Project
Epithelial-Mesenchymal Transition, human (Scheel, 2011)
Navigation
Downloads
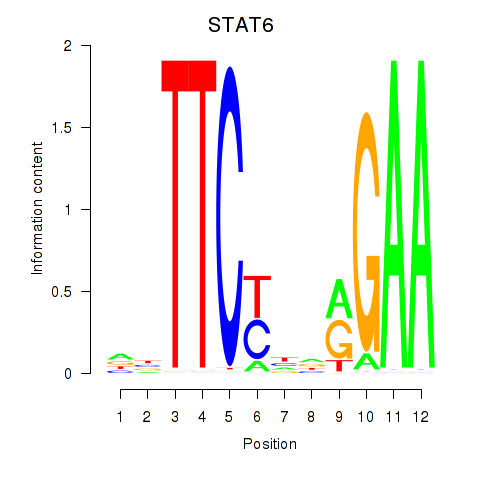
Results for STAT6
Z-value: 0.61
Transcription factors associated with STAT6
| Gene Symbol | Gene ID | Gene Info |
|---|---|---|
|
STAT6
|
ENSG00000166888.6 | STAT6 |
Activity-expression correlation:
| Gene | Promoter | Pearson corr. coef. | P-value | Plot |
|---|---|---|---|---|
| STAT6 | hg19_v2_chr12_-_57505121_57505216 | -0.26 | 5.3e-01 | Click! |
Activity profile of STAT6 motif
Sorted Z-values of STAT6 motif
Network of associatons between targets according to the STRING database.
First level regulatory network of STAT6
| Promoter | Score | Transcript | Gene | Gene Info |
|---|---|---|---|---|
| chr8_+_97506033 | 0.52 |
ENST00000518385.1 |
SDC2 |
syndecan 2 |
| chr9_+_90112741 | 0.42 |
ENST00000469640.2 |
DAPK1 |
death-associated protein kinase 1 |
| chr9_+_90112590 | 0.42 |
ENST00000472284.1 |
DAPK1 |
death-associated protein kinase 1 |
| chr9_+_90112767 | 0.41 |
ENST00000408954.3 |
DAPK1 |
death-associated protein kinase 1 |
| chr9_+_90112117 | 0.39 |
ENST00000358077.5 |
DAPK1 |
death-associated protein kinase 1 |
| chr1_-_76398077 | 0.33 |
ENST00000284142.6 |
ASB17 |
ankyrin repeat and SOCS box containing 17 |
| chrX_-_106960285 | 0.33 |
ENST00000503515.1 ENST00000372397.2 |
TSC22D3 |
TSC22 domain family, member 3 |
| chr2_-_190044480 | 0.29 |
ENST00000374866.3 |
COL5A2 |
collagen, type V, alpha 2 |
| chr12_+_7052974 | 0.27 |
ENST00000544681.1 ENST00000537087.1 |
C12orf57 |
chromosome 12 open reading frame 57 |
| chr6_-_24877490 | 0.27 |
ENST00000540914.1 ENST00000378023.4 |
FAM65B |
family with sequence similarity 65, member B |
| chr12_+_7053172 | 0.27 |
ENST00000229281.5 |
C12orf57 |
chromosome 12 open reading frame 57 |
| chrX_+_135251835 | 0.26 |
ENST00000456445.1 |
FHL1 |
four and a half LIM domains 1 |
| chr3_-_49851313 | 0.26 |
ENST00000333486.3 |
UBA7 |
ubiquitin-like modifier activating enzyme 7 |
| chr12_+_7053228 | 0.25 |
ENST00000540506.2 |
C12orf57 |
chromosome 12 open reading frame 57 |
| chr2_-_203735976 | 0.24 |
ENST00000435143.1 |
ICA1L |
islet cell autoantigen 1,69kDa-like |
| chr17_+_7461613 | 0.23 |
ENST00000438470.1 ENST00000436057.1 |
TNFSF13 |
tumor necrosis factor (ligand) superfamily, member 13 |
| chr11_+_77532155 | 0.23 |
ENST00000532481.1 ENST00000526415.1 ENST00000393427.2 ENST00000527134.1 ENST00000304716.8 |
AAMDC |
adipogenesis associated, Mth938 domain containing |
| chr6_+_32821924 | 0.22 |
ENST00000374859.2 ENST00000453265.2 |
PSMB9 |
proteasome (prosome, macropain) subunit, beta type, 9 |
| chr1_+_78383813 | 0.22 |
ENST00000342754.5 |
NEXN |
nexilin (F actin binding protein) |
| chr11_+_77532233 | 0.22 |
ENST00000525409.1 |
AAMDC |
adipogenesis associated, Mth938 domain containing |
| chr11_-_14379997 | 0.21 |
ENST00000526063.1 ENST00000532814.1 |
RRAS2 |
related RAS viral (r-ras) oncogene homolog 2 |
| chrX_+_135251783 | 0.21 |
ENST00000394153.2 |
FHL1 |
four and a half LIM domains 1 |
| chr1_+_150245177 | 0.21 |
ENST00000369098.3 |
C1orf54 |
chromosome 1 open reading frame 54 |
| chr1_+_202431859 | 0.21 |
ENST00000391959.3 ENST00000367270.4 |
PPP1R12B |
protein phosphatase 1, regulatory subunit 12B |
| chr12_+_12764773 | 0.20 |
ENST00000228865.2 |
CREBL2 |
cAMP responsive element binding protein-like 2 |
| chr2_+_202316392 | 0.20 |
ENST00000194530.3 ENST00000392249.2 |
STRADB |
STE20-related kinase adaptor beta |
| chr3_-_73673991 | 0.20 |
ENST00000308537.4 ENST00000263666.4 |
PDZRN3 |
