Project
Epithelial-Mesenchymal Transition, human (Scheel, 2011)
Navigation
Downloads
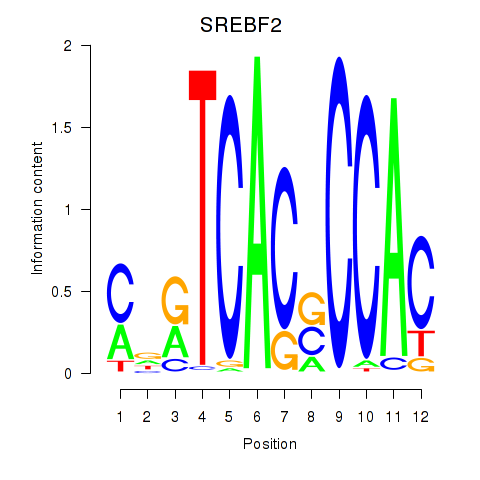
Results for SREBF2
Z-value: 0.45
Transcription factors associated with SREBF2
| Gene Symbol | Gene ID | Gene Info |
|---|---|---|
|
SREBF2
|
ENSG00000198911.7 | SREBF2 |
Activity-expression correlation:
| Gene | Promoter | Pearson corr. coef. | P-value | Plot |
|---|---|---|---|---|
| SREBF2 | hg19_v2_chr22_+_42229100_42229146 | -0.04 | 9.2e-01 | Click! |
Activity profile of SREBF2 motif
Sorted Z-values of SREBF2 motif
Network of associatons between targets according to the STRING database.
First level regulatory network of SREBF2
| Promoter | Score | Transcript | Gene | Gene Info |
|---|---|---|---|---|
| chr5_-_149535421 | 0.69 |
ENST00000261799.4 |
PDGFRB |
platelet-derived growth factor receptor, beta polypeptide |
| chr15_+_33010175 | 0.65 |
ENST00000300177.4 ENST00000560677.1 ENST00000560830.1 |
GREM1 |
gremlin 1, DAN family BMP antagonist |
| chr19_-_11308190 | 0.37 |
ENST00000586659.1 ENST00000592903.1 ENST00000589359.1 ENST00000588724.1 ENST00000432929.2 |
KANK2 |
KN motif and ankyrin repeat domains 2 |
| chr6_+_151561085 | 0.29 |
ENST00000402676.2 |
AKAP12 |
A kinase (PRKA) anchor protein 12 |
| chr20_+_33464238 | 0.25 |
ENST00000360596.2 |
ACSS2 |
acyl-CoA synthetase short-chain family member 2 |
| chr5_-_39425068 | 0.25 |
ENST00000515700.1 ENST00000339788.6 |
DAB2 |
Dab, mitogen-responsive phosphoprotein, homolog 2 (Drosophila) |
| chr20_+_33464407 | 0.22 |
ENST00000253382.5 |
ACSS2 |
acyl-CoA synthetase short-chain family member 2 |
| chr17_-_15168624 | 0.22 |
ENST00000312280.3 ENST00000494511.1 ENST00000580584.1 |
PMP22 |
peripheral myelin protein 22 |
| chr12_-_91505608 | 0.16 |
ENST00000266718.4 |
LUM |
lumican |
| chr8_+_26435359 | 0.15 |
ENST00000311151.5 |
DPYSL2 |
dihydropyrimidinase-like 2 |
| chr19_-_46285646 | 0.15 |
ENST00000458663.2 |
DMPK |
dystrophia myotonica-protein kinase |
| chr3_-_171528227 | 0.15 |
ENST00000356327.5 ENST00000342215.6 ENST00000340989.4 ENST00000351298.4 |
PLD1 |
phospholipase D1, phosphatidylcholine-specific |
| chr19_-_46285736 | 0.14 |
ENST00000291270.4 ENST00000447742.2 ENST00000354227.5 |
DMPK |
dystrophia myotonica-protein kinase |
| chr10_-_3827371 | 0.14 |
ENST00000469435.1 |
KLF6 |
Kruppel-like factor 6 |
| chr19_-_18392422 | 0.14 |
ENST00000252818.3 |
JUND |
jun D proto-oncogene |
| chr5_+_137774706 | 0.14 |
ENST00000378339.2 ENST00000254901.5 ENST00000506158.1 |
REEP2 |
receptor accessory protein 2 |
| chrX_-_103087136 | 0.12 |
ENST00000243298.2 |
RAB9B |
RAB9B, member RAS oncogene family |
| chr6_-_52860171 | 0.11 |
ENST00000370963.4 |
GSTA4 |
glutathione S-transferase alpha 4 |
| chr11_+_65479702 | 0.11 |
ENST00000530446.1 ENST00000534104.1 ENST00000530605.1 ENST00000528198.1 ENST00000531880.1 ENST00000534650.1 |
KAT5 |
K(lysine) acetyltransferase 5 |
| chr10_-_3827417 | 0.10 |
ENST00000497571.1 ENST00000542957.1 |
KLF6 |
Kruppel-like factor 6 |
| chr11_+_65479462 | 0.10 |
ENST00000377046.3 ENST00000352980.4 ENST00000341318.4 |
KAT5 |
K(lysine) acetyltransferase 5 |
| chr17_-_40075219 | 0.10 |
