Project
Epithelial-Mesenchymal Transition, human (Scheel, 2011)
Navigation
Downloads
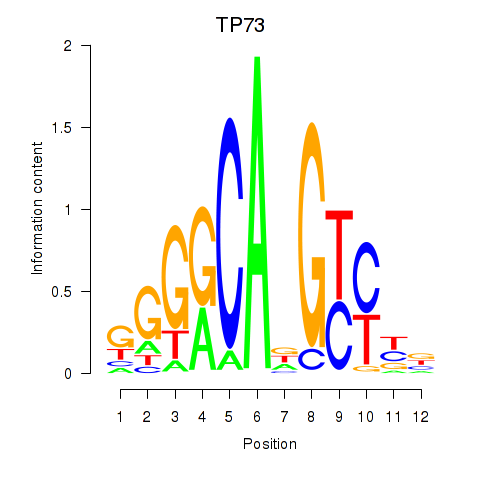
Results for TP73
Z-value: 0.56
Transcription factors associated with TP73
| Gene Symbol | Gene ID | Gene Info |
|---|---|---|
|
TP73
|
ENSG00000078900.10 | TP73 |
Activity-expression correlation:
| Gene | Promoter | Pearson corr. coef. | P-value | Plot |
|---|---|---|---|---|
| TP73 | hg19_v2_chr1_+_3614591_3614622 | 0.34 | 4.1e-01 | Click! |
Activity profile of TP73 motif
Sorted Z-values of TP73 motif
Network of associatons between targets according to the STRING database.
First level regulatory network of TP73
| Promoter | Score | Transcript | Gene | Gene Info |
|---|---|---|---|---|
| chr10_-_105212141 | 0.33 |
ENST00000369788.3 |
CALHM2 |
calcium homeostasis modulator 2 |
| chr10_-_105212059 | 0.32 |
ENST00000260743.5 |
CALHM2 |
calcium homeostasis modulator 2 |
| chr3_+_136676851 | 0.32 |
ENST00000309741.5 |
IL20RB |
interleukin 20 receptor beta |
| chr3_+_136676707 | 0.32 |
ENST00000329582.4 |
IL20RB |
interleukin 20 receptor beta |
| chr18_-_28681950 | 0.28 |
ENST00000251081.6 |
DSC2 |
desmocollin 2 |
| chr19_+_45281118 | 0.27 |
ENST00000270279.3 ENST00000341505.4 |
CBLC |
Cbl proto-oncogene C, E3 ubiquitin protein ligase |
| chr14_+_75746340 | 0.26 |
ENST00000555686.1 ENST00000555672.1 |
FOS |
FBJ murine osteosarcoma viral oncogene homolog |
| chr10_+_47746929 | 0.24 |
ENST00000340243.6 ENST00000374277.5 ENST00000449464.2 ENST00000538825.1 ENST00000335083.5 |
ANXA8L2 AL603965.1 |
annexin A8-like 2 Protein LOC100996760 |
| chr15_+_41136586 | 0.24 |
ENST00000431806.1 |
SPINT1 |
serine peptidase inhibitor, Kunitz type 1 |
| chr14_+_75745477 | 0.22 |
ENST00000303562.4 ENST00000554617.1 ENST00000554212.1 ENST00000535987.1 ENST00000555242.1 |
FOS |
FBJ murine osteosarcoma viral oncogene homolog |
| chr3_+_100120441 | 0.21 |
ENST00000489752.1 |
LNP1 |
leukemia NUP98 fusion partner 1 |
| chr11_-_17035943 | 0.20 |
ENST00000355661.3 ENST00000532079.1 ENST00000448080.2 ENST00000531066.1 |
PLEKHA7 |
pleckstrin homology domain containing, family A member 7 |
| chr18_-_53253323 | 0.18 |
ENST00000540999.1 ENST00000563888.2 |
TCF4 |
transcription factor 4 |
| chr18_-_53253112 | 0.17 |
ENST00000568673.1 ENST00000562847.1 ENST00000568147.1 |
TCF4 |
transcription factor 4 |
| chr15_+_41136216 | 0.17 |
ENST00000562057.1 ENST00000344051.4 |
SPINT1 |
serine peptidase inhibitor, Kunitz type 1 |
| chr11_+_60197069 | 0.16 |
ENST00000528905.1 ENST00000528093.1 |
MS4A5 |
membrane-spanning 4-domains, subfamily A, member 5 |
| chr2_+_95940186 | 0.14 |
ENST00000403131.2 ENST00000317668.4 ENST00000317620.9 |
PROM2 |
prominin 2 |
| chr10_-_47173994 | 0.14 |
ENST00000414655.2 ENST00000545298.1 ENST00000359178.4 ENST00000358140.4 ENST00000503031.1 |
ANXA8L1 LINC00842 |
annexin A8-like 1 long intergenic non-protein coding RNA 842 |
| chr22_-_20255212 | 0.13 |
ENST00000416372.1 |
RTN4R |
reticulon 4 receptor |
| chr6_-_46703069 | 0.13 |
ENST00000538237.1 ENST00000274793.7 |
