Project
Epithelial-Mesenchymal Transition, human (Scheel, 2011)
Navigation
Downloads
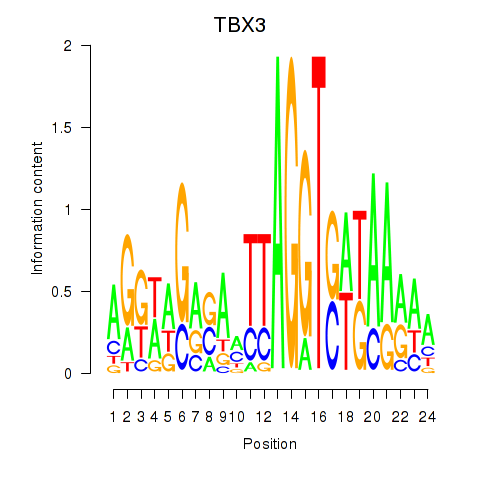
Results for TBX3
Z-value: 0.48
Transcription factors associated with TBX3
| Gene Symbol | Gene ID | Gene Info |
|---|---|---|
|
TBX3
|
ENSG00000135111.10 | TBX3 |
Activity-expression correlation:
| Gene | Promoter | Pearson corr. coef. | P-value | Plot |
|---|---|---|---|---|
| TBX3 | hg19_v2_chr12_-_115121962_115121987 | 0.14 | 7.4e-01 | Click! |
Activity profile of TBX3 motif
Sorted Z-values of TBX3 motif
Network of associatons between targets according to the STRING database.
First level regulatory network of TBX3
| Promoter | Score | Transcript | Gene | Gene Info |
|---|---|---|---|---|
| chr9_+_125281420 | 0.33 |
ENST00000340750.1 |
OR1J4 |
olfactory receptor, family 1, subfamily J, member 4 |
| chr21_+_25801041 | 0.33 |
ENST00000453784.2 ENST00000423581.1 |
AP000476.1 |
AP000476.1 |
| chr1_-_242612779 | 0.29 |
ENST00000427495.1 |
PLD5 |
phospholipase D family, member 5 |
| chr14_-_106471723 | 0.22 |
ENST00000390595.2 |
IGHV1-3 |
immunoglobulin heavy variable 1-3 |
| chr12_-_8815404 | 0.20 |
ENST00000359478.2 ENST00000396549.2 |
MFAP5 |
microfibrillar associated protein 5 |
| chr19_-_36004543 | 0.20 |
ENST00000339686.3 ENST00000447113.2 ENST00000440396.1 |
DMKN |
dermokine |
| chr9_-_39288092 | 0.17 |
ENST00000323947.7 ENST00000297668.6 ENST00000377656.2 ENST00000377659.1 |
CNTNAP3 |
contactin associated protein-like 3 |
| chr7_-_93519471 | 0.17 |
ENST00000451238.1 |
TFPI2 |
tissue factor pathway inhibitor 2 |
| chr3_+_124223586 | 0.16 |
ENST00000393496.1 |
KALRN |
kalirin, RhoGEF kinase |
| chr2_-_113594279 | 0.15 |
ENST00000416750.1 ENST00000418817.1 ENST00000263341.2 |
IL1B |
interleukin 1, beta |
| chr17_+_74372662 | 0.15 |
ENST00000591651.1 ENST00000545180.1 |
SPHK1 |
sphingosine kinase 1 |
| chr22_-_41258074 | 0.14 |
ENST00000307221.4 |
DNAJB7 |
DnaJ (Hsp40) homolog, subfamily B, member 7 |
| chr4_+_178230985 | 0.14 |
ENST00000264596.3 |
NEIL3 |
nei endonuclease VIII-like 3 (E. coli) |
| chr1_+_26869597 | 0.13 |
ENST00000530003.1 |
RPS6KA1 |
ribosomal protein S6 kinase, 90kDa, polypeptide 1 |
| chr7_+_98476095 | 0.13 |
ENST00000359863.4 ENST00000355540.3 |
TRRAP |
transformation/transcription domain-associated protein |
| chr3_-_121264848 | 0.13 |
ENST00000264233.5 |
POLQ |
polymerase (DNA directed), theta |
| chr4_-_80247162 | 0.13 |
ENST00000286794.4 |
NAA11 |
N(alpha)-acetyltransferase 11, NatA catalytic subunit |
| chr1_-_183560011 | 0.13 |
ENST00000367536.1 |
NCF2 |
neutrophil cytosolic factor 2 |
| chr19_-_33360647 | 0.13 |
ENST00000590341.1 ENST00000587772.1 ENST00000023064.4 |
SLC7A9 |
solute carrier family 7 (amino acid transporter light chain, bo,+ system), member 9 |
| chr11_+_19138670 | 0.13 |
ENST00000446113.2 ENST00000399351.3 |
ZDHHC13 |
zinc finger, DHHC-type containing 13 |
| chr19_-_51530916 | 0.12 |
ENST00000594768.1 |
KLK11 |
kallikrein-related peptidase 11 |
| chr19_-_51531272 | 0.12 |
ENST00000319720.7 |
KLK11 |
kallikrein-related peptidase 11 |
| chr1_+_212208919 | 0.12 |
ENST00000366991.4 ENST00000542077.1 |
DTL |
denticleless E3 ubiquitin protein ligase homolog (Drosophila) |
| chr6_-_32821599 | 0.12 |
ENST00000354258.4 |
TAP1 |
transporter 1, ATP-binding cassette, sub-family B (MDR/TAP) |
| chr6_-_29324054 | 0.12 |
ENST00000543825.1 |
OR5V1 |
olfactory receptor, family 5, subfamily V, member 1 |
| chr9_-_114090713 | 0.12 |
ENST00000302681.1 ENST00000374428.1 |
