00000416441.2 |
TAOK2 |
TAO kinase 2 |
| chr18_-_19284724 | 0.03 |
ENST00000580981.1 ENST00000289119.2 |
ABHD3 |
abhydrolase domain containing 3 |
| chr10_-_103578182 | 0.03 |
ENST00000439817.1 |
MGEA5 |
meningioma expressed antigen 5 (hyaluronidase) |
| chr20_+_3776371 | 0.03 |
ENST00000245960.5 |
CDC25B |
cell division cycle 25B |
| chrX_+_153770421 | 0.03 |
ENST00000369609.5 ENST00000369607.1 |
IKBKG |
inhibitor of kappa light polypeptide gene enhancer in B-cells, kinase gamma |
| chr3_+_46448648 | 0.03 |
ENST00000399036.3 |
CCRL2 |
chemokine (C-C motif) receptor-like 2 |
| chr7_-_127983877 | 0.03 |
ENST00000415472.2 ENST00000478061.1 ENST00000223073.2 ENST00000459726.1 |
RBM28 |
RNA binding motif protein 28 |
| chr12_+_58013693 | 0.03 |
ENST00000320442.4 ENST00000379218.2 |
SLC26A10 |
solute carrier family 26, member 10 |
| chr17_-_4458616 | 0.03 |
ENST00000381556.2 |
MYBBP1A |
MYB binding protein (P160) 1a |
| chr17_-_48785216 | 0.03 |
ENST00000285243.6 |
ANKRD40 |
ankyrin repeat domain 40 |
| chr10_+_76586348 | 0.03 |
ENST00000372724.1 ENST00000287239.4 ENST00000372714.1 |
KAT6B |
K(lysine) acetyltransferase 6B |
| chr1_-_157108266 | 0.03 |
ENST00000326786.4 |
ETV3 |
ets variant 3 |
| chr11_-_111741994 | 0.03 |
ENST00000398006.2 |
ALG9 |
ALG9, alpha-1,2-mannosyltransferase |
| chr6_+_107077435 | 0.03 |
ENST00000369046.4 |
QRSL1 |
glutaminyl-tRNA synthase (glutamine-hydrolyzing)-like 1 |
| chr17_-_49198216 | 0.03 |
ENST00000262013.7 ENST00000357122.4 |
SPAG9 |
sperm associated antigen 9 |
| chr10_+_94050913 | 0.03 |
ENST00000358935.2 |
MARCH5 |
membrane-associated ring finger (C3HC4) 5 |
| chr8_+_77318769 | 0.03 |
ENST00000518732.1 |
RP11-706J10.1 |
long intergenic non-protein coding RNA 1111 |
| chr10_-_14050522 | 0.03 |
ENST00000342409.2 |
FRMD4A |
FERM domain containing 4A |
| chr12_+_93963590 | 0.03 |
ENST00000340600.2 |
SOCS2 |
suppressor of cytokine signaling 2 |
| chr19_-_49552006 | 0.03 |
ENST00000391869.3 |
CGB1 |
chorionic gonadotropin, beta polypeptide 1 |
| chr6_+_36839616 | 0.03 |
ENST00000359359.2 ENST00000510325.2 |
C6orf89 |
chromosome 6 open reading frame 89 |
| chr5_-_87564620 | 0.03 |
ENST00000506536.1 ENST00000512429.1 ENST00000514135.1 ENST00000296595.6 ENST00000509387.1 |
TMEM161B |
transmembrane protein 161B |
| chr5_-_137071689 | 0.03 |
ENST00000505853.1 |
KLHL3 |
kelch-like family member 3 |
| chr3_-_27498235 | 0.03 |
ENST00000295736.5 ENST00000428386.1 ENST00000428179.1 |
SLC4A7 |
solute carrier family 4, sodium bicarbonate cotransporter, member 7 |
| chr5_+_169780485 | 0.02 |
ENST00000377360.4 |
KCNIP1 |
Kv channel interacting protein 1 |
| chr19_+_7011509 | 0.02 |
ENST00000377296.3 |
AC025278.1 |
Uncharacterized protein |