PLA2G7 |
phospholipase A2, group VII (platelet-activating factor acetylhydrolase, plasma) |
| chr2_+_95940220 | 0.13 |
ENST00000542147.1 |
PROM2 |
prominin 2 |
| chr17_+_73750699 | 0.12 |
ENST00000584939.1 |
ITGB4 |
integrin, beta 4 |
| chr11_+_60197040 | 0.11 |
ENST00000300190.2 |
MS4A5 |
membrane-spanning 4-domains, subfamily A, member 5 |
| chr1_-_182641367 | 0.10 |
ENST00000508450.1 |
RGS8 |
regulator of G-protein signaling 8 |
| chr7_-_27213893 | 0.10 |
ENST00000283921.4 |
HOXA10 |
homeobox A10 |
| chr17_-_42200996 | 0.10 |
ENST00000587135.1 ENST00000225983.6 ENST00000393622.2 ENST00000588703.1 |
HDAC5 |
histone deacetylase 5 |
| chr7_-_100171270 | 0.09 |
ENST00000538735.1 |
SAP25 |
Sin3A-associated protein, 25kDa |
| chr8_+_120885949 | 0.09 |
ENST00000523492.1 ENST00000286234.5 |
DEPTOR |
DEP domain containing MTOR-interacting protein |
| chr14_+_103566665 | 0.08 |
ENST00000559116.1 |
EXOC3L4 |
exocyst complex component 3-like 4 |
| chr19_-_35981358 | 0.08 |
ENST00000484218.2 ENST00000338897.3 |
KRTDAP |
keratinocyte differentiation-associated protein |
| chr14_+_105190514 | 0.08 |
ENST00000330877.2 |
ADSSL1 |
adenylosuccinate synthase like 1 |
| chr6_-_46703430 | 0.08 |
ENST00000537365.1 |
PLA2G7 |
phospholipase A2, group VII (platelet-activating factor acetylhydrolase, plasma) |
| chr19_-_19626838 | 0.07 |
ENST00000360913.3 |
TSSK6 |
testis-specific serine kinase 6 |
| chr19_+_44037546 | 0.07 |
ENST00000601282.1 |
ZNF575 |
zinc finger protein 575 |
| chr17_+_78388959 | 0.07 |
ENST00000518137.1 ENST00000520367.1 ENST00000523999.1 ENST00000323854.5 ENST00000522751.1 |
ENDOV |
endonuclease V |
| chr3_-_185538849 | 0.07 |
ENST00000421047.2 |
IGF2BP2 |
insulin-like growth factor 2 mRNA binding protein 2 |
| chr16_+_222846 | 0.06 |
ENST00000251595.6 ENST00000397806.1 |
HBA2 |
hemoglobin, alpha 2 |
| chr11_+_75479850 | 0.06 |
ENST00000376262.3 ENST00000604733.1 |
DGAT2 |
diacylglycerol O-acyltransferase 2 |
| chr12_-_57505121 | 0.06 |
ENST00000538913.2 ENST00000537215.2 ENST00000454075.3 ENST00000554825.1 ENST00000553275.1 ENST00000300134.3 |
STAT6 |
signal transducer and activator of transcription 6, interleukin-4 induced |
| chr1_-_203055129 | 0.06 |
ENST00000241651.4 |
MYOG |
myogenin (myogenic factor 4) |
| chr16_+_84178874 | 0.06 |
ENST00000378553.5 |
DNAAF1 |
dynein, axonemal, assembly factor 1 |
| chr2_-_73511559 | 0.06 |
ENST00000521871.1 |
FBXO41 |
F-box protein 41 |
| chr3_-_50340996 | 0.06 |
ENST00000266031.4 ENST00000395143.2 ENST00000457214.2 ENST00000447605.2 ENST00000418723.1 ENST00000395144.2 |
HYAL1 |
hyaluronoglucosaminidase 1 |
| chr20_-_52790055 | 0.05 |
ENST00000395955.3 |
CYP24A1 |
cytochrome P450, family 24, subfamily A, polypeptide 1 |
| chr17_-_42452063 | 0.05 |
ENST00000588098.1 |
ITGA2B |
integrin, alpha 2b (platelet glycoprotein IIb of IIb/IIIa complex, antigen CD41) |
| chr14_-_25045446 | 0.05 |
ENST00000216336.2 |
CTSG |
cathepsin G |
| chr16_-_20367584 | 0.05 |
ENST00000570689.1 |
UMOD |
uromodulin |
| chr13_-_48877795 | 0.05 |