ENST00000537919.1 ENST00000352035.2 ENST00000353196.1 ENST00000393896.2 |
ACLY |
ATP citrate lyase |
| chr14_-_35183886 | 0.09 |
ENST00000298159.6 |
CFL2 |
cofilin 2 (muscle) |
| chr1_-_42384343 | 0.09 |
ENST00000372584.1 |
HIVEP3 |
human immunodeficiency virus type I enhancer binding protein 3 |
| chr11_+_2923423 | 0.08 |
ENST00000312221.5 |
SLC22A18 |
solute carrier family 22, member 18 |
| chr16_+_2083265 | 0.08 |
ENST00000565855.1 ENST00000566198.1 |
SLC9A3R2 |
solute carrier family 9, subfamily A (NHE3, cation proton antiporter 3), member 3 regulator 2 |
| chr4_+_75230853 | 0.08 |
ENST00000244869.2 |
EREG |
epiregulin |
| chr2_-_219925189 | 0.08 |
ENST00000295731.6 |
IHH |
indian hedgehog |
| chr11_-_73309228 | 0.07 |
ENST00000356467.4 ENST00000064778.4 |
FAM168A |
family with sequence similarity 168, member A |
| chr15_+_47476275 | 0.07 |
ENST00000558014.1 |
SEMA6D |
sema domain, transmembrane domain (TM), and cytoplasmic domain, (semaphorin) 6D |
| chr11_+_2923619 | 0.07 |
ENST00000380574.1 |
SLC22A18 |
solute carrier family 22, member 18 |
| chr3_-_171527560 | 0.07 |
ENST00000331659.2 |
PP13439 |
PP13439 |
| chr11_+_2923499 | 0.07 |
ENST00000449793.2 |
SLC22A18 |
solute carrier family 22, member 18 |
| chr19_+_36266433 | 0.06 |
ENST00000314737.5 |
ARHGAP33 |
Rho GTPase activating protein 33 |
| chr6_-_52859968 | 0.06 |
ENST00000370959.1 |
GSTA4 |
glutathione S-transferase alpha 4 |
| chr17_-_40075197 | 0.06 |
ENST00000590770.1 ENST00000590151.1 |
ACLY |
ATP citrate lyase |
| chr17_-_56065484 | 0.06 |
ENST00000581208.1 |
VEZF1 |
vascular endothelial zinc finger 1 |
| chr5_-_107703556 | 0.06 |
ENST00000496714.1 |
FBXL17 |
F-box and leucine-rich repeat protein 17 |
| chr19_+_36266417 | 0.06 |
ENST00000378944.5 ENST00000007510.4 |
ARHGAP33 |
Rho GTPase activating protein 33 |
| chr3_+_5020801 | 0.06 |
ENST00000256495.3 |
BHLHE40 |
basic helix-loop-helix family, member e40 |
| chr10_-_74856608 | 0.06 |
ENST00000307116.2 ENST00000373008.2 ENST00000412021.2 ENST00000394890.2 ENST00000263556.3 ENST00000440381.1 |
P4HA1 |
prolyl 4-hydroxylase, alpha polypeptide I |
| chr15_-_73075964 | 0.06 |
ENST00000563907.1 |
ADPGK |
ADP-dependent glucokinase |
| chr1_+_39796810 | 0.05 |
ENST00000289893.4 |
MACF1 |
microtubule-actin crosslinking factor 1 |
| chr6_-_31926208 | 0.05 |
ENST00000454913.1 ENST00000436289.2 |
NELFE |
negative elongation factor complex member E |
| chr14_-_74959994 | 0.04 |
ENST00000238633.2 ENST00000434013.2 |
NPC2 |
Niemann-Pick disease, type C2 |
| chr3_-_160117301 | 0.04 |
ENST00000326448.7 ENST00000498409.1 ENST00000475677.1 ENST00000478536.1 |
IFT80 |
intraflagellar transport 80 homolog (Chlamydomonas) |
| chr16_+_31225337 | 0.04 |
ENST00000322122.3 |
TRIM72 |
tripartite motif containing 72 |
| chr19_+_11200038 | 0.04 |
ENST00000558518.1 ENST00000557933.1 ENST00000455727.2 ENST00000535915.1 ENST00000545707.1 ENST00000558013.1 |
LDLR |
low density lipoprotein receptor |
| chr14_-_74960030 | 0.04 |
ENST00000553490.1 ENST00000557510.1 |
NPC2 |
Niemann-Pick disease, type C2 |
| chr6_-_39902160 | 0.04 |
ENST00000340692.5 |
MOCS1 |
molybdenum cofactor synthesis 1 |
| chr14_-_74959978 | 0.03 |
