Project
Epithelial-Mesenchymal Transition, human (Scheel, 2011)
Navigation
Downloads
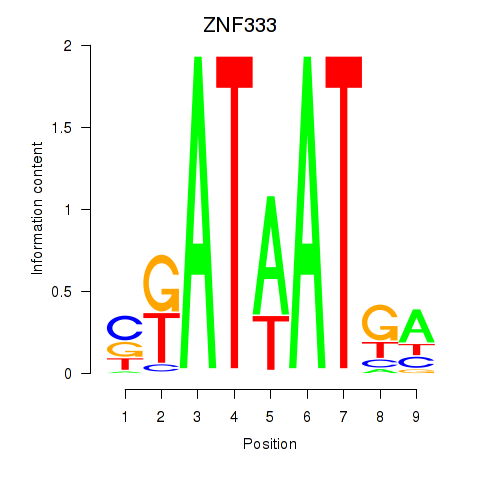
Results for ZNF333
Z-value: 0.66
Transcription factors associated with ZNF333
| Gene Symbol | Gene ID | Gene Info |
|---|---|---|
|
ZNF333
|
ENSG00000160961.7 | ZNF333 |
Activity-expression correlation:
| Gene | Promoter | Pearson corr. coef. | P-value | Plot |
|---|---|---|---|---|
| ZNF333 | hg19_v2_chr19_+_14800711_14800917 | -0.37 | 3.6e-01 | Click! |
Activity profile of ZNF333 motif
Sorted Z-values of ZNF333 motif
Network of associatons between targets according to the STRING database.
First level regulatory network of ZNF333
| Promoter | Score | Transcript | Gene | Gene Info |
|---|---|---|---|---|
| chr12_-_71031185 | 0.94 |
ENST00000548122.1 ENST00000551525.1 ENST00000550358.1 |
PTPRB |
protein tyrosine phosphatase, receptor type, B |
| chr12_-_71031220 | 0.86 |
ENST00000334414.6 |
PTPRB |
protein tyrosine phosphatase, receptor type, B |
| chr3_-_112564797 | 0.54 |
ENST00000398214.1 ENST00000448932.1 |
CD200R1L |
CD200 receptor 1-like |
| chr8_-_139926236 | 0.48 |
ENST00000303045.6 ENST00000435777.1 |
COL22A1 |
collagen, type XXII, alpha 1 |
| chr6_+_29274403 | 0.48 |
ENST00000377160.2 |
OR14J1 |
olfactory receptor, family 14, subfamily J, member 1 |
| chr7_-_83824169 | 0.46 |
ENST00000265362.4 |
SEMA3A |
sema domain, immunoglobulin domain (Ig), short basic domain, secreted, (semaphorin) 3A |
| chr8_+_18248755 | 0.39 |
ENST00000286479.3 |
NAT2 |
N-acetyltransferase 2 (arylamine N-acetyltransferase) |
| chr6_+_130339710 | 0.28 |
ENST00000526087.1 ENST00000533560.1 ENST00000361794.2 |
L3MBTL3 |
l(3)mbt-like 3 (Drosophila) |
| chr9_-_39239171 | 0.27 |
ENST00000358144.2 |
CNTNAP3 |
contactin associated protein-like 3 |
| chr18_+_28956740 | 0.26 |
ENST00000308128.4 ENST00000359747.4 |
DSG4 |
desmoglein 4 |
| chr5_-_16509101 | 0.26 |
ENST00000399793.2 |
FAM134B |
family with sequence similarity 134, member B |
| chrX_-_114253536 | 0.26 |
ENST00000371936.1 |
IL13RA2 |
interleukin 13 receptor, alpha 2 |
| chr5_+_140207536 | 0.24 |
ENST00000529310.1 ENST00000527624.1 |
PCDHA6 |
protocadherin alpha 6 |
| chr1_+_19638788 | 0.23 |
ENST00000375155.3 ENST00000375153.3 ENST00000400548.2 |
PQLC2 |
PQ loop repeat containing 2 |
| chr6_-_32908792 | 0.23 |
ENST00000418107.2 |
HLA-DMB |
major histocompatibility complex, class II, DM beta |
| chr13_-_46716969 | 0.22 |
ENST00000435666.2 |
LCP1 |
lymphocyte cytosolic protein 1 (L-plastin) |
| chr14_+_23842018 | 0.22 |
ENST00000397242.2 ENST00000329715.2 |
IL25 |
interleukin 25 |
| chr1_+_159409512 | 0.21 |
ENST00000423932.3 |
OR10J1 |
olfactory receptor, family 10, subfamily J, member 1 |
| chr11_-_85376121 | 0.20 |
ENST00000527447.1 |
CREBZF |
CREB/ATF bZIP transcription factor |
| chr1_-_11024258 | 0.20 |
ENST00000418570.2 |
C1orf127 |
chromosome 1 open reading frame 127 |
| chr6_-_32908765 | 0.19 |