| chr15_+_76352178 | 0.02 |
ENST00000388942.3 |
C15orf27 |
chromosome 15 open reading frame 27 |
| chr3_-_123339418 | 0.02 |
ENST00000583087.1 |
MYLK |
myosin light chain kinase |
| chr8_+_37553261 | 0.02 |
ENST00000331569.4 |
ZNF703 |
zinc finger protein 703 |
| chr5_+_140027355 | 0.02 |
ENST00000417647.2 ENST00000507593.1 ENST00000508301.1 |
IK |
IK cytokine, down-regulator of HLA II |
| chr3_-_113465065 | 0.02 |
ENST00000497255.1 ENST00000478020.1 ENST00000240922.3 ENST00000493900.1 |
NAA50 |
N(alpha)-acetyltransferase 50, NatE catalytic subunit |
| chr20_-_3748416 | 0.02 |
ENST00000399672.1 |
C20orf27 |
chromosome 20 open reading frame 27 |
| chr1_-_85156216 | 0.02 |
ENST00000342203.3 ENST00000370612.4 |
SSX2IP |
synovial sarcoma, X breakpoint 2 interacting protein |
| chr10_+_81107271 | 0.02 |
ENST00000448165.1 |
PPIF |
peptidylprolyl isomerase F |
| chrX_-_71526999 | 0.02 |
ENST00000453707.2 ENST00000373619.3 |
CITED1 |
Cbp/p300-interacting transactivator, with Glu/Asp-rich carboxy-terminal domain, 1 |
| chr5_-_131347583 | 0.02 |
ENST00000379255.1 ENST00000430403.1 ENST00000544770.1 ENST00000379246.1 ENST00000414078.1 ENST00000441995.1 |
ACSL6 |
acyl-CoA synthetase long-chain family member 6 |
| chr22_-_24110063 | 0.02 |
ENST00000520222.1 ENST00000401675.3 |
CHCHD10 |
coiled-coil-helix-coiled-coil-helix domain containing 10 |
| chr10_+_72575643 | 0.02 |
ENST00000373202.3 |
SGPL1 |
sphingosine-1-phosphate lyase 1 |
| chr5_-_131347501 | 0.02 |
ENST00000543479.1 |
ACSL6 |
acyl-CoA synthetase long-chain family member 6 |
| chr2_-_209118974 | 0.02 |
ENST00000415913.1 ENST00000415282.1 ENST00000446179.1 |
IDH1 |
isocitrate dehydrogenase 1 (NADP+), soluble |
| chr3_-_126236605 | 0.02 |
ENST00000290868.2 |
UROC1 |
urocanate hydratase 1 |
| chr9_-_72374848 | 0.02 |
ENST00000377200.5 ENST00000340434.4 ENST00000472967.2 |
PTAR1 |
protein prenyltransferase alpha subunit repeat containing 1 |
| chr9_-_116065551 | 0.02 |
ENST00000297894.5 |
RNF183 |
ring finger protein 183 |
| chr3_+_113465866 | 0.02 |
ENST00000273398.3 ENST00000538620.1 ENST00000496747.1 ENST00000475322.1 |
ATP6V1A |
ATPase, H+ transporting, lysosomal 70kDa, V1 subunit A |
| chr12_+_93964158 | 0.02 |
ENST00000549206.1 |
SOCS2 |
suppressor of cytokine signaling 2 |
| chr20_+_18125727 | 0.02 |
ENST00000489634.2 |
CSRP2BP |
CSRP2 binding protein |
| chr22_-_32341336 | 0.02 |
ENST00000248984.3 |
C22orf24 |
chromosome 22 open reading frame 24 |
| chr1_-_157108130 | 0.02 |
ENST00000368192.4 |
ETV3 |
ets variant 3 |
| chr3_-_123339343 | 0.02 |
ENST00000578202.1 |
MYLK |
myosin light chain kinase |
| chr6_-_41703296 | 0.02 |
ENST00000373033.1 |
TFEB |
transcription factor EB |
| chr1_-_32403903 | 0.02 |
