Project
Epithelial-Mesenchymal Transition, human (Scheel, 2011)
Navigation
Downloads
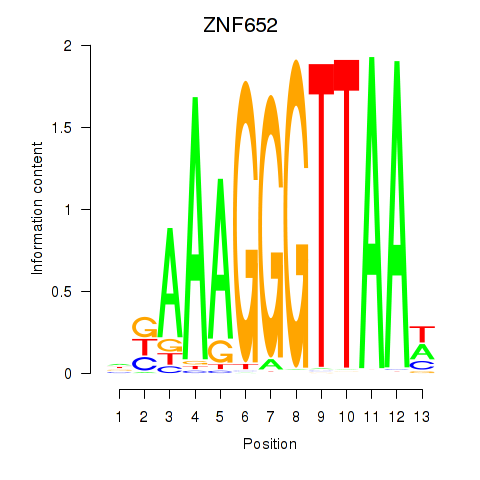
Results for ZNF652
Z-value: 1.26
Transcription factors associated with ZNF652
| Gene Symbol | Gene ID | Gene Info |
|---|---|---|
|
ZNF652
|
ENSG00000198740.4 | ZNF652 |
Activity profile of ZNF652 motif
Sorted Z-values of ZNF652 motif
Network of associatons between targets according to the STRING database.
First level regulatory network of ZNF652
| Promoter | Score | Transcript | Gene | Gene Info |
|---|---|---|---|---|
| chr17_-_19290117 | 1.15 |
ENST00000497081.2 |
MFAP4 |
microfibrillar-associated protein 4 |
| chr6_+_72926145 | 1.09 |
ENST00000425662.2 ENST00000453976.2 |
RIMS1 |
regulating synaptic membrane exocytosis 1 |
| chr4_-_90757364 | 1.00 |
ENST00000508895.1 |
SNCA |
synuclein, alpha (non A4 component of amyloid precursor) |
| chr11_-_35547572 | 0.86 |
ENST00000378880.2 |
PAMR1 |
peptidase domain containing associated with muscle regeneration 1 |
| chr11_-_35547151 | 0.83 |
ENST00000378878.3 ENST00000529303.1 ENST00000278360.3 |
PAMR1 |
peptidase domain containing associated with muscle regeneration 1 |
| chr3_+_148457585 | 0.80 |
ENST00000402260.1 |
AGTR1 |
angiotensin II receptor, type 1 |
| chr17_+_48172639 | 0.72 |
ENST00000503176.1 ENST00000503614.1 |
PDK2 |
pyruvate dehydrogenase kinase, isozyme 2 |
| chr4_-_90756769 | 0.70 |
ENST00000345009.4 ENST00000505199.1 ENST00000502987.1 |
SNCA |
synuclein, alpha (non A4 component of amyloid precursor) |
| chr11_-_111782484 | 0.69 |
ENST00000533971.1 |
CRYAB |
crystallin, alpha B |
| chr17_-_19290483 | 0.68 |
ENST00000395592.2 ENST00000299610.4 |
MFAP4 |
microfibrillar-associated protein 4 |
| chr11_-_111782696 | 0.68 |
ENST00000227251.3 ENST00000526180.1 |
CRYAB |
crystallin, alpha B |
| chrX_+_103031758 | 0.66 |
ENST00000303958.2 ENST00000361621.2 |
PLP1 |
proteolipid protein 1 |
| chr22_-_24641027 | 0.61 |
ENST00000398292.3 ENST00000263112.7 ENST00000418439.2 ENST00000424217.1 ENST00000327365.4 |
GGT5 |
gamma-glutamyltransferase 5 |
| chr16_-_79634595 | 0.60 |
ENST00000326043.4 ENST00000393350.1 |
MAF |
v-maf avian musculoaponeurotic fibrosarcoma oncogene homolog |
| chr1_-_168698433 | 0.58 |
ENST00000367817.3 |
DPT |
dermatopontin |
| chr2_-_68547061 | 0.57 |
ENST00000263655.3 |
CNRIP1 |
cannabinoid receptor interacting protein 1 |
| chr1_+_197881592 | 0.53 |
ENST00000367391.1 ENST00000367390.3 |
LHX9 |
LIM homeobox 9 |
| chr11_-_63381925 | 0.50 |
ENST00000415826.1 |
PLA2G16 |
phospholipase A2, group XVI |
| chrX_-_154250989 | 0.50 |
ENST00000360256.4 |
F8 |
coagulation factor VIII, procoagulant component |
| chr10_-_28623368 | 0.49 |
ENST00000441595.2 |
MPP7 |
membrane protein, palmitoylated 7 (MAGUK p55 subfamily member 7) |
| chr3_+_157154578 | 0.48 |
ENST00000295927.3 |
PTX3 |
pentraxin 3, long |
| chr3_+_51895621 | 0.48 |
ENST00000333127.3 |
IQCF2 |
IQ motif containing F2 |
| chr3_-_112360116 | 0.47 |
ENST00000206423.3 ENST00000439685.2 |
CCDC80 |
coiled-coil domain containing 80 |
| chr1_-_182641367 | 0.47 |
ENST00000508450.1 |
RGS8 |
regulator of G-protein signaling 8 |
| chr2_-_190044480 | 0.47 |
ENST00000374866.3 |
COL5A2 |
collagen, type V, alpha 2 |
| chr14_-_101034407 | 0.46 |
ENST00000443071.2 ENST00000557378.1 |
BEGAIN |
brain-enriched guanylate kinase-associated |
| chr13_-_38172863 | 0.46 |
ENST00000541481.1 ENST00000379743.4 ENST00000379742.4 ENST00000379749.4 ENST00000541179.1 ENST00000379747.4 |
POSTN |
periostin, osteoblast specific factor |
| chr1_-_156647189 | 0.45 |
ENST00000368223.3 |
NES |
nestin |
| chr9_-_123812542 | 0.41 |
ENST00000223642.1 |
C5 |
complement component 5 |
| chr7_+_80231466 | 0.40 |
ENST00000309881.7 ENST00000534394.1 |
CD36 |
CD36 molecule (thrombospondin receptor) |
| chr3_-_9994021 | 0.39 |
ENST00000411976.2 ENST00000412055.1 |
PRRT3 |
proline-rich transmembrane protein 3 |
| chr14_-_22005018 | 0.38 |
ENST00000546363.1 |
SALL2 |
spalt-like transcription factor 2 |
| chr14_+_59104741 | 0.37 |
ENST00000395153.3 ENST00000335867.4 |
DACT1 |
dishevelled-binding antagonist of beta-catenin 1 |
| chr5_-_178054105 | 0.35 |
ENST00000316308.4 |
CLK4 |
CDC-like kinase 4 |
| chr2_-_188378368 | 0.34 |
ENST00000392365.1 ENST00000435414.1 |
TFPI |
tissue factor pathway inhibitor (lipoprotein-associated coagulation inhibitor) |
| chr8_+_77593448 | 0.33 |
ENST00000521891.2 |
ZFHX4 |
zinc finger homeobox 4 |
| chr2_+_172378757 | 0.32 |
ENST00000409484.1 ENST00000321348.4 ENST00000375252.3 |
CYBRD1 |
cytochrome b reductase 1 |
| chr11_-_119293872 | 0.32 |
ENST00000524970.1 |
THY1 |
Thy-1 cell surface antigen |
| chr4_-_165898768 | 0.32 |
ENST00000329314.5 |
TRIM61 |
tripartite motif containing 61 |
| chr8_+_77593474 | 0.31 |
ENST00000455469.2 ENST00000050961.6 |
ZFHX4 |
zinc finger homeobox 4 |
| chr1_-_33168336 | 0.30 |
ENST00000373484.3 |
SYNC |
