Project
Epithelial-Mesenchymal Transition, human (Scheel, 2011)
Navigation
Downloads
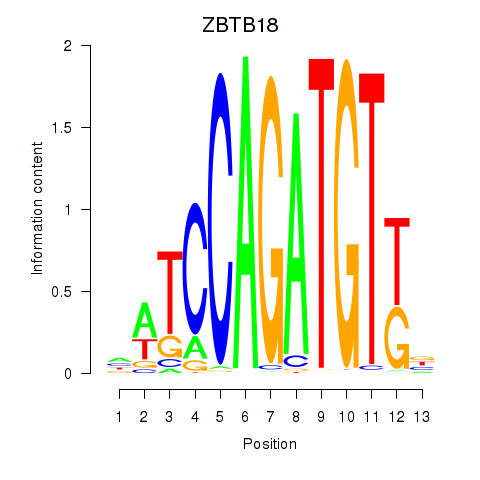
Results for ZBTB18
Z-value: 0.79
Transcription factors associated with ZBTB18
| Gene Symbol | Gene ID | Gene Info |
|---|---|---|
|
ZBTB18
|
ENSG00000179456.9 | ZBTB18 |
Activity-expression correlation:
| Gene | Promoter | Pearson corr. coef. | P-value | Plot |
|---|---|---|---|---|
| ZBTB18 | hg19_v2_chr1_+_244214577_244214593 | -0.47 | 2.4e-01 | Click! |
Activity profile of ZBTB18 motif
Sorted Z-values of ZBTB18 motif
Network of associatons between targets according to the STRING database.
First level regulatory network of ZBTB18
| Promoter | Score | Transcript | Gene | Gene Info |
|---|---|---|---|---|
| chr2_+_189839046 | 1.88 |
ENST00000304636.3 ENST00000317840.5 |
COL3A1 |
collagen, type III, alpha 1 |
| chr1_-_168698433 | 1.11 |
ENST00000367817.3 |
DPT |
dermatopontin |
| chr2_+_152214098 | 0.96 |
ENST00000243347.3 |
TNFAIP6 |
tumor necrosis factor, alpha-induced protein 6 |
| chr2_+_33359646 | 0.92 |
ENST00000390003.4 ENST00000418533.2 |
LTBP1 |
latent transforming growth factor beta binding protein 1 |
| chr2_+_33359687 | 0.91 |
ENST00000402934.1 ENST00000404525.1 ENST00000407925.1 |
LTBP1 |
latent transforming growth factor beta binding protein 1 |
| chr15_-_90358048 | 0.82 |
ENST00000300060.6 ENST00000560137.1 |
ANPEP |
alanyl (membrane) aminopeptidase |
| chr2_-_145278475 | 0.81 |
ENST00000558170.2 |
ZEB2 |
zinc finger E-box binding homeobox 2 |
| chr2_-_163100045 | 0.81 |
ENST00000188790.4 |
FAP |
fibroblast activation protein, alpha |
| chr2_-_163099885 | 0.79 |
ENST00000443424.1 |
FAP |
fibroblast activation protein, alpha |
| chr12_+_1738363 | 0.76 |
ENST00000397196.2 |
WNT5B |
wingless-type MMTV integration site family, member 5B |
| chr11_+_15136462 | 0.70 |
ENST00000379556.3 ENST00000424273.1 |
INSC |
inscuteable homolog (Drosophila) |
| chr11_+_5410607 | 0.66 |
ENST00000328611.3 |
OR51M1 |
olfactory receptor, family 51, subfamily M, member 1 |
| chr1_+_78354297 | 0.63 |
ENST00000334785.7 |
NEXN |
nexilin (F actin binding protein) |
| chr18_+_6834472 | 0.54 |
ENST00000581099.1 ENST00000419673.2 ENST00000531294.1 |
ARHGAP28 |
Rho GTPase activating protein 28 |
| chr17_-_19651668 | 0.52 |
ENST00000494157.2 ENST00000225740.6 |
ALDH3A1 |
aldehyde dehydrogenase 3 family, member A1 |
| chr1_+_78354175 | 0.52 |
ENST00000401035.3 ENST00000457030.1 ENST00000330010.8 |
NEXN |
nexilin (F actin binding protein) |
| chr17_-_19651654 | 0.51 |
ENST00000395555.3 |
ALDH3A1 |
aldehyde dehydrogenase 3 family, member A1 |
| chr17_-_67138015 | 0.48 |
ENST00000284425.2 ENST00000590645.1 |
ABCA6 |
ATP-binding cassette, sub-family A (ABC1), member 6 |
| chrX_-_101771645 | 0.48 |
ENST00000289373.4 |
TMSB15A |
thymosin beta 15a |
| chr3_-_114035026 | 0.46 |
ENST00000570269.1 |
RP11-553L6.5 |
RP11-553L6.5 |
| chr13_-_33780133 | 0.46 |
ENST00000399365.3 |
STARD13 |
StAR-related lipid transfer (START) domain containing 13 |
| chr1_-_153518270 | 0.45 |
ENST00000354332.4 ENST00000368716.4 |
S100A4 |
S100 calcium binding protein A4 |
| chr17_-_15168624 | 0.44 |
ENST00000312280.3 ENST00000494511.1 ENST00000580584.1 |
PMP22 |
peripheral myelin protein 22 |
| chrX_+_64808248 | 0.41 |
ENST00000609672.1 |
MSN |
moesin |
| chr6_+_155537771 | 0.39 |
ENST00000275246.7 |
TIAM2 |
T-cell lymphoma invasion and metastasis 2 |
| chr3_-_160167508 | 0.38 |
ENST00000479460.1 |
TRIM59 |
tripartite motif containing 59 |
| chr17_-_8263538 | 0.38 |