ENST00000344035.6 ENST00000356536.3 |
PTP4A2 |
protein tyrosine phosphatase type IVA, member 2 |
| chr19_-_50311896 | 0.02 |
ENST00000529634.2 |
FUZ |
fuzzy planar cell polarity protein |
| chr1_-_211752073 | 0.02 |
ENST00000367001.4 |
SLC30A1 |
solute carrier family 30 (zinc transporter), member 1 |
| chr9_+_134103496 | 0.02 |
ENST00000498010.1 ENST00000476004.1 ENST00000528406.1 |
NUP214 |
nucleoporin 214kDa |
| chr9_+_1051481 | 0.02 |
ENST00000358146.2 ENST00000259622.6 |
DMRT2 |
doublesex and mab-3 related transcription factor 2 |
| chr18_+_32073253 | 0.02 |
ENST00000283365.9 ENST00000596745.1 ENST00000315456.6 |
DTNA |
dystrobrevin, alpha |
| chr15_-_52263937 | 0.02 |
ENST00000315141.5 ENST00000299601.5 |
LEO1 |
Leo1, Paf1/RNA polymerase II complex component, homolog (S. cerevisiae) |
| chr3_+_130150307 | 0.02 |
ENST00000512836.1 |
COL6A5 |
collagen, type VI, alpha 5 |
| chr19_-_42931567 | 0.02 |
ENST00000244289.4 |
LIPE |
lipase, hormone-sensitive |
| chr1_-_205719295 | 0.02 |
ENST00000367142.4 |
NUCKS1 |
nuclear casein kinase and cyclin-dependent kinase substrate 1 |
| chr3_-_113464906 | 0.02 |
ENST00000477813.1 |
NAA50 |
N(alpha)-acetyltransferase 50, NatE catalytic subunit |
| chrX_-_30327495 | 0.02 |
ENST00000453287.1 |
NR0B1 |
nuclear receptor subfamily 0, group B, member 1 |
| chr10_-_128359074 | 0.02 |
ENST00000544758.1 |
C10orf90 |
chromosome 10 open reading frame 90 |
| chr11_+_65292884 | 0.02 |
ENST00000527009.1 |
SCYL1 |
SCY1-like 1 (S. cerevisiae) |
| chr12_-_58135903 | 0.02 |
ENST00000257897.3 |
AGAP2 |
ArfGAP with GTPase domain, ankyrin repeat and PH domain 2 |
| chr9_-_86322831 | 0.02 |
ENST00000257468.7 |
UBQLN1 |
ubiquilin 1 |
| chr10_-_65028938 | 0.02 |
ENST00000402544.1 |
JMJD1C |
jumonji domain containing 1C |
| chr7_-_25164969 | 0.02 |
ENST00000305786.2 |
CYCS |
cytochrome c, somatic |
| chr3_-_49158312 | 0.02 |
ENST00000398892.3 ENST00000453664.1 ENST00000398888.2 |
USP19 |
ubiquitin specific peptidase 19 |
| chr11_-_6462210 | 0.02 |
ENST00000265983.3 |
HPX |
hemopexin |
| chr1_+_160147176 | 0.02 |
ENST00000470705.1 |
ATP1A4 |
ATPase, Na+/K+ transporting, alpha 4 polypeptide |
| chr7_-_45151272 | 0.02 |
ENST00000461363.1 ENST00000495078.1 ENST00000494076.1 ENST00000478532.1 ENST00000258770.3 ENST00000361278.3 |
TBRG4 |
transforming growth factor beta regulator 4 |
| chr18_-_12377283 | 0.02 |
ENST00000269143.3 |
AFG3L2 |
AFG3-like AAA ATPase 2 |
| chr14_-_102552659 | 0.02 |
ENST00000441629.2 |
HSP90AA1 |
heat shock protein 90kDa alpha (cytosolic), class A member 1 |
| chr17_-_41132010 | 0.02 |
ENST00000409103.1 ENST00000360221.4 |
PTGES3L-AARSD1 |
PTGES3L-AARSD1 readthrough |
| chr3_-_49158218 | 0.01 |