syncoilin, intermediate filament protein |
| chr6_+_46655612 | 0.30 |
ENST00000544460.1 |
TDRD6 |
tudor domain containing 6 |
| chr7_+_12727250 | 0.29 |
ENST00000404894.1 |
ARL4A |
ADP-ribosylation factor-like 4A |
| chr17_-_63557759 | 0.29 |
ENST00000307078.5 |
AXIN2 |
axin 2 |
| chr22_-_39639021 | 0.29 |
ENST00000455790.1 |
PDGFB |
platelet-derived growth factor beta polypeptide |
| chr7_+_12726474 | 0.28 |
ENST00000396662.1 ENST00000356797.3 ENST00000396664.2 |
ARL4A |
ADP-ribosylation factor-like 4A |
| chr2_+_152214098 | 0.28 |
ENST00000243347.3 |
TNFAIP6 |
tumor necrosis factor, alpha-induced protein 6 |
| chr16_-_3355645 | 0.28 |
ENST00000396862.1 ENST00000573608.1 |
TIGD7 |
tigger transposable element derived 7 |
| chr6_+_52442083 | 0.28 |
ENST00000606714.1 |
TRAM2-AS1 |
TRAM2 antisense RNA 1 (head to head) |
| chr2_+_223536428 | 0.28 |
ENST00000446656.3 |
MOGAT1 |
monoacylglycerol O-acyltransferase 1 |
| chr6_+_126112074 | 0.28 |
ENST00000453302.1 ENST00000417494.1 ENST00000229634.9 |
NCOA7 |
nuclear receptor coactivator 7 |
| chr12_+_65672423 | 0.27 |
ENST00000355192.3 ENST00000308259.5 ENST00000540804.1 ENST00000535664.1 ENST00000541189.1 |
MSRB3 |
methionine sulfoxide reductase B3 |
| chr14_-_88459503 | 0.27 |
ENST00000393568.4 ENST00000261304.2 |
GALC |
galactosylceramidase |
| chr9_+_107526443 | 0.27 |
ENST00000374762.3 |
NIPSNAP3B |
nipsnap homolog 3B (C. elegans) |
| chr19_+_39903185 | 0.27 |
ENST00000409794.3 |
PLEKHG2 |
pleckstrin homology domain containing, family G (with RhoGef domain) member 2 |
| chr16_+_53242350 | 0.26 |
ENST00000565442.1 |
CHD9 |
chromodomain helicase DNA binding protein 9 |
| chr12_+_56546223 | 0.26 |
ENST00000550443.1 ENST00000207437.5 |
MYL6B |
myosin, light chain 6B, alkali, smooth muscle and non-muscle |
| chr12_+_56546363 | 0.26 |
ENST00000551834.1 ENST00000552568.1 |
MYL6B |
myosin, light chain 6B, alkali, smooth muscle and non-muscle |
| chr3_+_46616017 | 0.25 |
ENST00000542931.1 |
TDGF1 |
teratocarcinoma-derived growth factor 1 |
| chr11_+_19798964 | 0.25 |
ENST00000527559.2 |
NAV2 |
neuron navigator 2 |
| chr17_+_26800296 | 0.25 |
ENST00000444914.3 ENST00000314669.5 |
SLC13A2 |
solute carrier family 13 (sodium-dependent dicarboxylate transporter), member 2 |
| chr9_+_131683174 | 0.24 |
ENST00000372592.3 ENST00000428610.1 |
PHYHD1 |
phytanoyl-CoA dioxygenase domain containing 1 |
| chr16_+_3355472 | 0.24 |
ENST00000574298.1 |
ZNF75A |
zinc finger protein 75a |
| chr4_-_186732048 | 0.24 |
ENST00000448662.2 ENST00000439049.1 ENST00000420158.1 ENST00000431808.1 ENST00000319471.9 |
SORBS2 |
sorbin and SH3 domain containing 2 |
| chr16_+_31483451 | 0.24 |
ENST00000565360.1 ENST00000361773.3 |
TGFB1I1 |
transforming growth factor beta 1 induced transcript 1 |
| chr5_-_178054014 | 0.24 |
ENST00000520957.1 |
CLK4 |
CDC-like kinase 4 |
| chr1_+_201708992 | 0.23 |
ENST00000367295.1 |
NAV1 |
neuron navigator 1 |
| chr5_+_139505520 | 0.23 |
ENST00000333305.3 |
IGIP |
IgA-inducing protein |
| chr17_+_26800756 | 0.23 |
ENST00000537681.1 |
SLC13A2 |
solute carrier family 13 (sodium-dependent dicarboxylate transporter), member 2 |
| chr16_-_66583701 | 0.23 |
ENST00000527800.1 ENST00000525974.1 ENST00000563369.2 |
TK2 |
thymidine kinase 2, mitochondrial |
| chr11_-_86383157 | 0.23 |
ENST00000393324.3 |
ME3 |
malic enzyme 3, NADP(+)-dependent, mitochondrial |
| chr1_+_22351977 | 0.22 |
ENST00000420503.1 ENST00000416769.1 ENST00000404210.2 |
LINC00339 |
long intergenic non-protein coding RNA 339 |
| chr4_-_13546632 | 0.22 |
ENST00000382438.5 |
NKX3-2 |
NK3 homeobox 2 |
| chr17_+_26800648 | 0.22 |
ENST00000545060.1 |
SLC13A2 |
solute carrier family 13 (sodium-dependent dicarboxylate transporter), member 2 |
| chr4_+_71108300 | 0.22 |
ENST00000304954.3 |
CSN3 |
casein kappa |
| chr10_+_11060004 | 0.22 |
ENST00000542579.1 ENST00000399850.3 ENST00000417956.2 |
CELF2 |
CUGBP, Elav-like family member 2 |
| chrX_+_22291065 | 0.22 |
ENST00000323684.1 |
ZNF645 |
zinc finger protein 645 |
| chr7_-_150675372 | 0.21 |
ENST00000262186.5 |
KCNH2 |
potassium voltage-gated channel, subfamily H (eag-related), member 2 |
| chr16_+_31483374 | 0.21 |
ENST00000394863.3 |
TGFB1I1 |
transforming growth factor beta 1 induced transcript 1 |
| chr1_+_197871740 | 0.21 |
ENST00000367393.3 |
C1orf53 |
chromosome 1 open reading frame 53 |
| chr3_+_53528659 | 0.21 |
ENST00000350061.5 |
CACNA1D |
calcium channel, voltage-dependent, L type, alpha 1D subunit |
| chr8_-_110656995 | 0.21 |
ENST00000276646.9 ENST00000533065.1 |
SYBU |
syntabulin (syntaxin-interacting) |
| chr1_-_182641037 | 0.21 |
ENST00000483095.2 |
RGS8 |
regulator of G-protein signaling 8 |
| chr6_-_52441713 | 0.20 |
ENST00000182527.3 |
TRAM2 |
translocation associated membrane protein 2 |
| chr2_+_162016827 | 0.20 |
ENST00000429217.1 ENST00000406287.1 ENST00000402568.1 |
TANK |
TRAF family member-associated NFKB activator |