ENST00000436963.1 ENST00000433480.2 |
LINC00441 |
long intergenic non-protein coding RNA 441 |
| chr16_+_75032901 | 0.05 |
ENST00000335325.4 ENST00000320619.6 |
ZNRF1 |
zinc and ring finger 1, E3 ubiquitin protein ligase |
| chr11_-_75062829 | 0.05 |
ENST00000393505.4 |
ARRB1 |
arrestin, beta 1 |
| chr1_-_111148241 | 0.05 |
ENST00000440270.1 |
KCNA2 |
potassium voltage-gated channel, shaker-related subfamily, member 2 |
| chr11_+_236540 | 0.05 |
ENST00000532097.1 |
PSMD13 |
proteasome (prosome, macropain) 26S subunit, non-ATPase, 13 |
| chr11_+_236959 | 0.05 |
ENST00000431206.2 ENST00000528906.1 |
PSMD13 |
proteasome (prosome, macropain) 26S subunit, non-ATPase, 13 |
| chr2_-_26864228 | 0.04 |
ENST00000288861.4 |
CIB4 |
calcium and integrin binding family member 4 |
| chr11_+_237016 | 0.04 |
ENST00000352303.5 |
PSMD13 |
proteasome (prosome, macropain) 26S subunit, non-ATPase, 13 |
| chr14_-_105531759 | 0.04 |
ENST00000329797.3 ENST00000539291.2 ENST00000392585.2 |
GPR132 |
G protein-coupled receptor 132 |
| chr17_+_37356528 | 0.04 |
ENST00000225430.4 |
RPL19 |
ribosomal protein L19 |
| chr1_+_110082487 | 0.04 |
ENST00000527748.1 |
GPR61 |
G protein-coupled receptor 61 |
| chr22_+_29168652 | 0.04 |
ENST00000249064.4 ENST00000444523.1 ENST00000448492.2 ENST00000421503.2 |
CCDC117 |
coiled-coil domain containing 117 |
| chr17_+_21191341 | 0.04 |
ENST00000526076.2 ENST00000361818.5 ENST00000316920.6 |
MAP2K3 |
mitogen-activated protein kinase kinase 3 |
| chr17_+_37356586 | 0.04 |
ENST00000579260.1 ENST00000582193.1 |
RPL19 |
ribosomal protein L19 |
| chr1_-_78444738 | 0.04 |
ENST00000436586.2 ENST00000370768.2 |
FUBP1 |
far upstream element (FUSE) binding protein 1 |
| chr7_-_128045984 | 0.04 |
ENST00000470772.1 ENST00000480861.1 ENST00000496200.1 |
IMPDH1 |
IMP (inosine 5'-monophosphate) dehydrogenase 1 |
| chr19_-_39226045 | 0.04 |
ENST00000597987.1 ENST00000595177.1 |
CAPN12 |
calpain 12 |
| chr17_+_37356555 | 0.03 |
ENST00000579374.1 |
RPL19 |
ribosomal protein L19 |
| chr17_-_39023462 | 0.03 |
ENST00000251643.4 |
KRT12 |
keratin 12 |
| chr20_+_48807351 | 0.03 |
ENST00000303004.3 |
CEBPB |
CCAAT/enhancer binding protein (C/EBP), beta |
| chr21_+_43442100 | 0.03 |
ENST00000455701.1 ENST00000596595.1 |
ZNF295-AS1 |
ZNF295 antisense RNA 1 |
| chr1_-_78444776 | 0.03 |
ENST00000370767.1 ENST00000421641.1 |
FUBP1 |
far upstream element (FUSE) binding protein 1 |
| chr19_+_19627026 | 0.03 |
ENST00000608404.1 ENST00000555938.1 ENST00000503283.1 ENST00000512771.3 ENST00000428459.2 |
YJEFN3 CTC-260F20.3 NDUFA13 |
YjeF N-terminal domain containing 3 Uncharacterized protein NADH dehydrogenase (ubiquinone) 1 alpha subcomplex, 13 |
| chr17_+_7792101 | 0.03 |
ENST00000358181.4 ENST00000330494.7 |
CHD3 |
chromodomain helicase DNA binding protein 3 |
| chr16_+_8891670 | 0.03 |
ENST00000268261.4 ENST00000539622.1 ENST00000569958.1 ENST00000537352.1 |
PMM2 |
phosphomannomutase 2 |
| chr2_+_128293323 | 0.03 |
ENST00000389524.4 ENST00000428314.1 |
MYO7B |
myosin VIIB |
| chr16_-_8891481 | 0.03 |