ENST00000417901.1 ENST00000306026.5 ENST00000434032.2 |
USP19 |
ubiquitin specific peptidase 19 |
| chr1_+_236849754 | 0.01 |
ENST00000542672.1 ENST00000366578.4 |
ACTN2 |
actinin, alpha 2 |
| chr1_+_43148625 | 0.01 |
ENST00000436427.1 |
YBX1 |
Y box binding protein 1 |
| chr15_-_72767490 | 0.01 |
ENST00000565181.1 |
RP11-1007O24.3 |
RP11-1007O24.3 |
| chr6_-_134861089 | 0.01 |
ENST00000606039.1 |
RP11-557H15.4 |
RP11-557H15.4 |
| chr6_-_71012773 | 0.01 |
ENST00000370496.3 ENST00000357250.6 |
COL9A1 |
collagen, type IX, alpha 1 |
| chr5_-_140027357 | 0.01 |
ENST00000252102.4 |
NDUFA2 |
NADH dehydrogenase (ubiquinone) 1 alpha subcomplex, 2, 8kDa |
| chr9_-_35685452 | 0.01 |
ENST00000607559.1 |
TPM2 |
tropomyosin 2 (beta) |
| chr16_-_18812746 | 0.01 |
ENST00000546206.2 ENST00000562819.1 ENST00000562234.2 ENST00000304414.7 ENST00000567078.2 |
ARL6IP1 RP11-1035H13.3 |
ADP-ribosylation factor-like 6 interacting protein 1 Uncharacterized protein |
| chr17_+_39994032 | 0.01 |
ENST00000293303.4 ENST00000438813.1 |
KLHL10 |
kelch-like family member 10 |
| chr15_+_78441663 | 0.01 |
ENST00000299518.2 ENST00000558554.1 ENST00000557826.1 ENST00000561279.1 ENST00000559186.1 ENST00000560770.1 ENST00000559881.1 ENST00000559205.1 ENST00000441490.2 |
IDH3A |
isocitrate dehydrogenase 3 (NAD+) alpha |
| chr21_+_17553910 | 0.01 |
ENST00000428669.2 |
LINC00478 |
long intergenic non-protein coding RNA 478 |
| chr7_-_128415844 | 0.01 |
ENST00000249389.2 |
OPN1SW |
opsin 1 (cone pigments), short-wave-sensitive |
| chr12_+_109577202 | 0.01 |
ENST00000377848.3 ENST00000377854.5 |
ACACB |
acetyl-CoA carboxylase beta |
| chr1_-_91487770 | 0.01 |
ENST00000337393.5 |
ZNF644 |
zinc finger protein 644 |
| chr19_+_46850251 | 0.01 |
ENST00000012443.4 |
PPP5C |
protein phosphatase 5, catalytic subunit |
| chr19_-_51192661 | 0.01 |
ENST00000391813.1 |
SHANK1 |
SH3 and multiple ankyrin repeat domains 1 |
| chr6_-_89673280 | 0.01 |
ENST00000369475.3 ENST00000369485.4 ENST00000538899.1 ENST00000265607.6 |
RNGTT |
RNA guanylyltransferase and 5'-phosphatase |
| chr6_+_107077471 | 0.01 |
ENST00000369044.1 |
QRSL1 |
glutaminyl-tRNA synthase (glutamine-hydrolyzing)-like 1 |
| chr3_+_128968437 | 0.01 |
ENST00000314797.6 |
COPG1 |
coatomer protein complex, subunit gamma 1 |
| chrX_-_129299638 | 0.01 |
ENST00000535724.1 ENST00000346424.2 |
AIFM1 |
apoptosis-inducing factor, mitochondrion-associated, 1 |
| chr13_+_22245522 | 0.01 |
ENST00000382353.5 |
FGF9 |
fibroblast growth factor 9 |
| chr6_-_97345689 | 0.01 |
ENST00000316149.7 |
NDUFAF4 |
NADH dehydrogenase (ubiquinone) complex I, assembly factor 4 |
| chr7_-_123197733 | 0.01 |
ENST00000470123.1 ENST00000471770.1 |
NDUFA5 |
NADH dehydrogenase (ubiquinone) 1 alpha subcomplex, 5 |