ENST00000541064.1 |
NPC2 |
Niemann-Pick disease, type C2 |
| chr11_+_22688150 | 0.03 |
ENST00000454584.2 |
GAS2 |
growth arrest-specific 2 |
| chr9_+_131452239 | 0.03 |
ENST00000372688.4 ENST00000372686.5 |
SET |
SET nuclear oncogene |
| chrX_-_112084043 | 0.03 |
ENST00000304758.1 |
AMOT |
angiomotin |
| chr12_+_50344516 | 0.03 |
ENST00000199280.3 ENST00000550862.1 |
AQP2 |
aquaporin 2 (collecting duct) |
| chr6_-_42713792 | 0.02 |
ENST00000372876.1 |
TBCC |
tubulin folding cofactor C |
| chr3_+_160117418 | 0.02 |
ENST00000465903.1 ENST00000485645.1 ENST00000360111.2 ENST00000472991.1 ENST00000467468.1 ENST00000469762.1 ENST00000489573.1 ENST00000462787.1 ENST00000490207.1 ENST00000485867.1 |
SMC4 |
structural maintenance of chromosomes 4 |
| chr15_-_73076030 | 0.02 |
ENST00000311669.8 |
ADPGK |
ADP-dependent glucokinase |
| chr14_+_74960423 | 0.02 |
ENST00000556816.1 ENST00000298818.8 ENST00000554924.1 |
ISCA2 |
iron-sulfur cluster assembly 2 |
| chr5_+_159656437 | 0.02 |
ENST00000402432.3 |
FABP6 |
fatty acid binding protein 6, ileal |
| chr1_-_155232047 | 0.02 |
ENST00000302631.3 |
SCAMP3 |
secretory carrier membrane protein 3 |
| chr6_+_43968306 | 0.02 |
ENST00000442114.2 ENST00000336600.5 ENST00000439969.2 |
C6orf223 |
chromosome 6 open reading frame 223 |
| chr6_-_53213780 | 0.02 |
ENST00000304434.6 ENST00000370918.4 |
ELOVL5 |
ELOVL fatty acid elongase 5 |
| chr11_+_65383227 | 0.02 |
ENST00000355703.3 |
PCNXL3 |
pecanex-like 3 (Drosophila) |
| chr8_+_21882244 | 0.02 |
ENST00000289820.6 ENST00000381530.5 |
NPM2 |
nucleophosmin/nucleoplasmin 2 |
| chr15_-_66858298 | 0.02 |
ENST00000537670.1 |
LCTL |
lactase-like |
| chr1_-_155232221 | 0.02 |
ENST00000355379.3 |
SCAMP3 |
secretory carrier membrane protein 3 |
| chr22_+_42229100 | 0.01 |
ENST00000361204.4 |
SREBF2 |
sterol regulatory element binding transcription factor 2 |
| chr18_-_21166841 | 0.01 |
ENST00000269228.5 |
NPC1 |
Niemann-Pick disease, type C1 |
| chr1_+_206643787 | 0.01 |
ENST00000367120.3 |
IKBKE |
inhibitor of kappa light polypeptide gene enhancer in B-cells, kinase epsilon |
| chr6_+_69942298 | 0.01 |
ENST00000238918.8 |
BAI3 |
brain-specific angiogenesis inhibitor 3 |
| chr5_-_43313574 | 0.01 |
ENST00000325110.6 ENST00000433297.2 |
HMGCS1 |
3-hydroxy-3-methylglutaryl-CoA synthase 1 (soluble) |
| chr16_+_22103828 | 0.01 |
ENST00000567131.1 ENST00000568328.1 ENST00000389398.5 ENST00000389397.4 |
VWA3A |
von Willebrand factor A domain containing 3A |
| chrX_+_27826107 | 0.01 |
ENST00000356790.2 |
MAGEB10 |
melanoma antigen family B, 10 |
| chr13_+_24844819 | 0.00 |
ENST00000399949.2 |
SPATA13 |
spermatogenesis associated 13 |
Gene Ontology Analysis
Gene overrepresentation in biological_process category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.2 | 0.7 | GO:1900158 | negative regulation of osteoclast proliferation(GO:0090291) negative regulation of bone mineralization involved in bone maturation(GO:1900158) |
| 0.2 | 0.7 | GO:0035441 | cell migration involved in vasculogenesis(GO:0035441) metanephric glomerulus morphogenesis(GO:0072275) metanephric glomerulus vasculature morphogenesis(GO:0072276) metanephric glomerular capillary formation(GO:0072277) |