ENST00000416244.2 |
HLA-DMB |
major histocompatibility complex, class II, DM beta |
| chr1_+_36789335 | 0.18 |
ENST00000373137.2 |
RP11-268J15.5 |
RP11-268J15.5 |
| chr1_-_169599314 | 0.17 |
ENST00000367786.2 ENST00000458599.2 ENST00000367795.2 ENST00000263686.6 |
SELP |
selectin P (granule membrane protein 140kDa, antigen CD62) |
| chr1_-_169599353 | 0.16 |
ENST00000367793.2 ENST00000367794.2 ENST00000367792.2 ENST00000367791.2 ENST00000367788.2 |
SELP |
selectin P (granule membrane protein 140kDa, antigen CD62) |
| chr13_-_103719196 | 0.16 |
ENST00000245312.3 |
SLC10A2 |
solute carrier family 10 (sodium/bile acid cotransporter), member 2 |
| chr17_-_3337135 | 0.15 |
ENST00000248384.1 |
OR1E2 |
olfactory receptor, family 1, subfamily E, member 2 |
| chr6_+_160327974 | 0.15 |
ENST00000252660.4 |
MAS1 |
MAS1 oncogene |
| chr7_-_36406750 | 0.15 |
ENST00000453212.1 ENST00000415803.2 ENST00000440378.1 ENST00000431396.1 ENST00000317020.6 ENST00000436884.1 |
KIAA0895 |
KIAA0895 |
| chr11_-_72504681 | 0.14 |
ENST00000538536.1 ENST00000543304.1 ENST00000540587.1 ENST00000334805.6 |
STARD10 |
StAR-related lipid transfer (START) domain containing 10 |
| chr5_-_78281775 | 0.14 |
ENST00000396151.3 ENST00000565165.1 |
ARSB |
arylsulfatase B |
| chr4_+_110749143 | 0.14 |
ENST00000317735.4 |
RRH |
retinal pigment epithelium-derived rhodopsin homolog |
| chr1_+_218458625 | 0.14 |
ENST00000366932.3 |
RRP15 |
ribosomal RNA processing 15 homolog (S. cerevisiae) |
| chr3_+_101659682 | 0.13 |
ENST00000465215.1 |
RP11-221J22.1 |
RP11-221J22.1 |
| chr4_-_66536057 | 0.13 |
ENST00000273854.3 |
EPHA5 |
EPH receptor A5 |
| chrX_-_119445306 | 0.13 |
ENST00000371369.4 ENST00000440464.1 ENST00000519908.1 |
TMEM255A |
transmembrane protein 255A |
| chrX_-_107975917 | 0.13 |
ENST00000563887.1 |
RP6-24A23.6 |
Uncharacterized protein |
| chr3_-_194393206 | 0.12 |
ENST00000265245.5 |
LSG1 |
large 60S subunit nuclear export GTPase 1 |
| chr18_+_71815743 | 0.12 |
ENST00000169551.6 ENST00000580087.1 |
TIMM21 |
translocase of inner mitochondrial membrane 21 homolog (yeast) |
| chr8_-_11725549 | 0.12 |
ENST00000505496.2 ENST00000534636.1 ENST00000524500.1 ENST00000345125.3 ENST00000453527.2 ENST00000527215.2 ENST00000532392.1 ENST00000533455.1 ENST00000534510.1 ENST00000530640.2 ENST00000531089.1 ENST00000532656.2 ENST00000531502.1 ENST00000434271.1 ENST00000353047.6 |
CTSB |
cathepsin B |
| chr9_+_133971909 | 0.11 |
ENST00000247291.3 ENST00000372302.1 ENST00000372300.1 ENST00000372298.1 |
AIF1L |
allograft inflammatory factor 1-like |
| chr17_-_41623716 | 0.11 |
ENST00000319349.5 |
ETV4 |
ets variant 4 |
| chr9_+_133971863 | 0.11 |
ENST00000372309.3 |
AIF1L |
allograft inflammatory factor 1-like |
| chr1_+_95975672 | 0.11 |
ENST00000440116.2 ENST00000456933.1 |
RP11-286B14.1 |
RP11-286B14.1 |
| chr12_+_122880045 | 0.11 |
ENST00000539034.1 ENST00000535976.1 |
RP11-450K4.1 |
RP11-450K4.1 |