| chr3_-_49377446 | 0.01 |
ENST00000351842.4 ENST00000416417.1 ENST00000415188.1 |
USP4 |
ubiquitin specific peptidase 4 (proto-oncogene) |
| chr14_-_58893832 | 0.01 |
ENST00000556007.2 |
TIMM9 |
translocase of inner mitochondrial membrane 9 homolog (yeast) |
| chr1_-_153522562 | 0.01 |
ENST00000368714.1 |
S100A4 |
S100 calcium binding protein A4 |
| chr19_+_46850320 | 0.01 |
ENST00000391919.1 |
PPP5C |
protein phosphatase 5, catalytic subunit |
| chr14_-_23755297 | 0.01 |
ENST00000357460.5 |
HOMEZ |
homeobox and leucine zipper encoding |
| chr7_+_123241908 | 0.01 |
ENST00000434204.1 ENST00000437535.1 ENST00000451215.1 |
ASB15 |
ankyrin repeat and SOCS box containing 15 |
| chr2_-_207024134 | 0.01 |
ENST00000457011.1 ENST00000440274.1 ENST00000432169.1 |
NDUFS1 |
NADH dehydrogenase (ubiquinone) Fe-S protein 1, 75kDa (NADH-coenzyme Q reductase) |
| chr2_+_198365122 | 0.01 |
ENST00000604458.1 |
HSPE1-MOB4 |
HSPE1-MOB4 readthrough |
| chr12_+_56414851 | 0.01 |
ENST00000547167.1 |
IKZF4 |
IKAROS family zinc finger 4 (Eos) |
| chr2_+_171034646 | 0.01 |
ENST00000409044.3 ENST00000408978.4 |
MYO3B |
myosin IIIB |
| chr3_-_183735651 | 0.01 |
ENST00000427120.2 ENST00000392579.2 ENST00000382494.2 ENST00000446941.2 |
ABCC5 |
ATP-binding cassette, sub-family C (CFTR/MRP), member 5 |
| chr15_-_77988485 | 0.01 |
ENST00000561030.1 |
LINGO1 |
leucine rich repeat and Ig domain containing 1 |
| chr19_-_42498231 | 0.01 |
ENST00000602133.1 |
ATP1A3 |
ATPase, Na+/K+ transporting, alpha 3 polypeptide |
| chr19_-_42498369 | 0.01 |
ENST00000302102.5 ENST00000545399.1 |
ATP1A3 |
ATPase, Na+/K+ transporting, alpha 3 polypeptide |
| chr21_-_43816152 | 0.01 |
ENST00000433957.2 ENST00000398397.3 |
TMPRSS3 |
transmembrane protease, serine 3 |
| chr12_-_16761007 | 0.01 |
ENST00000354662.1 ENST00000441439.2 |
LMO3 |
LIM domain only 3 (rhombotin-like 2) |
| chr3_-_179322416 | 0.01 |
ENST00000259038.2 |
MRPL47 |
mitochondrial ribosomal protein L47 |
| chr16_-_47177874 | 0.01 |
ENST00000562435.1 |
NETO2 |
neuropilin (NRP) and tolloid (TLL)-like 2 |
| chr9_+_139717847 | 0.01 |
ENST00000436380.1 |
RABL6 |
RAB, member RAS oncogene family-like 6 |
| chrX_-_43741594 | 0.01 |
ENST00000536181.1 ENST00000378069.4 |
MAOB |
monoamine oxidase B |
| chr7_-_150777874 | 0.01 |
ENST00000540185.1 |
FASTK |
Fas-activated serine/threonine kinase |
| chr19_+_16059818 | 0.01 |
ENST00000322107.1 |
OR10H4 |
olfactory receptor, family 10, subfamily H, member 4 |
| chr3_-_179322436 | 0.01 |
ENST00000392659.2 ENST00000476781.1 |
MRPL47 |
mitochondrial ribosomal protein L47 |
| chr16_+_28303804 | 0.01 |
ENST00000341901.4 |
SBK1 |
SH3 domain binding kinase 1 |
| chr1_+_9294822 | 0.01 |
ENST00000377403.2 |