PDZ domain containing ring finger 3 |
| chr16_-_74455290 | 0.19 |
ENST00000339953.5 |
CLEC18B |
C-type lectin domain family 18, member B |
| chrX_+_135252050 | 0.19 |
ENST00000449474.1 ENST00000345434.3 |
FHL1 |
four and a half LIM domains 1 |
| chr1_-_179834311 | 0.18 |
ENST00000553856.1 |
IFRG15 |
Homo sapiens torsin A interacting protein 2 (TOR1AIP2), transcript variant 1, mRNA. |
| chr22_-_42342733 | 0.18 |
ENST00000402420.1 |
CENPM |
centromere protein M |
| chr6_+_30525051 | 0.17 |
ENST00000376557.3 |
PRR3 |
proline rich 3 |
| chr20_-_18477862 | 0.16 |
ENST00000337227.4 |
RBBP9 |
retinoblastoma binding protein 9 |
| chr1_-_203198790 | 0.16 |
ENST00000367229.1 ENST00000255427.3 ENST00000535569.1 |
CHIT1 |
chitinase 1 (chitotriosidase) |
| chr5_+_140602904 | 0.15 |
ENST00000515856.2 ENST00000239449.4 |
PCDHB14 |
protocadherin beta 14 |
| chr1_-_68299130 | 0.15 |
ENST00000370982.3 |
GNG12 |
guanine nucleotide binding protein (G protein), gamma 12 |
| chr7_-_75401513 | 0.15 |
ENST00000005180.4 |
CCL26 |
chemokine (C-C motif) ligand 26 |
| chr2_-_120124383 | 0.15 |
ENST00000334816.7 |
C2orf76 |
chromosome 2 open reading frame 76 |
| chr3_+_158787041 | 0.14 |
ENST00000471575.1 ENST00000476809.1 ENST00000485419.1 |
IQCJ-SCHIP1 |
IQCJ-SCHIP1 readthrough |
| chr15_+_25200108 | 0.14 |
ENST00000577949.1 ENST00000338094.6 ENST00000338327.4 ENST00000579070.1 ENST00000577565.1 |
SNURF SNRPN |
SNRPN upstream reading frame protein small nuclear ribonucleoprotein polypeptide N |
| chr14_+_55034599 | 0.14 |
ENST00000392067.3 ENST00000357634.3 |
SAMD4A |
sterile alpha motif domain containing 4A |
| chr2_-_120124258 | 0.14 |
ENST00000409877.1 ENST00000409523.1 ENST00000409466.2 ENST00000414534.1 |
C2orf76 |
chromosome 2 open reading frame 76 |
| chr1_+_61869748 | 0.13 |
ENST00000357977.5 |
NFIA |
nuclear factor I/A |
| chr14_-_24551195 | 0.13 |
ENST00000560550.1 |
NRL |
neural retina leucine zipper |
| chr14_-_25103388 | 0.13 |
ENST00000526004.1 ENST00000415355.3 |
GZMB |
granzyme B (granzyme 2, cytotoxic T-lymphocyte-associated serine esterase 1) |
| chr3_+_158991025 | 0.13 |
ENST00000337808.6 |
IQCJ-SCHIP1 |
IQCJ-SCHIP1 readthrough |
| chr2_-_175462456 | 0.13 |
ENST00000409891.1 ENST00000410117.1 |
WIPF1 |
WAS/WASL interacting protein family, member 1 |
| chr15_+_25200074 | 0.13 |
ENST00000390687.4 ENST00000584968.1 ENST00000346403.6 ENST00000554227.2 |
SNRPN |
small nuclear ribonucleoprotein polypeptide N |
| chr14_-_25103472 | 0.13 |
ENST00000216341.4 ENST00000382542.1 ENST00000382540.1 |
GZMB |
granzyme B (granzyme 2, cytotoxic T-lymphocyte-associated serine esterase 1) |
| chr16_+_70207686 | 0.13 |
ENST00000541793.2 ENST00000314151.8 ENST00000565806.1 ENST00000569347.2 ENST00000536907.2 |
CLEC18C |
C-type lectin domain family 18, member C |
| chr3_+_141121164 | 0.13 |
ENST00000510338.1 ENST00000504673.1 |
ZBTB38 |
zinc finger and BTB domain containing 38 |
| chr16_+_69985083 | 0.12 |
ENST00000288040.6 ENST00000449317.2 |
CLEC18A |
C-type lectin domain family 18, member A |
| chr4_-_186877806 | 0.12 |
ENST00000355634.5 |
SORBS2 |
sorbin and SH3 domain containing 2 |
| chr2_-_203735586 | 0.12 |
ENST00000454326.1 ENST00000432273.1 ENST00000450143.1 ENST00000411681.1 |
ICA1L |
islet cell autoantigen 1,69kDa-like |
| chr10_+_124907638 | 0.12 |
ENST00000339992.3 |
HMX2 |
H6 family homeobox 2 |
| chr2_-_202316260 | 0.12 |
ENST00000332624.3 |
TRAK2 |
trafficking protein, kinesin binding 2 |
| chr5_+_118691706 | 0.12 |
ENST00000415806.2 |
TNFAIP8 |
tumor necrosis factor, alpha-induced protein 8 |
