Project
Epithelial-Mesenchymal Transition, human (Scheel, 2011)
Navigation
Downloads
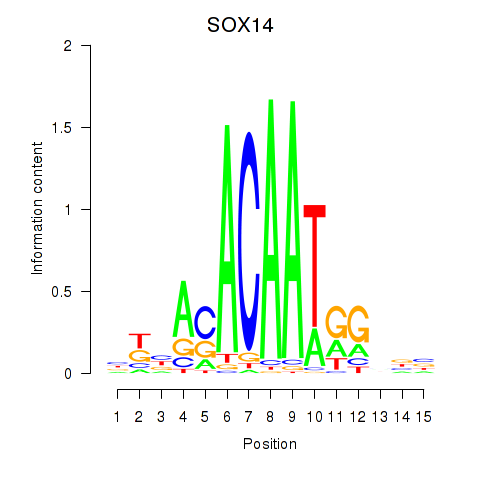
Results for SOX14
Z-value: 0.35
Transcription factors associated with SOX14
| Gene Symbol | Gene ID | Gene Info |
|---|---|---|
|
SOX14
|
ENSG00000168875.1 | SOX14 |
Activity-expression correlation:
| Gene | Promoter | Pearson corr. coef. | P-value | Plot |
|---|---|---|---|---|
| SOX14 | hg19_v2_chr3_+_137483579_137483579 | -0.41 | 3.1e-01 | Click! |
Activity profile of SOX14 motif
Sorted Z-values of SOX14 motif
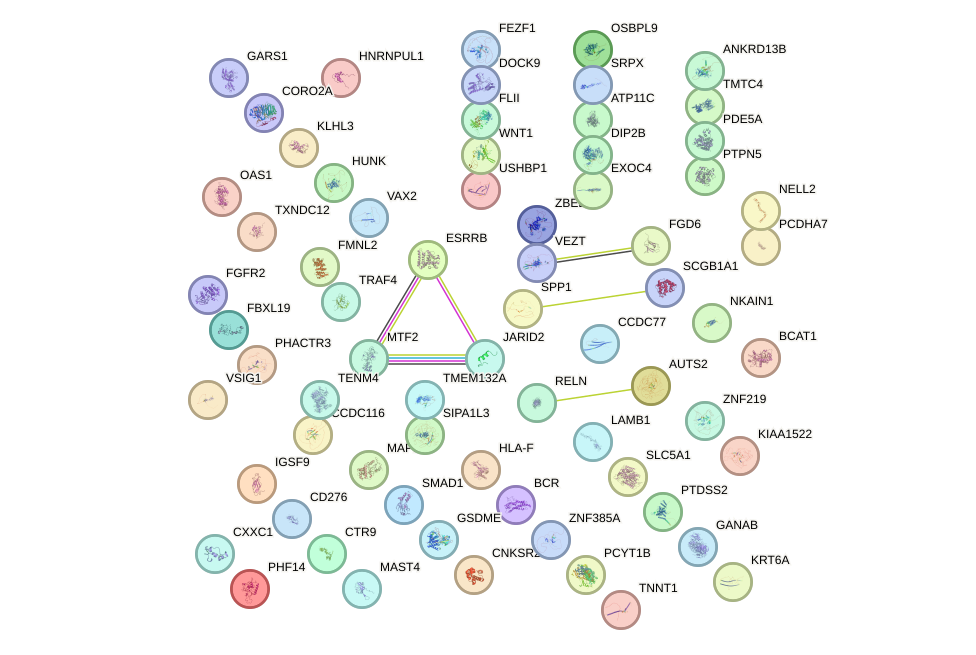
Network of associatons between targets according to the STRING database.
First level regulatory network of SOX14
| Promoter | Score | Transcript | Gene | Gene Info |
|---|---|---|---|---|
| chr12_-_52887034 | 0.49 |
ENST00000330722.6 |
KRT6A |
keratin 6A |
| chr1_+_209602156 | 0.47 |
ENST00000429156.1 ENST00000366437.3 ENST00000603283.1 ENST00000431096.1 |
MIR205HG |
MIR205 host gene (non-protein coding) |
| chr20_+_58179582 | 0.33 |
ENST00000371015.1 ENST00000395639.4 |
PHACTR3 |
phosphatase and actin regulator 3 |
| chr5_+_140213815 | 0.29 |
ENST00000525929.1 ENST00000378125.3 |
PCDHA7 |
protocadherin alpha 7 |
| chr1_-_159915386 | 0.28 |
ENST00000361509.3 ENST00000368094.1 |
IGSF9 |
immunoglobulin superfamily, member 9 |
| chr7_+_69064300 | 0.28 |
ENST00000342771.4 |
AUTS2 |
autism susceptibility candidate 2 |
| chr6_+_30848557 | 0.27 |
ENST00000460944.2 ENST00000324771.8 |
DDR1 |
discoidin domain receptor tyrosine kinase 1 |
| chr12_-_95611149 | 0.27 |
ENST00000549499.1 ENST00000343958.4 ENST00000546711.1 |
FGD6 |
FYVE, RhoGEF and PH domain containing 6 |
| chr17_-_39677971 | 0.26 |
ENST00000393976.2 |
KRT15 |
keratin 15 |
| chr5_+_66124590 | 0.26 |
ENST00000490016.2 ENST00000403666.1 ENST00000450827.1 |
MAST4 |
microtubule associated serine/threonine kinase family member 4 |
| chr6_+_138188551 | 0.21 |
ENST00000237289.4 ENST00000433680.1 |
TNFAIP3 |
tumor necrosis factor, alpha-induced protein 3 |
| chr11_+_60691924 | 0.17 |
ENST00000544065.1 ENST00000453848.2 ENST00000005286.4 |
TMEM132A |
transmembrane protein 132A |
| chr19_-_55653259 | 0.16 |
ENST00000593194.1 |
TNNT1 |
troponin T type 1 (skeletal, slow) |
| chr11_-_79151695 | 0.16 |
ENST00000278550.7 |
TENM4 |
teneurin transmembrane protein 4 |
| chr10_-_123274693 | 0.15 |
ENST00000429361.1 |
FGFR2 |
fibroblast growth factor receptor 2 |
| chr16_+_30934376 | 0.13 |
ENST00000562798.1 ENST00000471231.2 |
FBXL19 |
F-box and leucine-rich repeat protein 19 |
| chr15_+_73976545 | 0.12 |
ENST00000318443.5 ENST00000537340.2 ENST00000318424.5 |
CD276 |
CD276 molecule |
| chr11_-_44971702 | 0.12 |
ENST00000533940.1 ENST00000533937.1 |
TP53I11 |
tumor protein p53 inducible protein 11 |
| chr1_+_156119466 | 0.11 |
ENST00000414683.1 |
SEMA4A |
sema domain, immunoglobulin domain (Ig), transmembrane domain (TM) and short cytoplasmic domain, (semaphorin) 4A |
| chr17_-_8113542 | 0.11 |
ENST00000578549.1 ENST00000535053.1 ENST00000582368.1 |
AURKB |
aurora kinase B |
| chrX_-_24690771 | 0.11 |
ENST00000379145.1 |
PCYT1B |
phosphate cytidylyltransferase 1, choline, beta |
| chr14_-_35344093 | 0.11 |
ENST00000382422.2 |
BAZ1A |
bromodomain adjacent to zinc finger domain, 1A |
| chr12_+_95611516 | 0.11 |
ENST00000436874.1 |
VEZT |
vezatin, adherens junctions transmembrane protein |
| chr9_-_100954910 | 0.10 |
ENST00000375077.4 |
CORO2A |
coronin, actin binding protein, 2A |
| chr1_+_156119798 | 0.10 |
ENST00000355014.2 |
SEMA4A |
sema domain, immunoglobulin domain (Ig), transmembrane domain (TM) and short cytoplasmic domain, (semaphorin) 4A |