OR2K2 |
olfactory receptor, family 2, subfamily K, member 2 |
| chr15_-_75017711 | 0.11 |
ENST00000567032.1 ENST00000564596.1 ENST00000566503.1 ENST00000395049.4 ENST00000395048.2 ENST00000379727.3 |
CYP1A1 |
cytochrome P450, family 1, subfamily A, polypeptide 1 |
| chr12_-_88535842 | 0.11 |
ENST00000550962.1 ENST00000552810.1 |
CEP290 |
centrosomal protein 290kDa |
| chr19_-_51531210 | 0.11 |
ENST00000391804.3 |
KLK11 |
kallikrein-related peptidase 11 |
| chr4_+_103790462 | 0.10 |
ENST00000503643.1 |
CISD2 |
CDGSH iron sulfur domain 2 |
| chr16_+_50099852 | 0.10 |
ENST00000299192.7 ENST00000285767.4 |
HEATR3 |
HEAT repeat containing 3 |
| chr2_-_89310012 | 0.10 |
ENST00000493819.1 |
IGKV1-9 |
immunoglobulin kappa variable 1-9 |
| chr11_-_1629693 | 0.10 |
ENST00000399685.1 |
KRTAP5-3 |
keratin associated protein 5-3 |
| chr12_+_88536067 | 0.09 |
ENST00000549011.1 ENST00000266712.6 ENST00000551088.1 |
TMTC3 |
transmembrane and tetratricopeptide repeat containing 3 |
| chr1_+_220267429 | 0.09 |
ENST00000366922.1 ENST00000302637.5 |
IARS2 |
isoleucyl-tRNA synthetase 2, mitochondrial |
| chr2_+_17721920 | 0.09 |
ENST00000295156.4 |
VSNL1 |
visinin-like 1 |
| chr14_-_74416829 | 0.09 |
ENST00000534936.1 |
FAM161B |
family with sequence similarity 161, member B |
| chr11_+_34643600 | 0.09 |
ENST00000530286.1 ENST00000533754.1 |
EHF |
ets homologous factor |
| chr11_+_35198243 | 0.09 |
ENST00000528455.1 |
CD44 |
CD44 molecule (Indian blood group) |
| chr3_+_189349162 | 0.09 |
ENST00000264731.3 ENST00000382063.4 ENST00000418709.2 ENST00000320472.5 ENST00000392460.3 ENST00000440651.2 |
TP63 |
tumor protein p63 |
| chr19_-_47291843 | 0.09 |
ENST00000542575.2 |
SLC1A5 |
solute carrier family 1 (neutral amino acid transporter), member 5 |
| chr19_-_42931567 | 0.09 |
ENST00000244289.4 |
LIPE |
lipase, hormone-sensitive |
| chrX_+_149887090 | 0.08 |
ENST00000538506.1 |
MTMR1 |
myotubularin related protein 1 |
| chr16_-_67360662 | 0.08 |
ENST00000304372.5 |
KCTD19 |
potassium channel tetramerization domain containing 19 |
| chr4_+_15704573 | 0.08 |
ENST00000265016.4 |
BST1 |
bone marrow stromal cell antigen 1 |
| chr22_+_45680822 | 0.08 |
ENST00000216211.4 ENST00000396082.2 |
UPK3A |
uroplakin 3A |
| chr16_-_68057770 | 0.08 |
ENST00000332395.5 |
DDX28 |
DEAD (Asp-Glu-Ala-Asp) box polypeptide 28 |
| chr2_-_89266286 | 0.08 |
ENST00000464162.1 |
IGKV1-6 |
immunoglobulin kappa variable 1-6 |
| chrX_-_54824673 | 0.08 |
ENST00000218436.6 |
ITIH6 |
inter-alpha-trypsin inhibitor heavy chain family, member 6 |
| chrX_+_44703249 | 0.08 |
ENST00000339042.4 |
DUSP21 |
dual specificity phosphatase 21 |
| chr20_-_5931109 | 0.08 |
ENST00000203001.2 |
TRMT6 |
tRNA methyltransferase 6 homolog (S. cerevisiae) |
| chr16_+_66914264 | 0.08 |
ENST00000311765.2 ENST00000568869.1 ENST00000561704.1 ENST00000568398.1 ENST00000566776.1 |
PDP2 |
pyruvate dehyrogenase phosphatase catalytic subunit 2 |
| chr14_-_92333873 | 0.08 |
ENST00000435962.2 |
TC2N |
tandem C2 domains, nuclear |
| chr3_-_42452050 | 0.08 |
ENST00000441172.1 ENST00000287748.3 |
LYZL4 |
lysozyme-like 4 |
| chr11_+_65082289 | 0.07 |
ENST00000279249.2 |
CDC42EP2 |
CDC42 effector protein (Rho GTPase binding) 2 |
| chr22_+_41487711 | 0.07 |
ENST00000263253.7 |
EP300 |
E1A binding protein p300 |
| chr11_-_62607036 | 0.07 |
ENST00000311713.7 ENST00000278856.4 |
WDR74 |
WD repeat domain 74 |
| chr13_-_103719196 | 0.07 |
ENST00000245312.3 |
SLC10A2 |
solute carrier family 10 (sodium/bile acid cotransporter), member 2 |