ENST00000535173.1 |
AC135178.1 |
HCG1985372; Uncharacterized protein; cDNA FLJ37541 fis, clone BRCAN2026340 |
| chr8_-_11058847 | 0.36 |
ENST00000297303.4 ENST00000416569.2 |
XKR6 |
XK, Kell blood group complex subunit-related family, member 6 |
| chr5_-_180688105 | 0.35 |
ENST00000327767.4 |
TRIM52 |
tripartite motif containing 52 |
| chr2_-_230096756 | 0.34 |
ENST00000354069.6 |
PID1 |
phosphotyrosine interaction domain containing 1 |
| chr1_+_38022513 | 0.33 |
ENST00000296218.7 |
DNALI1 |
dynein, axonemal, light intermediate chain 1 |
| chr16_-_18470696 | 0.33 |
ENST00000427999.2 |
RP11-1212A22.4 |
LOC339047 protein; Nuclear pore complex-interacting protein family member A3; Nuclear pore complex-interacting protein family member A5; Protein PKD1P1 |
| chrX_+_43515467 | 0.32 |
ENST00000338702.3 ENST00000542639.1 |
MAOA |
monoamine oxidase A |
| chr12_-_104443890 | 0.31 |
ENST00000547583.1 ENST00000360814.4 ENST00000546851.1 |
GLT8D2 |
glycosyltransferase 8 domain containing 2 |
| chrX_+_103031421 | 0.31 |
ENST00000433491.1 ENST00000418604.1 ENST00000443502.1 |
PLP1 |
proteolipid protein 1 |
| chrX_+_103031758 | 0.31 |
ENST00000303958.2 ENST00000361621.2 |
PLP1 |
proteolipid protein 1 |
| chr3_-_105588231 | 0.29 |
ENST00000545639.1 ENST00000394027.3 ENST00000438603.1 ENST00000447441.1 ENST00000443752.1 |
CBLB |
Cbl proto-oncogene B, E3 ubiquitin protein ligase |
| chr7_+_100770328 | 0.29 |
ENST00000223095.4 ENST00000445463.2 |
SERPINE1 |
serpin peptidase inhibitor, clade E (nexin, plasminogen activator inhibitor type 1), member 1 |
| chr7_+_90225796 | 0.27 |
ENST00000380050.3 |
CDK14 |
cyclin-dependent kinase 14 |
| chr6_-_74363803 | 0.27 |
ENST00000355773.5 |
SLC17A5 |
solute carrier family 17 (acidic sugar transporter), member 5 |
| chrX_+_99899180 | 0.26 |
ENST00000373004.3 |
SRPX2 |
sushi-repeat containing protein, X-linked 2 |
| chr3_-_160167301 | 0.26 |
ENST00000494486.1 |
TRIM59 |
tripartite motif containing 59 |
| chr5_+_140792614 | 0.26 |
ENST00000398610.2 |
PCDHGA10 |
protocadherin gamma subfamily A, 10 |
| chr2_+_219247021 | 0.25 |
ENST00000539932.1 |
SLC11A1 |
solute carrier family 11 (proton-coupled divalent metal ion transporter), member 1 |
| chr12_-_56122426 | 0.25 |
ENST00000551173.1 |
CD63 |
CD63 molecule |
| chr9_+_97488939 | 0.25 |
ENST00000277198.2 ENST00000297979.5 |
C9orf3 |
chromosome 9 open reading frame 3 |
| chr3_-_178789220 | 0.25 |
ENST00000414084.1 |
ZMAT3 |
zinc finger, matrin-type 3 |
| chr5_+_140782351 | 0.23 |
ENST00000573521.1 |
PCDHGA9 |
protocadherin gamma subfamily A, 9 |
| chr1_+_211433275 | 0.22 |
ENST00000367005.4 |
RCOR3 |
REST corepressor 3 |
| chr1_-_156647189 | 0.22 |
ENST00000368223.3 |
NES |
nestin |
| chr1_-_32210275 | 0.22 |
ENST00000440175.2 |
BAI2 |
brain-specific angiogenesis inhibitor 2 |
| chr3_+_100211412 | 0.21 |
ENST00000323523.4 ENST00000403410.1 ENST00000449609.1 |
TMEM45A |
transmembrane protein 45A |
| chr2_-_85829780 | 0.20 |
ENST00000334462.5 |
TMEM150A |
transmembrane protein 150A |
| chr2_-_85829811 | 0.20 |
ENST00000306353.3 |
TMEM150A |
transmembrane protein 150A |
| chr9_+_118916082 | 0.19 |
ENST00000328252.3 |
PAPPA |
pregnancy-associated plasma protein A, pappalysin 1 |
| chr12_-_56121612 | 0.19 |
ENST00000546939.1 |
CD63 |
CD63 molecule |
| chr1_+_38022572 | 0.19 |
ENST00000541606.1 |
DNALI1 |
dynein, axonemal, light intermediate chain 1 |
| chr12_-_53893399 | 0.19 |
ENST00000267079.2 |
MAP3K12 |
mitogen-activated protein kinase kinase kinase 12 |
| chr19_+_39903185 | 0.19 |
