Project
Epithelial-Mesenchymal Transition, human (Scheel, 2011)
Navigation
Downloads
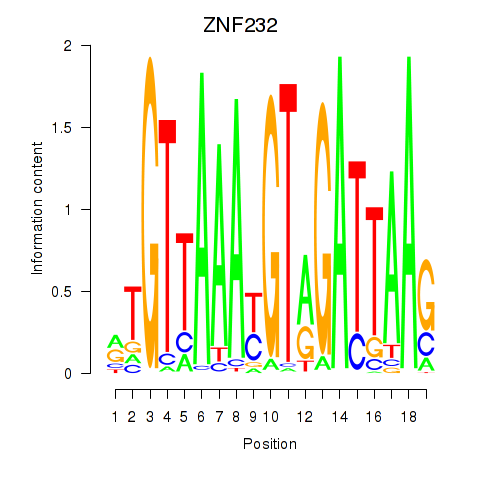
Results for ZNF232
Z-value: 0.21
Transcription factors associated with ZNF232
| Gene Symbol | Gene ID | Gene Info |
|---|---|---|
|
ZNF232
|
ENSG00000167840.9 | ZNF232 |
Activity-expression correlation:
| Gene | Promoter | Pearson corr. coef. | P-value | Plot |
|---|---|---|---|---|
| ZNF232 | hg19_v2_chr17_-_5026397_5026422 | 0.21 | 6.2e-01 | Click! |
Activity profile of ZNF232 motif
Sorted Z-values of ZNF232 motif
Network of associatons between targets according to the STRING database.
First level regulatory network of ZNF232
| Promoter | Score | Transcript | Gene | Gene Info |
|---|---|---|---|---|
| chr2_-_225811747 | 0.32 |
ENST00000409592.3 |
DOCK10 |
dedicator of cytokinesis 10 |
| chr3_+_8543393 | 0.21 |
ENST00000157600.3 ENST00000415597.1 ENST00000535732.1 |
LMCD1 |
LIM and cysteine-rich domains 1 |
| chr3_+_8543561 | 0.17 |
ENST00000397386.3 |
LMCD1 |
LIM and cysteine-rich domains 1 |
| chr3_+_8543533 | 0.16 |
ENST00000454244.1 |
LMCD1 |
LIM and cysteine-rich domains 1 |
| chr1_-_79472365 | 0.13 |
ENST00000370742.3 |
ELTD1 |
EGF, latrophilin and seven transmembrane domain containing 1 |
| chr17_+_39975455 | 0.12 |
ENST00000455106.1 |
FKBP10 |
FK506 binding protein 10, 65 kDa |
| chr17_+_39975544 | 0.11 |
ENST00000544340.1 |
FKBP10 |
FK506 binding protein 10, 65 kDa |
| chr7_+_142829162 | 0.07 |
ENST00000291009.3 |
PIP |
prolactin-induced protein |
| chr16_+_30662360 | 0.06 |
ENST00000542965.2 |
PRR14 |
proline rich 14 |
| chr9_+_5231413 | 0.06 |
ENST00000239316.4 |
INSL4 |
insulin-like 4 (placenta) |
| chr3_+_40566369 | 0.06 |
ENST00000403205.2 ENST00000310898.1 ENST00000339296.5 ENST00000431278.1 |
ZNF621 |
zinc finger protein 621 |
| chr16_+_30662050 | 0.06 |
ENST00000568754.1 |
PRR14 |
proline rich 14 |
| chr10_+_134150835 | 0.05 |
ENST00000432555.2 |
LRRC27 |
leucine rich repeat containing 27 |
| chr20_-_33732952 | 0.05 |
ENST00000541621.1 |
EDEM2 |
ER degradation enhancer, mannosidase alpha-like 2 |
| chr11_+_66276550 | 0.05 |
ENST00000419755.3 |
CTD-3074O7.11 |
Bardet-Biedl syndrome 1 protein |
| chr16_+_30662184 | 0.05 |
ENST00000300835.4 |
PRR14 |
proline rich 14 |
| chr15_-_56757329 | 0.04 |
ENST00000260453.3 |
MNS1 |
meiosis-specific nuclear structural 1 |
| chr2_-_74618964 | 0.04 |
ENST00000417090.1 ENST00000409868.1 |
DCTN1 |
dynactin 1 |
| chr2_+_241807870 | 0.04 |
ENST00000307503.3 |
AGXT |
alanine-glyoxylate aminotransferase |
| chr1_-_53686261 | 0.04 |
ENST00000294360.4 |
C1orf123 |
chromosome 1 open reading frame 123 |
| chr1_-_193074504 | 0.04 |
ENST00000367439.3 |
GLRX2 |
glutaredoxin 2 |
| chr7_+_1609694 | 0.04 |
ENST00000437621.2 ENST00000457484.2 |
PSMG3-AS1 |
PSMG3 antisense RNA 1 (head to head) |