| chr18_-_24445664 | 0.12 |
ENST00000578776.1 |
AQP4 |
aquaporin 4 |
| chr6_+_84222220 | 0.11 |
ENST00000369700.3 |
PRSS35 |
protease, serine, 35 |
| chr14_-_24551137 | 0.11 |
ENST00000396995.1 |
NRL |
neural retina leucine zipper |
| chr11_+_36616044 | 0.11 |
ENST00000334307.5 ENST00000531554.1 ENST00000347206.4 ENST00000534635.1 ENST00000446510.2 ENST00000530697.1 ENST00000527108.1 |
C11orf74 |
chromosome 11 open reading frame 74 |
| chr1_+_150245099 | 0.11 |
ENST00000369099.3 |
C1orf54 |
chromosome 1 open reading frame 54 |
| chr2_+_120124497 | 0.11 |
ENST00000355857.3 ENST00000535617.1 ENST00000535757.1 ENST00000409094.1 ENST00000311521.4 |
DBI |
diazepam binding inhibitor (GABA receptor modulator, acyl-CoA binding protein) |
| chr2_-_203736150 | 0.11 |
ENST00000457524.1 ENST00000421334.1 |
ICA1L |
islet cell autoantigen 1,69kDa-like |
| chr12_+_44152740 | 0.11 |
ENST00000440781.2 ENST00000431837.1 ENST00000550616.1 ENST00000448290.2 ENST00000551736.1 |
IRAK4 |
interleukin-1 receptor-associated kinase 4 |
| chr6_-_41168920 | 0.11 |
ENST00000483722.1 |
TREML2 |
triggering receptor expressed on myeloid cells-like 2 |
| chr15_+_91643442 | 0.11 |
ENST00000394232.1 |
SV2B |
synaptic vesicle glycoprotein 2B |
| chr13_-_74708372 | 0.10 |
ENST00000377666.4 |
KLF12 |
Kruppel-like factor 12 |
| chr8_+_39792474 | 0.10 |
ENST00000502986.2 |
IDO2 |
indoleamine 2,3-dioxygenase 2 |
| chr16_+_69984810 | 0.10 |
ENST00000393701.2 ENST00000568461.1 |
CLEC18A |
C-type lectin domain family 18, member A |
| chr17_+_40912764 | 0.10 |
ENST00000589683.1 ENST00000588928.1 |
RAMP2 |
receptor (G protein-coupled) activity modifying protein 2 |
| chr1_-_241803679 | 0.10 |
ENST00000331838.5 |
OPN3 |
opsin 3 |
| chr6_+_31540056 | 0.10 |
ENST00000418386.2 |
LTA |
lymphotoxin alpha |
| chr21_+_17791838 | 0.10 |
ENST00000453910.1 |
LINC00478 |
long intergenic non-protein coding RNA 478 |
| chr21_+_17791648 | 0.10 |
ENST00000602892.1 ENST00000418813.2 ENST00000435697.1 |
LINC00478 |
long intergenic non-protein coding RNA 478 |
| chrX_+_100878079 | 0.10 |
ENST00000471229.2 |
ARMCX3 |
armadillo repeat containing, X-linked 3 |
| chr1_+_26759295 | 0.10 |
ENST00000430232.1 |
DHDDS |
dehydrodolichyl diphosphate synthase |
| chr12_+_26164645 | 0.09 |
ENST00000542004.1 |
RASSF8 |
Ras association (RalGDS/AF-6) domain family (N-terminal) member 8 |
| chr4_+_166300084 | 0.09 |
ENST00000402744.4 |
CPE |
carboxypeptidase E |
| chr11_-_67980744 | 0.09 |
ENST00000401547.2 ENST00000453170.1 ENST00000304363.4 |
SUV420H1 |
suppressor of variegation 4-20 homolog 1 (Drosophila) |
| chr3_-_45957088 | 0.09 |
ENST00000539217.1 |
LZTFL1 |
leucine zipper transcription factor-like 1 |
| chr7_+_129015484 | 0.09 |
ENST00000490911.1 |
AHCYL2 |
adenosylhomocysteinase-like 2 |
| chr10_+_75541796 | 0.08 |
ENST00000372837.3 ENST00000372833.5 |
CHCHD1 |
coiled-coil-helix-coiled-coil-helix domain containing 1 |
| chr8_-_42623924 | 0.08 |
ENST00000276410.2 |
CHRNA6 |
cholinergic receptor, nicotinic, alpha 6 (neuronal) |
| chr21_+_17792672 | 0.08 |
ENST00000602620.1 |
LINC00478 |
long intergenic non-protein coding RNA 478 |
| chr2_-_204400013 | 0.08 |
ENST00000374489.2 ENST00000374488.2 ENST00000308091.4 ENST00000453034.1 ENST00000420371.1 |
RAPH1 |
Ras association (RalGDS/AF-6) and pleckstrin homology domains 1 |
| chr5_+_112227311 | 0.08 |
ENST00000391338.1 |
ZRSR1 |
zinc finger (CCCH type), RNA-binding motif and serine/arginine rich 1 |
| chr5_+_150051149 | 0.08 |
ENST00000523553.1 |
MYOZ3 |
myozenin 3 |
| chr19_+_15218180 | 0.08 |