| chr4_-_103266626 | 0.10 |
ENST00000356736.4 |
SLC39A8 |
solute carrier family 39 (zinc transporter), member 8 |
| chr11_+_128563652 | 0.09 |
ENST00000527786.2 |
FLI1 |
Fli-1 proto-oncogene, ETS transcription factor |
| chrX_+_107288239 | 0.09 |
ENST00000217957.5 |
VSIG1 |
V-set and immunoglobulin domain containing 1 |
| chr12_+_95611569 | 0.09 |
ENST00000261219.6 ENST00000551472.1 ENST00000552821.1 |
VEZT |
vezatin, adherens junctions transmembrane protein |
| chr14_+_24641062 | 0.09 |
ENST00000311457.3 ENST00000557806.1 ENST00000559919.1 |
REC8 |
REC8 meiotic recombination protein |
| chrX_-_138914394 | 0.09 |
ENST00000327569.3 ENST00000361648.2 ENST00000370543.1 ENST00000359686.2 |
ATP11C |
ATPase, class VI, type 11C |
| chr4_+_88896819 | 0.09 |
ENST00000237623.7 ENST00000395080.3 ENST00000508233.1 ENST00000360804.4 |
SPP1 |
secreted phosphoprotein 1 |
| chr11_+_128634589 | 0.08 |
ENST00000281428.8 |
FLI1 |
Fli-1 proto-oncogene, ETS transcription factor |
| chr12_+_113354341 | 0.08 |
ENST00000553152.1 |
OAS1 |
2'-5'-oligoadenylate synthetase 1, 40/46kDa |
| chr2_+_153191706 | 0.08 |
ENST00000288670.9 |
FMNL2 |
formin-like 2 |
| chr22_+_32439019 | 0.08 |
ENST00000266088.4 |
SLC5A1 |
solute carrier family 5 (sodium/glucose cotransporter), member 1 |
| chr14_+_76776957 | 0.08 |
ENST00000512784.1 |
ESRRB |
estrogen-related receptor beta |
| chr11_+_128563948 | 0.08 |
ENST00000534087.2 |
FLI1 |
Fli-1 proto-oncogene, ETS transcription factor |
| chr19_+_38397839 | 0.08 |
ENST00000222345.6 |
SIPA1L3 |
signal-induced proliferation-associated 1 like 3 |
| chr10_+_123922941 | 0.08 |
ENST00000360561.3 |
TACC2 |
transforming, acidic coiled-coil containing protein 2 |
| chr10_+_123923105 | 0.08 |
ENST00000368999.1 |
TACC2 |
transforming, acidic coiled-coil containing protein 2 |
| chr7_-_121944491 | 0.08 |
ENST00000331178.4 ENST00000427185.2 ENST00000442488.2 |
FEZF1 |
FEZ family zinc finger 1 |
| chr10_+_123923205 | 0.08 |
ENST00000369004.3 ENST00000260733.3 |
TACC2 |
transforming, acidic coiled-coil containing protein 2 |
| chr1_-_161014731 | 0.08 |
ENST00000368020.1 |
USF1 |
upstream transcription factor 1 |
| chrX_+_21392873 | 0.07 |
ENST00000379510.3 |
CNKSR2 |
connector enhancer of kinase suppressor of Ras 2 |
| chr19_+_41768561 | 0.07 |
ENST00000599719.1 ENST00000601309.1 |
HNRNPUL1 |
heterogeneous nuclear ribonucleoprotein U-like 1 |
| chr12_-_54778471 | 0.07 |
ENST00000550120.1 ENST00000394313.2 ENST00000547210.1 |
ZNF385A |
zinc finger protein 385A |
| chrX_-_38080077 | 0.07 |
ENST00000378533.3 ENST00000544439.1 ENST00000432886.2 ENST00000538295.1 |
SRPX |
sushi-repeat containing protein, X-linked |
| chr5_-_111754948 | 0.07 |
ENST00000261486.5 |
EPB41L4A |
erythrocyte membrane protein band 4.1 like 4A |
| chr1_+_33231268 | 0.07 |
ENST00000373480.1 |
KIAA1522 |
KIAA1522 |
| chr11_+_76494253 | 0.07 |
ENST00000333090.4 |
TSKU |
tsukushi, small leucine rich proteoglycan |
| chr6_+_15401075 | 0.07 |
ENST00000541660.1 |
JARID2 |
jumonji, AT rich interactive domain 2 |
| chr12_+_50898881 | 0.07 |
ENST00000301180.5 |
DIP2B |
DIP2 disco-interacting protein 2 homolog B (Drosophila) |
| chr13_-_99630233 | 0.07 |
ENST00000376460.1 ENST00000442173.1 |
DOCK9 |
dedicator of cytokinesis 9 |
| chr4_-_120548779 | 0.07 |
ENST00000264805.5 |
PDE5A |
phosphodiesterase 5A, cGMP-specific |
| chr12_-_25055177 | 0.07 |
ENST00000538118.1 |
BCAT1 |
branched chain amino-acid transaminase 1, cytosolic |
| chr3_+_49840685 | 0.06 |
ENST00000333323.4 |
FAM212A |
family with sequence similarity 212, member A |
| chrX_+_107288197 | 0.06 |
