Project
Epithelial-Mesenchymal Transition, human (Scheel, 2011)
Navigation
Downloads
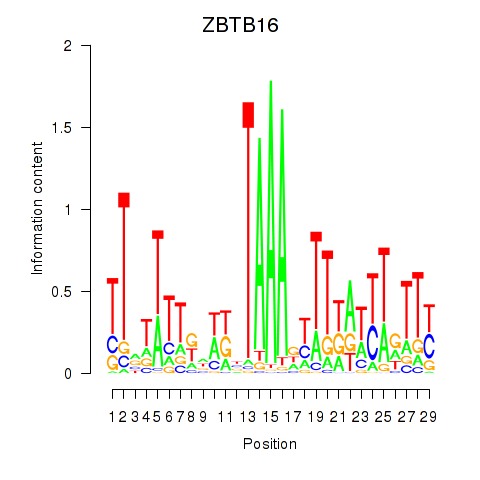
Results for ZBTB16
Z-value: 1.09
Transcription factors associated with ZBTB16
| Gene Symbol | Gene ID | Gene Info |
|---|---|---|
|
ZBTB16
|
ENSG00000109906.9 | ZBTB16 |
Activity-expression correlation:
| Gene | Promoter | Pearson corr. coef. | P-value | Plot |
|---|---|---|---|---|
| ZBTB16 | hg19_v2_chr11_+_113930291_113930339 | 0.89 | 3.1e-03 | Click! |
Activity profile of ZBTB16 motif
Sorted Z-values of ZBTB16 motif
Network of associatons between targets according to the STRING database.
First level regulatory network of ZBTB16
| Promoter | Score | Transcript | Gene | Gene Info |
|---|---|---|---|---|
| chr12_-_91539918 | 2.19 |
ENST00000548218.1 |
DCN |
decorin |
| chr4_+_14113592 | 2.03 |
ENST00000502759.1 ENST00000511200.1 ENST00000512754.1 ENST00000506739.1 |
LINC01085 |
long intergenic non-protein coding RNA 1085 |
| chr1_+_170633047 | 1.60 |
ENST00000239461.6 ENST00000497230.2 |
PRRX1 |
paired related homeobox 1 |
| chr2_+_233527443 | 1.45 |
ENST00000410095.1 |
EFHD1 |
EF-hand domain family, member D1 |
| chr3_+_148415571 | 1.14 |
ENST00000497524.1 ENST00000349243.3 ENST00000542281.1 ENST00000418473.2 ENST00000404754.2 |
AGTR1 |
angiotensin II receptor, type 1 |
| chr20_-_34042558 | 1.13 |
ENST00000374372.1 |
GDF5 |
growth differentiation factor 5 |
| chr2_-_188419078 | 1.02 |
ENST00000437725.1 ENST00000409676.1 ENST00000339091.4 ENST00000420747.1 |
TFPI |
tissue factor pathway inhibitor (lipoprotein-associated coagulation inhibitor) |
| chr14_+_22458631 | 0.93 |
ENST00000390444.1 |
TRAV16 |
T cell receptor alpha variable 16 |
| chr6_+_32812568 | 0.92 |
ENST00000414474.1 |
PSMB9 |
proteasome (prosome, macropain) subunit, beta type, 9 |
| chr4_-_159094194 | 0.87 |
ENST00000592057.1 ENST00000585682.1 ENST00000393807.5 |
FAM198B |
family with sequence similarity 198, member B |
| chr18_-_52626622 | 0.83 |
ENST00000591504.1 |
CCDC68 |
coiled-coil domain containing 68 |
| chr9_-_95244781 | 0.82 |
ENST00000375544.3 ENST00000375543.1 ENST00000395538.3 ENST00000450139.2 |
ASPN |
asporin |
| chr6_+_136172820 | 0.81 |
ENST00000308191.6 |
PDE7B |
phosphodiesterase 7B |
| chr17_-_295730 | 0.80 |
ENST00000329099.4 |
FAM101B |
family with sequence similarity 101, member B |
| chr21_-_31852663 | 0.76 |
ENST00000390689.2 |
KRTAP19-1 |
keratin associated protein 19-1 |
| chr5_+_67586465 | 0.76 |
ENST00000336483.5 |
PIK3R1 |
phosphoinositide-3-kinase, regulatory subunit 1 (alpha) |
| chr10_+_104614008 | 0.73 |
ENST00000369883.3 |
C10orf32 |
chromosome 10 open reading frame 32 |
| chr11_-_70672645 | 0.69 |
ENST00000423696.2 |
SHANK2 |
SH3 and multiple ankyrin repeat domains 2 |
| chr4_+_74347400 | 0.67 |
ENST00000226355.3 |
AFM |
afamin |
| chr20_-_18477862 | 0.65 |
ENST00000337227.4 |
RBBP9 |
retinoblastoma binding protein 9 |
| chr9_+_79074068 | 0.65 |
ENST00000444201.2 ENST00000376730.4 |
GCNT1 |
glucosaminyl (N-acetyl) transferase 1, core 2 |
| chr12_+_15699286 | 0.62 |
ENST00000442921.2 ENST00000542557.1 ENST00000445537.2 ENST00000544244.1 |
PTPRO |
protein tyrosine phosphatase, receptor type, O |
| chr7_+_7606497 | 0.62 |
ENST00000340080.4 ENST00000405785.1 ENST00000433635.1 |
MIOS |
missing oocyte, meiosis regulator, homolog (Drosophila) |
| chr10_+_111765562 | 0.62 |
ENST00000360162.3 |
ADD3 |
adducin 3 (gamma) |
| chr10_+_8096769 | 0.57 |
ENST00000346208.3 |
GATA3 |
GATA binding protein 3 |
| chr4_+_71108300 | 0.56 |
ENST00000304954.3 |
CSN3 |
casein kappa |
| chr2_+_42104692 | 0.56 |
ENST00000398796.2 ENST00000442214.1 |
AC104654.1 |
AC104654.1 |
| chr7_-_92777606 | 0.54 |
ENST00000437805.1 ENST00000446959.1 ENST00000439952.1 ENST00000414791.1 ENST00000446033.1 ENST00000411955.1 ENST00000318238.4 |
SAMD9L |
sterile alpha motif domain containing 9-like |
| chr14_+_23067146 | 0.53 |
ENST00000428304.2 |
ABHD4 |
abhydrolase domain containing 4 |
| chr22_+_38203898 | 0.52 |
ENST00000323205.6 ENST00000248924.6 ENST00000445195.1 |
GCAT |
glycine C-acetyltransferase |
| chr10_+_81891416 | 0.51 |
ENST00000372270.2 |
PLAC9 |
placenta-specific 9 |
| chr10_+_104613980 | 0.50 |
ENST00000339834.5 |
C10orf32 |
chromosome 10 open reading frame 32 |
| chr4_-_83769996 | 0.48 |
ENST00000511338.1 |
SEC31A |
SEC31 homolog A (S. cerevisiae) |
| chr12_-_71533055 | 0.47 |
ENST00000552128.1 |
TSPAN8 |
tetraspanin 8 |
| chr3_-_93692781 | 0.46 |
ENST00000394236.3 |
PROS1 |
protein S (alpha) |