ENST00000342784.2 ENST00000597977.1 ENST00000600440.1 |
SYDE1 |
synapse defective 1, Rho GTPase, homolog 1 (C. elegans) |
| chr6_+_26440700 | 0.08 |
ENST00000494393.1 ENST00000482451.1 ENST00000244519.2 ENST00000339789.4 ENST00000471353.1 ENST00000361232.3 ENST00000487627.1 ENST00000496719.1 ENST00000490254.1 ENST00000487272.1 |
BTN3A3 |
butyrophilin, subfamily 3, member A3 |
| chr2_-_203735484 | 0.08 |
ENST00000420558.1 ENST00000418208.1 |
ICA1L |
islet cell autoantigen 1,69kDa-like |
| chr8_-_42623747 | 0.08 |
ENST00000534622.1 |
CHRNA6 |
cholinergic receptor, nicotinic, alpha 6 (neuronal) |
| chr3_-_57233966 | 0.08 |
ENST00000473921.1 ENST00000295934.3 |
HESX1 |
HESX homeobox 1 |
| chr17_-_34890709 | 0.08 |
ENST00000544606.1 |
MYO19 |
myosin XIX |
| chr2_-_197226875 | 0.07 |
ENST00000409111.1 |
HECW2 |
HECT, C2 and WW domain containing E3 ubiquitin protein ligase 2 |
| chr3_-_179692042 | 0.07 |
ENST00000468741.1 |
PEX5L |
peroxisomal biogenesis factor 5-like |
| chr12_+_123949053 | 0.07 |
ENST00000350887.5 |
SNRNP35 |
small nuclear ribonucleoprotein 35kDa (U11/U12) |
| chr1_-_41328018 | 0.07 |
ENST00000372638.2 |
CITED4 |
Cbp/p300-interacting transactivator, with Glu/Asp-rich carboxy-terminal domain, 4 |
| chr15_+_67841330 | 0.07 |
ENST00000354498.5 |
MAP2K5 |
mitogen-activated protein kinase kinase 5 |
| chr22_+_44464923 | 0.07 |
ENST00000404989.1 |
PARVB |
parvin, beta |
| chr2_+_182756915 | 0.07 |
ENST00000428267.2 |
SSFA2 |
sperm specific antigen 2 |
| chrX_+_36254051 | 0.07 |
ENST00000378657.4 |
CXorf30 |
chromosome X open reading frame 30 |
| chr1_+_12123414 | 0.07 |
ENST00000263932.2 |
TNFRSF8 |
tumor necrosis factor receptor superfamily, member 8 |
| chr6_-_117150198 | 0.07 |
ENST00000310357.3 ENST00000368549.3 ENST00000530250.1 |
GPRC6A |
G protein-coupled receptor, family C, group 6, member A |
| chr6_-_30524951 | 0.07 |
ENST00000376621.3 |
GNL1 |
guanine nucleotide binding protein-like 1 |
| chr1_+_114472481 | 0.07 |
ENST00000369555.2 |
HIPK1 |
homeodomain interacting protein kinase 1 |
| chr11_+_73358594 | 0.07 |
ENST00000227214.6 ENST00000398494.4 ENST00000543085.1 |
PLEKHB1 |
pleckstrin homology domain containing, family B (evectins) member 1 |
| chr3_-_185655795 | 0.07 |
ENST00000342294.4 ENST00000382191.4 ENST00000453386.2 |
TRA2B |
transformer 2 beta homolog (Drosophila) |
| chr11_+_65101225 | 0.07 |
ENST00000528416.1 ENST00000415073.2 ENST00000252268.4 |
DPF2 |
D4, zinc and double PHD fingers family 2 |
| chr16_-_29757272 | 0.06 |
ENST00000329410.3 |
C16orf54 |
chromosome 16 open reading frame 54 |
| chrX_-_50386648 | 0.06 |
ENST00000460112.3 |
SHROOM4 |
shroom family member 4 |
| chr9_+_100000717 | 0.06 |
ENST00000375205.2 ENST00000357054.1 ENST00000395220.1 ENST00000375202.2 ENST00000411667.2 |
CCDC180 |
coiled-coil domain containing 180 |
| chr16_-_73093597 | 0.06 |
ENST00000397992.5 |
ZFHX3 |
zinc finger homeobox 3 |
| chr8_+_9953061 | 0.06 |
ENST00000522907.1 ENST00000528246.1 |
MSRA |
methionine sulfoxide reductase A |
| chr3_-_45957534 | 0.06 |
ENST00000536047.1 |
LZTFL1 |
leucine zipper transcription factor-like 1 |
| chr17_+_44588877 | 0.05 |
ENST00000576629.1 |
LRRC37A2 |
leucine rich repeat containing 37, member A2 |
| chr8_+_9953214 | 0.05 |
ENST00000382490.5 |
MSRA |
methionine sulfoxide reductase A |
| chr22_-_42343117 | 0.05 |
ENST00000407253.3 ENST00000215980.5 |
CENPM |
centromere protein M |
| chr15_+_64386261 | 0.05 |
ENST00000560829.1 |
SNX1 |
sorting nexin 1 |
| chr1_-_241683001 | 0.05 |
ENST00000366560.3 |
FH |