ENST00000415430.3 |
VSIG1 |
V-set and immunoglobulin domain containing 1 |
| chr17_-_46716647 | 0.06 |
ENST00000608940.1 |
RP11-357H14.17 |
RP11-357H14.17 |
| chr8_-_80993010 | 0.06 |
ENST00000537855.1 ENST00000520527.1 ENST00000517427.1 ENST00000448733.2 ENST00000379097.3 |
TPD52 |
tumor protein D52 |
| chr11_+_63753883 | 0.06 |
ENST00000538426.1 ENST00000543004.1 |
OTUB1 |
OTU domain, ubiquitin aldehyde binding 1 |
| chr11_-_18813110 | 0.06 |
ENST00000396168.1 |
PTPN5 |
protein tyrosine phosphatase, non-receptor type 5 (striatum-enriched) |
| chr1_+_169075554 | 0.06 |
ENST00000367815.4 |
ATP1B1 |
ATPase, Na+/K+ transporting, beta 1 polypeptide |
| chr2_+_102314161 | 0.05 |
ENST00000425019.1 |
MAP4K4 |
mitogen-activated protein kinase kinase kinase kinase 4 |
| chr8_+_144295067 | 0.05 |
ENST00000330824.2 |
GPIHBP1 |
glycosylphosphatidylinositol anchored high density lipoprotein binding protein 1 |
| chr14_-_21567009 | 0.05 |
ENST00000556174.1 ENST00000554478.1 ENST00000553980.1 ENST00000421093.2 |
ZNF219 |
zinc finger protein 219 |
| chr17_+_27920486 | 0.05 |
ENST00000394859.3 |
ANKRD13B |
ankyrin repeat domain 13B |
| chr2_+_71127699 | 0.05 |
ENST00000234392.2 |
VAX2 |
ventral anterior homeobox 2 |
| chr15_+_73976715 | 0.05 |
ENST00000558689.1 ENST00000560786.2 ENST00000561213.1 ENST00000563584.1 ENST00000561416.1 |
CD276 |
CD276 molecule |
| chr18_+_42260861 | 0.05 |
ENST00000282030.5 |
SETBP1 |
SET binding protein 1 |
| chr1_+_93544791 | 0.05 |
ENST00000545708.1 ENST00000540243.1 ENST00000370298.4 |
MTF2 |
metal response element binding transcription factor 2 |
| chr22_+_50247449 | 0.05 |
ENST00000216268.5 |
ZBED4 |
zinc finger, BED-type containing 4 |
| chr1_-_225616515 | 0.05 |
ENST00000338179.2 ENST00000425080.1 |
LBR |
lamin B receptor |
| chr15_+_52311398 | 0.05 |
ENST00000261845.5 |
MAPK6 |
mitogen-activated protein kinase 6 |
| chr13_+_111767650 | 0.04 |
ENST00000449979.1 ENST00000370623.3 |
ARHGEF7 |
Rho guanine nucleotide exchange factor (GEF) 7 |
| chr10_+_81466084 | 0.04 |
ENST00000342531.2 |
NUTM2B |
NUT family member 2B |
| chr22_+_23522552 | 0.04 |
ENST00000359540.3 ENST00000398512.5 |
BCR |
breakpoint cluster region |
| chr11_-_18813353 | 0.04 |
ENST00000358540.2 ENST00000396171.4 ENST00000396167.2 |
PTPN5 |
protein tyrosine phosphatase, non-receptor type 5 (striatum-enriched) |
| chr1_-_52521831 | 0.04 |
ENST00000371626.4 |
TXNDC12 |
thioredoxin domain containing 12 (endoplasmic reticulum) |
| chr19_-_17375527 | 0.04 |
ENST00000431146.2 ENST00000594190.1 |
USHBP1 |
Usher syndrome 1C binding protein 1 |
| chr4_+_146403912 | 0.04 |
ENST00000507367.1 ENST00000394092.2 ENST00000515385.1 |
SMAD1 |
SMAD family member 1 |
| chr12_-_45270151 | 0.04 |
ENST00000429094.2 |
NELL2 |
NEL-like 2 (chicken) |
| chr6_+_29691198 | 0.04 |
ENST00000440587.2 ENST00000434407.2 |
HLA-F |
major histocompatibility complex, class I, F |
| chr12_-_45270077 | 0.04 |
ENST00000551601.1 ENST00000549027.1 ENST00000452445.2 |
NELL2 |
NEL-like 2 (chicken) |
| chr21_+_33245548 | 0.04 |
ENST00000270112.2 |
HUNK |
hormonally up-regulated Neu-associated kinase |
| chr11_+_450255 | 0.04 |
ENST00000308020.5 |
PTDSS2 |
phosphatidylserine synthase 2 |
| chr9_+_112403059 | 0.04 |
ENST00000374531.2 |
PALM2 |
paralemmin 2 |
| chr17_-_8113886 | 0.04 |
ENST00000577833.1 ENST00000534871.1 ENST00000583915.1 ENST00000316199.6 ENST00000581511.1 ENST00000585124.1 |
AURKB |
aurora kinase B |
| chr17_+_27071002 | 0.04 |
ENST00000262395.5 ENST00000422344.1 ENST00000444415.3 ENST00000262396.6 |
TRAF4 |
TNF receptor-associated factor 4 |
| chr12_+_49372251 | 0.04 |
ENST00000293549.3 |
