Project
Epithelial-Mesenchymal Transition, human (Scheel, 2011)
Navigation
Downloads
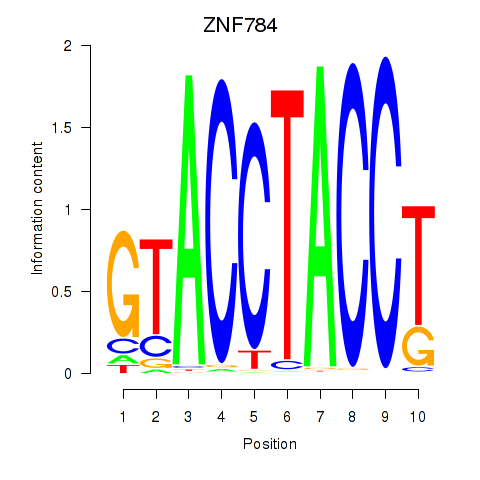
Results for ZNF784
Z-value: 0.54
Transcription factors associated with ZNF784
| Gene Symbol | Gene ID | Gene Info |
|---|---|---|
|
ZNF784
|
ENSG00000179922.5 | ZNF784 |
Activity-expression correlation:
| Gene | Promoter | Pearson corr. coef. | P-value | Plot |
|---|---|---|---|---|
| ZNF784 | hg19_v2_chr19_-_56135928_56135967 | -0.80 | 1.7e-02 | Click! |
Activity profile of ZNF784 motif
Sorted Z-values of ZNF784 motif
Network of associatons between targets according to the STRING database.
First level regulatory network of ZNF784
| Promoter | Score | Transcript | Gene | Gene Info |
|---|---|---|---|---|
| chr18_+_21529811 | 0.65 |
ENST00000588004.1 |
LAMA3 |
laminin, alpha 3 |
| chr7_+_26332645 | 0.47 |
ENST00000396376.1 |
SNX10 |
sorting nexin 10 |
| chr10_+_82168240 | 0.41 |
ENST00000372187.5 ENST00000372185.1 |
FAM213A |
family with sequence similarity 213, member A |
| chr15_+_40531621 | 0.40 |
ENST00000560346.1 |
PAK6 |
p21 protein (Cdc42/Rac)-activated kinase 6 |
| chr18_+_61254534 | 0.38 |
ENST00000269489.5 |
SERPINB13 |
serpin peptidase inhibitor, clade B (ovalbumin), member 13 |
| chr17_-_56494882 | 0.37 |
ENST00000584437.1 |
RNF43 |
ring finger protein 43 |
| chr17_-_56494713 | 0.37 |
ENST00000407977.2 |
RNF43 |
ring finger protein 43 |
| chr17_-_56494908 | 0.37 |
ENST00000577716.1 |
RNF43 |
ring finger protein 43 |
| chr15_+_40532058 | 0.36 |
ENST00000260404.4 |
PAK6 |
p21 protein (Cdc42/Rac)-activated kinase 6 |
| chr1_+_32042105 | 0.32 |
ENST00000457433.2 ENST00000441210.2 |
TINAGL1 |
tubulointerstitial nephritis antigen-like 1 |
| chr1_+_32042131 | 0.32 |
ENST00000271064.7 ENST00000537531.1 |
TINAGL1 |
tubulointerstitial nephritis antigen-like 1 |
| chr2_-_70780770 | 0.32 |
ENST00000444975.1 ENST00000445399.1 ENST00000418333.2 |
TGFA |
transforming growth factor, alpha |
| chr11_-_115375107 | 0.30 |
ENST00000545380.1 ENST00000452722.3 ENST00000537058.1 ENST00000536727.1 ENST00000542447.2 ENST00000331581.6 |
CADM1 |
cell adhesion molecule 1 |
| chr16_-_31147020 | 0.30 |
ENST00000568261.1 ENST00000567797.1 ENST00000317508.6 |
PRSS8 |
protease, serine, 8 |
| chr11_+_1861399 | 0.29 |
ENST00000381905.3 |
TNNI2 |
troponin I type 2 (skeletal, fast) |
| chr1_-_186649543 | 0.29 |
ENST00000367468.5 |
PTGS2 |
prostaglandin-endoperoxide synthase 2 (prostaglandin G/H synthase and cyclooxygenase) |
| chr18_+_61254570 | 0.28 |
ENST00000344731.5 |
SERPINB13 |
serpin peptidase inhibitor, clade B (ovalbumin), member 13 |
| chr4_-_6202247 | 0.27 |
ENST00000409021.3 ENST00000409371.3 |
JAKMIP1 |
janus kinase and microtubule interacting protein 1 |
| chr12_-_52911718 | 0.27 |
ENST00000548409.1 |
KRT5 |
keratin 5 |
| chr17_+_73717551 | 0.27 |
ENST00000450894.3 |
ITGB4 |
integrin, beta 4 |
| chr17_+_73717407 | 0.26 |
ENST00000579662.1 |
ITGB4 |
integrin, beta 4 |
| chr2_-_70780572 | 0.26 |
ENST00000450929.1 |
TGFA |
transforming growth factor, alpha |
| chr12_+_115800817 | 0.25 |
ENST00000547948.1 |
RP11-116D17.1 |
HCG2038717; Uncharacterized protein |
| chr6_-_47010061 | 0.24 |
ENST00000371253.2 |
GPR110 |
G protein-coupled receptor 110 |
| chr15_-_59665062 | 0.22 |
ENST00000288235.4 |
MYO1E |
myosin IE |
| chr17_+_73717516 | 0.22 |
ENST00000200181.3 ENST00000339591.3 |
ITGB4 |
integrin, beta 4 |
| chr1_-_175161890 | 0.21 |
ENST00000545251.2 ENST00000423313.1 |
KIAA0040 |
KIAA0040 |
| chr1_-_184943610 | 0.18 |
ENST00000367511.3 |
FAM129A |
family with sequence similarity 129, member A |
| chr16_+_82068830 | 0.18 |
ENST00000199936.4 |
HSD17B2 |
hydroxysteroid (17-beta) dehydrogenase 2 |
| chr17_-_46724186 | 0.18 |
ENST00000433510.2 |
RP11-357H14.17 |