| chr3_-_113918254 | 0.11 |
ENST00000460779.1 |
DRD3 |
dopamine receptor D3 |
| chr2_-_178753465 | 0.10 |
ENST00000389683.3 |
PDE11A |
phosphodiesterase 11A |
| chr22_+_22516550 | 0.10 |
ENST00000390284.2 |
IGLV4-60 |
immunoglobulin lambda variable 4-60 |
| chr17_-_38911580 | 0.10 |
ENST00000312150.4 |
KRT25 |
keratin 25 |
| chr20_-_44600810 | 0.10 |
ENST00000322927.2 ENST00000426788.1 |
ZNF335 |
zinc finger protein 335 |
| chrX_-_30595959 | 0.10 |
ENST00000378962.3 |
CXorf21 |
chromosome X open reading frame 21 |
| chr19_-_19626838 | 0.10 |
ENST00000360913.3 |
TSSK6 |
testis-specific serine kinase 6 |
| chr6_+_166945369 | 0.10 |
ENST00000598601.1 |
Z98049.1 |
CDNA FLJ25492 fis, clone CBR01389; Uncharacterized protein |
| chr2_+_191002486 | 0.10 |
ENST00000396974.2 |
C2orf88 |
chromosome 2 open reading frame 88 |
| chr4_-_100065440 | 0.09 |
ENST00000508393.1 ENST00000265512.7 |
ADH4 |
alcohol dehydrogenase 4 (class II), pi polypeptide |
| chr18_-_40857493 | 0.09 |
ENST00000255224.3 |
SYT4 |
synaptotagmin IV |
| chr14_+_85996471 | 0.09 |
ENST00000330753.4 |
FLRT2 |
fibronectin leucine rich transmembrane protein 2 |
| chr1_+_55013889 | 0.09 |
ENST00000343744.2 ENST00000371316.3 |
ACOT11 |
acyl-CoA thioesterase 11 |
| chr11_-_72504637 | 0.09 |
ENST00000536377.1 ENST00000359373.5 |
STARD10 ARAP1 |
StAR-related lipid transfer (START) domain containing 10 ArfGAP with RhoGAP domain, ankyrin repeat and PH domain 1 |
| chr11_-_66313699 | 0.09 |
ENST00000526986.1 ENST00000310442.3 |
ZDHHC24 |
zinc finger, DHHC-type containing 24 |
| chr7_-_130080977 | 0.09 |
ENST00000223208.5 |
CEP41 |
centrosomal protein 41kDa |
| chr11_+_10772534 | 0.09 |
ENST00000361367.2 |
CTR9 |
CTR9, Paf1/RNA polymerase II complex component |
| chr16_+_84328429 | 0.08 |
ENST00000568638.1 |
WFDC1 |
WAP four-disulfide core domain 1 |
| chr5_+_38403637 | 0.08 |
ENST00000336740.6 ENST00000397202.2 |
EGFLAM |
EGF-like, fibronectin type III and laminin G domains |
| chr4_-_177190364 | 0.08 |
ENST00000296525.3 |
ASB5 |
ankyrin repeat and SOCS box containing 5 |
| chr20_+_58515417 | 0.08 |
ENST00000360816.3 |
FAM217B |
family with sequence similarity 217, member B |
| chr3_-_112693865 | 0.08 |
ENST00000471858.1 ENST00000295863.4 ENST00000308611.3 |
CD200R1 |
CD200 receptor 1 |
| chrM_+_10758 | 0.08 |
ENST00000361381.2 |
MT-ND4 |
mitochondrially encoded NADH dehydrogenase 4 |
| chr16_-_28222797 | 0.08 |
ENST00000569951.1 ENST00000565698.1 |
XPO6 |
exportin 6 |
| chr21_-_46964306 | 0.08 |
ENST00000443742.1 ENST00000528477.1 ENST00000567670.1 |
SLC19A1 |
solute carrier family 19 (folate transporter), member 1 |
| chr16_+_84328252 | 0.08 |
ENST00000219454.5 |
WFDC1 |
WAP four-disulfide core domain 1 |
| chr1_-_198990166 | 0.07 |
ENST00000427439.1 |
RP11-16L9.3 |
RP11-16L9.3 |
| chrY_-_6742068 | 0.07 |
ENST00000215479.5 |
AMELY |
amelogenin, Y-linked |
| chr14_+_85996507 | 0.07 |
ENST00000554746.1 |
FLRT2 |