WNT1 |
wingless-type MMTV integration site family, member 1 |
| chr13_-_101327028 | 0.04 |
ENST00000328767.5 ENST00000342624.5 ENST00000376234.3 ENST00000423847.1 |
TMTC4 |
transmembrane and tetratricopeptide repeat containing 4 |
| chr1_+_93544821 | 0.04 |
ENST00000370303.4 |
MTF2 |
metal response element binding transcription factor 2 |
| chr7_+_11013491 | 0.04 |
ENST00000403050.3 ENST00000445996.2 |
PHF14 |
PHD finger protein 14 |
| chr7_-_24797032 | 0.04 |
ENST00000409970.1 ENST00000409775.3 |
DFNA5 |
deafness, autosomal dominant 5 |
| chr14_+_103589789 | 0.04 |
ENST00000558056.1 ENST00000560869.1 |
TNFAIP2 |
tumor necrosis factor, alpha-induced protein 2 |
| chr1_+_66458072 | 0.04 |
ENST00000423207.2 |
PDE4B |
phosphodiesterase 4B, cAMP-specific |
| chr6_+_170190417 | 0.04 |
ENST00000420557.2 |
LINC00574 |
long intergenic non-protein coding RNA 574 |
| chr1_-_201123586 | 0.04 |
ENST00000414605.2 ENST00000367334.5 ENST00000367332.1 |
TMEM9 |
transmembrane protein 9 |
| chr7_-_103629963 | 0.04 |
ENST00000428762.1 ENST00000343529.5 ENST00000424685.2 |
RELN |
reelin |
| chr12_+_498500 | 0.03 |
ENST00000540180.1 ENST00000422000.1 ENST00000535052.1 |
CCDC77 |
coiled-coil domain containing 77 |
| chr11_+_62186498 | 0.03 |
ENST00000278282.2 |
SCGB1A1 |
secretoglobin, family 1A, member 1 (uteroglobin) |
| chr11_-_62414070 | 0.03 |
ENST00000540933.1 ENST00000346178.4 ENST00000356638.3 ENST00000534779.1 ENST00000525994.1 |
GANAB |
glucosidase, alpha; neutral AB |
| chr1_-_31712401 | 0.03 |
ENST00000373736.2 |
NKAIN1 |
Na+/K+ transporting ATPase interacting 1 |
| chr6_+_32132360 | 0.03 |
ENST00000333845.6 ENST00000395512.1 ENST00000432129.1 |
EGFL8 |
EGF-like-domain, multiple 8 |
| chr9_-_116062045 | 0.03 |
ENST00000478815.1 |
RNF183 |
ring finger protein 183 |
| chr5_-_137071689 | 0.03 |
ENST00000505853.1 |
KLHL3 |
kelch-like family member 3 |
| chr11_+_10772534 | 0.03 |
ENST00000361367.2 |
CTR9 |
CTR9, Paf1/RNA polymerase II complex component |
| chr12_+_69633407 | 0.03 |
ENST00000551516.1 |
CPSF6 |
cleavage and polyadenylation specific factor 6, 68kDa |
| chr1_+_52082751 | 0.03 |
ENST00000447887.1 ENST00000435686.2 ENST00000428468.1 ENST00000453295.1 |
OSBPL9 |
oxysterol binding protein-like 9 |
| chr6_+_29691056 | 0.03 |
ENST00000414333.1 ENST00000334668.4 ENST00000259951.7 |
HLA-F |
major histocompatibility complex, class I, F |
| chr7_-_107643674 | 0.03 |
ENST00000222399.6 |
LAMB1 |
laminin, beta 1 |
| chr22_+_21987005 | 0.03 |
ENST00000607942.1 ENST00000425975.1 ENST00000292779.3 |
CCDC116 |
coiled-coil domain containing 116 |
| chr6_-_31509714 | 0.03 |
ENST00000456662.1 ENST00000431908.1 ENST00000456976.1 ENST00000428450.1 ENST00000453105.2 ENST00000418897.1 ENST00000415382.2 ENST00000449074.2 ENST00000419020.1 ENST00000428098.1 |
DDX39B |
DEAD (Asp-Glu-Ala-Asp) box polypeptide 39B |
| chr1_-_6546001 | 0.03 |
ENST00000400913.1 |
PLEKHG5 |
pleckstrin homology domain containing, family G (with RhoGef domain) member 5 |
| chr1_-_201123546 | 0.03 |
ENST00000435310.1 ENST00000485839.2 ENST00000367330.1 |
TMEM9 |
transmembrane protein 9 |
| chr5_-_41261540 | 0.03 |
ENST00000263413.3 |
C6 |
complement component 6 |
| chr11_-_63684316 | 0.03 |
ENST00000301459.4 |
RCOR2 |
REST corepressor 2 |
| chr1_+_23345930 | 0.03 |
ENST00000356634.3 |
KDM1A |
lysine (K)-specific demethylase 1A |
| chr19_+_58694396 | 0.03 |
ENST00000326804.4 ENST00000345813.3 ENST00000424679.2 |
ZNF274 |
zinc finger protein 274 |
| chr8_+_133787586 | 0.03 |
ENST00000395379.1 ENST00000395386.2 ENST00000337920.4 |
PHF20L1 |
PHD finger protein 20-like 1 |
| chr1_+_23345943 | 0.03 |