H6PD |
hexose-6-phosphate dehydrogenase (glucose 1-dehydrogenase) |
| chr11_+_82868030 | 0.01 |
ENST00000298281.4 ENST00000530660.1 |
PCF11 |
PCF11 cleavage and polyadenylation factor subunit |
| chr20_-_44485835 | 0.01 |
ENST00000457981.1 ENST00000426915.1 ENST00000217455.4 |
ACOT8 |
acyl-CoA thioesterase 8 |
| chr12_+_72080253 | 0.01 |
ENST00000549735.1 |
TMEM19 |
transmembrane protein 19 |
| chr20_+_49126881 | 0.01 |
ENST00000371621.3 ENST00000541713.1 |
PTPN1 |
protein tyrosine phosphatase, non-receptor type 1 |
| chr1_-_33502441 | 0.00 |
ENST00000548033.1 ENST00000487289.1 ENST00000373449.2 ENST00000480134.1 ENST00000467905.1 |
AK2 |
adenylate kinase 2 |
| chr17_-_1928621 | 0.00 |
ENST00000331238.6 |
RTN4RL1 |
reticulon 4 receptor-like 1 |
| chr14_-_70655684 | 0.00 |
ENST00000356921.2 ENST00000381269.2 ENST00000357887.3 |
SLC8A3 |
solute carrier family 8 (sodium/calcium exchanger), member 3 |
| chr16_-_30537839 | 0.00 |
ENST00000380412.5 |
ZNF768 |
zinc finger protein 768 |
| chr2_+_162016804 | 0.00 |
ENST00000392749.2 ENST00000440506.1 |
TANK |
TRAF family member-associated NFKB activator |
| chr6_+_52051171 | 0.00 |
ENST00000340057.1 |
IL17A |
interleukin 17A |
| chr2_+_162016827 | 0.00 |
ENST00000429217.1 ENST00000406287.1 ENST00000402568.1 |
TANK |
TRAF family member-associated NFKB activator |
| chr1_-_154164534 | 0.00 |
ENST00000271850.7 ENST00000368530.2 |
TPM3 |
tropomyosin 3 |
| chr3_+_50316458 | 0.00 |
ENST00000316436.3 |
LSMEM2 |
leucine-rich single-pass membrane protein 2 |
| chr2_+_162016916 | 0.00 |
ENST00000405852.1 |
TANK |
TRAF family member-associated NFKB activator |
| chr19_+_12949251 | 0.00 |
ENST00000251472.4 |
MAST1 |
microtubule associated serine/threonine kinase 1 |
| chr6_-_107077347 | 0.00 |
ENST00000369063.3 ENST00000539449.1 |
RTN4IP1 |
reticulon 4 interacting protein 1 |
| chr11_+_76571911 | 0.00 |
ENST00000534206.1 ENST00000532485.1 ENST00000526597.1 ENST00000533873.1 ENST00000538157.1 |
ACER3 |
alkaline ceramidase 3 |
| chr11_-_64511789 | 0.00 |
ENST00000419843.1 ENST00000394430.1 |
RASGRP2 |
RAS guanyl releasing protein 2 (calcium and DAG-regulated) |
| chr21_-_46330545 | 0.00 |
ENST00000320216.6 ENST00000397852.1 |
ITGB2 |
integrin, beta 2 (complement component 3 receptor 3 and 4 subunit) |
| chr2_+_219472637 | 0.00 |
ENST00000417849.1 |
PLCD4 |
phospholipase C, delta 4 |
| chr15_-_75017711 | 0.00 |
ENST00000567032.1 ENST00000564596.1 ENST00000566503.1 ENST00000395049.4 ENST00000395048.2 ENST00000379727.3 |
CYP1A1 |
cytochrome P450, family 1, subfamily A, polypeptide 1 |
| chr17_-_42144949 | 0.00 |
ENST00000591247.1 |
LSM12 |
LSM12 homolog (S. cerevisiae) |
| chr12_-_57039739 | 0.00 |