fibronectin leucine rich transmembrane protein 2 |
| chr9_-_21351377 | 0.07 |
ENST00000380210.1 |
IFNA6 |
interferon, alpha 6 |
| chrM_+_7586 | 0.07 |
ENST00000361739.1 |
MT-CO2 |
mitochondrially encoded cytochrome c oxidase II |
| chrX_+_11311533 | 0.07 |
ENST00000380714.3 ENST00000380712.3 ENST00000348912.4 |
AMELX |
amelogenin, X-linked |
| chr6_-_135375986 | 0.07 |
ENST00000525067.1 ENST00000367822.5 ENST00000367837.5 |
HBS1L |
HBS1-like (S. cerevisiae) |
| chr12_-_87232644 | 0.06 |
ENST00000549405.2 |
MGAT4C |
mannosyl (alpha-1,3-)-glycoprotein beta-1,4-N-acetylglucosaminyltransferase, isozyme C (putative) |
| chr7_-_105319536 | 0.06 |
ENST00000477775.1 |
ATXN7L1 |
ataxin 7-like 1 |
| chr4_-_100356551 | 0.06 |
ENST00000209665.4 |
ADH7 |
alcohol dehydrogenase 7 (class IV), mu or sigma polypeptide |
| chr14_+_105046094 | 0.06 |
ENST00000331952.2 |
C14orf180 |
chromosome 14 open reading frame 180 |
| chr3_+_35721182 | 0.06 |
ENST00000413378.1 ENST00000417925.1 |
ARPP21 |
cAMP-regulated phosphoprotein, 21kDa |
| chr5_+_140800638 | 0.06 |
ENST00000398587.2 ENST00000518882.1 |
PCDHGA11 |
protocadherin gamma subfamily A, 11 |
| chr19_-_45004556 | 0.06 |
ENST00000587047.1 ENST00000391956.4 ENST00000221327.4 ENST00000586637.1 ENST00000591064.1 ENST00000592529.1 |
ZNF180 |
zinc finger protein 180 |
| chr3_-_37216055 | 0.06 |
ENST00000336686.4 |
LRRFIP2 |
leucine rich repeat (in FLII) interacting protein 2 |
| chr8_-_114449112 | 0.05 |
ENST00000455883.2 ENST00000352409.3 ENST00000297405.5 |
CSMD3 |
CUB and Sushi multiple domains 3 |
| chr1_+_11994715 | 0.05 |
ENST00000449038.1 ENST00000376369.3 ENST00000429000.2 ENST00000196061.4 |
PLOD1 |
procollagen-lysine, 2-oxoglutarate 5-dioxygenase 1 |
| chr14_-_90798418 | 0.05 |
ENST00000354366.3 |
NRDE2 |
NRDE-2, necessary for RNA interference, domain containing |
| chr4_-_26492076 | 0.05 |
ENST00000295589.3 |
CCKAR |
cholecystokinin A receptor |
| chr10_+_50507181 | 0.05 |
ENST00000323868.4 |
C10orf71 |
chromosome 10 open reading frame 71 |
| chrM_+_10053 | 0.05 |
ENST00000361227.2 |
MT-ND3 |
mitochondrially encoded NADH dehydrogenase 3 |
| chr10_+_50507232 | 0.05 |
ENST00000374144.3 |
C10orf71 |
chromosome 10 open reading frame 71 |
| chr16_+_87636474 | 0.04 |
ENST00000284262.2 |
JPH3 |
junctophilin 3 |
| chr12_+_112563335 | 0.04 |
ENST00000549358.1 ENST00000257604.5 ENST00000548092.1 ENST00000552896.1 |
TRAFD1 |
TRAF-type zinc finger domain containing 1 |
| chr1_-_197036364 | 0.04 |
ENST00000367412.1 |
F13B |
coagulation factor XIII, B polypeptide |
| chr6_-_52705641 | 0.04 |
ENST00000370989.2 |
GSTA5 |
glutathione S-transferase alpha 5 |
| chr20_+_56964253 | 0.04 |
ENST00000395802.3 |
VAPB |
VAMP (vesicle-associated membrane protein)-associated protein B and C |
| chr6_+_46620676 | 0.04 |
ENST00000371347.5 ENST00000411689.2 |
SLC25A27 |
solute carrier family 25, member 27 |
| chr7_-_25268104 | 0.04 |
ENST00000222674.2 |