ENST00000400181.4 ENST00000542151.1 |
KDM1A |
lysine (K)-specific demethylase 1A |
| chr8_-_110620284 | 0.03 |
ENST00000529690.1 |
SYBU |
syntabulin (syntaxin-interacting) |
| chr4_-_121993673 | 0.03 |
ENST00000379692.4 |
NDNF |
neuron-derived neurotrophic factor |
| chrX_+_15525426 | 0.02 |
ENST00000342014.6 |
BMX |
BMX non-receptor tyrosine kinase |
| chr20_-_44991813 | 0.02 |
ENST00000372227.1 |
SLC35C2 |
solute carrier family 35 (GDP-fucose transporter), member C2 |
| chr12_+_69633372 | 0.02 |
ENST00000456847.3 ENST00000266679.8 |
CPSF6 |
cleavage and polyadenylation specific factor 6, 68kDa |
| chr8_+_24151553 | 0.02 |
ENST00000265769.4 ENST00000540823.1 ENST00000397649.3 |
ADAM28 |
ADAM metallopeptidase domain 28 |
| chr10_+_18429606 | 0.02 |
ENST00000324631.7 ENST00000352115.6 ENST00000377328.1 |
CACNB2 |
calcium channel, voltage-dependent, beta 2 subunit |
| chr21_-_32253874 | 0.02 |
ENST00000332378.4 |
KRTAP11-1 |
keratin associated protein 11-1 |
| chr13_-_24007815 | 0.02 |
ENST00000382298.3 |
SACS |
spastic ataxia of Charlevoix-Saguenay (sacsin) |
| chr5_-_137071756 | 0.02 |
ENST00000394937.3 ENST00000309755.4 |
KLHL3 |
kelch-like family member 3 |
| chr17_-_42441204 | 0.02 |
ENST00000293443.7 |
FAM171A2 |
family with sequence similarity 171, member A2 |
| chr17_+_25621102 | 0.02 |
ENST00000581440.1 ENST00000262394.2 ENST00000583742.1 ENST00000579733.1 ENST00000583193.1 ENST00000581185.1 ENST00000427287.2 ENST00000348811.2 |
WSB1 |
WD repeat and SOCS box containing 1 |
| chr19_-_58514129 | 0.02 |
ENST00000552184.1 ENST00000546715.1 ENST00000536132.1 ENST00000547828.1 ENST00000547121.1 ENST00000551380.1 |
ZNF606 |
zinc finger protein 606 |
| chr8_+_104427581 | 0.02 |
ENST00000521716.1 ENST00000521971.1 ENST00000519682.1 |
DCAF13 |
DDB1 and CUL4 associated factor 13 |
| chr17_+_68100989 | 0.02 |
ENST00000585558.1 ENST00000392670.1 |
KCNJ16 |
potassium inwardly-rectifying channel, subfamily J, member 16 |
| chr12_-_122907091 | 0.02 |
ENST00000358808.2 ENST00000361654.4 ENST00000539080.1 ENST00000537178.1 |
CLIP1 |
CAP-GLY domain containing linker protein 1 |
| chr2_-_55276320 | 0.02 |
ENST00000357376.3 |
RTN4 |
reticulon 4 |
| chr19_+_10765003 | 0.02 |
ENST00000407004.3 ENST00000589998.1 ENST00000589600.1 |
ILF3 |
interleukin enhancer binding factor 3, 90kDa |
| chr2_+_228337079 | 0.02 |
ENST00000409315.1 ENST00000373671.3 ENST00000409171.1 |
AGFG1 |
ArfGAP with FG repeats 1 |
| chr15_-_58571445 | 0.02 |
ENST00000558231.1 |
ALDH1A2 |
aldehyde dehydrogenase 1 family, member A2 |
| chr9_-_116061476 | 0.02 |
ENST00000441031.3 |
RNF183 |
ring finger protein 183 |
| chr20_-_43150601 | 0.02 |
ENST00000541235.1 ENST00000255175.1 ENST00000342374.4 |
SERINC3 |
serine incorporator 3 |
| chr11_+_66247880 | 0.02 |
ENST00000360510.2 ENST00000453114.1 ENST00000541961.1 ENST00000532019.1 ENST00000526515.1 ENST00000530165.1 ENST00000533725.1 |
DPP3 |
dipeptidyl-peptidase 3 |
| chr19_-_17375541 | 0.02 |
ENST00000252597.3 |
USHBP1 |
Usher syndrome 1C binding protein 1 |
| chr16_+_77233294 | 0.02 |
ENST00000378644.4 |
SYCE1L |
synaptonemal complex central element protein 1-like |
| chr5_-_179050660 | 0.02 |
ENST00000519056.1 ENST00000506721.1 ENST00000503105.1 ENST00000504348.1 ENST00000508103.1 ENST00000510431.1 ENST00000515158.1 ENST00000393432.4 ENST00000442819.2 |
HNRNPH1 |
heterogeneous nuclear ribonucleoprotein H1 (H) |
| chrX_+_140984021 | 0.02 |
ENST00000409007.1 |
MAGEC3 |
melanoma antigen family C, 3 |
| chr10_+_70480963 | 0.02 |
ENST00000265872.6 ENST00000535016.1 ENST00000538031.1 ENST00000543719.1 ENST00000539539.1 ENST00000543225.1 ENST00000536012.1 ENST00000494903.2 |