| chr11_-_86383370 | 0.20 |
ENST00000526834.1 ENST00000359636.2 |
ME3 |
malic enzyme 3, NADP(+)-dependent, mitochondrial |
| chr3_+_153202284 | 0.20 |
ENST00000446603.2 |
C3orf79 |
chromosome 3 open reading frame 79 |
| chr12_+_6982725 | 0.20 |
ENST00000433346.1 |
LRRC23 |
leucine rich repeat containing 23 |
| chr1_-_182640988 | 0.20 |
ENST00000367556.1 |
RGS8 |
regulator of G-protein signaling 8 |
| chr1_+_101185290 | 0.20 |
ENST00000370119.4 ENST00000347652.2 ENST00000294728.2 ENST00000370115.1 |
VCAM1 |
vascular cell adhesion molecule 1 |
| chr12_-_58329819 | 0.20 |
ENST00000551421.1 |
RP11-620J15.3 |
RP11-620J15.3 |
| chr16_-_87970122 | 0.20 |
ENST00000309893.2 |
CA5A |
carbonic anhydrase VA, mitochondrial |
| chr16_-_57570450 | 0.20 |
ENST00000258214.2 |
CCDC102A |
coiled-coil domain containing 102A |
| chr14_-_22005343 | 0.19 |
ENST00000327430.3 |
SALL2 |
spalt-like transcription factor 2 |
| chr2_-_73460334 | 0.19 |
ENST00000258083.2 |
PRADC1 |
protease-associated domain containing 1 |
| chr17_-_79139817 | 0.19 |
ENST00000326724.4 |
AATK |
apoptosis-associated tyrosine kinase |
| chr6_+_126102292 | 0.19 |
ENST00000368357.3 |
NCOA7 |
nuclear receptor coactivator 7 |
| chrX_-_100307076 | 0.19 |
ENST00000338687.7 ENST00000545398.1 ENST00000372931.5 |
TRMT2B |
tRNA methyltransferase 2 homolog B (S. cerevisiae) |
| chr14_-_22005062 | 0.19 |
ENST00000317492.5 |
SALL2 |
spalt-like transcription factor 2 |
| chr17_+_59489112 | 0.18 |
ENST00000335108.2 |
C17orf82 |
chromosome 17 open reading frame 82 |
| chrX_-_100307043 | 0.18 |
ENST00000372939.1 ENST00000372935.1 ENST00000372936.3 |
TRMT2B |
tRNA methyltransferase 2 homolog B (S. cerevisiae) |
| chr16_-_1843720 | 0.18 |
ENST00000415638.3 ENST00000215539.3 |
IGFALS |
insulin-like growth factor binding protein, acid labile subunit |
| chr19_+_17326191 | 0.18 |
ENST00000595101.1 ENST00000596136.1 ENST00000379776.4 |
USE1 |
unconventional SNARE in the ER 1 homolog (S. cerevisiae) |
| chr12_-_6982442 | 0.18 |
ENST00000523102.1 ENST00000524270.1 ENST00000519357.1 |
SPSB2 |
splA/ryanodine receptor domain and SOCS box containing 2 |
| chr7_+_1127723 | 0.17 |
ENST00000397088.3 |
GPER1 |
G protein-coupled estrogen receptor 1 |
| chr15_+_63569785 | 0.17 |
ENST00000380343.4 ENST00000560353.1 |
APH1B |
APH1B gamma secretase subunit |
| chr2_+_33701286 | 0.17 |
ENST00000403687.3 |
RASGRP3 |
RAS guanyl releasing protein 3 (calcium and DAG-regulated) |
| chr10_-_28571015 | 0.17 |
ENST00000375719.3 ENST00000375732.1 |
MPP7 |
membrane protein, palmitoylated 7 (MAGUK p55 subfamily member 7) |
| chr2_+_204193129 | 0.17 |
ENST00000417864.1 |
ABI2 |
abl-interactor 2 |
| chr3_+_183353356 | 0.17 |
ENST00000242810.6 ENST00000493074.1 ENST00000437402.1 ENST00000454495.2 ENST00000473045.1 ENST00000468101.1 ENST00000427201.2 ENST00000482138.1 ENST00000454652.2 |
KLHL24 |
kelch-like family member 24 |
| chr11_+_64058820 | 0.17 |
ENST00000422670.2 |
KCNK4 |
potassium channel, subfamily K, member 4 |
| chr2_+_162016804 | 0.17 |
ENST00000392749.2 ENST00000440506.1 |
TANK |
TRAF family member-associated NFKB activator |
| chr11_+_64058758 | 0.17 |
ENST00000538767.1 |
KCNK4 |
potassium channel, subfamily K, member 4 |
| chr15_+_25200074 | 0.16 |
ENST00000390687.4 ENST00000584968.1 ENST00000346403.6 ENST00000554227.2 |
SNRPN |
small nuclear ribonucleoprotein polypeptide N |
| chr2_+_204193101 | 0.16 |
ENST00000430418.1 ENST00000424558.1 ENST00000261016.6 |
ABI2 |
abl-interactor 2 |
| chr1_+_145293371 | 0.16 |
ENST00000342960.5 |
NBPF10 |
neuroblastoma breakpoint family, member 10 |
| chr2_+_162016916 | 0.15 |
ENST00000405852.1 |
TANK |
TRAF family member-associated NFKB activator |
| chr2_+_204193149 | 0.15 |
ENST00000422511.2 |
ABI2 |
abl-interactor 2 |
| chr3_-_48672859 | 0.15 |
ENST00000395550.2 ENST00000455886.2 ENST00000431739.1 ENST00000426599.1 ENST00000383733.3 ENST00000420764.2 ENST00000337000.8 |
SLC26A6 |
solute carrier family 26 (anion exchanger), member 6 |
| chr11_+_64059464 | 0.15 |
ENST00000394525.2 |
KCNK4 |
potassium channel, subfamily K, member 4 |
| chr2_+_161993465 | 0.15 |
ENST00000457476.1 |
TANK |
TRAF family member-associated NFKB activator |
| chr12_-_6809958 | 0.14 |
ENST00000320591.5 ENST00000534837.1 |
PIANP |
PILR alpha associated neural protein |
| chr3_+_138340067 | 0.14 |
ENST00000479848.1 |
FAIM |
Fas apoptotic inhibitory molecule |
| chr10_+_72164135 | 0.14 |
ENST00000373218.4 |
EIF4EBP2 |
eukaryotic translation initiation factor 4E binding protein 2 |
| chr2_+_38893047 | 0.14 |
ENST00000272252.5 |
GALM |
galactose mutarotase (aldose 1-epimerase) |
| chr3_+_56591184 | 0.14 |
ENST00000422222.1 ENST00000394672.3 ENST00000326595.7 |
CCDC66 |
coiled-coil domain containing 66 |
| chr17_+_7328925 | 0.14 |
ENST00000333870.3 ENST00000574034.1 |
C17orf74 |
chromosome 17 open reading frame 74 |
| chr5_+_81575281 | 0.14 |
ENST00000380167.4 |