| chr10_+_71561630 | 0.07 |
ENST00000398974.3 ENST00000398971.3 ENST00000398968.3 ENST00000398966.3 ENST00000398964.3 ENST00000398969.3 ENST00000356340.3 ENST00000398972.3 ENST00000398973.3 |
COL13A1 |
collagen, type XIII, alpha 1 |
| chr2_-_209118974 | 0.07 |
ENST00000415913.1 ENST00000415282.1 ENST00000446179.1 |
IDH1 |
isocitrate dehydrogenase 1 (NADP+), soluble |
| chr19_+_9296279 | 0.07 |
ENST00000344248.2 |
OR7D2 |
olfactory receptor, family 7, subfamily D, member 2 |
| chr12_+_12223867 | 0.07 |
ENST00000308721.5 |
BCL2L14 |
BCL2-like 14 (apoptosis facilitator) |
| chr20_-_5931051 | 0.07 |
ENST00000453074.2 |
TRMT6 |
tRNA methyltransferase 6 homolog (S. cerevisiae) |
| chr12_-_88535747 | 0.07 |
ENST00000309041.7 |
CEP290 |
centrosomal protein 290kDa |
| chrX_+_140984021 | 0.07 |
ENST00000409007.1 |
MAGEC3 |
melanoma antigen family C, 3 |
| chr3_+_118866222 | 0.07 |
ENST00000490594.1 |
RP11-484M3.5 |
Uncharacterized protein |
| chr7_+_150434430 | 0.07 |
ENST00000358647.3 |
GIMAP5 |
GTPase, IMAP family member 5 |
| chr1_-_157014865 | 0.07 |
ENST00000361409.2 |
ARHGEF11 |
Rho guanine nucleotide exchange factor (GEF) 11 |
| chr16_+_84801852 | 0.07 |
ENST00000569925.1 ENST00000567526.1 |
USP10 |
ubiquitin specific peptidase 10 |
| chrX_-_65259900 | 0.07 |
ENST00000412866.2 |
VSIG4 |
V-set and immunoglobulin domain containing 4 |
| chr19_-_3478477 | 0.07 |
ENST00000591708.1 |
C19orf77 |
chromosome 19 open reading frame 77 |
| chr15_-_74501310 | 0.07 |
ENST00000423167.2 ENST00000432245.2 |
STRA6 |
stimulated by retinoic acid 6 |
| chrX_-_65259914 | 0.07 |
ENST00000374737.4 ENST00000455586.2 |
VSIG4 |
V-set and immunoglobulin domain containing 4 |
| chr9_-_100954910 | 0.06 |
ENST00000375077.4 |
CORO2A |
coronin, actin binding protein, 2A |
| chr15_+_40861487 | 0.06 |
ENST00000315616.7 ENST00000559271.1 |
RPUSD2 |
RNA pseudouridylate synthase domain containing 2 |
| chr11_+_71259466 | 0.06 |
ENST00000528743.2 |
KRTAP5-9 |
keratin associated protein 5-9 |
| chr1_-_182921119 | 0.06 |
ENST00000423786.1 |
SHCBP1L |
SHC SH2-domain binding protein 1-like |
| chr10_+_71561649 | 0.06 |
ENST00000398978.3 ENST00000354547.3 ENST00000357811.3 |
COL13A1 |
collagen, type XIII, alpha 1 |
| chr12_+_1800179 | 0.06 |
ENST00000357103.4 |
ADIPOR2 |
adiponectin receptor 2 |
| chr3_+_149191723 | 0.06 |
ENST00000305354.4 |
TM4SF4 |
transmembrane 4 L six family member 4 |
| chr2_+_65215604 | 0.06 |
ENST00000531327.1 |
SLC1A4 |
solute carrier family 1 (glutamate/neutral amino acid transporter), member 4 |
| chr2_-_209119831 | 0.06 |
ENST00000345146.2 |
IDH1 |
isocitrate dehydrogenase 1 (NADP+), soluble |
| chr12_+_129028500 | 0.06 |
ENST00000315208.8 |
TMEM132C |
transmembrane protein 132C |
| chr12_+_133758115 | 0.06 |
ENST00000541009.2 ENST00000592241.1 |
ZNF268 |
zinc finger protein 268 |
| chr16_-_19896832 | 0.06 |
ENST00000537135.1 ENST00000564449.1 |
GPRC5B |
G protein-coupled receptor, family C, group 5, member B |
| chr1_-_154461642 | 0.06 |
ENST00000555188.1 |
SHE |
Src homology 2 domain containing E |
| chr21_-_43346790 | 0.05 |
ENST00000329623.7 |
C2CD2 |
C2 calcium-dependent domain containing 2 |
| chrX_+_73524020 | 0.05 |
ENST00000339534.2 |
ZCCHC13 |
zinc finger, CCHC domain containing 13 |
| chr6_-_35888905 | 0.05 |
ENST00000510290.1 ENST00000423325.2 ENST00000373822.1 |
SRPK1 |
SRSF protein kinase 1 |
| chr14_-_106642049 | 0.05 |
ENST00000390605.2 |
IGHV1-18 |
immunoglobulin heavy variable 1-18 |