ENST00000409794.3 |
PLEKHG2 |
pleckstrin homology domain containing, family G (with RhoGef domain) member 2 |
| chr19_-_10341948 | 0.19 |
ENST00000590320.1 ENST00000592342.1 ENST00000588952.1 |
S1PR2 DNMT1 |
sphingosine-1-phosphate receptor 2 DNA (cytosine-5-)-methyltransferase 1 |
| chr20_+_57875658 | 0.18 |
ENST00000371025.3 |
EDN3 |
endothelin 3 |
| chr16_-_31076332 | 0.18 |
ENST00000539836.3 ENST00000535577.1 ENST00000442862.2 |
ZNF668 |
zinc finger protein 668 |
| chr16_-_67185117 | 0.18 |
ENST00000449549.3 |
B3GNT9 |
UDP-GlcNAc:betaGal beta-1,3-N-acetylglucosaminyltransferase 9 |
| chr12_-_56121580 | 0.18 |
ENST00000550776.1 |
CD63 |
CD63 molecule |
| chr12_-_56122220 | 0.18 |
ENST00000552692.1 |
CD63 |
CD63 molecule |
| chr2_-_85829496 | 0.18 |
ENST00000409668.1 |
TMEM150A |
transmembrane protein 150A |
| chr9_+_131683174 | 0.18 |
ENST00000372592.3 ENST00000428610.1 |
PHYHD1 |
phytanoyl-CoA dioxygenase domain containing 1 |
| chr8_+_1772132 | 0.18 |
ENST00000349830.3 ENST00000520359.1 ENST00000518288.1 ENST00000398560.1 |
ARHGEF10 |
Rho guanine nucleotide exchange factor (GEF) 10 |
| chr16_-_49698136 | 0.17 |
ENST00000535559.1 |
ZNF423 |
zinc finger protein 423 |
| chr2_+_201994042 | 0.17 |
ENST00000417748.1 |
CFLAR |
CASP8 and FADD-like apoptosis regulator |
| chr17_-_62050278 | 0.17 |
ENST00000578147.1 ENST00000435607.1 |
SCN4A |
sodium channel, voltage-gated, type IV, alpha subunit |
| chr2_-_152146385 | 0.17 |
ENST00000414946.1 ENST00000243346.5 |
NMI |
N-myc (and STAT) interactor |
| chr1_-_113160826 | 0.16 |
ENST00000538187.1 ENST00000369664.1 |
ST7L |
suppression of tumorigenicity 7 like |
| chr15_-_74043816 | 0.16 |
ENST00000379822.4 |
C15orf59 |
chromosome 15 open reading frame 59 |
| chr8_+_38965048 | 0.16 |
ENST00000399831.3 ENST00000437682.2 ENST00000519315.1 ENST00000379907.4 ENST00000522506.1 |
ADAM32 |
ADAM metallopeptidase domain 32 |
| chr11_-_68780824 | 0.16 |
ENST00000441623.1 ENST00000309099.6 |
MRGPRF |
MAS-related GPR, member F |
| chr12_+_26126681 | 0.16 |
ENST00000542865.1 |
RASSF8 |
Ras association (RalGDS/AF-6) domain family (N-terminal) member 8 |
| chrX_+_153770421 | 0.15 |
ENST00000369609.5 ENST00000369607.1 |
IKBKG |
inhibitor of kappa light polypeptide gene enhancer in B-cells, kinase gamma |
| chr16_-_29910853 | 0.15 |
ENST00000308713.5 |
SEZ6L2 |
seizure related 6 homolog (mouse)-like 2 |
| chr5_+_49962495 | 0.15 |
ENST00000515175.1 |
PARP8 |
poly (ADP-ribose) polymerase family, member 8 |
| chr12_-_54121261 | 0.14 |
ENST00000549784.1 ENST00000262059.4 |
CALCOCO1 |
calcium binding and coiled-coil domain 1 |
| chr12_-_56122124 | 0.14 |
ENST00000552754.1 |
CD63 |
CD63 molecule |
| chr5_+_118690466 | 0.13 |
ENST00000503646.1 |
TNFAIP8 |
tumor necrosis factor, alpha-induced protein 8 |
| chr12_-_53893227 | 0.13 |
ENST00000547488.1 |
MAP3K12 |
mitogen-activated protein kinase kinase kinase 12 |
| chr22_+_42372931 | 0.13 |
ENST00000328414.8 ENST00000396425.3 |
SEPT3 |
septin 3 |
| chr4_+_156824840 | 0.13 |
ENST00000536354.2 |
TDO2 |
tryptophan 2,3-dioxygenase |
| chr14_+_64970662 | 0.12 |
ENST00000556965.1 ENST00000554015.1 |
ZBTB1 |
zinc finger and BTB domain containing 1 |
| chr3_-_160167540 | 0.12 |
ENST00000496222.1 ENST00000471396.1 ENST00000471155.1 ENST00000309784.4 |
TRIM59 |
tripartite motif containing 59 |
| chr16_-_30393752 | 0.12 |
ENST00000566517.1 ENST00000605106.1 |
SEPT1 SEPT1 |
septin 1 Uncharacterized protein |
| chr5_-_149324306 | 0.12 |
ENST00000255266.5 |
PDE6A |
phosphodiesterase 6A, cGMP-specific, rod, alpha |