| chr11_+_65657875 | 0.43 |
ENST00000312579.2 |
CCDC85B |
coiled-coil domain containing 85B |
| chr3_-_164796269 | 0.42 |
ENST00000264382.3 |
SI |
sucrase-isomaltase (alpha-glucosidase) |
| chr8_-_8318847 | 0.41 |
ENST00000521218.1 |
CTA-398F10.2 |
CTA-398F10.2 |
| chr5_-_61031495 | 0.40 |
ENST00000506550.1 ENST00000512882.2 |
CTD-2170G1.2 |
CTD-2170G1.2 |
| chr10_-_28623368 | 0.40 |
ENST00000441595.2 |
MPP7 |
membrane protein, palmitoylated 7 (MAGUK p55 subfamily member 7) |
| chr1_+_36335351 | 0.40 |
ENST00000373206.1 |
AGO1 |
argonaute RISC catalytic component 1 |
| chr15_+_75628419 | 0.39 |
ENST00000567377.1 ENST00000562789.1 ENST00000568301.1 |
COMMD4 |
COMM domain containing 4 |
| chr15_-_52944231 | 0.38 |
ENST00000546305.2 |
FAM214A |
family with sequence similarity 214, member A |
| chr1_+_104159999 | 0.38 |
ENST00000414303.2 ENST00000423678.1 |
AMY2A |
amylase, alpha 2A (pancreatic) |
| chr15_+_75628394 | 0.38 |
ENST00000564815.1 ENST00000338995.6 |
COMMD4 |
COMM domain containing 4 |
| chr15_+_75628232 | 0.38 |
ENST00000267935.8 ENST00000567195.1 |
COMMD4 |
COMM domain containing 4 |
| chr18_+_7946941 | 0.37 |
ENST00000444013.1 |
PTPRM |
protein tyrosine phosphatase, receptor type, M |
| chr11_-_66445219 | 0.37 |
ENST00000525754.1 ENST00000531969.1 ENST00000524637.1 ENST00000531036.2 ENST00000310046.4 |
RBM4B |
RNA binding motif protein 4B |
| chrX_-_47518527 | 0.36 |
ENST00000333119.3 |
UXT |
ubiquitously-expressed, prefoldin-like chaperone |
| chr21_+_40817749 | 0.33 |
ENST00000380637.3 ENST00000380634.1 ENST00000458295.1 ENST00000440288.2 ENST00000380631.1 |
SH3BGR |
SH3 domain binding glutamic acid-rich protein |
| chr16_+_20817761 | 0.33 |
ENST00000568046.1 ENST00000261377.6 |
AC004381.6 |
Putative RNA exonuclease NEF-sp |
| chr14_-_89878369 | 0.33 |
ENST00000553840.1 ENST00000556916.1 |
FOXN3 |
forkhead box N3 |
| chr10_+_8096631 | 0.33 |
ENST00000379328.3 |
GATA3 |
GATA binding protein 3 |
| chr8_+_11961898 | 0.33 |
ENST00000400085.3 |
ZNF705D |
zinc finger protein 705D |
| chr6_+_25963020 | 0.32 |
ENST00000357085.3 |
TRIM38 |
tripartite motif containing 38 |
| chr1_+_178310581 | 0.32 |
ENST00000462775.1 |
RASAL2 |
RAS protein activator like 2 |
| chr3_-_15382875 | 0.31 |
ENST00000408919.3 |
SH3BP5 |
SH3-domain binding protein 5 (BTK-associated) |
| chr17_-_39183452 | 0.31 |
ENST00000361883.5 |
KRTAP1-5 |
keratin associated protein 1-5 |
| chr15_-_55562582 | 0.31 |
ENST00000396307.2 |
RAB27A |
RAB27A, member RAS oncogene family |
| chr22_-_36236623 | 0.30 |
ENST00000405409.2 |
RBFOX2 |
RNA binding protein, fox-1 homolog (C. elegans) 2 |
| chr3_-_138312971 | 0.30 |
ENST00000485115.1 ENST00000484888.1 ENST00000468900.1 ENST00000542237.1 ENST00000481834.1 |
CEP70 |
centrosomal protein 70kDa |
| chr12_+_53836339 | 0.30 |
ENST00000549135.1 |
PRR13 |
proline rich 13 |
| chr7_-_107770794 | 0.30 |
ENST00000205386.4 ENST00000418464.1 ENST00000388781.3 ENST00000388780.3 ENST00000414450.2 |
LAMB4 |
laminin, beta 4 |
| chr21_-_35284635 | 0.30 |
ENST00000429238.1 |
AP000304.12 |
AP000304.12 |
| chr1_+_156095951 | 0.30 |
ENST00000448611.2 ENST00000368297.1 |
LMNA |
lamin A/C |
| chrX_+_12885183 | 0.29 |
ENST00000380659.3 |
TLR7 |
toll-like receptor 7 |
| chr18_+_11857439 | 0.29 |
ENST00000602628.1 |
GNAL |
guanine nucleotide binding protein (G protein), alpha activating activity polypeptide, olfactory type |
| chr10_+_124670121 | 0.29 |
ENST00000368894.1 |
FAM24A |
family with sequence similarity 24, member A |
| chr14_+_77228532 | 0.29 |
ENST00000167106.4 ENST00000554237.1 |
VASH1 |
vasohibin 1 |
| chr7_+_28725585 | 0.29 |
ENST00000396298.2 |
CREB5 |
cAMP responsive element binding protein 5 |
| chr14_-_105708942 | 0.28 |
ENST00000549655.1 |
BRF1 |
BRF1, RNA polymerase III transcription initiation factor 90 kDa subunit |
| chr3_+_40518599 | 0.28 |
ENST00000314686.5 ENST00000447116.2 ENST00000429348.2 ENST00000456778.1 |
ZNF619 |
zinc finger protein 619 |
| chr11_-_104972158 | 0.28 |
ENST00000598974.1 ENST00000593315.1 ENST00000594519.1 ENST00000415981.2 ENST00000525374.1 ENST00000375707.1 |
CASP1 CARD16 CARD17 |
caspase 1, apoptosis-related cysteine peptidase caspase recruitment domain family, member 16 caspase recruitment domain family, member 17 |
| chr1_+_210501589 | 0.27 |
ENST00000413764.2 ENST00000541565.1 |
HHAT |
hedgehog acyltransferase |
| chr6_-_34639733 | 0.27 |
ENST00000374021.1 |
C6orf106 |
chromosome 6 open reading frame 106 |
| chr22_-_36236265 | 0.27 |