ATP6AP1L |
ATPase, H+ transporting, lysosomal accessory protein 1-like |
| chr4_+_129730947 | 0.14 |
ENST00000452328.2 ENST00000504089.1 |
PHF17 |
jade family PHD finger 1 |
| chr14_+_79745682 | 0.14 |
ENST00000557594.1 |
NRXN3 |
neurexin 3 |
| chr9_+_117904097 | 0.14 |
ENST00000374016.1 |
DEC1 |
deleted in esophageal cancer 1 |
| chr8_-_140715294 | 0.13 |
ENST00000303015.1 ENST00000520439.1 |
KCNK9 |
potassium channel, subfamily K, member 9 |
| chr4_+_129730779 | 0.13 |
ENST00000226319.6 |
PHF17 |
jade family PHD finger 1 |
| chr2_+_204192942 | 0.13 |
ENST00000295851.5 ENST00000261017.5 |
ABI2 |
abl-interactor 2 |
| chr12_-_6809543 | 0.13 |
ENST00000540656.1 |
PIANP |
PILR alpha associated neural protein |
| chr1_+_78470530 | 0.13 |
ENST00000370763.5 |
DNAJB4 |
DnaJ (Hsp40) homolog, subfamily B, member 4 |
| chr19_-_47137942 | 0.13 |
ENST00000300873.4 |
GNG8 |
guanine nucleotide binding protein (G protein), gamma 8 |
| chr17_-_40264692 | 0.13 |
ENST00000591220.1 ENST00000251642.3 |
DHX58 |
DEXH (Asp-Glu-X-His) box polypeptide 58 |
| chr3_+_120626919 | 0.13 |
ENST00000273666.6 ENST00000471454.1 ENST00000472879.1 ENST00000497029.1 ENST00000492541.1 |
STXBP5L |
syntaxin binding protein 5-like |
| chr14_+_79745746 | 0.13 |
ENST00000281127.7 |
NRXN3 |
neurexin 3 |
| chr1_+_181003067 | 0.13 |
ENST00000434571.2 ENST00000367579.3 ENST00000282990.6 ENST00000367580.5 |
MR1 |
major histocompatibility complex, class I-related |
| chr11_+_57227981 | 0.13 |
ENST00000335099.3 |
RTN4RL2 |
reticulon 4 receptor-like 2 |
| chr16_-_66584059 | 0.13 |
ENST00000417693.3 ENST00000544898.1 ENST00000569718.1 ENST00000527284.1 ENST00000299697.7 ENST00000451102.2 |
TK2 |
thymidine kinase 2, mitochondrial |
| chr15_+_75639773 | 0.13 |
ENST00000567657.1 |
NEIL1 |
nei endonuclease VIII-like 1 (E. coli) |
| chr3_+_193853927 | 0.12 |
ENST00000232424.3 |
HES1 |
hes family bHLH transcription factor 1 |
| chr11_+_57365150 | 0.12 |
ENST00000457869.1 ENST00000340687.6 ENST00000378323.4 ENST00000378324.2 ENST00000403558.1 |
SERPING1 |
serpin peptidase inhibitor, clade G (C1 inhibitor), member 1 |
| chr5_-_153857819 | 0.12 |
ENST00000231121.2 |
HAND1 |
heart and neural crest derivatives expressed 1 |
| chr14_-_58332774 | 0.12 |
ENST00000556826.1 |
SLC35F4 |
solute carrier family 35, member F4 |
| chr2_-_86564776 | 0.12 |
ENST00000165698.5 ENST00000541910.1 ENST00000535845.1 |
REEP1 |
receptor accessory protein 1 |
| chr3_-_178976996 | 0.12 |
ENST00000485523.1 |
KCNMB3 |
potassium large conductance calcium-activated channel, subfamily M beta member 3 |
| chr2_+_161993412 | 0.11 |
ENST00000259075.2 ENST00000432002.1 |
TANK |
TRAF family member-associated NFKB activator |
| chr21_+_40817749 | 0.11 |
ENST00000380637.3 ENST00000380634.1 ENST00000458295.1 ENST00000440288.2 ENST00000380631.1 |
SH3BGR |
SH3 domain binding glutamic acid-rich protein |
| chr2_-_15701422 | 0.11 |
ENST00000441750.1 ENST00000281513.5 |
NBAS |
neuroblastoma amplified sequence |
| chr3_+_138340049 | 0.11 |
ENST00000464668.1 |
FAIM |
Fas apoptotic inhibitory molecule |
| chr19_+_17326141 | 0.11 |
ENST00000445667.2 ENST00000263897.5 |
USE1 |
unconventional SNARE in the ER 1 homolog (S. cerevisiae) |
| chr2_-_99224915 | 0.11 |
ENST00000328709.3 ENST00000409997.1 |
COA5 |
cytochrome c oxidase assembly factor 5 |
| chr6_-_28226984 | 0.11 |
ENST00000423974.2 |
ZKSCAN4 |
zinc finger with KRAB and SCAN domains 4 |
| chr6_-_16761678 | 0.11 |
ENST00000244769.4 ENST00000436367.1 |
ATXN1 |
ataxin 1 |
| chr12_+_108523133 | 0.11 |
ENST00000547525.1 |
WSCD2 |
WSC domain containing 2 |
| chr6_-_28220002 | 0.11 |
ENST00000377294.2 |
ZKSCAN4 |
zinc finger with KRAB and SCAN domains 4 |
| chr2_-_208989225 | 0.11 |
ENST00000264376.4 |
CRYGD |
crystallin, gamma D |
| chr1_-_221915418 | 0.11 |
ENST00000323825.3 ENST00000366899.3 |
DUSP10 |
dual specificity phosphatase 10 |
| chr20_-_43753104 | 0.10 |
ENST00000372785.3 |
WFDC12 |
WAP four-disulfide core domain 12 |
| chr9_-_35815013 | 0.10 |
ENST00000259667.5 |
HINT2 |
histidine triad nucleotide binding protein 2 |
| chr12_+_20848486 | 0.10 |
ENST00000545102.1 |
SLCO1C1 |
solute carrier organic anion transporter family, member 1C1 |
| chr4_+_129730839 | 0.10 |
ENST00000511647.1 |
PHF17 |
jade family PHD finger 1 |
| chr14_-_65769392 | 0.10 |
ENST00000555736.1 |
CTD-2509G16.5 |
CTD-2509G16.5 |
| chr7_+_101928380 | 0.10 |
ENST00000536178.1 |
SH2B2 |
SH2B adaptor protein 2 |
| chrX_+_52926322 | 0.10 |
ENST00000430150.2 ENST00000452021.2 ENST00000412319.1 |
FAM156B |
family with sequence similarity 156, member B |
| chr1_-_150780757 | 0.10 |
ENST00000271651.3 |
CTSK |
cathepsin K |
| chr14_+_79746249 | 0.10 |
ENST00000428277.2 |
NRXN3 |
neurexin 3 |
| chr10_-_61513146 | 0.10 |
ENST00000430431.1 |
LINC00948 |
long intergenic non-protein coding RNA 948 |
| chrX_-_134429952 | 0.10 |
ENST00000370764.1 |
ZNF75D |
zinc finger protein 75D |