NPVF |
neuropeptide VF precursor |
| chr9_+_109625378 | 0.04 |
ENST00000277225.5 ENST00000457913.1 ENST00000472574.1 |
ZNF462 |
zinc finger protein 462 |
| chr10_+_121410882 | 0.04 |
ENST00000369085.3 |
BAG3 |
BCL2-associated athanogene 3 |
| chr5_-_147162078 | 0.04 |
ENST00000507386.1 |
JAKMIP2 |
janus kinase and microtubule interacting protein 2 |
| chr17_-_2966901 | 0.04 |
ENST00000575751.1 |
OR1D5 |
olfactory receptor, family 1, subfamily D, member 5 |
| chrX_-_49121165 | 0.04 |
ENST00000376207.4 ENST00000376199.2 |
FOXP3 |
forkhead box P3 |
| chr10_+_90346519 | 0.04 |
ENST00000371939.3 |
LIPJ |
lipase, family member J |
| chr2_-_127977654 | 0.04 |
ENST00000409327.1 |
CYP27C1 |
cytochrome P450, family 27, subfamily C, polypeptide 1 |
| chr5_-_9630463 | 0.04 |
ENST00000382492.2 |
TAS2R1 |
taste receptor, type 2, member 1 |
| chr12_-_7848364 | 0.04 |
ENST00000329913.3 |
GDF3 |
growth differentiation factor 3 |
| chr10_-_125651258 | 0.03 |
ENST00000241305.3 |
CPXM2 |
carboxypeptidase X (M14 family), member 2 |
| chr13_-_99174252 | 0.03 |
ENST00000376547.3 |
STK24 |
serine/threonine kinase 24 |
| chr11_+_92085262 | 0.03 |
ENST00000298047.6 ENST00000409404.2 ENST00000541502.1 |
FAT3 |
FAT atypical cadherin 3 |
| chr6_-_46620522 | 0.03 |
ENST00000275016.2 |
CYP39A1 |
cytochrome P450, family 39, subfamily A, polypeptide 1 |
| chr1_-_205391178 | 0.03 |
ENST00000367153.4 ENST00000367151.2 ENST00000391936.2 ENST00000367149.3 |
LEMD1 |
LEM domain containing 1 |
| chr12_+_112563303 | 0.03 |
ENST00000412615.2 |
TRAFD1 |
TRAF-type zinc finger domain containing 1 |
| chr14_+_22748980 | 0.03 |
ENST00000390465.2 |
TRAV38-2DV8 |
T cell receptor alpha variable 38-2/delta variable 8 |
| chr10_+_114169299 | 0.03 |
ENST00000369410.3 |
ACSL5 |
acyl-CoA synthetase long-chain family member 5 |
| chr4_-_100065419 | 0.03 |
ENST00000504125.1 ENST00000505590.1 |
ADH4 |
alcohol dehydrogenase 4 (class II), pi polypeptide |
| chr12_-_100656134 | 0.03 |
ENST00000548313.1 |
DEPDC4 |
DEP domain containing 4 |
| chr2_+_228337079 | 0.02 |
ENST00000409315.1 ENST00000373671.3 ENST00000409171.1 |
AGFG1 |
ArfGAP with FG repeats 1 |
| chr19_-_19626351 | 0.02 |
ENST00000585580.3 |
TSSK6 |
testis-specific serine kinase 6 |
| chr17_-_60883993 | 0.02 |
ENST00000583803.1 ENST00000456609.2 |
MARCH10 |
membrane-associated ring finger (C3HC4) 10, E3 ubiquitin protein ligase |
| chr5_-_78281603 | 0.02 |
ENST00000264914.4 |
ARSB |
arylsulfatase B |
| chr11_+_74303575 | 0.02 |
ENST00000263681.2 |
POLD3 |
polymerase (DNA-directed), delta 3, accessory subunit |
| chrX_+_122318318 | 0.02 |
ENST00000371251.1 ENST00000371256.5 |
GRIA3 |
glutamate receptor, ionotropic, AMPA 3 |
| chr20_-_33413416 | 0.02 |
ENST00000359003.2 |
NCOA6 |
nuclear receptor coactivator 6 |
| chr10_-_128210005 | 0.02 |
ENST00000284694.7 ENST00000454341.1 ENST00000432642.1 ENST00000392694.1 |
C10orf90 |
chromosome 10 open reading frame 90 |