CCAR1 |
cell division cycle and apoptosis regulator 1 |
| chr19_-_51891209 | 0.02 |
ENST00000221973.3 ENST00000596399.1 |
LIM2 |
lens intrinsic membrane protein 2, 19kDa |
| chr11_-_130184470 | 0.02 |
ENST00000357899.4 ENST00000397753.1 |
ZBTB44 |
zinc finger and BTB domain containing 44 |
| chr5_-_180018540 | 0.02 |
ENST00000292641.3 |
SCGB3A1 |
secretoglobin, family 3A, member 1 |
| chr2_-_209028300 | 0.02 |
ENST00000304502.4 |
CRYGA |
crystallin, gamma A |
| chr3_+_51851612 | 0.02 |
ENST00000456080.1 |
IQCF3 |
IQ motif containing F3 |
| chr6_-_127840336 | 0.02 |
ENST00000525778.1 |
SOGA3 |
SOGA family member 3 |
| chr6_+_122720681 | 0.02 |
ENST00000368455.4 ENST00000452194.1 |
HSF2 |
heat shock transcription factor 2 |
| chr1_-_203155868 | 0.01 |
ENST00000255409.3 |
CHI3L1 |
chitinase 3-like 1 (cartilage glycoprotein-39) |
| chr4_+_113152978 | 0.01 |
ENST00000309703.6 |
AP1AR |
adaptor-related protein complex 1 associated regulatory protein |
| chr12_-_57400227 | 0.01 |
ENST00000300101.2 |
ZBTB39 |
zinc finger and BTB domain containing 39 |
| chr8_+_24151620 | 0.01 |
ENST00000437154.2 |
ADAM28 |
ADAM metallopeptidase domain 28 |
| chr1_+_32930647 | 0.01 |
ENST00000609129.1 |
ZBTB8B |
zinc finger and BTB domain containing 8B |
| chr11_-_47574664 | 0.01 |
ENST00000310513.5 ENST00000531165.1 |
CELF1 |
CUGBP, Elav-like family member 1 |
| chr17_-_37353950 | 0.01 |
ENST00000394310.3 ENST00000394303.3 ENST00000344140.5 |
CACNB1 |
calcium channel, voltage-dependent, beta 1 subunit |
| chr11_-_130184555 | 0.01 |
ENST00000525842.1 |
ZBTB44 |
zinc finger and BTB domain containing 44 |
| chr7_-_149470540 | 0.01 |
ENST00000302017.3 |
ZNF467 |
zinc finger protein 467 |
| chr2_-_209010874 | 0.01 |
ENST00000260988.4 |
CRYGB |
crystallin, gamma B |
| chr7_-_149470297 | 0.01 |
ENST00000484747.1 |
ZNF467 |
zinc finger protein 467 |
| chr6_-_127840021 | 0.01 |
ENST00000465909.2 |
SOGA3 |
SOGA family member 3 |
| chr17_+_4487294 | 0.01 |
ENST00000338859.4 |
SMTNL2 |
smoothelin-like 2 |
| chr20_-_35492048 | 0.01 |
ENST00000237536.4 |
SOGA1 |
suppressor of glucose, autophagy associated 1 |
| chr17_+_39411636 | 0.01 |
ENST00000394008.1 |
KRTAP9-9 |
keratin associated protein 9-9 |
| chr20_-_17662705 | 0.01 |
ENST00000455029.2 |
RRBP1 |
ribosome binding protein 1 |
| chr17_+_7608511 | 0.01 |
ENST00000226091.2 |
EFNB3 |
ephrin-B3 |
| chr6_+_30687978 | 0.01 |
ENST00000327892.8 ENST00000435534.1 |
TUBB |
tubulin, beta class I |
| chrX_-_2418936 | 0.01 |
ENST00000461691.1 ENST00000381223.4 ENST00000381222.2 ENST00000412516.2 ENST00000334651.5 |
ZBED1 DHRSX |
zinc finger, BED-type containing 1 dehydrogenase/reductase (SDR family) X-linked |
| chr16_-_52580920 | 0.01 |
ENST00000219746.9 |
TOX3 |
TOX high mobility group box family member 3 |
| chr5_+_79615790 | 0.01 |
ENST00000296739.4 |
SPZ1 |
spermatogenic leucine zipper 1 |
| chr2_+_115219171 | 0.01 |
ENST00000409163.1 |
DPP10 |
dipeptidyl-peptidase 10 (non-functional) |
| chr17_-_42100474 | 0.01 |
ENST00000585950.1 ENST00000592127.1 ENST00000589334.1 |
TMEM101 |
transmembrane protein 101 |
| chr11_+_63754294 | 0.01 |
ENST00000543988.1 |
OTUB1 |
OTU domain, ubiquitin aldehyde binding 1 |
| chr14_-_51297197 | 0.01 |
ENST00000382043.4 |
NIN |
ninein (GSK3B interacting protein) |
| chr15_+_71184931 | 0.01 |
ENST00000560369.1 ENST00000260382.5 |
LRRC49 |
leucine rich repeat containing 49 |
| chr2_-_157198860 | 0.01 |
ENST00000409572.1 |
NR4A2 |
nuclear receptor subfamily 4, group A, member 2 |
| chr7_+_30067973 | 0.01 |
ENST00000258679.7 ENST00000449726.1 ENST00000396257.2 ENST00000396259.1 |