fumarate hydratase |
| chr1_+_114472222 | 0.05 |
ENST00000369558.1 ENST00000369561.4 |
HIPK1 |
homeodomain interacting protein kinase 1 |
| chr3_-_10149907 | 0.05 |
ENST00000450660.2 ENST00000524279.1 |
FANCD2OS |
FANCD2 opposite strand |
| chr22_-_42342692 | 0.05 |
ENST00000404067.1 ENST00000402338.1 |
CENPM |
centromere protein M |
| chr1_-_94586651 | 0.05 |
ENST00000535735.1 ENST00000370225.3 |
ABCA4 |
ATP-binding cassette, sub-family A (ABC1), member 4 |
| chr17_-_34890665 | 0.05 |
ENST00000586007.1 |
MYO19 |
myosin XIX |
| chr1_-_160616804 | 0.05 |
ENST00000538290.1 |
SLAMF1 |
signaling lymphocytic activation molecule family member 1 |
| chr4_+_54966198 | 0.05 |
ENST00000326902.2 ENST00000503800.1 |
GSX2 |
GS homeobox 2 |
| chr15_+_34638066 | 0.05 |
ENST00000333756.4 |
NUTM1 |
NUT midline carcinoma, family member 1 |
| chr2_-_204400113 | 0.04 |
ENST00000319170.5 |
RAPH1 |
Ras association (RalGDS/AF-6) and pleckstrin homology domains 1 |
| chr18_-_24445729 | 0.04 |
ENST00000383168.4 |
AQP4 |
aquaporin 4 |
| chr1_-_227505826 | 0.04 |
ENST00000334218.5 ENST00000366766.2 ENST00000366764.2 |
CDC42BPA |
CDC42 binding protein kinase alpha (DMPK-like) |
| chr16_+_30075463 | 0.04 |
ENST00000562168.1 ENST00000569545.1 |
ALDOA |
aldolase A, fructose-bisphosphate |
| chr1_-_201915590 | 0.04 |
ENST00000367288.4 |
LMOD1 |
leiomodin 1 (smooth muscle) |
| chr15_+_96869165 | 0.04 |
ENST00000421109.2 |
NR2F2 |
nuclear receptor subfamily 2, group F, member 2 |
| chr4_+_71337834 | 0.04 |
ENST00000304887.5 |
MUC7 |
mucin 7, secreted |
| chr15_-_100882191 | 0.04 |
ENST00000268070.4 |
ADAMTS17 |
ADAM metallopeptidase with thrombospondin type 1 motif, 17 |
| chr3_+_148508845 | 0.04 |
ENST00000491148.1 |
CPB1 |
carboxypeptidase B1 (tissue) |
| chr9_-_100000957 | 0.04 |
ENST00000366109.2 ENST00000607322.1 |
RP11-498P14.5 |
RP11-498P14.5 |
| chr2_-_133427767 | 0.04 |
ENST00000397463.2 |
LYPD1 |
LY6/PLAUR domain containing 1 |
| chr1_+_109256067 | 0.03 |
ENST00000271311.2 |
FNDC7 |
fibronectin type III domain containing 7 |
| chr9_-_13175823 | 0.03 |
ENST00000545857.1 |
MPDZ |
multiple PDZ domain protein |
| chr5_+_102201687 | 0.03 |
ENST00000304400.7 |
PAM |
peptidylglycine alpha-amidating monooxygenase |
| chr16_+_30075595 | 0.03 |
ENST00000563060.2 |
ALDOA |
aldolase A, fructose-bisphosphate |
| chr8_+_49984894 | 0.03 |
ENST00000522267.1 ENST00000399653.4 ENST00000303202.8 |
C8orf22 |
chromosome 8 open reading frame 22 |
| chr5_-_146461027 | 0.03 |
ENST00000394410.2 ENST00000508267.1 ENST00000504198.1 |
PPP2R2B |
protein phosphatase 2, regulatory subunit B, beta |
| chr15_-_23932437 | 0.03 |
ENST00000331837.4 |
NDN |
necdin, melanoma antigen (MAGE) family member |
| chr22_-_29457832 | 0.03 |
ENST00000216071.4 |
C22orf31 |
chromosome 22 open reading frame 31 |
| chr7_-_150777949 | 0.03 |
ENST00000482571.1 |
FASTK |
Fas-activated serine/threonine kinase |
| chr3_+_179370517 | 0.03 |
ENST00000263966.3 |
USP13 |
ubiquitin specific peptidase 13 (isopeptidase T-3) |
| chr16_+_30075783 | 0.03 |
ENST00000412304.2 |
ALDOA |
aldolase A, fructose-bisphosphate |
| chr2_-_16804320 | 0.03 |
ENST00000355549.2 |
FAM49A |
family with sequence similarity 49, member A |
| chr7_-_76829125 | 0.03 |
ENST00000248598.5 |
FGL2 |
fibrinogen-like 2 |
| chr14_-_46185155 | 0.03 |
ENST00000555442.1 |
RP11-369C8.1 |
RP11-369C8.1 |
| chr12_+_80730292 | 0.03 |
ENST00000298820.3 |
OTOGL |
otogelin-like |
| chr19_-_7766991 | 0.02 |
ENST00000597921.1 ENST00000346664.5 |
FCER2 |
Fc fragment of IgE, low affinity II, receptor for (CD23) |