| chr6_-_153304148 | 0.11 |
ENST00000229758.3 |
FBXO5 |
F-box protein 5 |
| chr19_+_50936142 | 0.11 |
ENST00000357701.5 |
MYBPC2 |
myosin binding protein C, fast type |
| chr9_-_13165457 | 0.11 |
ENST00000542239.1 ENST00000538841.1 ENST00000433359.2 |
MPDZ |
multiple PDZ domain protein |
| chr20_+_30467600 | 0.11 |
ENST00000375934.4 ENST00000375922.4 |
TTLL9 |
tubulin tyrosine ligase-like family, member 9 |
| chr20_+_57875758 | 0.11 |
ENST00000395654.3 |
EDN3 |
endothelin 3 |
| chr15_+_67418047 | 0.10 |
ENST00000540846.2 |
SMAD3 |
SMAD family member 3 |
| chr11_+_94227129 | 0.10 |
ENST00000540349.1 ENST00000535502.1 ENST00000545130.1 ENST00000544253.1 ENST00000541144.1 |
ANKRD49 |
ankyrin repeat domain 49 |
| chr11_+_32112431 | 0.10 |
ENST00000054950.3 |
RCN1 |
reticulocalbin 1, EF-hand calcium binding domain |
| chr1_+_53527854 | 0.10 |
ENST00000371500.3 ENST00000395871.2 ENST00000312553.5 |
PODN |
podocan |
| chr9_+_72658490 | 0.09 |
ENST00000377182.4 |
MAMDC2 |
MAM domain containing 2 |
| chr3_+_183894566 | 0.09 |
ENST00000439647.1 |
AP2M1 |
adaptor-related protein complex 2, mu 1 subunit |
| chr12_-_48119301 | 0.09 |
ENST00000545824.2 ENST00000422538.3 |
ENDOU |
endonuclease, polyU-specific |
| chr17_+_48638371 | 0.09 |
ENST00000360761.4 ENST00000352832.5 ENST00000354983.4 |
CACNA1G |
calcium channel, voltage-dependent, T type, alpha 1G subunit |
| chr12_-_50222187 | 0.09 |
ENST00000335999.6 |
NCKAP5L |
NCK-associated protein 5-like |
| chr12_-_48119340 | 0.09 |
ENST00000229003.3 |
ENDOU |
endonuclease, polyU-specific |
| chr14_+_64971292 | 0.08 |
ENST00000358738.3 ENST00000394712.2 |
ZBTB1 |
zinc finger and BTB domain containing 1 |
| chr10_+_97733786 | 0.08 |
ENST00000371198.2 |
CC2D2B |
coiled-coil and C2 domain containing 2B |
| chr3_-_52273098 | 0.08 |
ENST00000499914.2 ENST00000305533.5 ENST00000597542.1 |
TWF2 TLR9 |
twinfilin actin-binding protein 2 toll-like receptor 9 |
| chr15_+_51669444 | 0.08 |
ENST00000396399.2 |
GLDN |
gliomedin |
| chr6_-_16761678 | 0.08 |
ENST00000244769.4 ENST00000436367.1 |
ATXN1 |
ataxin 1 |
| chr5_-_151304337 | 0.08 |
ENST00000455880.2 ENST00000545569.1 ENST00000274576.4 |
GLRA1 |
glycine receptor, alpha 1 |
| chr7_-_154863264 | 0.08 |
ENST00000395731.2 ENST00000543018.1 |
HTR5A-AS1 |
HTR5A antisense RNA 1 |
| chr17_+_32582293 | 0.08 |
ENST00000580907.1 ENST00000225831.4 |
CCL2 |
chemokine (C-C motif) ligand 2 |
| chr9_+_100174344 | 0.08 |
ENST00000422139.2 |
TDRD7 |
tudor domain containing 7 |
| chr1_-_224517823 | 0.08 |
ENST00000469968.1 ENST00000436927.1 ENST00000469075.1 ENST00000488718.1 ENST00000482491.1 ENST00000340871.4 ENST00000492281.1 ENST00000361463.3 ENST00000391875.2 ENST00000461546.1 |
NVL |
nuclear VCP-like |
| chr16_+_6069664 | 0.08 |
ENST00000422070.4 |
RBFOX1 |
RNA binding protein, fox-1 homolog (C. elegans) 1 |
| chr15_+_75335604 | 0.07 |
ENST00000563393.1 |
PPCDC |
phosphopantothenoylcysteine decarboxylase |
| chr17_+_77020224 | 0.07 |
ENST00000339142.2 |
C1QTNF1 |
C1q and tumor necrosis factor related protein 1 |
| chr17_+_77020325 | 0.07 |
ENST00000311661.4 |
C1QTNF1 |
C1q and tumor necrosis factor related protein 1 |
| chr6_+_163148973 | 0.07 |
ENST00000366888.2 |
PACRG |
PARK2 co-regulated |
| chr9_+_2029019 | 0.07 |
ENST00000382194.1 |
SMARCA2 |
SWI/SNF related, matrix associated, actin dependent regulator of chromatin, subfamily a, member 2 |
| chr16_-_87367879 | 0.07 |
ENST00000568879.1 |
RP11-178L8.4 |
RP11-178L8.4 |
| chr10_-_7829909 | 0.07 |