ENST00000333050.6 |
TMEM186 |
transmembrane protein 186 |
| chr20_-_52790512 | 0.03 |
ENST00000216862.3 |
CYP24A1 |
cytochrome P450, family 24, subfamily A, polypeptide 1 |
| chr15_+_75498355 | 0.03 |
ENST00000567617.1 |
C15orf39 |
chromosome 15 open reading frame 39 |
| chr14_+_105935396 | 0.03 |
ENST00000494981.1 |
MTA1 |
metastasis associated 1 |
| chr12_-_117319236 | 0.03 |
ENST00000257572.5 |
HRK |
harakiri, BCL2 interacting protein (contains only BH3 domain) |
| chrX_-_131228291 | 0.03 |
ENST00000370879.1 |
FRMD7 |
FERM domain containing 7 |
| chr5_+_38258511 | 0.02 |
ENST00000354891.3 ENST00000322350.5 |
EGFLAM |
EGF-like, fibronectin type III and laminin G domains |
| chr3_-_100120223 | 0.02 |
ENST00000284320.5 |
TOMM70A |
translocase of outer mitochondrial membrane 70 homolog A (S. cerevisiae) |
| chr5_+_78532003 | 0.02 |
ENST00000396137.4 |
JMY |
junction mediating and regulatory protein, p53 cofactor |
| chr20_-_62203808 | 0.02 |
ENST00000467148.1 |
HELZ2 |
helicase with zinc finger 2, transcriptional coactivator |
| chr10_-_79686284 | 0.02 |
ENST00000372391.2 ENST00000372388.2 |
DLG5 |
discs, large homolog 5 (Drosophila) |
| chr11_+_66234216 | 0.02 |
ENST00000349459.6 ENST00000320740.7 ENST00000524466.1 ENST00000526296.1 |
PELI3 |
pellino E3 ubiquitin protein ligase family member 3 |
| chr3_+_42201653 | 0.02 |
ENST00000341421.3 ENST00000396175.1 |
TRAK1 |
trafficking protein, kinesin binding 1 |
| chr6_+_31515337 | 0.02 |
ENST00000376148.4 ENST00000376145.4 |
NFKBIL1 |
nuclear factor of kappa light polypeptide gene enhancer in B-cells inhibitor-like 1 |
| chr17_-_2614927 | 0.02 |
ENST00000435359.1 |
CLUH |
clustered mitochondria (cluA/CLU1) homolog |
| chr14_+_103566481 | 0.02 |
ENST00000380069.3 |
EXOC3L4 |
exocyst complex component 3-like 4 |
| chr5_-_135528822 | 0.02 |
ENST00000607574.1 |
AC009014.3 |
AC009014.3 |
| chr19_+_41856816 | 0.02 |
ENST00000539627.1 |
TMEM91 |
transmembrane protein 91 |
| chr1_-_220220000 | 0.02 |
ENST00000366923.3 |
EPRS |
glutamyl-prolyl-tRNA synthetase |
| chr10_+_70661014 | 0.01 |
ENST00000373585.3 |
DDX50 |
DEAD (Asp-Glu-Ala-Asp) box polypeptide 50 |
| chr1_-_26197744 | 0.01 |
ENST00000374296.3 |
PAQR7 |
progestin and adipoQ receptor family member VII |
| chr18_-_4455260 | 0.01 |
ENST00000581527.1 |
DLGAP1 |
discs, large (Drosophila) homolog-associated protein 1 |
| chr2_+_114163945 | 0.01 |
ENST00000453673.3 |
IGKV1OR2-108 |
immunoglobulin kappa variable 1/OR2-108 (non-functional) |
| chr20_+_35504522 | 0.01 |
ENST00000602922.1 ENST00000217320.3 |
TLDC2 |
TBC/LysM-associated domain containing 2 |
| chr6_-_112194484 | 0.01 |
ENST00000518295.1 ENST00000484067.2 ENST00000229470.5 ENST00000356013.2 ENST00000368678.4 ENST00000523238.1 ENST00000354650.3 |
FYN |
FYN oncogene related to SRC, FGR, YES |
| chr22_-_37976082 | 0.01 |
ENST00000215886.4 |
LGALS2 |
lectin, galactoside-binding, soluble, 2 |
| chr2_-_27357479 | 0.01 |
ENST00000406567.3 ENST00000260643.2 |
PREB |
prolactin regulatory element binding |
| chr9_+_87285257 | 0.01 |