RP11-357H14.17 |
| chr11_-_11747257 | 0.18 |
ENST00000601641.1 |
AC131935.1 |
AC131935.1 |
| chr8_+_80523321 | 0.17 |
ENST00000518111.1 |
STMN2 |
stathmin-like 2 |
| chr1_+_24646002 | 0.17 |
ENST00000356046.2 |
GRHL3 |
grainyhead-like 3 (Drosophila) |
| chr4_-_90758227 | 0.17 |
ENST00000506691.1 ENST00000394986.1 ENST00000506244.1 ENST00000394989.2 ENST00000394991.3 |
SNCA |
synuclein, alpha (non A4 component of amyloid precursor) |
| chr1_-_228135599 | 0.17 |
ENST00000272164.5 |
WNT9A |
wingless-type MMTV integration site family, member 9A |
| chr4_-_90756769 | 0.17 |
ENST00000345009.4 ENST00000505199.1 ENST00000502987.1 |
SNCA |
synuclein, alpha (non A4 component of amyloid precursor) |
| chr1_+_24645807 | 0.17 |
ENST00000361548.4 |
GRHL3 |
grainyhead-like 3 (Drosophila) |
| chr1_+_24645865 | 0.17 |
ENST00000342072.4 |
GRHL3 |
grainyhead-like 3 (Drosophila) |
| chr12_+_29376673 | 0.17 |
ENST00000547116.1 |
FAR2 |
fatty acyl CoA reductase 2 |
| chr11_+_76494253 | 0.17 |
ENST00000333090.4 |
TSKU |
tsukushi, small leucine rich proteoglycan |
| chr7_-_22234381 | 0.16 |
ENST00000458533.1 |
RAPGEF5 |
Rap guanine nucleotide exchange factor (GEF) 5 |
| chr11_+_45944190 | 0.16 |
ENST00000401752.1 ENST00000389968.3 ENST00000325468.5 ENST00000536139.1 |
GYLTL1B |
glycosyltransferase-like 1B |
| chr12_+_29376592 | 0.16 |
ENST00000182377.4 |
FAR2 |
fatty acyl CoA reductase 2 |
| chrX_-_80065146 | 0.15 |
ENST00000373275.4 |
BRWD3 |
bromodomain and WD repeat domain containing 3 |
| chr1_+_33219592 | 0.15 |
ENST00000373481.3 |
KIAA1522 |
KIAA1522 |
| chr4_-_90758118 | 0.15 |
ENST00000420646.2 |
SNCA |
synuclein, alpha (non A4 component of amyloid precursor) |
| chr19_+_6464243 | 0.15 |
ENST00000600229.1 ENST00000356762.3 |
CRB3 |
crumbs homolog 3 (Drosophila) |
| chr7_-_558876 | 0.15 |
ENST00000354513.5 ENST00000402802.3 |
PDGFA |
platelet-derived growth factor alpha polypeptide |
| chr5_+_55147205 | 0.15 |
ENST00000396836.2 ENST00000396834.1 ENST00000447346.2 ENST00000359040.5 |
IL31RA |
interleukin 31 receptor A |
| chr1_-_153029980 | 0.15 |
ENST00000392653.2 |
SPRR2A |
small proline-rich protein 2A |
| chr4_-_90757364 | 0.15 |
ENST00000508895.1 |
SNCA |
synuclein, alpha (non A4 component of amyloid precursor) |
| chr21_-_43916296 | 0.14 |
ENST00000398352.3 |
RSPH1 |
radial spoke head 1 homolog (Chlamydomonas) |
| chr14_+_22739823 | 0.14 |
ENST00000390464.2 |
TRAV38-1 |
T cell receptor alpha variable 38-1 |
| chr21_-_43916433 | 0.14 |
ENST00000291536.3 |
RSPH1 |
radial spoke head 1 homolog (Chlamydomonas) |
| chr8_-_16859690 | 0.14 |
ENST00000180166.5 |
FGF20 |
fibroblast growth factor 20 |
| chr19_+_18284477 | 0.14 |
ENST00000407280.3 |
IFI30 |
interferon, gamma-inducible protein 30 |
| chr6_+_151042224 | 0.13 |
ENST00000358517.2 |
PLEKHG1 |
pleckstrin homology domain containing, family G (with RhoGef domain) member 1 |
| chr17_+_77020224 | 0.13 |
ENST00000339142.2 |
C1QTNF1 |
C1q and tumor necrosis factor related protein 1 |
| chr10_+_28966271 | 0.13 |
ENST00000375533.3 |
BAMBI |
BMP and activin membrane-bound inhibitor |
| chr20_-_6103666 | 0.12 |
ENST00000536936.1 |
FERMT1 |
fermitin family member 1 |
| chr7_+_142636440 | 0.12 |
ENST00000458732.1 |
C7orf34 |
chromosome 7 open reading frame 34 |
| chr17_+_77020325 | 0.12 |
ENST00000311661.4 |
C1QTNF1 |
C1q and tumor necrosis factor related protein 1 |
| chr4_-_6202291 | 0.12 |
ENST00000282924.5 |
JAKMIP1 |
janus kinase and microtubule interacting protein 1 |
| chr14_-_106330458 | 0.12 |
ENST00000461719.1 |
IGHJ4 |
immunoglobulin heavy joining 4 |
| chr2_-_73869508 | 0.12 |
ENST00000272425.3 |
NAT8 |
N-acetyltransferase 8 (GCN5-related, putative) |
| chrX_-_32173579 | 0.12 |