| chr11_+_71249071 | 0.05 |
ENST00000398534.3 |
KRTAP5-8 |
keratin associated protein 5-8 |
| chr7_+_73106926 | 0.05 |
ENST00000453316.1 |
WBSCR22 |
Williams Beuren syndrome chromosome region 22 |
| chr1_+_171750776 | 0.05 |
ENST00000458517.1 ENST00000362019.3 ENST00000367737.5 ENST00000361735.3 |
METTL13 |
methyltransferase like 13 |
| chr7_-_86595190 | 0.05 |
ENST00000398276.2 ENST00000416314.1 ENST00000425689.1 |
KIAA1324L |
KIAA1324-like |
| chr2_-_165698662 | 0.05 |
ENST00000194871.6 ENST00000445474.2 |
COBLL1 |
cordon-bleu WH2 repeat protein-like 1 |
| chr1_-_241799232 | 0.05 |
ENST00000366553.1 |
CHML |
choroideremia-like (Rab escort protein 2) |
| chr2_-_4021609 | 0.05 |
ENST00000425949.1 ENST00000453880.1 ENST00000588796.1 |
AC107070.1 |
AC107070.1 |
| chr9_+_273038 | 0.05 |
ENST00000487230.1 ENST00000469391.1 |
DOCK8 |
dedicator of cytokinesis 8 |
| chr15_-_59665062 | 0.05 |
ENST00000288235.4 |
MYO1E |
myosin IE |
| chr11_+_71238313 | 0.05 |
ENST00000398536.4 |
KRTAP5-7 |
keratin associated protein 5-7 |
| chr7_-_134896234 | 0.05 |
ENST00000354475.4 ENST00000344400.5 |
WDR91 |
WD repeat domain 91 |
| chr1_-_110613276 | 0.05 |
ENST00000369792.4 |
ALX3 |
ALX homeobox 3 |
| chr5_+_145826867 | 0.05 |
ENST00000296702.5 ENST00000394421.2 |
TCERG1 |
transcription elongation regulator 1 |
| chr2_-_4021643 | 0.05 |
ENST00000454455.1 |
AC107070.1 |
AC107070.1 |
| chr8_-_22014339 | 0.05 |
ENST00000306317.2 |
LGI3 |
leucine-rich repeat LGI family, member 3 |
| chr9_-_95087604 | 0.05 |
ENST00000542613.1 ENST00000542053.1 ENST00000358855.4 ENST00000545558.1 ENST00000432670.2 ENST00000433029.2 ENST00000411621.2 |
NOL8 |
nucleolar protein 8 |
| chr17_-_41132410 | 0.05 |
ENST00000409446.3 ENST00000453594.1 ENST00000409399.1 ENST00000421990.2 |
PTGES3L PTGES3L-AARSD1 |
prostaglandin E synthase 3 (cytosolic)-like PTGES3L-AARSD1 readthrough |
| chr11_-_104893863 | 0.05 |
ENST00000260315.3 ENST00000526056.1 ENST00000531367.1 ENST00000456094.1 ENST00000444749.2 ENST00000393141.2 ENST00000418434.1 ENST00000393139.2 |
CASP5 |
caspase 5, apoptosis-related cysteine peptidase |
| chr4_-_48908822 | 0.05 |
ENST00000508632.1 |
OCIAD2 |
OCIA domain containing 2 |
| chr10_+_32856764 | 0.05 |
ENST00000375030.2 ENST00000375028.3 |
C10orf68 |
Homo sapiens coiled-coil domain containing 7 (CCDC7), transcript variant 5, mRNA. |
| chr12_+_16500599 | 0.04 |
ENST00000535309.1 ENST00000540056.1 ENST00000396209.1 ENST00000540126.1 |
MGST1 |
microsomal glutathione S-transferase 1 |
| chr12_-_53729525 | 0.04 |
ENST00000303846.3 |
SP7 |
Sp7 transcription factor |
| chr8_-_20161466 | 0.04 |
ENST00000381569.1 |
LZTS1 |
leucine zipper, putative tumor suppressor 1 |
| chr10_+_47894572 | 0.04 |
ENST00000355876.5 |
FAM21B |
family with sequence similarity 21, member B |
| chr7_+_99971129 | 0.04 |
ENST00000394000.2 ENST00000350573.2 |
PILRA |
paired immunoglobin-like type 2 receptor alpha |
| chr3_+_57741957 | 0.04 |
ENST00000295951.3 |
SLMAP |
sarcolemma associated protein |
| chr4_-_48908737 | 0.04 |
ENST00000381464.2 |
OCIAD2 |
OCIA domain containing 2 |
| chr19_-_55791058 | 0.04 |
ENST00000587959.1 ENST00000585927.1 ENST00000587922.1 ENST00000585698.1 |
HSPBP1 |
HSPA (heat shock 70kDa) binding protein, cytoplasmic cochaperone 1 |
| chr11_+_77774897 | 0.04 |
ENST00000281030.2 |
THRSP |
thyroid hormone responsive |
| chr2_+_90153696 | 0.04 |
ENST00000417279.2 |
IGKV3D-15 |
immunoglobulin kappa variable 3D-15 (gene/pseudogene) |
| chr12_+_16500571 | 0.04 |