ENST00000323115.4 |
NTRK2 |
neurotrophic tyrosine kinase, receptor, type 2 |
| chr5_-_156593273 | 0.01 |
ENST00000302938.4 |
FAM71B |
family with sequence similarity 71, member B |
| chr1_+_26485511 | 0.01 |
ENST00000374268.3 |
FAM110D |
family with sequence similarity 110, member D |
| chr8_-_70745575 | 0.01 |
ENST00000524945.1 |
SLCO5A1 |
solute carrier organic anion transporter family, member 5A1 |
| chr11_-_65686496 | 0.01 |
ENST00000449692.3 |
C11orf68 |
chromosome 11 open reading frame 68 |
| chr19_+_17326521 | 0.01 |
ENST00000593597.1 |
USE1 |
unconventional SNARE in the ER 1 homolog (S. cerevisiae) |
| chr11_+_65154070 | 0.01 |
ENST00000317568.5 ENST00000531296.1 ENST00000533782.1 ENST00000355991.5 ENST00000416776.2 ENST00000526201.1 |
FRMD8 |
FERM domain containing 8 |
| chr17_-_79818354 | 0.01 |
ENST00000576541.1 ENST00000576380.1 ENST00000571617.1 ENST00000576052.1 ENST00000576390.1 ENST00000573778.2 ENST00000439918.2 ENST00000574914.1 ENST00000331483.4 |
P4HB |
prolyl 4-hydroxylase, beta polypeptide |
| chr19_+_49622646 | 0.01 |
ENST00000334186.4 |
PPFIA3 |
protein tyrosine phosphatase, receptor type, f polypeptide (PTPRF), interacting protein (liprin), alpha 3 |
| chr9_+_6757634 | 0.01 |
ENST00000543771.1 ENST00000401787.3 ENST00000381306.3 ENST00000381309.3 |
KDM4C |
lysine (K)-specific demethylase 4C |
| chr14_+_22554680 | 0.01 |
ENST00000390451.2 |
TRAV23DV6 |
T cell receptor alpha variable 23/delta variable 6 |
| chr12_-_82752565 | 0.00 |
ENST00000256151.7 |
CCDC59 |
coiled-coil domain containing 59 |
| chr17_-_40835076 | 0.00 |
ENST00000591765.1 |
CCR10 |
chemokine (C-C motif) receptor 10 |
| chr7_+_116502527 | 0.00 |
ENST00000361183.3 |
CAPZA2 |
capping protein (actin filament) muscle Z-line, alpha 2 |
| chr16_-_2581409 | 0.00 |
ENST00000567119.1 ENST00000565480.1 ENST00000382350.1 |
CEMP1 |
cementum protein 1 |
| chr12_+_82752275 | 0.00 |
ENST00000248306.3 |
METTL25 |
methyltransferase like 25 |
| chr17_-_47492236 | 0.00 |
ENST00000434917.2 ENST00000300408.3 ENST00000511832.1 ENST00000419140.2 |
PHB |
prohibitin |
| chr17_-_47492164 | 0.00 |
ENST00000512041.2 ENST00000446735.1 ENST00000504124.1 |
PHB |
prohibitin |
| chrX_-_138724677 | 0.00 |
ENST00000370573.4 ENST00000338585.6 ENST00000370576.4 |
MCF2 |
MCF.2 cell line derived transforming sequence |
| chr14_+_96858433 | 0.00 |
ENST00000267584.4 |
AK7 |
adenylate kinase 7 |
Gene Ontology Analysis
Gene overrepresentation in biological_process category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.2 | 0.6 | GO:0001808 | negative regulation of type IV hypersensitivity(GO:0001808) |
| 0.1 | 0.3 | GO:2001287 | negative regulation of caveolin-mediated endocytosis(GO:2001287) |
| 0.1 | 0.5 | GO:0001661 | conditioned taste aversion(GO:0001661) |
| 0.0 | 0.2 | GO:0034441 | plasma lipoprotein particle oxidation(GO:0034441) |
| 0.0 | 0.2 | GO:0045218 | zonula adherens maintenance(GO:0045218) |
| 0.0 | 0.3 | GO:0086073 | bundle of His cell-Purkinje myocyte adhesion involved in cell communication(GO:0086073) |