ENST00000359836.1 ENST00000343523.2 ENST00000378707.3 ENST00000541735.1 ENST00000474231.1 |
DMD |
dystrophin |
| chr6_-_47009996 | 0.11 |
ENST00000371243.2 |
GPR110 |
G protein-coupled receptor 110 |
| chr2_-_72375167 | 0.11 |
ENST00000001146.2 |
CYP26B1 |
cytochrome P450, family 26, subfamily B, polypeptide 1 |
| chr9_+_88556036 | 0.11 |
ENST00000361671.5 ENST00000416045.1 |
NAA35 |
N(alpha)-acetyltransferase 35, NatC auxiliary subunit |
| chr1_+_90287480 | 0.11 |
ENST00000394593.3 |
LRRC8D |
leucine rich repeat containing 8 family, member D |
| chr1_-_25256368 | 0.10 |
ENST00000308873.6 |
RUNX3 |
runt-related transcription factor 3 |
| chr5_+_137225158 | 0.10 |
ENST00000290431.5 |
PKD2L2 |
polycystic kidney disease 2-like 2 |
| chr1_-_203320617 | 0.10 |
ENST00000354955.4 |
FMOD |
fibromodulin |
| chr2_-_74692473 | 0.10 |
ENST00000535045.1 ENST00000409065.1 ENST00000414701.1 ENST00000448666.1 ENST00000233616.4 ENST00000452063.2 |
MOGS |
mannosyl-oligosaccharide glucosidase |
| chr5_-_1524015 | 0.10 |
ENST00000283415.3 |
LPCAT1 |
lysophosphatidylcholine acyltransferase 1 |
| chr19_+_6464502 | 0.10 |
ENST00000308243.7 |
CRB3 |
crumbs homolog 3 (Drosophila) |
| chr9_-_112260531 | 0.09 |
ENST00000374541.2 ENST00000262539.3 |
PTPN3 |
protein tyrosine phosphatase, non-receptor type 3 |
| chr4_-_130692631 | 0.09 |
ENST00000500092.2 ENST00000509105.1 |
RP11-519M16.1 |
RP11-519M16.1 |
| chr2_+_85661918 | 0.09 |
ENST00000340326.2 |
SH2D6 |
SH2 domain containing 6 |
| chr13_-_96705624 | 0.09 |
ENST00000376747.3 ENST00000376712.4 ENST00000397618.3 ENST00000376714.3 |
UGGT2 |
UDP-glucose glycoprotein glucosyltransferase 2 |
| chr5_-_178772424 | 0.09 |
ENST00000251582.7 ENST00000274609.5 |
ADAMTS2 |
ADAM metallopeptidase with thrombospondin type 1 motif, 2 |
| chr3_+_124303539 | 0.09 |
ENST00000428018.2 |
KALRN |
kalirin, RhoGEF kinase |
| chr7_-_140098257 | 0.09 |
ENST00000340308.3 ENST00000447932.2 ENST00000429996.2 ENST00000469193.1 ENST00000326232.9 |
SLC37A3 |
solute carrier family 37, member 3 |
| chr16_+_19179549 | 0.08 |
ENST00000355377.2 ENST00000568115.1 |
SYT17 |
synaptotagmin XVII |
| chr3_+_124303472 | 0.08 |
ENST00000291478.5 |
KALRN |
kalirin, RhoGEF kinase |
| chr11_+_3829691 | 0.08 |
ENST00000278243.4 ENST00000463452.2 ENST00000479072.1 ENST00000496834.2 ENST00000469307.2 |
PGAP2 |
post-GPI attachment to proteins 2 |
| chr17_-_48546232 | 0.08 |
ENST00000258969.4 |
CHAD |
chondroadherin |
| chr7_+_26677490 | 0.08 |
ENST00000409974.3 |
C7orf71 |
chromosome 7 open reading frame 71 |
| chr11_+_64008525 | 0.08 |
ENST00000449942.2 |
FKBP2 |
FK506 binding protein 2, 13kDa |
| chr9_+_6328338 | 0.07 |
ENST00000344545.5 |
TPD52L3 |
tumor protein D52-like 3 |
| chr12_+_41221794 | 0.07 |
ENST00000547849.1 |
CNTN1 |
contactin 1 |
| chr6_+_30294612 | 0.07 |
ENST00000440271.1 ENST00000396551.3 ENST00000376656.4 ENST00000540416.1 ENST00000428728.1 ENST00000396548.1 ENST00000428404.1 |
TRIM39 |
tripartite motif containing 39 |
| chr11_+_64008443 | 0.07 |
ENST00000309366.4 |
FKBP2 |
FK506 binding protein 2, 13kDa |
| chr2_-_56150184 | 0.07 |
ENST00000394554.1 |
EFEMP1 |
EGF containing fibulin-like extracellular matrix protein 1 |
| chr11_-_8290263 | 0.07 |
ENST00000428101.2 |
LMO1 |
LIM domain only 1 (rhombotin 1) |
| chr2_-_134326009 | 0.07 |
ENST00000409261.1 ENST00000409213.1 |
NCKAP5 |
NCK-associated protein 5 |
| chr5_+_137225125 | 0.07 |
ENST00000350250.4 ENST00000508638.1 ENST00000502810.1 ENST00000508883.1 |
PKD2L2 |
polycystic kidney disease 2-like 2 |
| chr1_-_115238207 | 0.07 |
ENST00000520113.2 ENST00000369538.3 ENST00000353928.6 |
AMPD1 |
adenosine monophosphate deaminase 1 |
| chr19_+_3880581 | 0.07 |
ENST00000450849.2 ENST00000301260.6 ENST00000398448.3 |
ATCAY |
ataxia, cerebellar, Cayman type |