PLEKHA8 |
pleckstrin homology domain containing, family A (phosphoinositide binding specific) member 8 |
| chr11_-_62476965 | 0.01 |
ENST00000405837.1 ENST00000531524.1 |
BSCL2 |
Berardinelli-Seip congenital lipodystrophy 2 (seipin) |
| chr2_+_223536428 | 0.01 |
ENST00000446656.3 |
MOGAT1 |
monoacylglycerol O-acyltransferase 1 |
| chr22_-_19512893 | 0.01 |
ENST00000403084.1 ENST00000413119.2 |
CLDN5 |
claudin 5 |
| chr12_+_48722763 | 0.01 |
ENST00000335017.1 |
H1FNT |
H1 histone family, member N, testis-specific |
| chr14_-_106478603 | 0.01 |
ENST00000390596.2 |
IGHV4-4 |
immunoglobulin heavy variable 4-4 |
| chr1_+_14075903 | 0.01 |
ENST00000343137.4 ENST00000503842.1 ENST00000407521.3 ENST00000505823.1 |
PRDM2 |
PR domain containing 2, with ZNF domain |
| chr10_+_30722866 | 0.00 |
ENST00000263056.1 |
MAP3K8 |
mitogen-activated protein kinase kinase kinase 8 |
| chr3_+_182983090 | 0.00 |
ENST00000465010.1 |
B3GNT5 |
UDP-GlcNAc:betaGal beta-1,3-N-acetylglucosaminyltransferase 5 |
| chr17_-_39041479 | 0.00 |
ENST00000167588.3 |
KRT20 |
keratin 20 |
| chr10_-_33247124 | 0.00 |
ENST00000414670.1 ENST00000302278.3 ENST00000374956.4 ENST00000488494.1 ENST00000417122.2 ENST00000474568.1 |
ITGB1 |
integrin, beta 1 (fibronectin receptor, beta polypeptide, antigen CD29 includes MDF2, MSK12) |
| chr10_-_32217717 | 0.00 |
ENST00000396144.4 ENST00000375245.4 ENST00000344936.2 ENST00000375250.5 |
ARHGAP12 |
Rho GTPase activating protein 12 |
| chr2_-_9771075 | 0.00 |
ENST00000446619.1 ENST00000238081.3 |
YWHAQ |
tyrosine 3-monooxygenase/tryptophan 5-monooxygenase activation protein, theta |
| chr7_+_30068260 | 0.00 |
ENST00000440706.2 |
PLEKHA8 |
pleckstrin homology domain containing, family A (phosphoinositide binding specific) member 8 |
| chr5_-_146435572 | 0.00 |
ENST00000394414.1 |
PPP2R2B |
protein phosphatase 2, regulatory subunit B, beta |
| chrX_+_103357202 | 0.00 |
ENST00000537356.3 |
ZCCHC18 |
zinc finger, CCHC domain containing 18 |
| chr16_-_49890016 | 0.00 |
ENST00000563137.2 |
ZNF423 |
zinc finger protein 423 |
Gene Ontology Analysis
Gene overrepresentation in biological_process category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.1 | 0.5 | GO:0051710 | cytolysis by symbiont of host cells(GO:0001897) regulation of cytolysis in other organism(GO:0051710) |
| 0.1 | 0.3 | GO:0098582 | innate vocalization behavior(GO:0098582) |
| 0.1 | 0.2 | GO:0071947 | B-1 B cell homeostasis(GO:0001922) toll-like receptor 5 signaling pathway(GO:0034146) regulation of nucleotide-binding oligomerization domain containing 1 signaling pathway(GO:0070428) protein deubiquitination involved in ubiquitin-dependent protein catabolic process(GO:0071947) negative regulation of osteoclast proliferation(GO:0090291) |
| 0.1 | 0.2 | GO:0032289 | central nervous system myelin formation(GO:0032289) |
| 0.0 | 0.1 | GO:0035604 | fibroblast growth factor receptor signaling pathway involved in negative regulation of apoptotic process in bone marrow(GO:0035602) fibroblast growth factor receptor signaling pathway involved in hemopoiesis(GO:0035603) fibroblast growth factor receptor signaling pathway involved in positive regulation of cell proliferation in bone marrow(GO:0035604) |
| 0.0 | 0.3 | GO:0061302 | smooth muscle cell-matrix adhesion(GO:0061302) |
| 0.0 | 0.2 | GO:0043988 | histone H3-S28 phosphorylation(GO:0043988) |
| 0.0 | 0.1 | GO:2000866 | positive regulation of estrogen secretion(GO:2000863) positive regulation of estradiol secretion(GO:2000866) |
| 0.0 | 0.1 | GO:0000117 | regulation of transcription involved in G2/M transition of mitotic cell cycle(GO:0000117) |