| chr15_+_75639951 | 0.10 |
ENST00000564784.1 ENST00000569035.1 |
NEIL1 |
nei endonuclease VIII-like 1 (E. coli) |
| chr8_-_57359131 | 0.10 |
ENST00000518974.1 ENST00000523051.1 ENST00000518770.1 ENST00000451791.2 |
PENK |
proenkephalin |
| chr3_+_119499331 | 0.10 |
ENST00000393716.2 ENST00000466380.1 |
NR1I2 |
nuclear receptor subfamily 1, group I, member 2 |
| chr8_-_7309887 | 0.10 |
ENST00000458665.1 ENST00000528168.1 |
SPAG11B |
sperm associated antigen 11B |
| chr15_-_63449663 | 0.10 |
ENST00000439025.1 |
RPS27L |
ribosomal protein S27-like |
| chr13_-_81801115 | 0.10 |
ENST00000567258.1 |
LINC00564 |
long intergenic non-protein coding RNA 564 |
| chr9_+_131174024 | 0.10 |
ENST00000420034.1 ENST00000372842.1 |
CERCAM |
cerebral endothelial cell adhesion molecule |
| chr6_-_33385870 | 0.10 |
ENST00000488034.1 |
CUTA |
cutA divalent cation tolerance homolog (E. coli) |
| chr4_+_81118647 | 0.10 |
ENST00000415738.2 |
PRDM8 |
PR domain containing 8 |
| chr12_-_49393092 | 0.09 |
ENST00000421952.2 |
DDN |
dendrin |
| chr1_+_202431859 | 0.09 |
ENST00000391959.3 ENST00000367270.4 |
PPP1R12B |
protein phosphatase 1, regulatory subunit 12B |
| chr3_+_113616317 | 0.09 |
ENST00000440446.2 ENST00000488680.1 |
GRAMD1C |
GRAM domain containing 1C |
| chr17_-_26694979 | 0.09 |
ENST00000438614.1 |
VTN |
vitronectin |
| chr14_+_24590560 | 0.09 |
ENST00000558325.1 |
RP11-468E2.6 |
RP11-468E2.6 |
| chr7_-_127255982 | 0.09 |
ENST00000338516.3 ENST00000378740.2 |
PAX4 |
paired box 4 |
| chr1_-_193155729 | 0.09 |
ENST00000367434.4 |
B3GALT2 |
UDP-Gal:betaGlcNAc beta 1,3-galactosyltransferase, polypeptide 2 |
| chr4_+_183370146 | 0.09 |
ENST00000510504.1 |
TENM3 |
teneurin transmembrane protein 3 |
| chr17_+_12569306 | 0.09 |
ENST00000425538.1 |
MYOCD |
myocardin |
| chr17_-_26695013 | 0.09 |
ENST00000555059.2 |
CTB-96E2.2 |
Homeobox protein SEBOX |
| chr15_-_34880646 | 0.09 |
ENST00000543376.1 |
GOLGA8A |
golgin A8 family, member A |
| chr5_+_147258266 | 0.09 |
ENST00000296694.4 |
SCGB3A2 |
secretoglobin, family 3A, member 2 |
| chr12_-_76477707 | 0.09 |
ENST00000551992.1 |
NAP1L1 |
nucleosome assembly protein 1-like 1 |
| chr8_-_23261589 | 0.09 |
ENST00000524168.1 ENST00000523833.2 ENST00000519243.1 ENST00000389131.3 |
LOXL2 |
lysyl oxidase-like 2 |
| chr11_+_72975578 | 0.09 |
ENST00000393592.2 |
P2RY6 |
pyrimidinergic receptor P2Y, G-protein coupled, 6 |
| chr1_+_145293114 | 0.08 |
ENST00000369338.1 |
NBPF10 |
neuroblastoma breakpoint family, member 10 |
| chr10_-_61513201 | 0.08 |
ENST00000414264.1 ENST00000594536.1 |
LINC00948 |
long intergenic non-protein coding RNA 948 |
| chr17_-_6554877 | 0.08 |
ENST00000225728.3 ENST00000575197.1 |
MED31 |
mediator complex subunit 31 |
Gene Ontology Analysis
Gene overrepresentation in biological_process category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.3 | 1.7 | GO:1903284 | negative regulation of dopamine uptake involved in synaptic transmission(GO:0051585) norepinephrine uptake(GO:0051620) regulation of norepinephrine uptake(GO:0051621) negative regulation of norepinephrine uptake(GO:0051622) negative regulation of catecholamine uptake involved in synaptic transmission(GO:0051945) regulation of glutathione peroxidase activity(GO:1903282) positive regulation of glutathione peroxidase activity(GO:1903284) positive regulation of hydrogen peroxide catabolic process(GO:1903285) positive regulation of peroxidase activity(GO:2000470) |
| 0.2 | 0.8 | GO:0086097 | phospholipase C-activating angiotensin-activated signaling pathway(GO:0086097) |
| 0.2 | 0.5 | GO:1903225 | negative regulation of endodermal cell differentiation(GO:1903225) |
| 0.1 | 1.8 | GO:0048251 | elastic fiber assembly(GO:0048251) |
| 0.1 | 0.5 | GO:0035262 | gonad morphogenesis(GO:0035262) |
| 0.1 | 0.8 | GO:2000158 | positive regulation of ubiquitin-specific protease activity(GO:2000158) |
| 0.1 | 0.5 | GO:0071469 | detection of mechanical stimulus involved in sensory perception of touch(GO:0050976) cellular response to alkaline pH(GO:0071469) |
| 0.1 | 0.3 | GO:2000298 | regulation of Rho-dependent protein serine/threonine kinase activity(GO:2000298) |
| 0.1 | 0.3 | GO:0061181 | regulation of chondrocyte development(GO:0061181) |
| 0.1 | 0.5 | GO:0032571 | response to vitamin K(GO:0032571) bone regeneration(GO:1990523) |
| 0.1 | 0.4 | GO:0046104 | thymidine metabolic process(GO:0046104) pyrimidine deoxyribonucleoside metabolic process(GO:0046125) |
| 0.1 | 0.4 | GO:0010760 | negative regulation of macrophage chemotaxis(GO:0010760) |
| 0.1 | 0.5 | GO:0052199 | negative regulation of catalytic activity in other organism involved in symbiotic interaction(GO:0052199) |
| 0.1 | 1.4 | GO:0007021 | tubulin complex assembly(GO:0007021) |
| 0.1 | 0.7 | GO:0071896 | protein localization to adherens junction(GO:0071896) |