| chr7_+_90032667 | 0.06 |
ENST00000496677.1 ENST00000287916.4 ENST00000535571.1 ENST00000394604.1 ENST00000394605.2 |
CLDN12 |
claudin 12 |
| chr11_-_134281812 | 0.06 |
ENST00000392580.1 ENST00000312527.4 |
B3GAT1 |
beta-1,3-glucuronyltransferase 1 (glucuronosyltransferase P) |
| chr10_-_96122682 | 0.06 |
ENST00000371361.3 |
NOC3L |
nucleolar complex associated 3 homolog (S. cerevisiae) |
| chr7_+_22766766 | 0.06 |
ENST00000426291.1 ENST00000401651.1 ENST00000258743.5 ENST00000420258.2 ENST00000407492.1 ENST00000401630.3 ENST00000406575.1 |
IL6 |
interleukin 6 (interferon, beta 2) |
| chr12_+_57828521 | 0.06 |
ENST00000309668.2 |
INHBC |
inhibin, beta C |
| chr11_-_10590118 | 0.06 |
ENST00000529598.1 |
LYVE1 |
lymphatic vessel endothelial hyaluronan receptor 1 |
| chr12_+_14518598 | 0.06 |
ENST00000261168.4 ENST00000538511.1 ENST00000545723.1 ENST00000543189.1 ENST00000536444.1 |
ATF7IP |
activating transcription factor 7 interacting protein |
| chr16_+_2564254 | 0.06 |
ENST00000565223.1 |
ATP6V0C |
ATPase, H+ transporting, lysosomal 16kDa, V0 subunit c |
| chr6_+_139456226 | 0.06 |
ENST00000367658.2 |
HECA |
headcase homolog (Drosophila) |
| chr17_+_42429493 | 0.06 |
ENST00000586242.1 |
GRN |
granulin |
| chr11_-_5248294 | 0.06 |
ENST00000335295.4 |
HBB |
hemoglobin, beta |
| chr11_+_20409070 | 0.06 |
ENST00000331079.6 |
PRMT3 |
protein arginine methyltransferase 3 |
| chr19_-_18548921 | 0.06 |
ENST00000545187.1 ENST00000578352.1 |
ISYNA1 |
inositol-3-phosphate synthase 1 |
| chr3_+_139654018 | 0.06 |
ENST00000458420.3 |
CLSTN2 |
calsyntenin 2 |
| chr4_+_86396321 | 0.06 |
ENST00000503995.1 |
ARHGAP24 |
Rho GTPase activating protein 24 |
| chr1_+_205473720 | 0.06 |
ENST00000429964.2 ENST00000506784.1 ENST00000360066.2 |
CDK18 |
cyclin-dependent kinase 18 |
| chr1_-_25291475 | 0.06 |
ENST00000338888.3 ENST00000399916.1 |
RUNX3 |
runt-related transcription factor 3 |
| chr22_+_23522552 | 0.06 |
ENST00000359540.3 ENST00000398512.5 |
BCR |
breakpoint cluster region |
| chr6_-_150039249 | 0.06 |
ENST00000543571.1 |
LATS1 |
large tumor suppressor kinase 1 |
| chrX_-_54384425 | 0.05 |
ENST00000375169.3 ENST00000354646.2 |
WNK3 |
WNK lysine deficient protein kinase 3 |
| chr21_-_35831880 | 0.05 |
ENST00000399289.3 ENST00000432085.1 |
KCNE1 |
potassium voltage-gated channel, Isk-related family, member 1 |
| chr12_+_7060432 | 0.05 |
ENST00000318974.9 ENST00000456013.1 |
PTPN6 |
protein tyrosine phosphatase, non-receptor type 6 |
| chr6_-_45983549 | 0.05 |
ENST00000544153.1 |
CLIC5 |
chloride intracellular channel 5 |
| chr20_+_30598231 | 0.05 |
ENST00000300415.8 ENST00000262659.8 |
CCM2L |
cerebral cavernous malformation 2-like |
| chr6_-_109804412 | 0.05 |
ENST00000230122.3 |
ZBTB24 |
zinc finger and BTB domain containing 24 |
| chr2_-_56150910 | 0.05 |
ENST00000424836.2 ENST00000438672.1 ENST00000440439.1 ENST00000429909.1 ENST00000424207.1 ENST00000452337.1 ENST00000355426.3 ENST00000439193.1 ENST00000421664.1 |
EFEMP1 |
EGF containing fibulin-like extracellular matrix protein 1 |
| chr6_+_30295036 | 0.05 |
ENST00000376659.5 ENST00000428555.1 |
TRIM39 |
tripartite motif containing 39 |
| chr19_-_14629224 | 0.05 |
ENST00000254322.2 |
DNAJB1 |
DnaJ (Hsp40) homolog, subfamily B, member 1 |
| chr3_-_122512619 | 0.05 |
ENST00000383659.1 ENST00000306103.2 |
HSPBAP1 |
HSPB (heat shock 27kDa) associated protein 1 |
| chr15_+_84322827 | 0.05 |
ENST00000286744.5 ENST00000567476.1 |
ADAMTSL3 |
ADAMTS-like 3 |
| chr16_-_3661578 | 0.05 |
ENST00000294008.3 |
SLX4 |
SLX4 structure-specific endonuclease subunit |
| chr4_+_48833160 | 0.05 |
ENST00000506801.1 |
OCIAD1 |
OCIA domain containing 1 |
| chr1_-_207095324 | 0.05 |
ENST00000530505.1 ENST00000367091.3 ENST00000442471.2 |