ENST00000543076.1 ENST00000396210.3 |
MGST1 |
microsomal glutathione S-transferase 1 |
| chr11_-_18258342 | 0.04 |
ENST00000278222.4 |
SAA4 |
serum amyloid A4, constitutive |
| chr4_-_6202291 | 0.04 |
ENST00000282924.5 |
JAKMIP1 |
janus kinase and microtubule interacting protein 1 |
| chr19_+_53868940 | 0.04 |
ENST00000474037.1 ENST00000491101.1 ENST00000467003.1 ENST00000475179.1 ENST00000593918.1 |
ZNF525 |
zinc finger protein 525 |
| chr2_+_202937972 | 0.04 |
ENST00000541917.1 ENST00000295844.3 |
AC079354.1 |
uncharacterized protein KIAA2012 |
| chr12_+_124155652 | 0.04 |
ENST00000426174.2 ENST00000303372.5 |
TCTN2 |
tectonic family member 2 |
| chr13_-_73356234 | 0.04 |
ENST00000545453.1 |
DIS3 |
DIS3 mitotic control homolog (S. cerevisiae) |
| chr4_-_69215699 | 0.04 |
ENST00000510746.1 ENST00000344157.4 ENST00000355665.3 |
YTHDC1 |
YTH domain containing 1 |
| chr10_-_112064665 | 0.04 |
ENST00000369603.5 |
SMNDC1 |
survival motor neuron domain containing 1 |
| chr15_-_50411412 | 0.04 |
ENST00000284509.6 |
ATP8B4 |
ATPase, class I, type 8B, member 4 |
| chr21_-_15918618 | 0.04 |
ENST00000400564.1 ENST00000400566.1 |
SAMSN1 |
SAM domain, SH3 domain and nuclear localization signals 1 |
| chr15_-_74504560 | 0.04 |
ENST00000449139.2 |
STRA6 |
stimulated by retinoic acid 6 |
| chr15_-_74504597 | 0.04 |
ENST00000416286.3 |
STRA6 |
stimulated by retinoic acid 6 |
| chr7_-_138386097 | 0.04 |
ENST00000421622.1 |
SVOPL |
SVOP-like |
| chr22_-_42915788 | 0.04 |
ENST00000323013.6 |
RRP7A |
ribosomal RNA processing 7 homolog A (S. cerevisiae) |
| chr10_-_90611566 | 0.04 |
ENST00000371930.4 |
ANKRD22 |
ankyrin repeat domain 22 |
| chr8_-_22014255 | 0.04 |
ENST00000424267.2 |
LGI3 |
leucine-rich repeat LGI family, member 3 |
| chr4_+_123300664 | 0.04 |
ENST00000388725.2 |
ADAD1 |
adenosine deaminase domain containing 1 (testis-specific) |
| chr10_+_51827648 | 0.04 |
ENST00000351071.6 ENST00000314664.7 ENST00000282633.5 |
FAM21A |
family with sequence similarity 21, member A |
| chr3_-_46068969 | 0.04 |
ENST00000542109.1 ENST00000395946.2 |
XCR1 |
chemokine (C motif) receptor 1 |
| chr14_+_77924204 | 0.04 |
ENST00000555133.1 |
AHSA1 |
AHA1, activator of heat shock 90kDa protein ATPase homolog 1 (yeast) |
| chr7_-_19748640 | 0.04 |
ENST00000222567.5 |
TWISTNB |
TWIST neighbor |
| chrX_+_51942963 | 0.04 |
ENST00000375625.3 |
RP11-363G10.2 |
RP11-363G10.2 |
| chr4_+_187148556 | 0.04 |
ENST00000264690.6 ENST00000446598.2 ENST00000414291.1 ENST00000513864.1 |
KLKB1 |
kallikrein B, plasma (Fletcher factor) 1 |
| chr19_-_51538118 | 0.04 |
ENST00000529888.1 |
KLK12 |
kallikrein-related peptidase 12 |
| chr7_-_112726393 | 0.03 |
ENST00000449591.1 ENST00000449735.1 ENST00000438062.1 ENST00000424100.1 |
GPR85 |
G protein-coupled receptor 85 |
| chr19_-_10491130 | 0.03 |
ENST00000530829.1 ENST00000529370.1 |
TYK2 |
tyrosine kinase 2 |
| chr19_-_16682987 | 0.03 |
ENST00000431408.1 ENST00000436553.2 ENST00000595753.1 |
SLC35E1 |
solute carrier family 35, member E1 |
| chr2_+_216974020 | 0.03 |
ENST00000392132.2 ENST00000417391.1 |
XRCC5 |
X-ray repair complementing defective repair in Chinese hamster cells 5 (double-strand-break rejoining) |
| chr14_+_96722152 | 0.03 |
ENST00000216629.6 |
BDKRB1 |
bradykinin receptor B1 |
| chr15_+_79603404 | 0.03 |
ENST00000299705.5 |
TMED3 |
transmembrane emp24 protein transport domain containing 3 |
| chr22_+_23063100 | 0.03 |
ENST00000390309.2 |
IGLV3-19 |
immunoglobulin lambda variable 3-19 |
| chr1_+_36396313 | 0.03 |