| 0.1 | 0.3 | GO:1905174 | metanephric glomerular mesangial cell development(GO:0072255) reversible differentiation(GO:0090677) cell dedifferentiation involved in phenotypic switching(GO:0090678) positive regulation of phenotypic switching(GO:1900241) regulation of vascular smooth muscle cell dedifferentiation(GO:1905174) positive regulation of vascular smooth muscle cell dedifferentiation(GO:1905176) vascular smooth muscle cell dedifferentiation(GO:1990936) |
| 0.1 | 0.2 | GO:1902303 | regulation of heart rate by hormone(GO:0003064) negative regulation of potassium ion export(GO:1902303) |
| 0.1 | 0.3 | GO:2000724 | positive regulation of cardiac vascular smooth muscle cell differentiation(GO:2000724) |
| 0.1 | 0.4 | GO:0048619 | embryonic hindgut morphogenesis(GO:0048619) |
| 0.1 | 0.4 | GO:0072564 | lipoprotein particle mediated signaling(GO:0055095) low-density lipoprotein particle mediated signaling(GO:0055096) blood microparticle formation(GO:0072564) regulation of blood microparticle formation(GO:2000332) positive regulation of blood microparticle formation(GO:2000334) |
| 0.1 | 1.1 | GO:0098828 | positive regulation of inhibitory postsynaptic potential(GO:0097151) modulation of inhibitory postsynaptic potential(GO:0098828) |
| 0.1 | 0.6 | GO:2000601 | positive regulation of Arp2/3 complex-mediated actin nucleation(GO:2000601) |
| 0.1 | 0.5 | GO:0046485 | ether lipid metabolic process(GO:0046485) |
| 0.1 | 0.2 | GO:0042938 | dipeptide transport(GO:0042938) |
| 0.0 | 0.4 | GO:0030579 | ubiquitin-dependent SMAD protein catabolic process(GO:0030579) |
| 0.0 | 0.3 | GO:0007598 | blood coagulation, extrinsic pathway(GO:0007598) |
| 0.0 | 0.1 | GO:0033499 | galactose catabolic process via UDP-galactose(GO:0033499) |
| 0.0 | 0.3 | GO:0006651 | diacylglycerol biosynthetic process(GO:0006651) |
| 0.0 | 0.1 | GO:2000974 | trochlear nerve development(GO:0021558) auditory receptor cell fate determination(GO:0042668) negative regulation of auditory receptor cell differentiation(GO:0045608) regulation of timing of neuron differentiation(GO:0060164) negative regulation of pro-B cell differentiation(GO:2000974) |
| 0.0 | 0.7 | GO:0050812 | regulation of acyl-CoA biosynthetic process(GO:0050812) |
| 0.0 | 0.9 | GO:0060159 | regulation of dopamine receptor signaling pathway(GO:0060159) |
| 0.0 | 0.1 | GO:0003218 | cardiac left ventricle formation(GO:0003218) |
| 0.0 | 0.3 | GO:0072592 | oxygen metabolic process(GO:0072592) |
| 0.0 | 0.6 | GO:1901750 | leukotriene D4 metabolic process(GO:1901748) leukotriene D4 biosynthetic process(GO:1901750) |
| 0.0 | 0.2 | GO:2000312 | regulation of kainate selective glutamate receptor activity(GO:2000312) |
| 0.0 | 0.1 | GO:0039534 | negative regulation of MDA-5 signaling pathway(GO:0039534) positive regulation of MDA-5 signaling pathway(GO:1900245) |
| 0.0 | 0.2 | GO:0021564 | vagus nerve development(GO:0021564) |
| 0.0 | 0.6 | GO:0007597 | blood coagulation, intrinsic pathway(GO:0007597) |
| 0.0 | 0.2 | GO:0086046 | membrane depolarization during SA node cell action potential(GO:0086046) |
| 0.0 | 0.2 | GO:0022614 | membrane to membrane docking(GO:0022614) |
| 0.0 | 0.7 | GO:0032291 | central nervous system myelination(GO:0022010) axon ensheathment in central nervous system(GO:0032291) |
| 0.0 | 0.3 | GO:0046477 | glycosylceramide catabolic process(GO:0046477) |
| 0.0 | 0.2 | GO:0045048 | protein insertion into ER membrane(GO:0045048) |
| 0.0 | 0.3 | GO:0030091 | protein repair(GO:0030091) |
| 0.0 | 0.1 | GO:0051866 | general adaptation syndrome(GO:0051866) |
| 0.0 | 0.5 | GO:1902083 | negative regulation of peptidyl-cysteine S-nitrosylation(GO:1902083) |
| 0.0 | 0.3 | GO:0032074 | negative regulation of nuclease activity(GO:0032074) |
| 0.0 | 0.1 | GO:2001271 | negative regulation of cysteine-type endopeptidase activity involved in execution phase of apoptosis(GO:2001271) |
| 0.0 | 0.1 | GO:0070681 | glutaminyl-tRNAGln biosynthesis via transamidation(GO:0070681) |
| 0.0 | 0.1 | GO:0060266 | negative regulation of respiratory burst involved in inflammatory response(GO:0060266) |
| 0.0 | 0.1 | GO:0044240 | multicellular organism lipid catabolic process(GO:0044240) |
| 0.0 | 0.2 | GO:0030321 | transepithelial chloride transport(GO:0030321) |
| 0.0 | 0.1 | GO:0001957 | intramembranous ossification(GO:0001957) direct ossification(GO:0036072) |
| 0.0 | 0.1 | GO:0019236 | response to pheromone(GO:0019236) |
| 0.0 | 0.1 | GO:0045897 | positive regulation of transcription during mitosis(GO:0045897) |
| 0.0 | 0.1 | GO:0031914 | negative regulation of synaptic plasticity(GO:0031914) |
| 0.0 | 0.2 | GO:0035279 | mRNA cleavage involved in gene silencing by miRNA(GO:0035279) mRNA cleavage involved in gene silencing(GO:0098795) |
| 0.0 | 0.1 | GO:2000412 | positive regulation of thymocyte migration(GO:2000412) positive regulation of CD8-positive, alpha-beta T cell proliferation(GO:2000566) |