FAIM3 |
Fas apoptotic inhibitory molecule 3 |
| chr12_+_57998400 | 0.05 |
ENST00000548804.1 ENST00000550596.1 ENST00000551835.1 ENST00000549583.1 |
DTX3 |
deltex homolog 3 (Drosophila) |
| chr6_-_161695042 | 0.05 |
ENST00000366908.5 ENST00000366911.5 ENST00000366905.3 |
AGPAT4 |
1-acylglycerol-3-phosphate O-acyltransferase 4 |
| chr1_-_92951607 | 0.05 |
ENST00000427103.1 |
GFI1 |
growth factor independent 1 transcription repressor |
| chr5_-_74062930 | 0.05 |
ENST00000509430.1 ENST00000345239.2 ENST00000427854.2 ENST00000506778.1 |
GFM2 |
G elongation factor, mitochondrial 2 |
| chr7_+_128577972 | 0.05 |
ENST00000357234.5 ENST00000477535.1 ENST00000479582.1 ENST00000464557.1 ENST00000402030.2 |
IRF5 |
interferon regulatory factor 5 |
| chrX_-_135056216 | 0.05 |
ENST00000305963.2 |
MMGT1 |
membrane magnesium transporter 1 |
| chr12_+_57998595 | 0.05 |
ENST00000337737.3 ENST00000548198.1 ENST00000551632.1 |
DTX3 |
deltex homolog 3 (Drosophila) |
| chr11_-_36619771 | 0.05 |
ENST00000311485.3 ENST00000527033.1 ENST00000532616.1 |
RAG2 |
recombination activating gene 2 |
| chr7_+_112063192 | 0.05 |
ENST00000005558.4 |
IFRD1 |
interferon-related developmental regulator 1 |
| chr4_+_48833183 | 0.05 |
ENST00000503016.1 |
OCIAD1 |
OCIA domain containing 1 |
| chr9_-_116861337 | 0.04 |
ENST00000374118.3 |
KIF12 |
kinesin family member 12 |
| chr17_-_46716647 | 0.04 |
ENST00000608940.1 |
RP11-357H14.17 |
RP11-357H14.17 |
| chr8_+_38831683 | 0.04 |
ENST00000302495.4 |
HTRA4 |
HtrA serine peptidase 4 |
| chr8_-_145016692 | 0.04 |
ENST00000357649.2 |
PLEC |
plectin |
| chr17_+_4736627 | 0.04 |
ENST00000355280.6 ENST00000347992.7 |
MINK1 |
misshapen-like kinase 1 |
| chr22_+_20067738 | 0.04 |
ENST00000351989.3 ENST00000383024.2 |
DGCR8 |
DGCR8 microprocessor complex subunit |
| chr17_+_55055466 | 0.04 |
ENST00000262288.3 ENST00000572710.1 ENST00000575395.1 |
SCPEP1 |
serine carboxypeptidase 1 |
| chr17_+_42264556 | 0.04 |
ENST00000319511.6 ENST00000589785.1 ENST00000592825.1 ENST00000589184.1 |
TMUB2 |
transmembrane and ubiquitin-like domain containing 2 |
| chr8_-_140715294 | 0.04 |
ENST00000303015.1 ENST00000520439.1 |
KCNK9 |
potassium channel, subfamily K, member 9 |
| chr22_+_42196666 | 0.04 |
ENST00000402061.3 ENST00000255784.5 |
CCDC134 |
coiled-coil domain containing 134 |
| chr19_-_51537982 | 0.04 |
ENST00000525263.1 |
KLK12 |
kallikrein-related peptidase 12 |
| chr4_+_48833312 | 0.04 |
ENST00000508293.1 ENST00000513391.2 |
OCIAD1 |
OCIA domain containing 1 |
| chr19_+_49622646 | 0.04 |
ENST00000334186.4 |
PPFIA3 |
protein tyrosine phosphatase, receptor type, f polypeptide (PTPRF), interacting protein (liprin), alpha 3 |
| chr4_+_48833070 | 0.04 |
ENST00000511662.1 ENST00000508996.1 ENST00000507210.1 ENST00000264312.7 ENST00000396448.2 ENST00000512236.1 ENST00000509164.1 ENST00000511102.1 ENST00000381473.3 |
OCIAD1 |
OCIA domain containing 1 |
| chr1_-_204165610 | 0.04 |
ENST00000367194.4 |
KISS1 |
KiSS-1 metastasis-suppressor |
| chr7_-_130418888 | 0.04 |
ENST00000310992.4 |
KLF14 |
Kruppel-like factor 14 |
| chr7_-_103629963 | 0.04 |
ENST00000428762.1 ENST00000343529.5 ENST00000424685.2 |
RELN |
reelin |
| chr11_+_125439298 | 0.04 |
ENST00000278903.6 ENST00000343678.4 ENST00000524723.1 ENST00000527842.2 |
EI24 |
etoposide induced 2.4 |
| chr7_+_107384579 | 0.04 |
ENST00000222597.2 ENST00000415884.2 |
CBLL1 |
Cbl proto-oncogene-like 1, E3 ubiquitin protein ligase |
| chr12_-_96390063 | 0.04 |
ENST00000541929.1 |
HAL |
histidine ammonia-lyase |
| chr4_+_48833234 | 0.04 |
ENST00000510824.1 ENST00000425583.2 |
OCIAD1 |
OCIA domain containing 1 |
| chr11_-_119066545 | 0.03 |
ENST00000415318.1 |
CCDC153 |
coiled-coil domain containing 153 |
| chr6_-_161695074 | 0.03 |