ENST00000414461.2 ENST00000416721.2 ENST00000449924.2 ENST00000262829.7 ENST00000397305.3 |
RBFOX2 |
RNA binding protein, fox-1 homolog (C. elegans) 2 |
| chr19_+_20959098 | 0.27 |
ENST00000360204.5 ENST00000594534.1 |
ZNF66 |
zinc finger protein 66 |
| chrX_+_71401526 | 0.27 |
ENST00000218432.5 ENST00000423432.2 ENST00000373669.2 |
PIN4 |
protein (peptidylprolyl cis/trans isomerase) NIMA-interacting, 4 (parvulin) |
| chr3_-_138313161 | 0.27 |
ENST00000489254.1 ENST00000474781.1 ENST00000462419.1 ENST00000264982.3 |
CEP70 |
centrosomal protein 70kDa |
| chr2_+_242289502 | 0.26 |
ENST00000451310.1 |
SEPT2 |
septin 2 |
| chr21_-_30391636 | 0.26 |
ENST00000493196.1 |
RWDD2B |
RWD domain containing 2B |
| chr1_+_33546714 | 0.26 |
ENST00000294517.6 ENST00000358680.3 ENST00000373443.3 ENST00000398167.1 |
ADC |
arginine decarboxylase |
| chr14_+_52164820 | 0.26 |
ENST00000554167.1 |
FRMD6 |
FERM domain containing 6 |
| chr16_+_22501658 | 0.26 |
ENST00000415833.2 |
NPIPB5 |
nuclear pore complex interacting protein family, member B5 |
| chr10_-_4285923 | 0.25 |
ENST00000418372.1 ENST00000608792.1 |
LINC00702 |
long intergenic non-protein coding RNA 702 |
| chr4_+_86699834 | 0.25 |
ENST00000395183.2 |
ARHGAP24 |
Rho GTPase activating protein 24 |
| chr3_+_183353356 | 0.25 |
ENST00000242810.6 ENST00000493074.1 ENST00000437402.1 ENST00000454495.2 ENST00000473045.1 ENST00000468101.1 ENST00000427201.2 ENST00000482138.1 ENST00000454652.2 |
KLHL24 |
kelch-like family member 24 |
| chr10_+_5135981 | 0.25 |
ENST00000380554.3 |
AKR1C3 |
aldo-keto reductase family 1, member C3 |
| chr19_-_58662139 | 0.25 |
ENST00000598312.1 |
ZNF329 |
zinc finger protein 329 |
| chr10_-_100995603 | 0.24 |
ENST00000370552.3 ENST00000370549.1 |
HPSE2 |
heparanase 2 |
| chr9_-_36400213 | 0.24 |
ENST00000259605.6 ENST00000353739.4 |
RNF38 |
ring finger protein 38 |
| chr11_-_327537 | 0.24 |
ENST00000602735.1 |
IFITM3 |
interferon induced transmembrane protein 3 |
| chr10_-_100995540 | 0.23 |
ENST00000370546.1 ENST00000404542.1 |
HPSE2 |
heparanase 2 |
| chr12_+_10460549 | 0.23 |
ENST00000543420.1 ENST00000543777.1 |
KLRD1 |
killer cell lectin-like receptor subfamily D, member 1 |
| chr3_-_146262488 | 0.22 |
ENST00000487389.1 |
PLSCR1 |
phospholipid scramblase 1 |
| chr12_-_11062161 | 0.22 |
ENST00000390677.2 |
TAS2R13 |
taste receptor, type 2, member 13 |
| chr4_-_48082192 | 0.21 |
ENST00000507351.1 |
TXK |
TXK tyrosine kinase |
| chr19_-_23869970 | 0.21 |
ENST00000601010.1 |
ZNF675 |
zinc finger protein 675 |
| chr1_-_44818599 | 0.20 |
ENST00000537474.1 |
ERI3 |
ERI1 exoribonuclease family member 3 |
| chr17_+_46908350 | 0.20 |
ENST00000258947.3 ENST00000509507.1 ENST00000448105.2 ENST00000570513.1 ENST00000509415.1 ENST00000513119.1 ENST00000416445.2 ENST00000508679.1 ENST00000505071.1 |
CALCOCO2 |
calcium binding and coiled-coil domain 2 |
| chr8_-_48651648 | 0.20 |
ENST00000408965.3 |
CEBPD |
CCAAT/enhancer binding protein (C/EBP), delta |
| chr16_-_4588822 | 0.20 |
ENST00000564828.1 |
CDIP1 |
cell death-inducing p53 target 1 |
| chr12_+_10460417 | 0.20 |
ENST00000381908.3 ENST00000336164.4 ENST00000350274.5 |
KLRD1 |
killer cell lectin-like receptor subfamily D, member 1 |
| chr2_+_131862872 | 0.20 |
ENST00000439822.2 |
PLEKHB2 |
pleckstrin homology domain containing, family B (evectins) member 2 |
| chr12_+_26164645 | 0.20 |
ENST00000542004.1 |
RASSF8 |
Ras association (RalGDS/AF-6) domain family (N-terminal) member 8 |
| chr9_-_15472730 | 0.19 |
ENST00000481862.1 |
PSIP1 |
PC4 and SFRS1 interacting protein 1 |
| chr17_-_9479128 | 0.19 |
ENST00000574431.1 |
STX8 |
syntaxin 8 |
| chr3_-_165555200 | 0.19 |
ENST00000479451.1 ENST00000540653.1 ENST00000488954.1 ENST00000264381.3 |
BCHE |
butyrylcholinesterase |
| chr8_-_30585439 | 0.19 |
ENST00000221130.5 |
GSR |
glutathione reductase |
| chr19_+_36142147 | 0.19 |
ENST00000590618.1 |
COX6B1 |
cytochrome c oxidase subunit VIb polypeptide 1 (ubiquitous) |
| chr20_-_44298878 | 0.19 |
ENST00000324384.3 ENST00000356562.2 |
WFDC11 |
WAP four-disulfide core domain 11 |
| chr16_-_4524543 | 0.19 |
ENST00000573571.1 ENST00000404295.3 ENST00000574425.1 |
NMRAL1 |
NmrA-like family domain containing 1 |
| chr11_+_120255997 | 0.19 |
ENST00000532993.1 |
ARHGEF12 |
Rho guanine nucleotide exchange factor (GEF) 12 |
| chr19_+_13135386 | 0.18 |
ENST00000360105.4 ENST00000588228.1 ENST00000591028.1 |
NFIX |
nuclear factor I/X (CCAAT-binding transcription factor) |
| chr5_+_180682720 | 0.17 |
ENST00000599439.1 |
AC008443.1 |