| 0.0 | 0.4 | GO:0090129 | positive regulation of synapse maturation(GO:0090129) |
| 0.0 | 0.2 | GO:0090154 | positive regulation of sphingolipid biosynthetic process(GO:0090154) positive regulation of ceramide biosynthetic process(GO:2000304) |
| 0.0 | 0.1 | GO:2000622 | regulation of nuclear-transcribed mRNA catabolic process, nonsense-mediated decay(GO:2000622) negative regulation of nuclear-transcribed mRNA catabolic process, nonsense-mediated decay(GO:2000623) |
| 0.0 | 0.4 | GO:1901881 | positive regulation of protein depolymerization(GO:1901881) |
| 0.0 | 0.1 | GO:1904158 | axonemal central apparatus assembly(GO:1904158) |
| 0.0 | 0.7 | GO:0070306 | lens fiber cell differentiation(GO:0070306) |
| 0.0 | 0.3 | GO:0007351 | tripartite regional subdivision(GO:0007351) anterior/posterior axis specification, embryo(GO:0008595) |
| 0.0 | 0.2 | GO:0042738 | exogenous drug catabolic process(GO:0042738) |
| 0.0 | 0.1 | GO:0043951 | negative regulation of cAMP-mediated signaling(GO:0043951) |
| 0.0 | 0.6 | GO:0050873 | brown fat cell differentiation(GO:0050873) |
| 0.0 | 0.1 | GO:0061302 | smooth muscle cell-matrix adhesion(GO:0061302) |
| 0.0 | 0.1 | GO:0019375 | galactosylceramide biosynthetic process(GO:0006682) galactolipid biosynthetic process(GO:0019375) |
Gene overrepresentation in cellular_component category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.2 | 1.8 | GO:0071953 | elastic fiber(GO:0071953) |
| 0.2 | 0.7 | GO:0005967 | mitochondrial pyruvate dehydrogenase complex(GO:0005967) |
| 0.2 | 0.5 | GO:0005588 | collagen type V trimer(GO:0005588) |
| 0.1 | 1.4 | GO:0097512 | cardiac myofibril(GO:0097512) |
| 0.1 | 0.2 | GO:0071065 | alpha9-beta1 integrin-vascular cell adhesion molecule-1 complex(GO:0071065) |
| 0.1 | 2.1 | GO:0031092 | platelet alpha granule membrane(GO:0031092) |
| 0.0 | 0.7 | GO:0097025 | MPP7-DLG1-LIN7 complex(GO:0097025) |
| 0.0 | 0.4 | GO:0005579 | membrane attack complex(GO:0005579) |
| 0.0 | 0.2 | GO:1902937 | inward rectifier potassium channel complex(GO:1902937) |
| 0.0 | 0.6 | GO:0031209 | SCAR complex(GO:0031209) |
| 0.0 | 0.1 | GO:0070939 | Dsl1p complex(GO:0070939) |
| 0.0 | 0.1 | GO:0071062 | alphav-beta3 integrin-vitronectin complex(GO:0071062) |
| 0.0 | 0.5 | GO:0030877 | beta-catenin destruction complex(GO:0030877) |
| 0.0 | 0.7 | GO:0005614 | interstitial matrix(GO:0005614) |
| 0.0 | 0.5 | GO:0016461 | unconventional myosin complex(GO:0016461) |
| 0.0 | 0.1 | GO:0005594 | collagen type IX trimer(GO:0005594) |
| 0.0 | 0.1 | GO:1990716 | axonemal central apparatus(GO:1990716) |
| 0.0 | 1.2 | GO:0044298 | neuronal cell body membrane(GO:0032809) cell body membrane(GO:0044298) |
| 0.0 | 0.8 | GO:0016581 | NuRD complex(GO:0016581) CHD-type complex(GO:0090545) |
| 0.0 | 0.1 | GO:0030956 | glutamyl-tRNA(Gln) amidotransferase complex(GO:0030956) |
| 0.0 | 0.2 | GO:0097433 | dense body(GO:0097433) |
| 0.0 | 0.2 | GO:0042567 | insulin-like growth factor ternary complex(GO:0042567) |
| 0.0 | 0.1 | GO:0070847 | core mediator complex(GO:0070847) |
| 0.0 | 0.2 | GO:0044327 | dendritic spine head(GO:0044327) |
| 0.0 | 0.2 | GO:1990454 | L-type voltage-gated calcium channel complex(GO:1990454) |
| 0.0 | 0.3 | GO:0033391 | chromatoid body(GO:0033391) |
| 0.0 | 0.1 | GO:0032280 | symmetric synapse(GO:0032280) |
| 0.0 | 0.1 | GO:0042272 | nuclear RNA export factor complex(GO:0042272) |
| 0.0 | 0.2 | GO:0070765 | gamma-secretase complex(GO:0070765) |
| 0.0 | 0.7 | GO:0048786 | presynaptic active zone(GO:0048786) |
| 0.0 | 0.1 | GO:0036021 | endolysosome lumen(GO:0036021) |
| 0.0 | 0.1 | GO:0042612 | MHC class I protein complex(GO:0042612) |
| 0.0 | 0.3 | GO:0043194 | axon initial segment(GO:0043194) |
Gene overrepresentation in molecular_function category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.3 | 1.7 | GO:0060961 | phospholipase D inhibitor activity(GO:0060961) |
| 0.3 | 0.8 | GO:0001596 | angiotensin type I receptor activity(GO:0001596) |
| 0.2 | 0.7 | GO:0015361 | low-affinity sodium:dicarboxylate symporter activity(GO:0015361) |
| 0.2 | 0.7 | GO:0004740 | pyruvate dehydrogenase (acetyl-transferring) kinase activity(GO:0004740) |
| 0.2 | 0.5 | GO:0098782 | mechanically-gated potassium channel activity(GO:0098782) |
| 0.1 | 0.8 | GO:0035800 | deubiquitinase activator activity(GO:0035800) |
| 0.1 | 0.4 | GO:0004137 | deoxycytidine kinase activity(GO:0004137) thymidine kinase activity(GO:0004797) |
| 0.1 | 0.5 | GO:0052740 | 1-acyl-2-lysophosphatidylserine acylhydrolase activity(GO:0052740) |
| 0.1 | 0.4 | GO:0004471 | malate dehydrogenase (decarboxylating) (NAD+) activity(GO:0004471) malate dehydrogenase (decarboxylating) (NADP+) activity(GO:0004473) |
| 0.1 | 0.4 | GO:0070892 | lipoteichoic acid receptor activity(GO:0070892) |
| 0.1 | 0.2 | GO:0017082 | mineralocorticoid receptor activity(GO:0017082) |
| 0.1 | 0.3 | GO:0004144 | diacylglycerol O-acyltransferase activity(GO:0004144) |