| 0.0 | 0.1 | GO:0061073 | ciliary body morphogenesis(GO:0061073) |
| 0.0 | 0.1 | GO:0021797 | forebrain anterior/posterior pattern specification(GO:0021797) |
| 0.0 | 0.2 | GO:0031444 | slow-twitch skeletal muscle fiber contraction(GO:0031444) |
| 0.0 | 0.0 | GO:0021812 | neuronal-glial interaction involved in cerebral cortex radial glia guided migration(GO:0021812) |
| 0.0 | 0.1 | GO:0044861 | protein transport into plasma membrane raft(GO:0044861) |
| 0.0 | 0.1 | GO:1902164 | mRNA localization resulting in posttranscriptional regulation of gene expression(GO:0010609) regulation of DNA damage response, signal transduction by p53 class mediator resulting in transcription of p21 class mediator(GO:1902162) positive regulation of DNA damage response, signal transduction by p53 class mediator resulting in transcription of p21 class mediator(GO:1902164) |
| 0.0 | 0.0 | GO:0043314 | negative regulation of neutrophil degranulation(GO:0043314) |
| 0.0 | 0.1 | GO:0072156 | distal tubule morphogenesis(GO:0072156) |
| 0.0 | 0.2 | GO:0045078 | positive regulation of interferon-gamma biosynthetic process(GO:0045078) |
| 0.0 | 0.0 | GO:2000583 | regulation of platelet-derived growth factor receptor-alpha signaling pathway(GO:2000583) negative regulation of platelet-derived growth factor receptor-alpha signaling pathway(GO:2000584) |
| 0.0 | 0.1 | GO:0033184 | positive regulation of histone ubiquitination(GO:0033184) |
| 0.0 | 0.1 | GO:0000430 | regulation of transcription from RNA polymerase II promoter by glucose(GO:0000430) positive regulation of transcription from RNA polymerase II promoter by glucose(GO:0000432) |
| 0.0 | 0.1 | GO:0001951 | intestinal D-glucose absorption(GO:0001951) |
| 0.0 | 0.1 | GO:0009082 | branched-chain amino acid biosynthetic process(GO:0009082) leucine biosynthetic process(GO:0009098) valine biosynthetic process(GO:0009099) |
| 0.0 | 0.1 | GO:0010986 | positive regulation of lipoprotein particle clearance(GO:0010986) |
| 0.0 | 0.3 | GO:0035855 | megakaryocyte development(GO:0035855) |
Gene overrepresentation in cellular_component category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.0 | 0.2 | GO:0032133 | chromosome passenger complex(GO:0032133) |
| 0.0 | 0.1 | GO:0034991 | nuclear meiotic cohesin complex(GO:0034991) |
| 0.0 | 0.3 | GO:0060077 | inhibitory synapse(GO:0060077) |
| 0.0 | 0.2 | GO:0031010 | ISWI-type complex(GO:0031010) |
| 0.0 | 0.0 | GO:0005607 | laminin-2 complex(GO:0005607) |
Gene overrepresentation in molecular_function category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.1 | 0.3 | GO:0038062 | protein tyrosine kinase collagen receptor activity(GO:0038062) |
| 0.0 | 0.1 | GO:0004105 | choline-phosphate cytidylyltransferase activity(GO:0004105) |
| 0.0 | 0.1 | GO:0061752 | telomeric repeat-containing RNA binding(GO:0061752) |
| 0.0 | 0.2 | GO:0035174 | histone serine kinase activity(GO:0035174) |
| 0.0 | 0.0 | GO:0050613 | delta14-sterol reductase activity(GO:0050613) |
| 0.0 | 0.2 | GO:0061578 | Lys63-specific deubiquitinase activity(GO:0061578) |
| 0.0 | 0.1 | GO:0005007 | fibroblast growth factor-activated receptor activity(GO:0005007) |
| 0.0 | 0.2 | GO:0031014 | troponin T binding(GO:0031014) |
| 0.0 | 0.0 | GO:0019834 | phospholipase A2 inhibitor activity(GO:0019834) |
| 0.0 | 0.1 | GO:0004084 | branched-chain-amino-acid transaminase activity(GO:0004084) L-leucine transaminase activity(GO:0052654) L-valine transaminase activity(GO:0052655) L-isoleucine transaminase activity(GO:0052656) |
| 0.0 | 0.1 | GO:0046979 | TAP2 binding(GO:0046979) |