ENST00000324350.5 |
AGO3 |
argonaute RISC catalytic component 3 |
| chr21_-_46340807 | 0.03 |
ENST00000397846.3 ENST00000524251.1 ENST00000522688.1 |
ITGB2 |
integrin, beta 2 (complement component 3 receptor 3 and 4 subunit) |
| chr20_+_37209820 | 0.03 |
ENST00000537425.1 ENST00000373348.3 ENST00000416116.1 |
ADIG |
adipogenin |
| chr1_-_152552980 | 0.03 |
ENST00000368787.3 |
LCE3D |
late cornified envelope 3D |
| chrX_+_140968348 | 0.03 |
ENST00000536088.1 ENST00000443323.2 |
MAGEC3 |
melanoma antigen family C, 3 |
| chr12_+_48876275 | 0.03 |
ENST00000314014.2 |
C12orf54 |
chromosome 12 open reading frame 54 |
| chr4_-_48908805 | 0.03 |
ENST00000273860.4 |
OCIAD2 |
OCIA domain containing 2 |
| chr2_+_90192768 | 0.03 |
ENST00000390275.2 |
IGKV1D-13 |
immunoglobulin kappa variable 1D-13 |
| chr1_-_155959853 | 0.03 |
ENST00000462460.2 ENST00000368316.1 |
ARHGEF2 |
Rho/Rac guanine nucleotide exchange factor (GEF) 2 |
| chr4_+_155484155 | 0.03 |
ENST00000509493.1 |
FGB |
fibrinogen beta chain |
| chr10_+_71561704 | 0.03 |
ENST00000520267.1 |
COL13A1 |
collagen, type XIII, alpha 1 |
| chr22_-_36031181 | 0.03 |
ENST00000594060.1 |
AL049747.1 |
AL049747.1 |
| chr11_-_122932730 | 0.03 |
ENST00000532182.1 ENST00000524590.1 ENST00000528292.1 ENST00000533540.1 ENST00000525463.1 |
HSPA8 |
heat shock 70kDa protein 8 |
| chr2_+_66666432 | 0.03 |
ENST00000495021.2 |
MEIS1 |
Meis homeobox 1 |
| chr19_-_51538148 | 0.03 |
ENST00000319590.4 ENST00000250351.4 |
KLK12 |
kallikrein-related peptidase 12 |
| chr4_-_190948409 | 0.03 |
ENST00000504750.1 ENST00000378763.1 |
FRG2 |
FSHD region gene 2 |
| chr19_+_19322758 | 0.03 |
ENST00000252575.6 |
NCAN |
neurocan |
| chr7_+_150264365 | 0.03 |
ENST00000255945.2 ENST00000461940.1 |
GIMAP4 |
GTPase, IMAP family member 4 |
| chr20_+_34359905 | 0.03 |
ENST00000374012.3 ENST00000439301.1 ENST00000339089.6 ENST00000374000.4 |
PHF20 |
PHD finger protein 20 |
| chr9_+_35490101 | 0.03 |
ENST00000361226.3 |
RUSC2 |
RUN and SH3 domain containing 2 |
| chr11_+_130542851 | 0.03 |
ENST00000317019.2 |
C11orf44 |
chromosome 11 open reading frame 44 |
| chr17_-_7017559 | 0.03 |
ENST00000446679.2 |
ASGR2 |
asialoglycoprotein receptor 2 |
| chr10_+_46222648 | 0.03 |
ENST00000336378.4 ENST00000540872.1 ENST00000537517.1 ENST00000374362.2 ENST00000359860.4 ENST00000420848.1 |
FAM21C |
family with sequence similarity 21, member C |
| chr10_+_47894023 | 0.03 |
ENST00000358474.5 |
FAM21B |
family with sequence similarity 21, member B |
| chr17_+_20483037 | 0.03 |
ENST00000399044.1 |
CDRT15L2 |
CMT1A duplicated region transcript 15-like 2 |
| chr8_+_23386305 | 0.03 |
ENST00000519973.1 |
SLC25A37 |
solute carrier family 25 (mitochondrial iron transporter), member 37 |
| chr7_-_99381798 | 0.03 |
ENST00000415003.1 ENST00000354593.2 |
CYP3A4 |
cytochrome P450, family 3, subfamily A, polypeptide 4 |
| chrX_-_101726732 | 0.03 |
ENST00000457521.2 ENST00000412230.2 ENST00000453326.2 |
NXF2B TCP11X2 |
nuclear RNA export factor 2B t-complex 11 family, X-linked 2 |
| chrX_+_101470280 | 0.03 |
ENST00000395088.2 ENST00000330252.5 ENST00000333110.5 |
NXF2 TCP11X1 |
nuclear RNA export factor 2 t-complex 11 family, X-linked 1 |
| chr22_+_23237555 | 0.03 |
ENST00000390321.2 |
IGLC1 |
immunoglobulin lambda constant 1 (Mcg marker) |
| chr14_+_90722886 | 0.03 |
ENST00000543772.2 |
PSMC1 |
proteasome (prosome, macropain) 26S subunit, ATPase, 1 |
| chr19_+_1077393 | 0.03 |
ENST00000590577.1 |
HMHA1 |
histocompatibility (minor) HA-1 |