| chr6_-_27860956 | 0.02 |
ENST00000359611.2 |
HIST1H2AM |
histone cluster 1, H2am |
| chr17_+_7462031 | 0.02 |
ENST00000380535.4 |
TNFSF13 |
tumor necrosis factor (ligand) superfamily, member 13 |
| chr7_-_150777920 | 0.02 |
ENST00000353841.2 ENST00000297532.6 |
FASTK |
Fas-activated serine/threonine kinase |
| chr20_+_57209828 | 0.02 |
ENST00000598340.1 |
MGC4294 |
HCG1785561; MGC4294 protein; Uncharacterized protein |
| chr16_+_67694849 | 0.02 |
ENST00000602551.1 ENST00000458121.2 ENST00000219255.3 |
PARD6A |
par-6 family cell polarity regulator alpha |
| chr17_+_32582293 | 0.02 |
ENST00000580907.1 ENST00000225831.4 |
CCL2 |
chemokine (C-C motif) ligand 2 |
| chr11_-_77531752 | 0.02 |
ENST00000440064.2 ENST00000528095.1 |
RSF1 |
remodeling and spacing factor 1 |
| chr17_+_7461580 | 0.02 |
ENST00000483039.1 ENST00000396542.1 |
TNFSF13 |
tumor necrosis factor (ligand) superfamily, member 13 |
| chr2_-_109605663 | 0.02 |
ENST00000409271.1 ENST00000258443.2 ENST00000376651.1 |
EDAR |
ectodysplasin A receptor |
| chr17_-_17740325 | 0.02 |
ENST00000338854.5 |
SREBF1 |
sterol regulatory element binding transcription factor 1 |
| chr6_-_11232891 | 0.02 |
ENST00000379433.5 ENST00000379446.5 |
NEDD9 |
neural precursor cell expressed, developmentally down-regulated 9 |
| chr13_-_45915221 | 0.02 |
ENST00000309246.5 ENST00000379060.4 ENST00000379055.1 ENST00000527226.1 ENST00000379056.1 |
TPT1 |
tumor protein, translationally-controlled 1 |
| chrX_-_139866723 | 0.02 |
ENST00000370532.2 |
CDR1 |
cerebellar degeneration-related protein 1, 34kDa |
| chr14_-_37642016 | 0.02 |
ENST00000331299.5 |
SLC25A21 |
solute carrier family 25 (mitochondrial oxoadipate carrier), member 21 |
| chr2_+_173600514 | 0.02 |
ENST00000264111.6 |
RAPGEF4 |
Rap guanine nucleotide exchange factor (GEF) 4 |
| chr10_-_134121438 | 0.02 |
ENST00000298630.3 |
STK32C |
serine/threonine kinase 32C |
| chr6_-_41254403 | 0.02 |
ENST00000589614.1 ENST00000334475.6 ENST00000591620.1 ENST00000244709.4 |
TREM1 |
triggering receptor expressed on myeloid cells 1 |
| chr13_+_53226963 | 0.02 |
ENST00000343788.6 ENST00000535397.1 ENST00000310528.8 |
SUGT1 |
SGT1, suppressor of G2 allele of SKP1 (S. cerevisiae) |
| chr5_+_102201722 | 0.02 |
ENST00000274392.9 ENST00000455264.2 |
PAM |
peptidylglycine alpha-amidating monooxygenase |
| chr11_-_77532050 | 0.02 |
ENST00000308488.6 |
RSF1 |
remodeling and spacing factor 1 |
| chr7_-_150777874 | 0.02 |
ENST00000540185.1 |
FASTK |
Fas-activated serine/threonine kinase |
| chr7_-_15726296 | 0.01 |
ENST00000262041.5 |
MEOX2 |
mesenchyme homeobox 2 |
| chr1_-_116383738 | 0.01 |
ENST00000320238.3 |
NHLH2 |
nescient helix loop helix 2 |
| chrX_-_100546314 | 0.01 |
ENST00000356784.1 |
TAF7L |
TAF7-like RNA polymerase II, TATA box binding protein (TBP)-associated factor, 50kDa |
| chr11_-_119252359 | 0.01 |
ENST00000455332.2 |
USP2 |
ubiquitin specific peptidase 2 |
| chr7_+_31569365 | 0.01 |
ENST00000454513.1 ENST00000451887.2 |
CCDC129 |
coiled-coil domain containing 129 |
| chr5_-_88119580 | 0.01 |
ENST00000539796.1 |
MEF2C |
myocyte enhancer factor 2C |
| chr1_+_32608566 | 0.01 |
ENST00000545542.1 |
KPNA6 |
karyopherin alpha 6 (importin alpha 7) |
| chr7_-_122840015 | 0.01 |
ENST00000194130.2 |
SLC13A1 |
solute carrier family 13 (sodium/sulfate symporter), member 1 |
| chr2_+_173600565 | 0.01 |
ENST00000397081.3 |
RAPGEF4 |
Rap guanine nucleotide exchange factor (GEF) 4 |
| chr1_+_152691998 | 0.01 |
ENST00000368775.2 |
C1orf68 |
chromosome 1 open reading frame 68 |