ENST00000379562.4 ENST00000543003.1 ENST00000535925.1 |
KIN |
KIN, antigenic determinant of recA protein homolog (mouse) |
| chr5_+_118812237 | 0.06 |
ENST00000513628.1 |
HSD17B4 |
hydroxysteroid (17-beta) dehydrogenase 4 |
| chr11_+_71900703 | 0.06 |
ENST00000393681.2 |
FOLR1 |
folate receptor 1 (adult) |
| chr5_+_118812294 | 0.06 |
ENST00000509514.1 |
HSD17B4 |
hydroxysteroid (17-beta) dehydrogenase 4 |
| chr1_-_151882031 | 0.06 |
ENST00000489410.1 |
THEM4 |
thioesterase superfamily member 4 |
| chr15_+_57210961 | 0.06 |
ENST00000557843.1 |
TCF12 |
transcription factor 12 |
| chr21_-_35883541 | 0.05 |
ENST00000399284.1 |
KCNE1 |
potassium voltage-gated channel, Isk-related family, member 1 |
| chr13_+_113777105 | 0.05 |
ENST00000409306.1 ENST00000375551.3 ENST00000375559.3 |
F10 |
coagulation factor X |
| chr7_-_135433460 | 0.05 |
ENST00000415751.1 |
FAM180A |
family with sequence similarity 180, member A |
| chr2_-_89399845 | 0.05 |
ENST00000479981.1 |
IGKV1-16 |
immunoglobulin kappa variable 1-16 |
| chr9_+_100174232 | 0.05 |
ENST00000355295.4 |
TDRD7 |
tudor domain containing 7 |
| chr19_+_7584088 | 0.05 |
ENST00000394341.2 |
ZNF358 |
zinc finger protein 358 |
| chrX_-_48056199 | 0.05 |
ENST00000311798.1 ENST00000347757.1 |
SSX5 |
synovial sarcoma, X breakpoint 5 |
| chr16_-_30394143 | 0.05 |
ENST00000321367.3 ENST00000571393.1 |
SEPT1 |
septin 1 |
| chr11_-_5462744 | 0.05 |
ENST00000380211.1 |
OR51I1 |
olfactory receptor, family 51, subfamily I, member 1 |
| chr19_+_41119323 | 0.05 |
ENST00000599724.1 ENST00000597071.1 ENST00000243562.9 |
LTBP4 |
latent transforming growth factor beta binding protein 4 |
| chr1_-_99470368 | 0.05 |
ENST00000263177.4 |
LPPR5 |
Lipid phosphate phosphatase-related protein type 5 |
| chr15_-_57210769 | 0.05 |
ENST00000559000.1 |
ZNF280D |
zinc finger protein 280D |
| chr6_+_160542870 | 0.05 |
ENST00000324965.4 ENST00000457470.2 |
SLC22A1 |
solute carrier family 22 (organic cation transporter), member 1 |
| chr1_-_99470558 | 0.04 |
ENST00000370188.3 |
LPPR5 |
Lipid phosphate phosphatase-related protein type 5 |
| chr9_+_116263639 | 0.04 |
ENST00000343817.5 |
RGS3 |
regulator of G-protein signaling 3 |
| chr15_-_31453359 | 0.04 |
ENST00000542188.1 |
TRPM1 |
transient receptor potential cation channel, subfamily M, member 1 |
| chr22_-_37213554 | 0.04 |
ENST00000443735.1 |
PVALB |
parvalbumin |
| chr2_+_108994466 | 0.04 |
ENST00000272452.2 |
SULT1C4 |
sulfotransferase family, cytosolic, 1C, member 4 |
| chr2_+_177053307 | 0.04 |
ENST00000331462.4 |
HOXD1 |
homeobox D1 |
| chrY_-_2655644 | 0.04 |
ENST00000525526.2 ENST00000534739.2 ENST00000383070.1 |
SRY |
sex determining region Y |
| chr11_-_62323702 | 0.04 |
ENST00000530285.1 |
AHNAK |
AHNAK nucleoprotein |
| chr9_+_116263778 | 0.03 |
ENST00000394646.3 |
RGS3 |
regulator of G-protein signaling 3 |
| chr3_+_159570722 | 0.03 |
ENST00000482804.1 |
SCHIP1 |
schwannomin interacting protein 1 |
| chr15_-_74501360 | 0.03 |
ENST00000323940.5 |
STRA6 |
stimulated by retinoic acid 6 |
| chr11_-_5537920 | 0.03 |
ENST00000380184.1 |
UBQLNL |
ubiquilin-like |
| chr15_-_43559055 | 0.02 |
ENST00000220420.5 ENST00000349114.4 |
TGM5 |
transglutaminase 5 |
| chr4_-_100009856 | 0.02 |
ENST00000296412.8 |
ADH5 |
alcohol dehydrogenase 5 (class III), chi polypeptide |
| chr2_-_89417335 | 0.02 |
ENST00000490686.1 |
IGKV1-17 |
immunoglobulin kappa variable 1-17 |
| chr14_+_67707826 | 0.02 |
ENST00000261681.4 |
MPP5 |
membrane protein, palmitoylated 5 (MAGUK p55 subfamily member 5) |
| chr5_+_159895275 | 0.02 |