| chr11_-_33744256 | 0.03 |
ENST00000415002.2 ENST00000437761.2 ENST00000445143.2 |
CD59 |
CD59 molecule, complement regulatory protein |
| chr19_+_58281014 | 0.03 |
ENST00000391702.3 ENST00000598885.1 ENST00000598183.1 ENST00000396154.2 ENST00000599802.1 ENST00000396150.4 |
ZNF586 |
zinc finger protein 586 |
| chr11_-_33744487 | 0.03 |
ENST00000426650.2 |
CD59 |
CD59 molecule, complement regulatory protein |
| chr12_-_54758251 | 0.03 |
ENST00000267015.3 ENST00000551809.1 |
GPR84 |
G protein-coupled receptor 84 |
| chr2_-_26569611 | 0.03 |
ENST00000541401.1 ENST00000433584.1 ENST00000333478.6 |
GPR113 |
G protein-coupled receptor 113 |
| chr1_+_154229547 | 0.03 |
ENST00000428595.1 |
UBAP2L |
ubiquitin associated protein 2-like |
| chrX_+_46306714 | 0.02 |
ENST00000298190.6 ENST00000377919.2 ENST00000476762.1 ENST00000478600.1 |
KRBOX4 |
KRAB box domain containing 4 |
| chr14_+_90722839 | 0.02 |
ENST00000261303.8 ENST00000553835.1 |
PSMC1 |
proteasome (prosome, macropain) 26S subunit, ATPase, 1 |
| chr1_-_11865351 | 0.02 |
ENST00000413656.1 ENST00000376585.1 ENST00000423400.1 ENST00000431243.1 |
MTHFR |
methylenetetrahydrofolate reductase (NAD(P)H) |
| chr6_+_6588316 | 0.02 |
ENST00000379953.2 |
LY86 |
lymphocyte antigen 86 |
| chr17_+_39411636 | 0.02 |
ENST00000394008.1 |
KRTAP9-9 |
keratin associated protein 9-9 |
| chr17_+_41132564 | 0.02 |
ENST00000361677.1 ENST00000589705.1 |
RUNDC1 |
RUN domain containing 1 |
| chr6_-_134861089 | 0.02 |
ENST00000606039.1 |
RP11-557H15.4 |
RP11-557H15.4 |
| chr11_-_122931881 | 0.02 |
ENST00000526110.1 ENST00000227378.3 |
HSPA8 |
heat shock 70kDa protein 8 |
| chr20_-_1472029 | 0.02 |
ENST00000359801.3 |
SIRPB2 |
signal-regulatory protein beta 2 |
| chr4_-_46911248 | 0.02 |
ENST00000355591.3 ENST00000505102.1 |
COX7B2 |
cytochrome c oxidase subunit VIIb2 |
Gene Ontology Analysis
Gene overrepresentation in biological_process category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.0 | 0.1 | GO:0060557 | positive regulation of vitamin metabolic process(GO:0046136) positive regulation of vitamin D biosynthetic process(GO:0060557) positive regulation of calcidiol 1-monooxygenase activity(GO:0060559) |
| 0.0 | 0.1 | GO:0046521 | sphingoid catabolic process(GO:0046521) |
| 0.0 | 0.1 | GO:0006097 | glyoxylate cycle(GO:0006097) |
| 0.0 | 0.1 | GO:0018894 | dibenzo-p-dioxin metabolic process(GO:0018894) cellular alkene metabolic process(GO:0043449) |
| 0.0 | 0.2 | GO:1903575 | cornified envelope assembly(GO:1903575) |
| 0.0 | 0.1 | GO:0017198 | N-terminal peptidyl-serine acetylation(GO:0017198) N-terminal peptidyl-glutamic acid acetylation(GO:0018002) peptidyl-serine acetylation(GO:0030920) |
| 0.0 | 0.2 | GO:0015811 | L-cystine transport(GO:0015811) |
| 0.0 | 0.1 | GO:0006428 | isoleucyl-tRNA aminoacylation(GO:0006428) |
| 0.0 | 0.1 | GO:0046967 | cytosol to ER transport(GO:0046967) |
| 0.0 | 0.1 | GO:0071449 | cellular response to lipid hydroperoxide(GO:0071449) |
| 0.0 | 0.1 | GO:0061143 | alveolar primary septum development(GO:0061143) |
| 0.0 | 0.1 | GO:0045897 | positive regulation of transcription during mitosis(GO:0045897) |
| 0.0 | 0.2 | GO:0060125 | negative regulation of growth hormone secretion(GO:0060125) |
| 0.0 | 0.1 | GO:0007499 | ectoderm and mesoderm interaction(GO:0007499) |
| 0.0 | 0.1 | GO:0010585 | glutamine secretion(GO:0010585) L-glutamine import(GO:0036229) L-glutamine import into cell(GO:1903803) |
| 0.0 | 0.1 | GO:2000491 | positive regulation of hepatic stellate cell activation(GO:2000491) |