CDNA: FLJ23158 fis, clone LNG09623; Uncharacterized protein |
| chr4_-_140223670 | 0.17 |
ENST00000394228.1 ENST00000539387.1 |
NDUFC1 |
NADH dehydrogenase (ubiquinone) 1, subcomplex unknown, 1, 6kDa |
| chr22_+_24198890 | 0.17 |
ENST00000345044.6 |
SLC2A11 |
solute carrier family 2 (facilitated glucose transporter), member 11 |
| chr12_+_48166978 | 0.17 |
ENST00000442218.2 |
SLC48A1 |
solute carrier family 48 (heme transporter), member 1 |
| chr6_+_37897735 | 0.17 |
ENST00000373389.5 |
ZFAND3 |
zinc finger, AN1-type domain 3 |
| chr6_+_42749759 | 0.17 |
ENST00000314073.5 |
GLTSCR1L |
GLTSCR1-like |
| chr4_-_140223614 | 0.16 |
ENST00000394223.1 |
NDUFC1 |
NADH dehydrogenase (ubiquinone) 1, subcomplex unknown, 1, 6kDa |
| chr8_+_42552533 | 0.16 |
ENST00000289957.2 |
CHRNB3 |
cholinergic receptor, nicotinic, beta 3 (neuronal) |
| chr3_+_46395219 | 0.16 |
ENST00000445132.2 ENST00000292301.4 |
CCR2 |
chemokine (C-C motif) receptor 2 |
| chr18_+_616711 | 0.16 |
ENST00000579494.1 |
CLUL1 |
clusterin-like 1 (retinal) |
| chr12_-_74686314 | 0.16 |
ENST00000551210.1 ENST00000515416.2 ENST00000549905.1 |
RP11-81H3.2 |
RP11-81H3.2 |
| chr3_+_184529948 | 0.16 |
ENST00000436792.2 ENST00000446204.2 ENST00000422105.1 |
VPS8 |
vacuolar protein sorting 8 homolog (S. cerevisiae) |
| chr1_+_12916941 | 0.16 |
ENST00000240189.2 |
PRAMEF2 |
PRAME family member 2 |
| chr3_+_184529929 | 0.15 |
ENST00000287546.4 ENST00000437079.3 |
VPS8 |
vacuolar protein sorting 8 homolog (S. cerevisiae) |
| chr19_+_3762645 | 0.15 |
ENST00000330133.4 |
MRPL54 |
mitochondrial ribosomal protein L54 |
| chr1_-_115292591 | 0.15 |
ENST00000438362.2 |
CSDE1 |
cold shock domain containing E1, RNA-binding |
| chr19_+_48248779 | 0.14 |
ENST00000246802.5 |
GLTSCR2 |
glioma tumor suppressor candidate region gene 2 |
| chr3_-_146262637 | 0.14 |
ENST00000472349.1 ENST00000342435.4 |
PLSCR1 |
phospholipid scramblase 1 |
| chr5_+_68860949 | 0.14 |
ENST00000507595.1 |
GTF2H2C |
general transcription factor IIH, polypeptide 2C |
| chr20_-_31592113 | 0.14 |
ENST00000420875.1 ENST00000375519.2 ENST00000375523.3 ENST00000356173.3 |
SUN5 |
Sad1 and UNC84 domain containing 5 |
| chr1_+_198608146 | 0.14 |
ENST00000367376.2 ENST00000352140.3 ENST00000594404.1 ENST00000598951.1 ENST00000530727.1 ENST00000442510.2 ENST00000367367.4 ENST00000348564.6 ENST00000367364.1 ENST00000413409.2 |
PTPRC |
protein tyrosine phosphatase, receptor type, C |
| chr20_+_31407692 | 0.14 |
ENST00000375571.5 |
MAPRE1 |
microtubule-associated protein, RP/EB family, member 1 |
| chr1_-_246357029 | 0.13 |
ENST00000391836.2 |
SMYD3 |
SET and MYND domain containing 3 |
| chr1_+_12851545 | 0.13 |
ENST00000332296.7 |
PRAMEF1 |
PRAME family member 1 |
| chr11_-_13517565 | 0.13 |
ENST00000282091.1 ENST00000529816.1 |
PTH |
parathyroid hormone |
| chr18_-_48351743 | 0.13 |
ENST00000588444.1 ENST00000256425.2 ENST00000428869.2 |
MRO |
maestro |
| chr1_+_247712383 | 0.12 |
ENST00000366488.4 ENST00000536561.1 |
GCSAML |
germinal center-associated, signaling and motility-like |
| chr3_+_51663407 | 0.12 |
ENST00000432863.1 ENST00000296477.3 |
RAD54L2 |
RAD54-like 2 (S. cerevisiae) |
| chr11_+_5009424 | 0.12 |
ENST00000300762.1 |
MMP26 |
matrix metallopeptidase 26 |
| chr7_+_134464414 | 0.11 |
ENST00000361901.2 |
CALD1 |
caldesmon 1 |
| chr10_-_61495760 | 0.11 |
ENST00000395347.1 |
SLC16A9 |
solute carrier family 16, member 9 |
| chr22_+_29168652 | 0.11 |
ENST00000249064.4 ENST00000444523.1 ENST00000448492.2 ENST00000421503.2 |
CCDC117 |
coiled-coil domain containing 117 |
| chr12_-_53601055 | 0.11 |
ENST00000552972.1 ENST00000422257.3 ENST00000267082.5 |
ITGB7 |
integrin, beta 7 |
| chr12_-_53601000 | 0.11 |
ENST00000338737.4 ENST00000549086.2 |
ITGB7 |
integrin, beta 7 |
| chr5_+_159848854 | 0.11 |
ENST00000517480.1 ENST00000520452.1 ENST00000393964.1 |
PTTG1 |
pituitary tumor-transforming 1 |
| chr18_+_39535152 | 0.11 |
ENST00000262039.4 ENST00000398870.3 ENST00000586545.1 |
PIK3C3 |
phosphatidylinositol 3-kinase, catalytic subunit type 3 |
| chr12_+_133657461 | 0.10 |
ENST00000412146.2 ENST00000544426.1 ENST00000440984.2 ENST00000319849.3 ENST00000440550.2 |
ZNF140 |
zinc finger protein 140 |
| chr19_+_7011509 | 0.10 |
ENST00000377296.3 |
AC025278.1 |
Uncharacterized protein |
| chr19_-_14992264 | 0.10 |
ENST00000327462.2 |
OR7A17 |
olfactory receptor, family 7, subfamily A, member 17 |
| chr22_+_24407642 | 0.10 |
ENST00000454754.1 ENST00000263119.5 |
CABIN1 |
calcineurin binding protein 1 |
| chr3_+_150126101 | 0.10 |