ENST00000517927.1 |
MIR146A |
microRNA 146a |
| chr7_+_62809239 | 0.02 |
ENST00000456890.1 |
AC006455.1 |
AC006455.1 |
| chr9_+_125137565 | 0.02 |
ENST00000373698.5 |
PTGS1 |
prostaglandin-endoperoxide synthase 1 (prostaglandin G/H synthase and cyclooxygenase) |
| chr15_+_55611128 | 0.02 |
ENST00000164305.5 ENST00000539642.1 |
PIGB |
phosphatidylinositol glycan anchor biosynthesis, class B |
| chr6_+_160542821 | 0.02 |
ENST00000366963.4 |
SLC22A1 |
solute carrier family 22 (organic cation transporter), member 1 |
| chr19_+_42806250 | 0.01 |
ENST00000598490.1 ENST00000341747.3 |
PRR19 |
proline rich 19 |
| chr4_-_105416039 | 0.01 |
ENST00000394767.2 |
CXXC4 |
CXXC finger protein 4 |
| chr15_+_57210818 | 0.01 |
ENST00000438423.2 ENST00000267811.5 ENST00000452095.2 ENST00000559609.1 ENST00000333725.5 |
TCF12 |
transcription factor 12 |
| chr11_-_63684316 | 0.01 |
ENST00000301459.4 |
RCOR2 |
REST corepressor 2 |
| chr8_-_49834299 | 0.01 |
ENST00000396822.1 |
SNAI2 |
snail family zinc finger 2 |
| chr1_+_9005917 | 0.01 |
ENST00000549778.1 ENST00000480186.3 ENST00000377443.2 ENST00000377436.3 ENST00000377442.2 |
CA6 |
carbonic anhydrase VI |
| chr8_-_49833978 | 0.01 |
ENST00000020945.1 |
SNAI2 |
snail family zinc finger 2 |
| chr10_+_105253661 | 0.01 |
ENST00000369780.4 |
NEURL |
neuralized E3 ubiquitin protein ligase 1 |
| chr19_+_9251052 | 0.01 |
ENST00000247956.6 ENST00000360385.3 |
ZNF317 |
zinc finger protein 317 |
| chr14_-_38725573 | 0.00 |
ENST00000342213.2 |
CLEC14A |
C-type lectin domain family 14, member A |
| chr2_-_7005785 | 0.00 |
ENST00000256722.5 ENST00000404168.1 ENST00000458098.1 |
CMPK2 |
cytidine monophosphate (UMP-CMP) kinase 2, mitochondrial |
| chr10_+_7830125 | 0.00 |
ENST00000335698.4 ENST00000541227.1 |
ATP5C1 |
ATP synthase, H+ transporting, mitochondrial F1 complex, gamma polypeptide 1 |
| chr11_-_34379546 | 0.00 |
ENST00000435224.2 |
ABTB2 |
ankyrin repeat and BTB (POZ) domain containing 2 |
| chr4_-_152149033 | 0.00 |
ENST00000514152.1 |
SH3D19 |
SH3 domain containing 19 |
| chr2_+_90139056 | 0.00 |
ENST00000492446.1 |
IGKV1D-16 |
immunoglobulin kappa variable 1D-16 |
Gene Ontology Analysis
Gene overrepresentation in biological_process category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.4 | 1.6 | GO:0097325 | melanocyte proliferation(GO:0097325) |
| 0.2 | 1.9 | GO:0060414 | aorta smooth muscle tissue morphogenesis(GO:0060414) |
| 0.2 | 1.8 | GO:0035583 | sequestering of TGFbeta in extracellular matrix(GO:0035583) |
| 0.2 | 0.5 | GO:1904425 | negative regulation of GTP binding(GO:1904425) |
| 0.1 | 0.8 | GO:1902748 | positive regulation of lens fiber cell differentiation(GO:1902748) |
| 0.1 | 0.3 | GO:2000097 | chronological cell aging(GO:0001300) regulation of smooth muscle cell-matrix adhesion(GO:2000097) |
| 0.1 | 0.3 | GO:2001170 | negative regulation of mitochondrial DNA replication(GO:0090298) negative regulation of mitochondrial DNA metabolic process(GO:1901859) negative regulation of ATP biosynthetic process(GO:2001170) |
| 0.1 | 0.3 | GO:0055073 | cadmium ion homeostasis(GO:0055073) divalent metal ion export(GO:0070839) |
| 0.1 | 0.3 | GO:2000672 | negative regulation of motor neuron apoptotic process(GO:2000672) |
| 0.1 | 0.5 | GO:0097498 | endothelial tube lumen extension(GO:0097498) |
| 0.1 | 0.9 | GO:2000680 | rubidium ion transport(GO:0035826) regulation of rubidium ion transport(GO:2000680) |
| 0.1 | 1.2 | GO:0048739 | cardiac muscle fiber development(GO:0048739) |