ENST00000552959.1 ENST00000551020.1 ENST00000553007.2 ENST00000552919.1 ENST00000552104.1 ENST00000262030.3 |
ATP5B |
ATP synthase, H+ transporting, mitochondrial F1 complex, beta polypeptide |
| chr5_+_38258511 | 0.00 |
ENST00000354891.3 ENST00000322350.5 |
EGFLAM |
EGF-like, fibronectin type III and laminin G domains |
| chr17_-_46716647 | 0.00 |
ENST00000608940.1 |
RP11-357H14.17 |
RP11-357H14.17 |
| chr17_+_48046538 | 0.00 |
ENST00000240306.3 |
DLX4 |
distal-less homeobox 4 |
| chr1_+_165600436 | 0.00 |
ENST00000367888.4 ENST00000367885.1 ENST00000367884.2 |
MGST3 |
microsomal glutathione S-transferase 3 |
| chr5_-_126409159 | 0.00 |
ENST00000607731.1 ENST00000535381.1 ENST00000296662.5 ENST00000509733.3 |
C5orf63 |
chromosome 5 open reading frame 63 |
| chr7_-_43965937 | 0.00 |
ENST00000455877.1 ENST00000223341.7 ENST00000447717.3 ENST00000426198.1 |
URGCP |
upregulator of cell proliferation |
| chr21_-_43816052 | 0.00 |
ENST00000398405.1 |
TMPRSS3 |
transmembrane protease, serine 3 |
| chr7_+_120969045 | 0.00 |
ENST00000222462.2 |
WNT16 |
wingless-type MMTV integration site family, member 16 |
Gene Ontology Analysis
Gene overrepresentation in biological_process category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.0 | 0.1 | GO:1903300 | negative regulation of glucokinase activity(GO:0033132) negative regulation of hexokinase activity(GO:1903300) |
| 0.0 | 0.0 | GO:0007518 | myoblast fate determination(GO:0007518) |
| 0.0 | 0.0 | GO:0070681 | glutaminyl-tRNAGln biosynthesis via transamidation(GO:0070681) |
| 0.0 | 0.1 | GO:2001271 | negative regulation of cysteine-type endopeptidase activity involved in execution phase of apoptosis(GO:2001271) |
| 0.0 | 0.1 | GO:0021590 | cerebellum maturation(GO:0021590) cerebellar cortex maturation(GO:0021699) |
| 0.0 | 0.1 | GO:0030091 | protein repair(GO:0030091) |
Gene overrepresentation in cellular_component category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.0 | 0.0 | GO:0030956 | glutamyl-tRNA(Gln) amidotransferase complex(GO:0030956) |
| 0.0 | 0.0 | GO:0090571 | RNA polymerase II transcription repressor complex(GO:0090571) |
| 0.0 | 0.0 | GO:0005607 | laminin-2 complex(GO:0005607) |
| 0.0 | 0.0 | GO:0005968 | Rab-protein geranylgeranyltransferase complex(GO:0005968) |
Gene overrepresentation in molecular_function category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.0 | 0.1 | GO:0033743 | peptide-methionine (R)-S-oxide reductase activity(GO:0033743) |
| 0.0 | 0.0 | GO:0050613 | delta14-sterol reductase activity(GO:0050613) |
| 0.0 | 0.0 | GO:0050567 | glutaminyl-tRNA synthase (glutamine-hydrolyzing) activity(GO:0050567) |
| 0.0 | 0.0 | GO:0000026 | alpha-1,2-mannosyltransferase activity(GO:0000026) |