ENST00000361875.3 ENST00000361136.2 |
TSC22D2 |
TSC22 domain family, member 2 |
| chr4_+_36283213 | 0.10 |
ENST00000357504.3 |
DTHD1 |
death domain containing 1 |
| chr19_-_46195029 | 0.10 |
ENST00000588599.1 ENST00000585392.1 ENST00000590212.1 ENST00000587367.1 ENST00000391932.3 |
SNRPD2 |
small nuclear ribonucleoprotein D2 polypeptide 16.5kDa |
| chr12_+_21284118 | 0.10 |
ENST00000256958.2 |
SLCO1B1 |
solute carrier organic anion transporter family, member 1B1 |
| chr19_-_23869999 | 0.10 |
ENST00000601935.1 ENST00000359788.4 ENST00000600313.1 ENST00000596211.1 ENST00000599168.1 |
ZNF675 |
zinc finger protein 675 |
| chr2_+_131862900 | 0.09 |
ENST00000438882.2 ENST00000538982.1 ENST00000404460.1 |
PLEKHB2 |
pleckstrin homology domain containing, family B (evectins) member 2 |
| chrX_+_36053908 | 0.09 |
ENST00000378660.2 |
CHDC2 |
calponin homology domain containing 2 |
| chr19_+_10123925 | 0.09 |
ENST00000591589.1 ENST00000171214.1 |
RDH8 |
retinol dehydrogenase 8 (all-trans) |
| chr2_-_228244013 | 0.09 |
ENST00000304568.3 |
TM4SF20 |
transmembrane 4 L six family member 20 |
| chr19_+_57640011 | 0.09 |
ENST00000598197.1 |
USP29 |
ubiquitin specific peptidase 29 |
| chr10_+_47894023 | 0.09 |
ENST00000358474.5 |
FAM21B |
family with sequence similarity 21, member B |
| chr2_-_61245363 | 0.09 |
ENST00000316752.6 |
PUS10 |
pseudouridylate synthase 10 |
| chr16_+_81070792 | 0.08 |
ENST00000564241.1 ENST00000565237.1 |
ATMIN |
ATM interactor |
| chr12_-_122107549 | 0.08 |
ENST00000355329.3 |
MORN3 |
MORN repeat containing 3 |
| chr7_+_1727755 | 0.08 |
ENST00000424383.2 |
ELFN1 |
extracellular leucine-rich repeat and fibronectin type III domain containing 1 |
| chr18_+_45778672 | 0.08 |
ENST00000600091.1 |
AC091150.1 |
HCG1818186; Uncharacterized protein |
| chrX_-_24045303 | 0.08 |
ENST00000328046.8 |
KLHL15 |
kelch-like family member 15 |
| chr4_+_158493642 | 0.08 |
ENST00000507108.1 ENST00000455598.1 ENST00000509450.1 |
RP11-364P22.1 |
RP11-364P22.1 |
| chr19_+_24269981 | 0.07 |
ENST00000339642.6 ENST00000357002.4 |
ZNF254 |
zinc finger protein 254 |
| chr22_-_30901637 | 0.07 |
ENST00000381982.3 ENST00000255858.7 ENST00000540456.1 ENST00000392772.2 |
SEC14L4 |
SEC14-like 4 (S. cerevisiae) |
| chr2_-_32490859 | 0.07 |
ENST00000404025.2 |
NLRC4 |
NLR family, CARD domain containing 4 |
| chr18_+_616672 | 0.07 |
ENST00000338387.7 |
CLUL1 |
clusterin-like 1 (retinal) |
| chr18_+_33709834 | 0.07 |
ENST00000358232.6 ENST00000351393.6 ENST00000442325.2 ENST00000423854.2 ENST00000350494.6 ENST00000542824.1 |
ELP2 |
elongator acetyltransferase complex subunit 2 |
| chr12_-_7656357 | 0.07 |
ENST00000396620.3 ENST00000432237.2 ENST00000359156.4 |
CD163 |
CD163 molecule |
| chr5_-_175395283 | 0.07 |
ENST00000513482.1 ENST00000265097.4 |
THOC3 |
THO complex 3 |
| chr12_+_133757995 | 0.07 |
ENST00000536435.2 ENST00000228289.5 ENST00000541211.2 ENST00000500625.3 ENST00000539248.2 ENST00000542711.2 ENST00000536899.2 ENST00000542986.2 ENST00000537565.1 ENST00000541975.2 |
ZNF268 |
zinc finger protein 268 |
| chr19_+_3762703 | 0.07 |
ENST00000589174.1 |
MRPL54 |
mitochondrial ribosomal protein L54 |
| chr7_+_120591170 | 0.06 |
ENST00000431467.1 |
ING3 |
inhibitor of growth family, member 3 |
| chr1_-_228353112 | 0.06 |
ENST00000366713.1 |
IBA57-AS1 |
IBA57 antisense RNA 1 (head to head) |
| chr1_+_241695670 | 0.06 |
ENST00000366557.4 |
KMO |
kynurenine 3-monooxygenase (kynurenine 3-hydroxylase) |
| chrX_+_65382381 | 0.06 |
ENST00000519389.1 |
HEPH |
hephaestin |
| chr19_-_44079611 | 0.06 |
ENST00000594107.1 ENST00000543982.1 |
XRCC1 |
X-ray repair complementing defective repair in Chinese hamster cells 1 |
| chr5_-_34919094 | 0.06 |
ENST00000341754.4 |
RAD1 |
RAD1 homolog (S. pombe) |
| chr5_+_159848807 | 0.06 |
ENST00000352433.5 |
PTTG1 |
pituitary tumor-transforming 1 |
| chr17_-_39341594 | 0.06 |
ENST00000398472.1 |
KRTAP4-1 |
keratin associated protein 4-1 |
| chr7_-_32534850 | 0.05 |
ENST00000409952.3 ENST00000409909.3 |
LSM5 |
LSM5 homolog, U6 small nuclear RNA associated (S. cerevisiae) |
| chr3_-_126327398 | 0.05 |
ENST00000383572.2 |
TXNRD3NB |
thioredoxin reductase 3 neighbor |
| chr19_-_13617247 | 0.05 |
ENST00000573710.2 |
CACNA1A |
calcium channel, voltage-dependent, P/Q type, alpha 1A subunit |
| chr4_+_37962018 | 0.05 |
ENST00000504686.1 |
PTTG2 |
pituitary tumor-transforming 2 |
| chrX_+_118602363 | 0.05 |
ENST00000317881.8 |
SLC25A5 |
solute carrier family 25 (mitochondrial carrier; adenine nucleotide translocator), member 5 |