| 0.1 | 0.4 | GO:0022614 | membrane to membrane docking(GO:0022614) |
| 0.1 | 0.2 | GO:0002572 | pro-T cell differentiation(GO:0002572) |
| 0.1 | 0.8 | GO:0070307 | lens fiber cell development(GO:0070307) |
| 0.0 | 0.3 | GO:0014826 | cellular magnesium ion homeostasis(GO:0010961) vein smooth muscle contraction(GO:0014826) |
| 0.0 | 0.1 | GO:0019442 | tryptophan catabolic process to acetyl-CoA(GO:0019442) |
| 0.0 | 0.2 | GO:0045349 | interferon-alpha biosynthetic process(GO:0045349) regulation of interferon-alpha biosynthetic process(GO:0045354) |
| 0.0 | 0.1 | GO:0007057 | spindle assembly involved in female meiosis I(GO:0007057) |
| 0.0 | 0.2 | GO:0090309 | positive regulation of methylation-dependent chromatin silencing(GO:0090309) |
| 0.0 | 0.6 | GO:0042759 | long-chain fatty acid biosynthetic process(GO:0042759) |
| 0.0 | 0.3 | GO:0090050 | positive regulation of cell migration involved in sprouting angiogenesis(GO:0090050) |
| 0.0 | 0.2 | GO:0034334 | adherens junction maintenance(GO:0034334) |
| 0.0 | 1.0 | GO:0030728 | ovulation(GO:0030728) |
| 0.0 | 0.1 | GO:2000502 | negative regulation of natural killer cell chemotaxis(GO:2000502) |
| 0.0 | 0.8 | GO:0046597 | negative regulation of viral entry into host cell(GO:0046597) |
| 0.0 | 1.1 | GO:0030199 | collagen fibril organization(GO:0030199) |
| 0.0 | 0.4 | GO:0032060 | bleb assembly(GO:0032060) |
| 0.0 | 0.1 | GO:0061713 | neural crest cell migration involved in heart formation(GO:0003147) anterior neural tube closure(GO:0061713) |
| 0.0 | 0.1 | GO:0061767 | negative regulation of lung blood pressure(GO:0061767) |
| 0.0 | 0.1 | GO:0036111 | very long-chain fatty-acyl-CoA metabolic process(GO:0036111) |
| 0.0 | 0.3 | GO:0042420 | dopamine catabolic process(GO:0042420) |
| 0.0 | 0.2 | GO:1903943 | regulation of hepatocyte apoptotic process(GO:1903943) negative regulation of hepatocyte apoptotic process(GO:1903944) |
| 0.0 | 0.8 | GO:0043171 | peptide catabolic process(GO:0043171) |
| 0.0 | 0.3 | GO:0007175 | negative regulation of epidermal growth factor-activated receptor activity(GO:0007175) |
| 0.0 | 0.1 | GO:0051970 | negative regulation of transmission of nerve impulse(GO:0051970) |
| 0.0 | 0.1 | GO:0086046 | membrane depolarization during SA node cell action potential(GO:0086046) |
Gene overrepresentation in cellular_component category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.1 | 1.6 | GO:0071438 | invadopodium membrane(GO:0071438) |
| 0.1 | 1.9 | GO:0098643 | fibrillar collagen trimer(GO:0005583) banded collagen fibril(GO:0098643) |
| 0.1 | 1.8 | GO:0001527 | microfibril(GO:0001527) fibril(GO:0043205) |
| 0.1 | 0.9 | GO:0097487 | multivesicular body, internal vesicle(GO:0097487) |
| 0.1 | 0.3 | GO:0000308 | cytoplasmic cyclin-dependent protein kinase holoenzyme complex(GO:0000308) |
| 0.0 | 0.8 | GO:0030992 | intraciliary transport particle B(GO:0030992) |
| 0.0 | 0.7 | GO:0005938 | cell cortex(GO:0005938) |
| 0.0 | 0.4 | GO:0071437 | invadopodium(GO:0071437) |
| 0.0 | 0.6 | GO:0043218 | compact myelin(GO:0043218) |
| 0.0 | 0.1 | GO:0071144 | SMAD2-SMAD3 protein complex(GO:0071144) |
| 0.0 | 0.1 | GO:0042272 | nuclear RNA export factor complex(GO:0042272) |
| 0.0 | 1.0 | GO:1904724 | tertiary granule lumen(GO:1904724) |
Gene overrepresentation in molecular_function category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.4 | 1.8 | GO:0050436 | microfibril binding(GO:0050436) |
| 0.2 | 1.0 | GO:0018479 | benzaldehyde dehydrogenase (NAD+) activity(GO:0018479) |