| chr10_-_115904361 | 0.04 |
ENST00000428953.1 ENST00000543782.1 |
C10orf118 |
chromosome 10 open reading frame 118 |
| chr9_-_115480303 | 0.03 |
ENST00000374234.1 ENST00000374238.1 ENST00000374236.1 ENST00000374242.4 |
INIP |
INTS3 and NABP interacting protein |
| chr3_+_153202284 | 0.03 |
ENST00000446603.2 |
C3orf79 |
chromosome 3 open reading frame 79 |
| chr1_+_156105878 | 0.03 |
ENST00000508500.1 |
LMNA |
lamin A/C |
| chrX_+_100805496 | 0.03 |
ENST00000372829.3 |
ARMCX1 |
armadillo repeat containing, X-linked 1 |
| chr2_-_74619152 | 0.03 |
ENST00000440727.1 ENST00000409240.1 |
DCTN1 |
dynactin 1 |
| chr17_+_26800756 | 0.03 |
ENST00000537681.1 |
SLC13A2 |
solute carrier family 13 (sodium-dependent dicarboxylate transporter), member 2 |
| chr5_+_177435986 | 0.03 |
ENST00000398106.2 |
FAM153C |
family with sequence similarity 153, member C |
| chr4_+_185570871 | 0.03 |
ENST00000512834.1 |
PRIMPOL |
primase and polymerase (DNA-directed) |
| chr17_+_26800648 | 0.03 |
ENST00000545060.1 |
SLC13A2 |
solute carrier family 13 (sodium-dependent dicarboxylate transporter), member 2 |
| chr8_-_143999259 | 0.03 |
ENST00000323110.2 |
CYP11B2 |
cytochrome P450, family 11, subfamily B, polypeptide 2 |
| chr14_-_107131560 | 0.02 |
ENST00000390632.2 |
IGHV3-66 |
immunoglobulin heavy variable 3-66 |
| chr17_+_26800296 | 0.02 |
ENST00000444914.3 ENST00000314669.5 |
SLC13A2 |
solute carrier family 13 (sodium-dependent dicarboxylate transporter), member 2 |
| chrX_+_30261847 | 0.02 |
ENST00000378981.3 ENST00000397550.1 |
MAGEB1 |
melanoma antigen family B, 1 |
| chr10_-_27389392 | 0.02 |
ENST00000376087.4 |
ANKRD26 |
ankyrin repeat domain 26 |
| chrX_+_36065053 | 0.02 |
ENST00000313548.4 |
CHDC2 |
calponin homology domain containing 2 |
| chr15_-_60771128 | 0.02 |
ENST00000558512.1 ENST00000561114.1 |
NARG2 |
NMDA receptor regulated 2 |
| chr19_+_18043810 | 0.02 |
ENST00000445755.2 |
CCDC124 |
coiled-coil domain containing 124 |
| chr1_+_182419261 | 0.02 |
ENST00000294854.8 ENST00000542961.1 |
RGSL1 |
regulator of G-protein signaling like 1 |
| chr19_-_50380536 | 0.02 |
ENST00000391832.3 ENST00000391834.2 ENST00000344175.5 |
AKT1S1 |
AKT1 substrate 1 (proline-rich) |
| chr6_-_26056695 | 0.01 |
ENST00000343677.2 |
HIST1H1C |
histone cluster 1, H1c |
| chr4_+_96761238 | 0.01 |
ENST00000295266.4 |
PDHA2 |
pyruvate dehydrogenase (lipoamide) alpha 2 |
| chr21_+_38792602 | 0.01 |
ENST00000398960.2 ENST00000398956.2 |
DYRK1A |
dual-specificity tyrosine-(Y)-phosphorylation regulated kinase 1A |
| chr15_-_60771280 | 0.01 |
ENST00000560072.1 ENST00000560406.1 ENST00000560520.1 ENST00000261520.4 ENST00000439632.1 |
NARG2 |
NMDA receptor regulated 2 |
| chr17_-_6915646 | 0.01 |
ENST00000574377.1 ENST00000399541.2 ENST00000399540.2 ENST00000575727.1 ENST00000573939.1 |
AC027763.2 |