| chr17_-_18266797 | 0.05 |
ENST00000316694.3 ENST00000539052.1 |
SHMT1 |
serine hydroxymethyltransferase 1 (soluble) |
| chr7_+_106505912 | 0.04 |
ENST00000359195.3 |
PIK3CG |
phosphatidylinositol-4,5-bisphosphate 3-kinase, catalytic subunit gamma |
| chr10_-_75226166 | 0.04 |
ENST00000544628.1 |
PPP3CB |
protein phosphatase 3, catalytic subunit, beta isozyme |
| chr15_-_54051831 | 0.04 |
ENST00000557913.1 ENST00000360509.5 |
WDR72 |
WD repeat domain 72 |
| chr2_-_73520667 | 0.04 |
ENST00000545030.1 ENST00000436467.2 |
EGR4 |
early growth response 4 |
| chrX_-_14047996 | 0.04 |
ENST00000380523.4 ENST00000398355.3 |
GEMIN8 |
gem (nuclear organelle) associated protein 8 |
Gene Ontology Analysis
Gene overrepresentation in biological_process category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.3 | 1.1 | GO:0086097 | phospholipase C-activating angiotensin-activated signaling pathway(GO:0086097) |
| 0.2 | 0.9 | GO:0072204 | pro-T cell differentiation(GO:0002572) cell-cell signaling involved in kidney development(GO:0060995) Wnt signaling pathway involved in kidney development(GO:0061289) canonical Wnt signaling pathway involved in metanephric kidney development(GO:0061290) cell-cell signaling involved in metanephros development(GO:0072204) negative regulation of DNA demethylation(GO:1901536) |
| 0.2 | 0.6 | GO:0036060 | filtration diaphragm assembly(GO:0036058) slit diaphragm assembly(GO:0036060) |
| 0.2 | 0.8 | GO:0070171 | negative regulation of tooth mineralization(GO:0070171) |
| 0.2 | 0.5 | GO:0006566 | threonine metabolic process(GO:0006566) |
| 0.2 | 1.1 | GO:0043932 | ossification involved in bone remodeling(GO:0043932) |
| 0.1 | 2.2 | GO:1900747 | negative regulation of vascular endothelial growth factor signaling pathway(GO:1900747) |
| 0.1 | 1.6 | GO:0048664 | neuron fate determination(GO:0048664) |
| 0.1 | 1.0 | GO:0007598 | blood coagulation, extrinsic pathway(GO:0007598) |
| 0.1 | 0.8 | GO:0044245 | polysaccharide digestion(GO:0044245) |
| 0.1 | 0.3 | GO:1901491 | negative regulation of lymphangiogenesis(GO:1901491) |
| 0.1 | 0.3 | GO:0045356 | positive regulation of interferon-alpha biosynthetic process(GO:0045356) |
| 0.1 | 0.8 | GO:0034058 | endosomal vesicle fusion(GO:0034058) |
| 0.1 | 0.3 | GO:0050717 | positive regulation of interleukin-1 alpha secretion(GO:0050717) |
| 0.1 | 0.2 | GO:0014016 | neuroblast differentiation(GO:0014016) |
| 0.1 | 0.6 | GO:0010724 | regulation of definitive erythrocyte differentiation(GO:0010724) |
| 0.1 | 0.3 | GO:0016095 | polyprenol catabolic process(GO:0016095) |
| 0.1 | 0.3 | GO:0035105 | sterol regulatory element binding protein import into nucleus(GO:0035105) |
| 0.1 | 0.4 | GO:0090625 | mRNA cleavage involved in gene silencing by siRNA(GO:0090625) |
| 0.1 | 0.3 | GO:1903435 | positive regulation of constitutive secretory pathway(GO:1903435) |
| 0.1 | 0.2 | GO:0003366 | cell-matrix adhesion involved in ameboidal cell migration(GO:0003366) |
| 0.1 | 0.6 | GO:0060352 | cell adhesion molecule production(GO:0060352) |
| 0.1 | 0.2 | GO:2000439 | negative regulation of eosinophil activation(GO:1902567) positive regulation of monocyte extravasation(GO:2000439) |
| 0.1 | 0.3 | GO:2000312 | regulation of kainate selective glutamate receptor activity(GO:2000312) |
| 0.0 | 0.1 | GO:1903031 | regulation of microtubule plus-end binding(GO:1903031) positive regulation of microtubule plus-end binding(GO:1903033) |
| 0.0 | 0.4 | GO:0071896 | protein localization to adherens junction(GO:0071896) |
| 0.0 | 0.1 | GO:1900158 | positive regulation of osteoclast proliferation(GO:0090290) negative regulation of bone mineralization involved in bone maturation(GO:1900158) |
| 0.0 | 0.4 | GO:0006659 | phosphatidylserine biosynthetic process(GO:0006659) regulation of DNA topoisomerase (ATP-hydrolyzing) activity(GO:2000371) positive regulation of DNA topoisomerase (ATP-hydrolyzing) activity(GO:2000373) |
| 0.0 | 0.2 | GO:0060332 | positive regulation of response to interferon-gamma(GO:0060332) positive regulation of interferon-gamma-mediated signaling pathway(GO:0060335) |
| 0.0 | 0.2 | GO:0034334 | adherens junction maintenance(GO:0034334) |
| 0.0 | 0.3 | GO:0003383 | apical constriction(GO:0003383) |
| 0.0 | 0.8 | GO:0051531 | NFAT protein import into nucleus(GO:0051531) |
| 0.0 | 0.5 | GO:0017187 | peptidyl-glutamic acid carboxylation(GO:0017187) protein carboxylation(GO:0018214) |
| 0.0 | 0.3 | GO:0009304 | tRNA transcription(GO:0009304) |