ENST00000457520.2 ENST00000366906.5 ENST00000320285.4 |
AGPAT4 |
1-acylglycerol-3-phosphate O-acyltransferase 4 |
| chr20_+_20348740 | 0.03 |
ENST00000310227.1 |
INSM1 |
insulinoma-associated 1 |
| chr4_+_48833015 | 0.03 |
ENST00000509122.1 ENST00000509664.1 ENST00000505922.2 ENST00000514981.1 |
OCIAD1 |
OCIA domain containing 1 |
| chr21_-_36421535 | 0.03 |
ENST00000416754.1 ENST00000437180.1 ENST00000455571.1 |
RUNX1 |
runt-related transcription factor 1 |
| chrX_+_70798261 | 0.03 |
ENST00000373696.3 |
ACRC |
acidic repeat containing |
| chr3_+_16926441 | 0.03 |
ENST00000418129.2 ENST00000396755.2 |
PLCL2 |
phospholipase C-like 2 |
| chr3_+_85008089 | 0.03 |
ENST00000383699.3 |
CADM2 |
cell adhesion molecule 2 |
| chr6_+_123100620 | 0.03 |
ENST00000368444.3 |
FABP7 |
fatty acid binding protein 7, brain |
| chr15_+_71839566 | 0.03 |
ENST00000357769.4 |
THSD4 |
thrombospondin, type I, domain containing 4 |
| chr4_+_48833119 | 0.03 |
ENST00000444354.2 ENST00000509963.1 ENST00000509246.1 |
OCIAD1 |
OCIA domain containing 1 |
| chrX_+_106769876 | 0.03 |
ENST00000439554.1 |
FRMPD3 |
FERM and PDZ domain containing 3 |
| chr11_+_129939811 | 0.03 |
ENST00000345598.5 ENST00000338167.5 |
APLP2 |
amyloid beta (A4) precursor-like protein 2 |
| chr1_+_55446465 | 0.03 |
ENST00000371268.3 |
TMEM61 |
transmembrane protein 61 |
| chr1_-_120190396 | 0.03 |
ENST00000421812.2 |
ZNF697 |
zinc finger protein 697 |
| chr20_-_56285595 | 0.03 |
ENST00000395816.3 ENST00000347215.4 |
PMEPA1 |
prostate transmembrane protein, androgen induced 1 |
| chr6_-_31550192 | 0.03 |
ENST00000429299.2 ENST00000446745.2 |
LTB |
lymphotoxin beta (TNF superfamily, member 3) |
| chr19_-_10213335 | 0.03 |
ENST00000592641.1 ENST00000253109.4 |
ANGPTL6 |
angiopoietin-like 6 |
| chr15_+_89346657 | 0.03 |
ENST00000439576.2 |
ACAN |
aggrecan |
| chr11_+_65154070 | 0.03 |
ENST00000317568.5 ENST00000531296.1 ENST00000533782.1 ENST00000355991.5 ENST00000416776.2 ENST00000526201.1 |
FRMD8 |
FERM domain containing 8 |
| chr2_+_14772810 | 0.03 |
ENST00000295092.2 ENST00000331243.4 |
FAM84A |
family with sequence similarity 84, member A |
| chr17_+_36452989 | 0.03 |
ENST00000312513.5 ENST00000582535.1 |
MRPL45 |
mitochondrial ribosomal protein L45 |
| chr2_-_96874553 | 0.03 |
ENST00000337288.5 ENST00000443962.1 |
STARD7 |
StAR-related lipid transfer (START) domain containing 7 |
| chr17_-_10372875 | 0.03 |
ENST00000255381.2 |
MYH4 |
myosin, heavy chain 4, skeletal muscle |
| chr12_+_50497784 | 0.03 |
ENST00000548814.1 |
GPD1 |
glycerol-3-phosphate dehydrogenase 1 (soluble) |
| chr15_+_40226304 | 0.03 |
ENST00000559624.1 ENST00000382727.2 ENST00000263791.5 ENST00000560648.1 |
EIF2AK4 |
eukaryotic translation initiation factor 2 alpha kinase 4 |
| chr19_-_51538118 | 0.03 |
ENST00000529888.1 |
KLK12 |
kallikrein-related peptidase 12 |
| chr2_+_85811525 | 0.03 |
ENST00000306384.4 |
VAMP5 |
vesicle-associated membrane protein 5 |
| chr11_+_129939779 | 0.03 |
ENST00000533195.1 ENST00000533713.1 ENST00000528499.1 ENST00000539648.1 ENST00000263574.5 |
APLP2 |
amyloid beta (A4) precursor-like protein 2 |
| chr21_-_36421626 | 0.03 |
ENST00000300305.3 |
RUNX1 |
runt-related transcription factor 1 |
| chr6_+_123100853 | 0.03 |
ENST00000356535.4 |
FABP7 |
fatty acid binding protein 7, brain |
| chr9_-_127533519 | 0.03 |
ENST00000487099.2 ENST00000344523.4 ENST00000373584.3 |
NR6A1 |
nuclear receptor subfamily 6, group A, member 1 |
| chr2_-_68384603 | 0.03 |
ENST00000406245.2 ENST00000409164.1 ENST00000295121.6 |
WDR92 |
WD repeat domain 92 |
| chr17_-_64216748 | 0.03 |
ENST00000585162.1 |
APOH |
apolipoprotein H (beta-2-glycoprotein I) |
| chr11_-_18765389 | 0.03 |
ENST00000477854.1 |
PTPN5 |
protein tyrosine phosphatase, non-receptor type 5 (striatum-enriched) |