| chr5_-_27038683 | 0.02 |
ENST00000511822.1 ENST00000231021.4 |
CDH9 |
cadherin 9, type 2 (T1-cadherin) |
| chr6_+_50786414 | 0.02 |
ENST00000344788.3 ENST00000393655.3 ENST00000263046.4 |
TFAP2B |
transcription factor AP-2 beta (activating enhancer binding protein 2 beta) |
| chr20_+_59654146 | 0.02 |
ENST00000441660.1 |
RP5-827L5.1 |
RP5-827L5.1 |
| chr19_-_42721819 | 0.02 |
ENST00000336034.4 ENST00000598200.1 ENST00000598727.1 ENST00000596251.1 |
DEDD2 |
death effector domain containing 2 |
| chr10_+_54074033 | 0.01 |
ENST00000373970.3 |
DKK1 |
dickkopf WNT signaling pathway inhibitor 1 |
| chr11_-_18062872 | 0.01 |
ENST00000250018.2 |
TPH1 |
tryptophan hydroxylase 1 |
| chr6_+_17110726 | 0.01 |
ENST00000354384.5 |
STMND1 |
stathmin domain containing 1 |
| chr5_+_137225158 | 0.01 |
ENST00000290431.5 |
PKD2L2 |
polycystic kidney disease 2-like 2 |
| chr12_-_75905374 | 0.01 |
ENST00000438169.2 ENST00000229214.4 |
KRR1 |
KRR1, small subunit (SSU) processome component, homolog (yeast) |
| chr3_+_35721106 | 0.01 |
ENST00000474696.1 ENST00000412048.1 ENST00000396482.2 ENST00000432682.1 |
ARPP21 |
cAMP-regulated phosphoprotein, 21kDa |
| chr12_-_52995210 | 0.01 |
ENST00000398066.3 |
KRT72 |
keratin 72 |
| chr12_-_52995242 | 0.01 |
ENST00000537672.2 |
KRT72 |
keratin 72 |
| chr11_+_18433840 | 0.01 |
ENST00000541669.1 ENST00000280704.4 |
LDHC |
lactate dehydrogenase C |
| chr12_-_49393092 | 0.01 |
ENST00000421952.2 |
DDN |
dendrin |
| chr22_-_29107919 | 0.01 |
ENST00000434810.1 ENST00000456369.1 |
CHEK2 |
checkpoint kinase 2 |
| chr8_+_40010989 | 0.01 |
ENST00000315792.3 |
C8orf4 |
chromosome 8 open reading frame 4 |
| chr6_+_46620705 | 0.01 |
ENST00000452689.2 |
SLC25A27 |
solute carrier family 25, member 27 |
| chr16_+_81772633 | 0.01 |
ENST00000566191.1 ENST00000565272.1 ENST00000563954.1 ENST00000565054.1 |
RP11-960L18.1 PLCG2 |
RP11-960L18.1 phospholipase C, gamma 2 (phosphatidylinositol-specific) |
| chr1_-_51763661 | 0.01 |
ENST00000530004.1 |
TTC39A |
tetratricopeptide repeat domain 39A |
| chr16_-_20702578 | 0.00 |
ENST00000307493.4 ENST00000219151.4 |
ACSM1 |
acyl-CoA synthetase medium-chain family member 1 |
| chr11_+_94300474 | 0.00 |
ENST00000299001.6 |
PIWIL4 |
piwi-like RNA-mediated gene silencing 4 |
| chr4_-_40632605 | 0.00 |
ENST00000514014.1 |
RBM47 |
RNA binding motif protein 47 |
| chr5_+_137225125 | 0.00 |
ENST00000350250.4 ENST00000508638.1 ENST00000502810.1 ENST00000508883.1 |
PKD2L2 |
polycystic kidney disease 2-like 2 |
| chr15_+_69857515 | 0.00 |
ENST00000559477.1 |
RP11-279F6.1 |
RP11-279F6.1 |
| chr15_+_38746307 | 0.00 |
ENST00000397609.2 ENST00000491535.1 |
FAM98B |
family with sequence similarity 98, member B |
| chr10_+_118350468 | 0.00 |
ENST00000358834.4 ENST00000528052.1 ENST00000442761.1 |
PNLIPRP1 |
pancreatic lipase-related protein 1 |
| chr3_+_107318157 | 0.00 |
ENST00000406780.1 |
BBX |
bobby sox homolog (Drosophila) |