| 0.0 | 0.1 | GO:0002378 | immunoglobulin biosynthetic process(GO:0002378) |
| 0.0 | 0.3 | GO:0046598 | positive regulation of viral entry into host cell(GO:0046598) |
| 0.0 | 0.2 | GO:1901098 | positive regulation of autophagosome maturation(GO:1901098) |
| 0.0 | 0.1 | GO:0070966 | nuclear-transcribed mRNA catabolic process, no-go decay(GO:0070966) |
| 0.0 | 0.5 | GO:0090110 | cargo loading into COPII-coated vesicle(GO:0090110) |
| 0.0 | 0.2 | GO:0000395 | mRNA 5'-splice site recognition(GO:0000395) |
| 0.0 | 0.1 | GO:0051598 | meiotic recombination checkpoint(GO:0051598) |
| 0.0 | 0.7 | GO:0071625 | vocalization behavior(GO:0071625) |
| 0.0 | 0.3 | GO:0097094 | craniofacial suture morphogenesis(GO:0097094) |
| 0.0 | 0.8 | GO:0006198 | cAMP catabolic process(GO:0006198) |
| 0.0 | 0.2 | GO:0015886 | heme transport(GO:0015886) |
| 0.0 | 0.0 | GO:1904481 | response to tetrahydrofolate(GO:1904481) cellular response to tetrahydrofolate(GO:1904482) |
| 0.0 | 0.1 | GO:1903516 | base-excision repair, DNA ligation(GO:0006288) regulation of single strand break repair(GO:1903516) |
| 0.0 | 0.1 | GO:0030242 | pexophagy(GO:0030242) |
| 0.0 | 0.6 | GO:0032008 | positive regulation of TOR signaling(GO:0032008) |
| 0.0 | 0.7 | GO:0051180 | vitamin transport(GO:0051180) |
| 0.0 | 0.0 | GO:0045897 | positive regulation of transcription during mitosis(GO:0045897) |
| 0.0 | 0.4 | GO:0047497 | establishment of mitochondrion localization, microtubule-mediated(GO:0034643) mitochondrion transport along microtubule(GO:0047497) |
| 0.0 | 0.1 | GO:0071630 | nucleus-associated proteasomal ubiquitin-dependent protein catabolic process(GO:0071630) |
| 0.0 | 0.2 | GO:0000050 | urea cycle(GO:0000050) |
| 0.0 | 0.4 | GO:0043153 | entrainment of circadian clock by photoperiod(GO:0043153) |
| 0.0 | 0.3 | GO:0007191 | adenylate cyclase-activating dopamine receptor signaling pathway(GO:0007191) |
| 0.0 | 0.4 | GO:0010842 | retina layer formation(GO:0010842) |
Gene overrepresentation in cellular_component category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.2 | 0.8 | GO:1990578 | perinuclear endoplasmic reticulum membrane(GO:1990578) |
| 0.2 | 2.2 | GO:0098647 | collagen type VI trimer(GO:0005589) collagen beaded filament(GO:0098647) |
| 0.1 | 0.9 | GO:1990111 | spermatoproteasome complex(GO:1990111) |
| 0.1 | 0.3 | GO:0000126 | transcription factor TFIIIB complex(GO:0000126) |
| 0.1 | 0.2 | GO:0034669 | integrin alpha4-beta7 complex(GO:0034669) |
| 0.1 | 0.3 | GO:0033263 | CORVET complex(GO:0033263) |
| 0.0 | 0.6 | GO:0061700 | GATOR2 complex(GO:0061700) |
| 0.0 | 0.3 | GO:0072558 | NLRP1 inflammasome complex(GO:0072558) protease inhibitor complex(GO:0097179) |
| 0.0 | 0.7 | GO:0005883 | neurofilament(GO:0005883) |
| 0.0 | 0.1 | GO:0034272 | phosphatidylinositol 3-kinase complex, class III, type I(GO:0034271) phosphatidylinositol 3-kinase complex, class III, type II(GO:0034272) |
| 0.0 | 0.3 | GO:0005638 | lamin filament(GO:0005638) |
| 0.0 | 0.4 | GO:0097025 | MPP7-DLG1-LIN7 complex(GO:0097025) |
| 0.0 | 0.4 | GO:0035068 | micro-ribonucleoprotein complex(GO:0035068) |
| 0.0 | 0.3 | GO:0032009 | early phagosome(GO:0032009) |
| 0.0 | 0.2 | GO:0098560 | cytoplasmic side of late endosome membrane(GO:0098560) |
| 0.0 | 0.3 | GO:0033093 | Weibel-Palade body(GO:0033093) |
| 0.0 | 0.1 | GO:0030981 | cortical microtubule cytoskeleton(GO:0030981) |
| 0.0 | 0.5 | GO:0030127 | COPII vesicle coat(GO:0030127) |
| 0.0 | 0.3 | GO:0097227 | sperm annulus(GO:0097227) |
| 0.0 | 0.2 | GO:0035327 | transcriptionally active chromatin(GO:0035327) |
| 0.0 | 0.1 | GO:0070937 | CRD-mediated mRNA stability complex(GO:0070937) |
| 0.0 | 0.2 | GO:0005641 | nuclear envelope lumen(GO:0005641) |
| 0.0 | 0.1 | GO:0000439 | core TFIIH complex(GO:0000439) |
| 0.0 | 0.1 | GO:0034715 | pICln-Sm protein complex(GO:0034715) |
| 0.0 | 0.3 | GO:0031235 | intrinsic component of the cytoplasmic side of the plasma membrane(GO:0031235) |
| 0.0 | 0.4 | GO:0000930 | gamma-tubulin complex(GO:0000930) |
| 0.0 | 0.1 | GO:0030666 | endocytic vesicle membrane(GO:0030666) |
| 0.0 | 0.0 | GO:0005944 | phosphatidylinositol 3-kinase complex, class IB(GO:0005944) |
| 0.0 | 0.1 | GO:0072557 | IPAF inflammasome complex(GO:0072557) |
Gene overrepresentation in molecular_function category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.4 | 1.1 | GO:0001596 | angiotensin type I receptor activity(GO:0001596) |
| 0.2 | 0.6 | GO:0003829 | beta-1,3-galactosyl-O-glycosyl-glycoprotein beta-1,6-N-acetylglucosaminyltransferase activity(GO:0003829) |