Uncharacterized protein |
| chr20_+_4129426 | 0.01 |
ENST00000339123.6 ENST00000305958.4 ENST00000278795.3 |
SMOX |
spermine oxidase |
| chr1_-_16556038 | 0.01 |
ENST00000375605.2 |
C1orf134 |
chromosome 1 open reading frame 134 |
| chr19_-_10491234 | 0.01 |
ENST00000524462.1 ENST00000531836.1 ENST00000525621.1 |
TYK2 |
tyrosine kinase 2 |
| chr11_-_117800080 | 0.01 |
ENST00000524993.1 ENST00000528626.1 ENST00000445164.2 ENST00000430170.2 ENST00000526090.1 |
TMPRSS13 |
transmembrane protease, serine 13 |
| chr15_-_34635314 | 0.01 |
ENST00000557912.1 ENST00000328848.4 |
NOP10 |
NOP10 ribonucleoprotein |
| chr6_+_109416313 | 0.01 |
ENST00000521277.1 ENST00000517392.1 ENST00000407272.1 ENST00000336977.4 ENST00000519286.1 ENST00000518329.1 ENST00000522461.1 ENST00000518853.1 |
CEP57L1 |
centrosomal protein 57kDa-like 1 |
| chr5_+_31532373 | 0.01 |
ENST00000325366.9 ENST00000355907.3 ENST00000507818.2 |
C5orf22 |
chromosome 5 open reading frame 22 |
| chr19_-_38210622 | 0.01 |
ENST00000591664.1 ENST00000355202.4 |
ZNF607 |
zinc finger protein 607 |
| chr7_-_56101826 | 0.01 |
ENST00000421626.1 |
PSPH |
phosphoserine phosphatase |
| chr11_-_18127566 | 0.00 |
ENST00000532452.1 ENST00000530180.1 ENST00000300013.4 ENST00000529318.1 ENST00000524803.1 |
SAAL1 |
serum amyloid A-like 1 |
| chr4_-_88450612 | 0.00 |
ENST00000418378.1 ENST00000282470.6 |
SPARCL1 |
SPARC-like 1 (hevin) |
| chr17_-_3301704 | 0.00 |
ENST00000322608.2 |
OR1E1 |
olfactory receptor, family 1, subfamily E, member 1 |
| chr1_+_237205476 | 0.00 |
ENST00000366574.2 |
RYR2 |
ryanodine receptor 2 (cardiac) |
| chr3_+_51863433 | 0.00 |
ENST00000444293.1 |
IQCF3 |
IQ motif containing F3 |
| chr12_-_9885888 | 0.00 |
ENST00000327839.3 |
CLECL1 |
C-type lectin-like 1 |
| chr14_-_106816253 | 0.00 |
ENST00000390615.2 |
IGHV3-33 |
immunoglobulin heavy variable 3-33 |
| chr14_-_24711865 | 0.00 |
ENST00000399423.4 ENST00000267415.7 |
TINF2 |
TERF1 (TRF1)-interacting nuclear factor 2 |
Gene Ontology Analysis
Gene overrepresentation in biological_process category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.1 | 0.2 | GO:0000412 | histone peptidyl-prolyl isomerization(GO:0000412) |
| 0.0 | 0.3 | GO:0002315 | marginal zone B cell differentiation(GO:0002315) |
| 0.0 | 0.5 | GO:0070886 | positive regulation of calcineurin-NFAT signaling cascade(GO:0070886) |
| 0.0 | 0.0 | GO:0036511 | trimming of terminal mannose on B branch(GO:0036509) trimming of first mannose on A branch(GO:0036511) trimming of second mannose on A branch(GO:0036512) |
| 0.0 | 0.0 | GO:0009436 | glyoxylate catabolic process(GO:0009436) |
Gene overrepresentation in molecular_function category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.0 | 0.1 | GO:0015361 | low-affinity sodium:dicarboxylate symporter activity(GO:0015361) |