| chr4_+_106473768 | 0.00 |
ENST00000265154.2 ENST00000420470.2 |
ARHGEF38 |
Rho guanine nucleotide exchange factor (GEF) 38 |
Gene Ontology Analysis
Gene overrepresentation in biological_process category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.1 | 0.4 | GO:2001190 | positive regulation of T cell activation via T cell receptor contact with antigen bound to MHC molecule on antigen presenting cell(GO:2001190) |
| 0.1 | 0.5 | GO:0048880 | sensory system development(GO:0048880) |
| 0.1 | 0.2 | GO:0061582 | intestinal epithelial cell migration(GO:0061582) |
| 0.0 | 0.2 | GO:1903401 | L-lysine transmembrane transport(GO:1903401) |
| 0.0 | 0.3 | GO:0043305 | negative regulation of mast cell degranulation(GO:0043305) |
| 0.0 | 0.2 | GO:0009624 | response to nematode(GO:0009624) eosinophil differentiation(GO:0030222) |
| 0.0 | 0.1 | GO:2001160 | regulation of histone H3-K79 methylation(GO:2001160) positive regulation of histone H3-K79 methylation(GO:2001162) |
| 0.0 | 0.2 | GO:0061343 | cell adhesion involved in heart morphogenesis(GO:0061343) |
| 0.0 | 0.1 | GO:0050883 | negative regulation of sodium:proton antiporter activity(GO:0032416) musculoskeletal movement, spinal reflex action(GO:0050883) |
| 0.0 | 0.1 | GO:0098838 | reduced folate transmembrane transport(GO:0098838) |
| 0.0 | 0.3 | GO:0010572 | positive regulation of platelet activation(GO:0010572) |
| 0.0 | 0.1 | GO:0030037 | actin filament reorganization involved in cell cycle(GO:0030037) |
| 0.0 | 0.3 | GO:0043249 | erythrocyte maturation(GO:0043249) |
| 0.0 | 0.1 | GO:0010430 | fatty acid omega-oxidation(GO:0010430) |
| 0.0 | 0.0 | GO:0032831 | regulation of CD4-positive, CD25-positive, alpha-beta regulatory T cell differentiation(GO:0032829) positive regulation of CD4-positive, CD25-positive, alpha-beta regulatory T cell differentiation(GO:0032831) |
| 0.0 | 0.1 | GO:0046946 | hydroxylysine metabolic process(GO:0046946) hydroxylysine biosynthetic process(GO:0046947) |
| 0.0 | 0.1 | GO:0038188 | cholecystokinin signaling pathway(GO:0038188) |
| 0.0 | 0.1 | GO:0031339 | negative regulation of vesicle fusion(GO:0031339) |
| 0.0 | 0.1 | GO:0032782 | bile acid secretion(GO:0032782) |
| 0.0 | 0.2 | GO:0051639 | actin filament network formation(GO:0051639) |
| 0.0 | 0.0 | GO:0070507 | microtubule polymerization or depolymerization(GO:0031109) regulation of microtubule polymerization or depolymerization(GO:0031110) regulation of microtubule cytoskeleton organization(GO:0070507) |
| 0.0 | 0.0 | GO:0051562 | negative regulation of mitochondrial calcium ion concentration(GO:0051562) |
| 0.0 | 1.6 | GO:0035335 | peptidyl-tyrosine dephosphorylation(GO:0035335) |
Gene overrepresentation in cellular_component category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.0 | 0.1 | GO:0032127 | dense core granule membrane(GO:0032127) |
| 0.0 | 0.2 | GO:0046581 | intercellular canaliculus(GO:0046581) |