| chr4_-_153274078 | 0.02 |
ENST00000263981.5 |
FBXW7 |
F-box and WD repeat domain containing 7, E3 ubiquitin protein ligase |
| chr1_+_47901689 | 0.02 |
ENST00000334793.5 |
FOXD2 |
forkhead box D2 |
Gene Ontology Analysis
Gene overrepresentation in biological_process category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.2 | 1.1 | GO:0038018 | Wnt receptor catabolic process(GO:0038018) |
| 0.2 | 0.9 | GO:0032227 | negative regulation of synaptic transmission, dopaminergic(GO:0032227) |
| 0.1 | 0.3 | GO:0042271 | susceptibility to natural killer cell mediated cytotoxicity(GO:0042271) |
| 0.1 | 1.7 | GO:0031581 | hemidesmosome assembly(GO:0031581) |
| 0.1 | 0.2 | GO:0061073 | ciliary body morphogenesis(GO:0061073) |
| 0.1 | 0.7 | GO:0097283 | keratinocyte apoptotic process(GO:0097283) regulation of keratinocyte apoptotic process(GO:1902172) |
| 0.1 | 0.3 | GO:0010166 | wax biosynthetic process(GO:0010025) wax metabolic process(GO:0010166) |
| 0.0 | 0.2 | GO:0031117 | positive regulation of microtubule depolymerization(GO:0031117) |
| 0.0 | 0.1 | GO:0035981 | tongue muscle cell differentiation(GO:0035981) positive regulation of skeletal muscle fiber differentiation(GO:1902811) regulation of tongue muscle cell differentiation(GO:2001035) positive regulation of tongue muscle cell differentiation(GO:2001037) |
| 0.0 | 0.2 | GO:0071072 | negative regulation of phospholipid biosynthetic process(GO:0071072) |
| 0.0 | 0.6 | GO:0045741 | positive regulation of epidermal growth factor-activated receptor activity(GO:0045741) |
| 0.0 | 0.2 | GO:2000857 | positive regulation of mineralocorticoid secretion(GO:2000857) positive regulation of aldosterone secretion(GO:2000860) |
| 0.0 | 0.1 | GO:0097359 | UDP-glucosylation(GO:0097359) |
| 0.0 | 0.5 | GO:0090179 | planar cell polarity pathway involved in neural tube closure(GO:0090179) |
| 0.0 | 0.1 | GO:0090271 | positive regulation of fibroblast growth factor production(GO:0090271) |
| 0.0 | 0.1 | GO:0007181 | transforming growth factor beta receptor complex assembly(GO:0007181) |
| 0.0 | 0.1 | GO:0051884 | negative regulation of hair follicle maturation(GO:0048817) regulation of anagen(GO:0051884) |
| 0.0 | 0.0 | GO:1904431 | positive regulation of t-circle formation(GO:1904431) |
| 0.0 | 0.2 | GO:0060125 | negative regulation of growth hormone secretion(GO:0060125) |
| 0.0 | 0.1 | GO:1904338 | regulation of dopaminergic neuron differentiation(GO:1904338) |
| 0.0 | 0.1 | GO:0002384 | hepatic immune response(GO:0002384) regulation of STAT protein import into nucleus(GO:2000364) positive regulation of STAT protein import into nucleus(GO:2000366) |
| 0.0 | 0.1 | GO:0035166 | post-embryonic hemopoiesis(GO:0035166) |
| 0.0 | 0.1 | GO:1900154 | regulation of bone trabecula formation(GO:1900154) negative regulation of bone trabecula formation(GO:1900155) |
| 0.0 | 0.1 | GO:0043314 | negative regulation of neutrophil degranulation(GO:0043314) |
| 0.0 | 0.1 | GO:0046013 | regulation of T cell homeostatic proliferation(GO:0046013) |
| 0.0 | 0.1 | GO:0033277 | abortive mitotic cell cycle(GO:0033277) |
| 0.0 | 0.1 | GO:2000211 | regulation of glutamate metabolic process(GO:2000211) |
| 0.0 | 0.0 | GO:0070105 | positive regulation of interleukin-6-mediated signaling pathway(GO:0070105) |
| 0.0 | 0.0 | GO:0007518 | myoblast fate determination(GO:0007518) |
| 0.0 | 0.1 | GO:0030185 | nitric oxide transport(GO:0030185) |
| 0.0 | 0.1 | GO:0048050 | post-embryonic eye morphogenesis(GO:0048050) |
| 0.0 | 0.1 | GO:2000686 | regulation of rubidium ion transmembrane transporter activity(GO:2000686) |
| 0.0 | 0.2 | GO:0061303 | cornea development in camera-type eye(GO:0061303) |