| 0.1 | 0.4 | GO:0070563 | negative regulation of vitamin D receptor signaling pathway(GO:0070563) |
| 0.1 | 0.5 | GO:0019427 | acetate biosynthetic process(GO:0019413) acetyl-CoA biosynthetic process from acetate(GO:0019427) propionate metabolic process(GO:0019541) propionate biosynthetic process(GO:0019542) |
| 0.1 | 0.3 | GO:0035026 | leading edge cell differentiation(GO:0035026) |
| 0.0 | 0.1 | GO:0050916 | sensory perception of sweet taste(GO:0050916) |
| 0.0 | 0.1 | GO:0016539 | intein-mediated protein splicing(GO:0016539) protein splicing(GO:0030908) |
| 0.0 | 0.2 | GO:0046618 | drug export(GO:0046618) |
| 0.0 | 0.3 | GO:0014722 | regulation of skeletal muscle contraction by calcium ion signaling(GO:0014722) |
| 0.0 | 0.1 | GO:0019747 | regulation of isoprenoid metabolic process(GO:0019747) |
| 0.0 | 0.2 | GO:0032914 | positive regulation of transforming growth factor beta1 production(GO:0032914) |
| 0.0 | 0.3 | GO:0010739 | positive regulation of protein kinase A signaling(GO:0010739) |
| 0.0 | 0.1 | GO:0032534 | regulation of microvillus assembly(GO:0032534) |
| 0.0 | 0.1 | GO:0048165 | ovarian cumulus expansion(GO:0001550) fused antrum stage(GO:0048165) |
Gene overrepresentation in cellular_component category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.0 | 0.2 | GO:0032777 | Piccolo NuA4 histone acetyltransferase complex(GO:0032777) |
| 0.0 | 0.3 | GO:0070022 | transforming growth factor beta receptor homodimeric complex(GO:0070022) |
| 0.0 | 0.1 | GO:0035976 | AP1 complex(GO:0035976) |
| 0.0 | 0.1 | GO:0016222 | procollagen-proline 4-dioxygenase complex(GO:0016222) |
| 0.0 | 0.0 | GO:0019008 | molybdopterin synthase complex(GO:0019008) |
Gene overrepresentation in molecular_function category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.2 | 0.7 | GO:0005017 | platelet-derived growth factor-activated receptor activity(GO:0005017) |
| 0.1 | 0.5 | GO:0003987 | acetate-CoA ligase activity(GO:0003987) |
| 0.1 | 0.7 | GO:0016015 | morphogen activity(GO:0016015) |
| 0.1 | 0.2 | GO:0004157 | dihydropyrimidinase activity(GO:0004157) |
| 0.0 | 0.2 | GO:0015307 | drug:proton antiporter activity(GO:0015307) |
| 0.0 | 0.2 | GO:0046912 | transferase activity, transferring acyl groups, acyl groups converted into alkyl on transfer(GO:0046912) |
| 0.0 | 0.1 | GO:0070290 | N-acylphosphatidylethanolamine-specific phospholipase D activity(GO:0070290) |
| 0.0 | 0.3 | GO:0008179 | adenylate cyclase binding(GO:0008179) |
| 0.0 | 0.3 | GO:0035615 | clathrin adaptor activity(GO:0035615) endocytic adaptor activity(GO:0098748) |
Gene overrepresentation in C2:CP category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.0 | 0.7 | PID S1P S1P1 PATHWAY | S1P1 pathway |
| 0.0 | 0.5 | PID HDAC CLASSIII PATHWAY | Signaling events mediated by HDAC Class III |
Gene overrepresentation in C2:CP:REACTOME category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.0 | 0.5 | REACTOME ETHANOL OXIDATION | Genes involved in Ethanol oxidation |
| 0.0 | 0.3 | REACTOME GAP JUNCTION DEGRADATION | Genes involved in Gap junction degradation |