| 0.0 | 0.1 | GO:0018076 | N-terminal peptidyl-lysine acetylation(GO:0018076) |
| 0.0 | 0.1 | GO:0046340 | diacylglycerol catabolic process(GO:0046340) |
| 0.0 | 0.1 | GO:0060157 | urinary bladder development(GO:0060157) |
| 0.0 | 0.1 | GO:1902775 | mitochondrial large ribosomal subunit assembly(GO:1902775) |
| 0.0 | 0.1 | GO:0001971 | negative regulation of activation of membrane attack complex(GO:0001971) |
| 0.0 | 0.2 | GO:0030916 | otic vesicle formation(GO:0030916) pronephros development(GO:0048793) |
| 0.0 | 0.0 | GO:0002541 | activation of plasma proteins involved in acute inflammatory response(GO:0002541) |
| 0.0 | 0.1 | GO:1900625 | regulation of monocyte aggregation(GO:1900623) positive regulation of monocyte aggregation(GO:1900625) |
| 0.0 | 0.1 | GO:1904764 | negative regulation of fibril organization(GO:1902904) chaperone-mediated autophagy translocation complex disassembly(GO:1904764) |
| 0.0 | 0.1 | GO:2000042 | negative regulation of double-strand break repair via homologous recombination(GO:2000042) |
Gene overrepresentation in cellular_component category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.0 | 0.2 | GO:0005600 | collagen type XIII trimer(GO:0005600) transmembrane collagen trimer(GO:0030936) |
| 0.0 | 0.1 | GO:0031515 | tRNA (m1A) methyltransferase complex(GO:0031515) |
| 0.0 | 0.1 | GO:0042825 | TAP complex(GO:0042825) |
| 0.0 | 0.1 | GO:0031415 | NatA complex(GO:0031415) |
| 0.0 | 0.1 | GO:0032010 | phagolysosome(GO:0032010) |
| 0.0 | 0.1 | GO:0031465 | Cul4B-RING E3 ubiquitin ligase complex(GO:0031465) |
| 0.0 | 0.1 | GO:0000125 | PCAF complex(GO:0000125) |
| 0.0 | 0.1 | GO:0005968 | Rab-protein geranylgeranyltransferase complex(GO:0005968) |
| 0.0 | 0.1 | GO:0035692 | macrophage migration inhibitory factor receptor complex(GO:0035692) |
| 0.0 | 0.2 | GO:0036038 | MKS complex(GO:0036038) |
Gene overrepresentation in molecular_function category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.0 | 0.1 | GO:0004450 | isocitrate dehydrogenase (NADP+) activity(GO:0004450) (R)-2-hydroxyglutarate dehydrogenase activity(GO:0051990) |
| 0.0 | 0.1 | GO:1990190 | peptide-glutamate-N-acetyltransferase activity(GO:1990190) |
| 0.0 | 0.2 | GO:0015184 | L-cystine transmembrane transporter activity(GO:0015184) |
| 0.0 | 0.1 | GO:0004822 | isoleucine-tRNA ligase activity(GO:0004822) |
| 0.0 | 0.1 | GO:0033878 | hormone-sensitive lipase activity(GO:0033878) |
| 0.0 | 0.1 | GO:0050135 | NAD(P)+ nucleosidase activity(GO:0050135) |
| 0.0 | 0.1 | GO:0070576 | vitamin D 24-hydroxylase activity(GO:0070576) |
| 0.0 | 0.1 | GO:0008481 | sphinganine kinase activity(GO:0008481) D-erythro-sphingosine kinase activity(GO:0017050) |
| 0.0 | 0.1 | GO:0008508 | bile acid:sodium symporter activity(GO:0008508) |
| 0.0 | 0.1 | GO:0015186 | L-glutamine transmembrane transporter activity(GO:0015186) |
| 0.0 | 0.1 | GO:0000405 | bubble DNA binding(GO:0000405) |
| 0.0 | 0.1 | GO:0046979 | TAP2 binding(GO:0046979) |
| 0.0 | 0.2 | GO:0051575 | 5'-deoxyribose-5-phosphate lyase activity(GO:0051575) |
| 0.0 | 0.1 | GO:0004741 | [pyruvate dehydrogenase (lipoamide)] phosphatase activity(GO:0004741) |
| 0.0 | 0.1 | GO:0001515 | opioid peptide activity(GO:0001515) |
| 0.0 | 0.2 | GO:0051011 | microtubule minus-end binding(GO:0051011) |
| 0.0 | 0.0 | GO:0004947 | bradykinin receptor activity(GO:0004947) |
| 0.0 | 0.0 | GO:0008336 | gamma-butyrobetaine dioxygenase activity(GO:0008336) |