| 0.0 | 0.3 | GO:0031088 | platelet dense granule membrane(GO:0031088) |
| 0.0 | 0.4 | GO:0042613 | MHC class II protein complex(GO:0042613) |
| 0.0 | 0.1 | GO:0036021 | endolysosome lumen(GO:0036021) |
| 0.0 | 1.8 | GO:0070821 | tertiary granule membrane(GO:0070821) |
Gene overrepresentation in molecular_function category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.1 | 0.4 | GO:0004060 | arylamine N-acetyltransferase activity(GO:0004060) |
| 0.1 | 1.6 | GO:0019198 | transmembrane receptor protein tyrosine phosphatase activity(GO:0005001) transmembrane receptor protein phosphatase activity(GO:0019198) |
| 0.1 | 0.3 | GO:0042806 | fucose binding(GO:0042806) |
| 0.1 | 0.2 | GO:0003943 | N-acetylgalactosamine-4-sulfatase activity(GO:0003943) |
| 0.0 | 0.2 | GO:0008508 | bile acid:sodium symporter activity(GO:0008508) |
| 0.0 | 0.2 | GO:0005503 | all-trans retinal binding(GO:0005503) |
| 0.0 | 0.6 | GO:0045499 | chemorepellent activity(GO:0045499) |
| 0.0 | 0.2 | GO:0015189 | L-lysine transmembrane transporter activity(GO:0015189) |
| 0.0 | 0.5 | GO:0005549 | odorant binding(GO:0005549) |
| 0.0 | 0.1 | GO:0008518 | reduced folate carrier activity(GO:0008518) |
| 0.0 | 0.1 | GO:0046848 | hydroxyapatite binding(GO:0046848) |
| 0.0 | 0.2 | GO:0001595 | angiotensin receptor activity(GO:0001595) angiotensin type II receptor activity(GO:0004945) |
| 0.0 | 0.1 | GO:0033823 | procollagen-lysine 5-dioxygenase activity(GO:0008475) procollagen glucosyltransferase activity(GO:0033823) |
| 0.0 | 0.1 | GO:0004951 | cholecystokinin receptor activity(GO:0004951) |
| 0.0 | 0.1 | GO:0004118 | cGMP-stimulated cyclic-nucleotide phosphodiesterase activity(GO:0004118) |
| 0.0 | 0.1 | GO:0005005 | transmembrane-ephrin receptor activity(GO:0005005) |
| 0.0 | 0.1 | GO:0030348 | syntaxin-3 binding(GO:0030348) |
| 0.0 | 0.1 | GO:0001591 | dopamine neurotransmitter receptor activity, coupled via Gi/Go(GO:0001591) |
| 0.0 | 0.1 | GO:0035276 | aldehyde oxidase activity(GO:0004031) ethanol binding(GO:0035276) |
| 0.0 | 0.4 | GO:0023026 | MHC class II protein complex binding(GO:0023026) |
| 0.0 | 0.1 | GO:0030021 | extracellular matrix structural constituent conferring compression resistance(GO:0030021) structural constituent of tooth enamel(GO:0030345) |
| 0.0 | 0.1 | GO:0008454 | alpha-1,3-mannosylglycoprotein 4-beta-N-acetylglucosaminyltransferase activity(GO:0008454) |
Gene overrepresentation in C2:CP category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.0 | 0.3 | ST IL 13 PATHWAY | Interleukin 13 (IL-13) Pathway |
Gene overrepresentation in C2:CP:REACTOME category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.0 | 0.5 | REACTOME SEMA3A PLEXIN REPULSION SIGNALING BY INHIBITING INTEGRIN ADHESION | Genes involved in SEMA3A-Plexin repulsion signaling by inhibiting Integrin adhesion |
| 0.0 | 0.2 | REACTOME ETHANOL OXIDATION | Genes involved in Ethanol oxidation |