| 0.2 | 0.7 | GO:0008431 | vitamin E binding(GO:0008431) |
| 0.1 | 0.4 | GO:0004574 | oligo-1,6-glucosidase activity(GO:0004574) |
| 0.1 | 0.5 | GO:0030305 | heparanase activity(GO:0030305) |
| 0.1 | 0.9 | GO:0005134 | interleukin-2 receptor binding(GO:0005134) |
| 0.1 | 0.5 | GO:0016453 | C-acetyltransferase activity(GO:0016453) |
| 0.1 | 0.8 | GO:0043559 | insulin binding(GO:0043559) |
| 0.1 | 0.4 | GO:0016160 | amylase activity(GO:0016160) |
| 0.1 | 0.3 | GO:0001026 | TFIIIB-type transcription factor activity(GO:0001026) |
| 0.1 | 0.7 | GO:0030160 | GKAP/Homer scaffold activity(GO:0030160) |
| 0.1 | 0.4 | GO:0023024 | MHC class I protein complex binding(GO:0023024) |
| 0.1 | 1.1 | GO:0036122 | BMP binding(GO:0036122) |
| 0.1 | 1.6 | GO:0071837 | HMG box domain binding(GO:0071837) |
| 0.1 | 0.2 | GO:0004362 | glutathione-disulfide reductase activity(GO:0004362) |
| 0.1 | 0.3 | GO:0035671 | enone reductase activity(GO:0035671) |
| 0.0 | 0.2 | GO:0003990 | acetylcholinesterase activity(GO:0003990) |
| 0.0 | 1.1 | GO:0005001 | transmembrane receptor protein tyrosine phosphatase activity(GO:0005001) transmembrane receptor protein phosphatase activity(GO:0019198) |
| 0.0 | 0.1 | GO:0031727 | CCR2 chemokine receptor binding(GO:0031727) |
| 0.0 | 0.9 | GO:0070003 | threonine-type endopeptidase activity(GO:0004298) threonine-type peptidase activity(GO:0070003) |
| 0.0 | 2.2 | GO:0050840 | extracellular matrix binding(GO:0050840) |
| 0.0 | 0.4 | GO:0035198 | miRNA binding(GO:0035198) |
| 0.0 | 0.4 | GO:0017128 | phospholipid scramblase activity(GO:0017128) |
| 0.0 | 0.8 | GO:0004115 | 3',5'-cyclic-AMP phosphodiesterase activity(GO:0004115) |
| 0.0 | 0.3 | GO:0035197 | siRNA binding(GO:0035197) |
| 0.0 | 0.2 | GO:0019869 | chloride channel inhibitor activity(GO:0019869) |
| 0.0 | 0.2 | GO:0097100 | supercoiled DNA binding(GO:0097100) |
| 0.0 | 0.1 | GO:0001162 | RNA polymerase II intronic transcription regulatory region sequence-specific DNA binding(GO:0001162) |
| 0.0 | 0.2 | GO:0015232 | heme transporter activity(GO:0015232) |
| 0.0 | 0.3 | GO:0050700 | CARD domain binding(GO:0050700) |
| 0.0 | 0.3 | GO:0031489 | myosin V binding(GO:0031489) |
| 0.0 | 0.0 | GO:0015207 | ATP:ADP antiporter activity(GO:0005471) adenine transmembrane transporter activity(GO:0015207) |
| 0.0 | 0.1 | GO:0015349 | thyroid hormone transmembrane transporter activity(GO:0015349) |
| 0.0 | 0.0 | GO:0008386 | cholesterol monooxygenase (side-chain-cleaving) activity(GO:0008386) |
Gene overrepresentation in C2:CP category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.0 | 3.0 | NABA PROTEOGLYCANS | Genes encoding proteoglycans |
| 0.0 | 0.8 | PID IL5 PATHWAY | IL5-mediated signaling events |
| 0.0 | 0.9 | PID IL27 PATHWAY | IL27-mediated signaling events |
| 0.0 | 1.1 | PID ARF6 PATHWAY | Arf6 signaling events |
| 0.0 | 1.0 | PID SYNDECAN 4 PATHWAY | Syndecan-4-mediated signaling events |
| 0.0 | 0.1 | ST INTERFERON GAMMA PATHWAY | Interferon gamma pathway. |
| 0.0 | 0.4 | PID NECTIN PATHWAY | Nectin adhesion pathway |
| 0.0 | 0.3 | PID ANTHRAX PATHWAY | Cellular roles of Anthrax toxin |
Gene overrepresentation in C2:CP:REACTOME category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.1 | 2.2 | REACTOME CS DS DEGRADATION | Genes involved in CS/DS degradation |
| 0.1 | 0.8 | REACTOME DIGESTION OF DIETARY CARBOHYDRATE | Genes involved in Digestion of dietary carbohydrate |
| 0.0 | 1.5 | REACTOME FORMATION OF FIBRIN CLOT CLOTTING CASCADE | Genes involved in Formation of Fibrin Clot (Clotting Cascade) |
| 0.0 | 0.8 | REACTOME NEPHRIN INTERACTIONS | Genes involved in Nephrin interactions |
| 0.0 | 0.2 | REACTOME APOBEC3G MEDIATED RESISTANCE TO HIV1 INFECTION | Genes involved in APOBEC3G mediated resistance to HIV-1 infection |
| 0.0 | 0.3 | REACTOME TRAFFICKING AND PROCESSING OF ENDOSOMAL TLR | Genes involved in Trafficking and processing of endosomal TLR |
| 0.0 | 0.5 | REACTOME HS GAG DEGRADATION | Genes involved in HS-GAG degradation |
| 0.0 | 0.3 | REACTOME THE NLRP3 INFLAMMASOME | Genes involved in The NLRP3 inflammasome |
| 0.0 | 0.5 | REACTOME ANTIGEN PRESENTATION FOLDING ASSEMBLY AND PEPTIDE LOADING OF CLASS I MHC | Genes involved in Antigen Presentation: Folding, assembly and peptide loading of class I MHC |
| 0.0 | 0.9 | REACTOME CROSS PRESENTATION OF SOLUBLE EXOGENOUS ANTIGENS ENDOSOMES | Genes involved in Cross-presentation of soluble exogenous antigens (endosomes) |