| chr11_-_77531858 | 0.01 |
ENST00000360355.2 |
RSF1 |
remodeling and spacing factor 1 |
| chr8_+_123793633 | 0.01 |
ENST00000314393.4 |
ZHX2 |
zinc fingers and homeoboxes 2 |
| chr14_-_24553834 | 0.01 |
ENST00000397002.2 |
NRL |
neural retina leucine zipper |
| chr13_-_28545276 | 0.01 |
ENST00000381020.7 |
CDX2 |
caudal type homeobox 2 |
| chr12_-_53097247 | 0.00 |
ENST00000341809.3 ENST00000537195.1 |
KRT77 |
keratin 77 |
| chr15_-_64385981 | 0.00 |
ENST00000557835.1 ENST00000380290.3 ENST00000559950.1 |
FAM96A |
family with sequence similarity 96, member A |
| chr17_-_62308087 | 0.00 |
ENST00000583097.1 |
TEX2 |
testis expressed 2 |
| chr4_-_10686373 | 0.00 |
ENST00000442825.2 |
CLNK |
cytokine-dependent hematopoietic cell linker |
| chr12_+_56435637 | 0.00 |
ENST00000356464.5 ENST00000552361.1 |
RPS26 |
ribosomal protein S26 |
| chr15_-_88799661 | 0.00 |
ENST00000360948.2 ENST00000357724.2 ENST00000355254.2 ENST00000317501.3 |
NTRK3 |
neurotrophic tyrosine kinase, receptor, type 3 |
| chr4_-_46392290 | 0.00 |
ENST00000515082.1 |
GABRA2 |
gamma-aminobutyric acid (GABA) A receptor, alpha 2 |
| chr12_-_82152420 | 0.00 |
ENST00000552948.1 ENST00000548586.1 |
PPFIA2 |
protein tyrosine phosphatase, receptor type, f polypeptide (PTPRF), interacting protein (liprin), alpha 2 |
| chr15_-_64386120 | 0.00 |
ENST00000300030.3 |
FAM96A |
family with sequence similarity 96, member A |
| chr3_+_51575596 | 0.00 |
ENST00000409535.2 |
RAD54L2 |
RAD54-like 2 (S. cerevisiae) |
Gene Ontology Analysis
Gene overrepresentation in biological_process category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.1 | 0.8 | GO:0036343 | psychomotor behavior(GO:0036343) |
| 0.1 | 0.3 | GO:1903225 | negative regulation of endodermal cell differentiation(GO:1903225) |
| 0.1 | 0.5 | GO:0008218 | bioluminescence(GO:0008218) |
| 0.1 | 1.6 | GO:0071447 | cellular response to hydroperoxide(GO:0071447) |
| 0.1 | 0.3 | GO:0045872 | regulation of rhodopsin gene expression(GO:0007468) positive regulation of rhodopsin gene expression(GO:0045872) |
| 0.0 | 0.1 | GO:1903568 | negative regulation of protein localization to cilium(GO:1903565) regulation of protein localization to ciliary membrane(GO:1903567) negative regulation of protein localization to ciliary membrane(GO:1903568) |
| 0.0 | 0.1 | GO:0098935 | dendritic transport(GO:0098935) anterograde dendritic transport(GO:0098937) |
| 0.0 | 0.3 | GO:0008626 | granzyme-mediated apoptotic signaling pathway(GO:0008626) |
| 0.0 | 0.3 | GO:0070236 | negative regulation of activation-induced cell death of T cells(GO:0070236) |
| 0.0 | 0.1 | GO:0002874 | regulation of chronic inflammatory response to antigenic stimulus(GO:0002874) |
| 0.0 | 0.1 | GO:0016094 | polyprenol biosynthetic process(GO:0016094) |
| 0.0 | 0.3 | GO:0032020 | ISG15-protein conjugation(GO:0032020) |
| 0.0 | 0.1 | GO:0032639 | TRAIL production(GO:0032639) regulation of TRAIL production(GO:0032679) positive regulation of TRAIL production(GO:0032759) |
| 0.0 | 0.1 | GO:0030070 | insulin processing(GO:0030070) |
| 0.0 | 0.2 | GO:0044245 | polysaccharide digestion(GO:0044245) |
| 0.0 | 0.1 | GO:1901165 | positive regulation of trophoblast cell migration(GO:1901165) |
| 0.0 | 0.1 | GO:0034334 | adherens junction maintenance(GO:0034334) |
| 0.0 | 0.2 | GO:0048298 | positive regulation of isotype switching to IgA isotypes(GO:0048298) |
| 0.0 | 0.1 | GO:2000342 | negative regulation of chemokine (C-X-C motif) ligand 2 production(GO:2000342) |
| 0.0 | 0.1 | GO:0006106 | fumarate metabolic process(GO:0006106) |