| 0.0 | 0.1 | GO:0042369 | vitamin D catabolic process(GO:0042369) |
| 0.0 | 0.4 | GO:0060670 | branching involved in labyrinthine layer morphogenesis(GO:0060670) |
| 0.0 | 0.1 | GO:0071400 | cellular response to oleic acid(GO:0071400) |
| 0.0 | 0.1 | GO:0002296 | T-helper 1 cell lineage commitment(GO:0002296) |
| 0.0 | 0.1 | GO:0044208 | 'de novo' AMP biosynthetic process(GO:0044208) |
| 0.0 | 0.1 | GO:1900106 | hyaluranon cable assembly(GO:0036118) regulation of hyaluranon cable assembly(GO:1900104) positive regulation of hyaluranon cable assembly(GO:1900106) |
| 0.0 | 0.3 | GO:0007175 | negative regulation of epidermal growth factor-activated receptor activity(GO:0007175) |
| 0.0 | 0.1 | GO:1903862 | regulation of muscle atrophy(GO:0014735) response to electrical stimulus involved in regulation of muscle adaptation(GO:0014878) positive regulation of oxidative phosphorylation(GO:1903862) |
Gene overrepresentation in cellular_component category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.1 | 0.5 | GO:0035976 | AP1 complex(GO:0035976) |
| 0.1 | 0.3 | GO:0044393 | microspike(GO:0044393) |
| 0.0 | 0.2 | GO:0005915 | zonula adherens(GO:0005915) |
| 0.0 | 0.1 | GO:0008541 | proteasome regulatory particle, lid subcomplex(GO:0008541) |
Gene overrepresentation in molecular_function category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.1 | 0.6 | GO:0042015 | interleukin-20 binding(GO:0042015) |
| 0.0 | 0.1 | GO:0038131 | neuregulin receptor activity(GO:0038131) |
| 0.0 | 0.3 | GO:0086083 | cell adhesive protein binding involved in bundle of His cell-Purkinje myocyte communication(GO:0086083) |
| 0.0 | 0.1 | GO:0070180 | large ribosomal subunit rRNA binding(GO:0070180) |
| 0.0 | 0.3 | GO:0001011 | transcription factor activity, sequence-specific DNA binding, RNA polymerase recruiting(GO:0001011) transcription factor activity, TFIIB-class binding(GO:0001087) |
| 0.0 | 0.1 | GO:0004019 | adenylosuccinate synthase activity(GO:0004019) |
| 0.0 | 0.2 | GO:0070097 | delta-catenin binding(GO:0070097) |
| 0.0 | 0.2 | GO:0003847 | 1-alkyl-2-acetylglycerophosphocholine esterase activity(GO:0003847) |
| 0.0 | 0.1 | GO:0050252 | retinol O-fatty-acyltransferase activity(GO:0050252) |
| 0.0 | 0.1 | GO:0001025 | RNA polymerase III transcription factor binding(GO:0001025) |
| 0.0 | 0.1 | GO:0070576 | vitamin D 24-hydroxylase activity(GO:0070576) |
| 0.0 | 0.0 | GO:0031896 | V2 vasopressin receptor binding(GO:0031896) |
| 0.0 | 0.1 | GO:0050501 | hyaluronan synthase activity(GO:0050501) |
| 0.0 | 0.1 | GO:0070051 | fibrinogen binding(GO:0070051) |
Gene overrepresentation in C2:CP category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.0 | 0.5 | PID TCR RAS PATHWAY | Ras signaling in the CD4+ TCR pathway |
Gene overrepresentation in C2:CP:REACTOME category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.0 | 0.4 | REACTOME ACTIVATION OF THE AP1 FAMILY OF TRANSCRIPTION FACTORS | Genes involved in Activation of the AP-1 family of transcription factors |