| 0.1 | 0.3 | GO:0000293 | ferric-chelate reductase activity(GO:0000293) |
| 0.1 | 0.5 | GO:0001849 | complement component C1q binding(GO:0001849) |
| 0.1 | 0.4 | GO:0016300 | tRNA (uracil) methyltransferase activity(GO:0016300) |
| 0.0 | 0.4 | GO:0048495 | Roundabout binding(GO:0048495) |
| 0.0 | 1.5 | GO:0005212 | structural constituent of eye lens(GO:0005212) |
| 0.0 | 0.4 | GO:0070097 | delta-catenin binding(GO:0070097) |
| 0.0 | 0.3 | GO:0033743 | peptide-methionine (R)-S-oxide reductase activity(GO:0033743) |
| 0.0 | 0.2 | GO:0045029 | UDP-activated nucleotide receptor activity(GO:0045029) |
| 0.0 | 0.2 | GO:0015563 | uptake transmembrane transporter activity(GO:0015563) |
| 0.0 | 0.3 | GO:0034235 | GPI anchor binding(GO:0034235) |
| 0.0 | 0.4 | GO:0019215 | intermediate filament binding(GO:0019215) |
| 0.0 | 0.6 | GO:0036374 | glutathione hydrolase activity(GO:0036374) |
| 0.0 | 0.1 | GO:0047275 | glucosaminylgalactosylglucosylceramide beta-galactosyltransferase activity(GO:0047275) |
| 0.0 | 0.1 | GO:0031849 | olfactory receptor binding(GO:0031849) |
| 0.0 | 0.8 | GO:0008093 | cytoskeletal adaptor activity(GO:0008093) |
| 0.0 | 0.2 | GO:0000155 | phosphorelay sensor kinase activity(GO:0000155) protein histidine kinase activity(GO:0004673) |
| 0.0 | 0.2 | GO:0008131 | primary amine oxidase activity(GO:0008131) |
| 0.0 | 0.1 | GO:0043120 | tumor necrosis factor binding(GO:0043120) |
| 0.0 | 0.4 | GO:0019911 | structural constituent of myelin sheath(GO:0019911) |
| 0.0 | 0.1 | GO:0050567 | glutaminyl-tRNA synthase (glutamine-hydrolyzing) activity(GO:0050567) |
| 0.0 | 0.3 | GO:0016176 | superoxide-generating NADPH oxidase activator activity(GO:0016176) |
| 0.0 | 0.2 | GO:0086007 | voltage-gated calcium channel activity involved in cardiac muscle cell action potential(GO:0086007) |
| 0.0 | 0.3 | GO:0097109 | neuroligin family protein binding(GO:0097109) |
| 0.0 | 0.1 | GO:0034046 | poly(G) binding(GO:0034046) |
| 0.0 | 0.6 | GO:0001968 | fibronectin binding(GO:0001968) |
| 0.0 | 0.1 | GO:0004720 | protein-lysine 6-oxidase activity(GO:0004720) |
| 0.0 | 0.1 | GO:0001515 | opioid peptide activity(GO:0001515) |
| 0.0 | 0.3 | GO:0070411 | I-SMAD binding(GO:0070411) |
| 0.0 | 0.1 | GO:0030020 | extracellular matrix structural constituent conferring tensile strength(GO:0030020) |
| 0.0 | 0.3 | GO:0003906 | DNA-(apurinic or apyrimidinic site) lyase activity(GO:0003906) |
| 0.0 | 0.0 | GO:0030305 | heparanase activity(GO:0030305) |
| 0.0 | 0.2 | GO:0004089 | carbonate dehydratase activity(GO:0004089) |
| 0.0 | 0.1 | GO:0032393 | MHC class I receptor activity(GO:0032393) |
Gene overrepresentation in C2:CP category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.0 | 1.6 | PID ALPHA SYNUCLEIN PATHWAY | Alpha-synuclein signaling |
| 0.0 | 0.7 | ST WNT BETA CATENIN PATHWAY | Wnt/beta-catenin Pathway |
| 0.0 | 0.8 | PID ARF6 PATHWAY | Arf6 signaling events |
| 0.0 | 0.3 | PID S1P S1P1 PATHWAY | S1P1 pathway |
| 0.0 | 0.4 | ST MYOCYTE AD PATHWAY | Myocyte Adrenergic Pathway is a specific case of the generalized Adrenergic Pathway. |
| 0.0 | 3.1 | NABA ECM GLYCOPROTEINS | Genes encoding structural ECM glycoproteins |
| 0.0 | 0.6 | NABA COLLAGENS | Genes encoding collagen proteins |
Gene overrepresentation in C2:CP:REACTOME category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.1 | 1.1 | REACTOME NOREPINEPHRINE NEUROTRANSMITTER RELEASE CYCLE | Genes involved in Norepinephrine Neurotransmitter Release Cycle |
| 0.0 | 0.7 | REACTOME REGULATION OF PYRUVATE DEHYDROGENASE PDH COMPLEX | Genes involved in Regulation of pyruvate dehydrogenase (PDH) complex |
| 0.0 | 0.7 | REACTOME BILE SALT AND ORGANIC ANION SLC TRANSPORTERS | Genes involved in Bile salt and organic anion SLC transporters |
| 0.0 | 0.6 | REACTOME TANDEM PORE DOMAIN POTASSIUM CHANNELS | Genes involved in Tandem pore domain potassium channels |
| 0.0 | 0.6 | REACTOME INTRINSIC PATHWAY | Genes involved in Intrinsic Pathway |
| 0.0 | 0.5 | REACTOME ACYL CHAIN REMODELLING OF PS | Genes involved in Acyl chain remodelling of PS |
| 0.0 | 0.8 | REACTOME TRAF6 MEDIATED IRF7 ACTIVATION | Genes involved in TRAF6 mediated IRF7 activation |
| 0.0 | 0.3 | REACTOME FORMATION OF FIBRIN CLOT CLOTTING CASCADE | Genes involved in Formation of Fibrin Clot (Clotting Cascade) |
| 0.0 | 1.5 | REACTOME AMYLOIDS | Genes involved in Amyloids |
| 0.0 | 0.6 | REACTOME GLUTATHIONE CONJUGATION | Genes involved in Glutathione conjugation |
| 0.0 | 0.2 | REACTOME REVERSIBLE HYDRATION OF CARBON DIOXIDE | Genes involved in Reversible Hydration of Carbon Dioxide |
| 0.0 | 0.4 | REACTOME G1 S SPECIFIC TRANSCRIPTION | Genes involved in G1/S-Specific Transcription |
| 0.0 | 0.2 | REACTOME P2Y RECEPTORS | Genes involved in P2Y receptors |
| 0.0 | 0.2 | REACTOME REGULATED PROTEOLYSIS OF P75NTR | Genes involved in Regulated proteolysis of p75NTR |