| 0.0 | 0.0 | GO:0002277 | myeloid dendritic cell activation involved in immune response(GO:0002277) |
| 0.0 | 0.0 | GO:0021798 | forebrain dorsal/ventral pattern formation(GO:0021798) |
| 0.0 | 0.1 | GO:1904059 | regulation of locomotor rhythm(GO:1904059) |
| 0.0 | 0.2 | GO:0048739 | cardiac muscle fiber development(GO:0048739) |
| 0.0 | 0.1 | GO:0060235 | lens induction in camera-type eye(GO:0060235) |
| 0.0 | 0.0 | GO:0018032 | protein amidation(GO:0018032) |
| 0.0 | 0.1 | GO:0033353 | S-adenosylmethionine cycle(GO:0033353) |
| 0.0 | 0.1 | GO:0097646 | calcitonin family receptor signaling pathway(GO:0097646) amylin receptor signaling pathway(GO:0097647) |
Gene overrepresentation in cellular_component category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.1 | 0.3 | GO:0005588 | collagen type V trimer(GO:0005588) |
| 0.0 | 0.2 | GO:1990111 | spermatoproteasome complex(GO:1990111) |
| 0.0 | 0.3 | GO:0005687 | U4 snRNP(GO:0005687) |
| 0.0 | 0.1 | GO:1903439 | calcitonin family receptor complex(GO:1903439) amylin receptor complex(GO:1903440) |
| 0.0 | 0.0 | GO:0097679 | other organism cytoplasm(GO:0097679) |
| 0.0 | 0.1 | GO:0089701 | U2AF(GO:0089701) |
| 0.0 | 0.0 | GO:0031213 | RSF complex(GO:0031213) |
Gene overrepresentation in molecular_function category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.0 | 1.6 | GO:0017075 | syntaxin-1 binding(GO:0017075) |
| 0.0 | 0.3 | GO:0004839 | ubiquitin activating enzyme activity(GO:0004839) |
| 0.0 | 0.3 | GO:0043426 | MRF binding(GO:0043426) |
| 0.0 | 0.1 | GO:0033754 | indoleamine 2,3-dioxygenase activity(GO:0033754) |
| 0.0 | 0.1 | GO:0002094 | polyprenyltransferase activity(GO:0002094) |
| 0.0 | 0.2 | GO:0031728 | CCR3 chemokine receptor binding(GO:0031728) |
| 0.0 | 0.1 | GO:0033867 | Fas-activated serine/threonine kinase activity(GO:0033867) |
| 0.0 | 0.1 | GO:0008113 | peptide-methionine (S)-S-oxide reductase activity(GO:0008113) |
| 0.0 | 0.1 | GO:0097643 | amylin receptor activity(GO:0097643) |
| 0.0 | 0.1 | GO:0004013 | adenosylhomocysteinase activity(GO:0004013) trialkylsulfonium hydrolase activity(GO:0016802) |
| 0.0 | 0.1 | GO:0060002 | plus-end directed microfilament motor activity(GO:0060002) |
| 0.0 | 0.1 | GO:0051373 | FATZ binding(GO:0051373) |
| 0.0 | 0.3 | GO:0043522 | leucine zipper domain binding(GO:0043522) |
| 0.0 | 0.4 | GO:0001871 | pattern binding(GO:0001871) polysaccharide binding(GO:0030247) |
| 0.0 | 0.0 | GO:0004504 | peptidylglycine monooxygenase activity(GO:0004504) peptidylamidoglycolate lyase activity(GO:0004598) |
| 0.0 | 0.2 | GO:0004568 | chitinase activity(GO:0004568) |
| 0.0 | 0.0 | GO:0030550 | acetylcholine receptor inhibitor activity(GO:0030550) |
| 0.0 | 0.1 | GO:0036042 | long-chain fatty acyl-CoA binding(GO:0036042) |
| 0.0 | 0.1 | GO:1990460 | leptin receptor binding(GO:1990460) |
| 0.0 | 0.1 | GO:0030628 | pre-mRNA 3'-splice site binding(GO:0030628) |
| 0.0 | 0.2 | GO:0042301 | phosphate ion binding(GO:0042301) |
Gene overrepresentation in C2:CP category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.0 | 1.6 | PID IFNG PATHWAY | IFN-gamma pathway |
Gene overrepresentation in C2:CP:REACTOME category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.1 | 1.6 | REACTOME ROLE OF DCC IN REGULATING APOPTOSIS | Genes involved in Role of DCC in regulating apoptosis |
| 0.0 | 0.5 | REACTOME HS GAG DEGRADATION | Genes involved in HS-GAG degradation |
| 0.0 | 0.2 | REACTOME PASSIVE TRANSPORT BY AQUAPORINS | Genes involved in Passive Transport by Aquaporins |