| 0.1 | 1.9 | GO:0048407 | platelet-derived growth factor binding(GO:0048407) |
| 0.1 | 0.3 | GO:0051139 | metal ion:proton antiporter activity(GO:0051139) |
| 0.1 | 1.6 | GO:0008239 | dipeptidyl-peptidase activity(GO:0008239) |
| 0.1 | 0.3 | GO:0031708 | endothelin B receptor binding(GO:0031708) |
| 0.0 | 0.3 | GO:0008131 | primary amine oxidase activity(GO:0008131) |
| 0.0 | 0.6 | GO:0019911 | structural constituent of myelin sheath(GO:0019911) |
| 0.0 | 0.8 | GO:0070006 | metalloaminopeptidase activity(GO:0070006) |
| 0.0 | 0.1 | GO:0043027 | cysteine-type endopeptidase inhibitor activity involved in apoptotic process(GO:0043027) |
| 0.0 | 0.1 | GO:0004833 | tryptophan 2,3-dioxygenase activity(GO:0004833) |
| 0.0 | 0.2 | GO:0003886 | DNA (cytosine-5-)-methyltransferase activity(GO:0003886) |
| 0.0 | 0.1 | GO:0008332 | low voltage-gated calcium channel activity(GO:0008332) |
| 0.0 | 1.0 | GO:0005540 | hyaluronic acid binding(GO:0005540) |
| 0.0 | 0.1 | GO:0045322 | unmethylated CpG binding(GO:0045322) |
| 0.0 | 0.5 | GO:0045504 | dynein heavy chain binding(GO:0045504) |
| 0.0 | 0.5 | GO:0050786 | RAGE receptor binding(GO:0050786) |
| 0.0 | 0.1 | GO:0031962 | mineralocorticoid receptor binding(GO:0031962) |
| 0.0 | 0.1 | GO:0033989 | 3alpha,7alpha,12alpha-trihydroxy-5beta-cholest-24-enoyl-CoA hydratase activity(GO:0033989) 17-beta-hydroxysteroid dehydrogenase (NAD+) activity(GO:0044594) |
| 0.0 | 0.1 | GO:0031727 | CCR2 chemokine receptor binding(GO:0031727) |
| 0.0 | 0.8 | GO:0070412 | R-SMAD binding(GO:0070412) |
| 0.0 | 0.1 | GO:0005277 | acetylcholine transmembrane transporter activity(GO:0005277) secondary active organic cation transmembrane transporter activity(GO:0008513) acetate ester transmembrane transporter activity(GO:1901375) |
| 0.0 | 0.1 | GO:0016933 | extracellular-glycine-gated ion channel activity(GO:0016933) extracellular-glycine-gated chloride channel activity(GO:0016934) |
| 0.0 | 0.1 | GO:0061714 | folic acid receptor activity(GO:0061714) |
| 0.0 | 0.6 | GO:0005109 | frizzled binding(GO:0005109) |
| 0.0 | 0.2 | GO:0019215 | intermediate filament binding(GO:0019215) |
| 0.0 | 0.1 | GO:0034046 | poly(G) binding(GO:0034046) |
| 0.0 | 0.3 | GO:0005351 | sugar:proton symporter activity(GO:0005351) cation:sugar symporter activity(GO:0005402) |
| 0.0 | 0.4 | GO:0070530 | K63-linked polyubiquitin binding(GO:0070530) |
| 0.0 | 1.3 | GO:0008307 | structural constituent of muscle(GO:0008307) |
| 0.0 | 0.1 | GO:0055104 | ligase inhibitor activity(GO:0055104) ubiquitin ligase inhibitor activity(GO:1990948) |
| 0.0 | 0.5 | GO:0003785 | actin monomer binding(GO:0003785) |
Gene overrepresentation in C2:CP category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.0 | 1.9 | NABA COLLAGENS | Genes encoding collagen proteins |
| 0.0 | 4.0 | NABA ECM GLYCOPROTEINS | Genes encoding structural ECM glycoproteins |
| 0.0 | 0.4 | SA FAS SIGNALING | The TNF-type receptor Fas induces apoptosis on ligand binding. |
Gene overrepresentation in C2:CP:REACTOME category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.0 | 1.7 | REACTOME NCAM1 INTERACTIONS | Genes involved in NCAM1 interactions |
| 0.0 | 0.5 | REACTOME ABCA TRANSPORTERS IN LIPID HOMEOSTASIS | Genes involved in ABCA transporters in lipid homeostasis |
| 0.0 | 0.3 | REACTOME NOREPINEPHRINE NEUROTRANSMITTER RELEASE CYCLE | Genes involved in Norepinephrine Neurotransmitter Release Cycle |
| 0.0 | 0.2 | REACTOME IKK COMPLEX RECRUITMENT MEDIATED BY RIP1 | Genes involved in IKK complex recruitment mediated by RIP1 |