| 0.0 | 0.0 | GO:0002331 | pre-B cell allelic exclusion(GO:0002331) |
| 0.0 | 0.4 | GO:0071539 | protein localization to centrosome(GO:0071539) |
| 0.0 | 0.0 | GO:0003358 | noradrenergic neuron development(GO:0003358) |
| 0.0 | 0.0 | GO:0002337 | B-1a B cell differentiation(GO:0002337) |
| 0.0 | 0.1 | GO:0014809 | regulation of skeletal muscle contraction by regulation of release of sequestered calcium ion(GO:0014809) |
| 0.0 | 0.2 | GO:0006703 | estrogen biosynthetic process(GO:0006703) |
Gene overrepresentation in cellular_component category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.1 | 0.6 | GO:0005610 | laminin-5 complex(GO:0005610) |
| 0.1 | 0.5 | GO:0031313 | extrinsic component of endosome membrane(GO:0031313) |
| 0.0 | 0.8 | GO:0030056 | hemidesmosome(GO:0030056) |
| 0.0 | 0.1 | GO:0031417 | NatC complex(GO:0031417) |
| 0.0 | 0.3 | GO:0072687 | meiotic spindle(GO:0072687) |
| 0.0 | 0.7 | GO:0031092 | platelet alpha granule membrane(GO:0031092) |
| 0.0 | 0.1 | GO:0005896 | interleukin-6 receptor complex(GO:0005896) |
| 0.0 | 0.1 | GO:0031838 | haptoglobin-hemoglobin complex(GO:0031838) |
| 0.0 | 0.0 | GO:0048476 | Holliday junction resolvase complex(GO:0048476) |
| 0.0 | 0.3 | GO:0005861 | troponin complex(GO:0005861) |
| 0.0 | 0.1 | GO:0016013 | syntrophin complex(GO:0016013) |
| 0.0 | 0.3 | GO:0031231 | integral component of peroxisomal membrane(GO:0005779) intrinsic component of peroxisomal membrane(GO:0031231) |
Gene overrepresentation in molecular_function category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.1 | 0.6 | GO:0060961 | phospholipase D inhibitor activity(GO:0060961) |
| 0.1 | 0.3 | GO:0004666 | prostaglandin-endoperoxide synthase activity(GO:0004666) |
| 0.1 | 0.3 | GO:0080019 | fatty-acyl-CoA reductase (alcohol-forming) activity(GO:0080019) |
| 0.1 | 0.7 | GO:0038132 | neuregulin binding(GO:0038132) |
| 0.0 | 0.2 | GO:0047006 | 17-alpha,20-alpha-dihydroxypregn-4-en-3-one dehydrogenase activity(GO:0047006) |
| 0.0 | 0.1 | GO:0047192 | 1-alkylglycerophosphocholine O-acetyltransferase activity(GO:0047192) |
| 0.0 | 1.4 | GO:0005109 | frizzled binding(GO:0005109) |
| 0.0 | 0.1 | GO:0003980 | UDP-glucose:glycoprotein glucosyltransferase activity(GO:0003980) |
| 0.0 | 0.7 | GO:0030247 | pattern binding(GO:0001871) polysaccharide binding(GO:0030247) |
| 0.0 | 0.1 | GO:0005006 | epidermal growth factor-activated receptor activity(GO:0005006) |
| 0.0 | 0.1 | GO:0008401 | retinoic acid 4-hydroxylase activity(GO:0008401) |
| 0.0 | 0.3 | GO:0031014 | troponin T binding(GO:0031014) |
| 0.0 | 0.4 | GO:0050811 | GABA receptor binding(GO:0050811) |
| 0.0 | 0.5 | GO:0005545 | 1-phosphatidylinositol binding(GO:0005545) |
| 0.0 | 0.1 | GO:0004468 | lysine N-acetyltransferase activity, acting on acetyl phosphate as donor(GO:0004468) |
| 0.0 | 0.2 | GO:0017034 | Rap guanyl-nucleotide exchange factor activity(GO:0017034) |
| 0.0 | 0.1 | GO:0015119 | hexose phosphate transmembrane transporter activity(GO:0015119) organophosphate:inorganic phosphate antiporter activity(GO:0015315) hexose-phosphate:inorganic phosphate antiporter activity(GO:0015526) glucose 6-phosphate:inorganic phosphate antiporter activity(GO:0061513) |
Gene overrepresentation in C2:CP category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.1 | 1.4 | PID INTEGRIN4 PATHWAY | Alpha6 beta4 integrin-ligand interactions |
| 0.0 | 0.6 | PID ERBB NETWORK PATHWAY | ErbB receptor signaling network |
| 0.0 | 0.8 | SIG REGULATION OF THE ACTIN CYTOSKELETON BY RHO GTPASES | Genes related to regulation of the actin cytoskeleton |
Gene overrepresentation in C2:CP:REACTOME category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.0 | 0.7 | REACTOME ACTIVATION OF RAC | Genes involved in Activation of Rac |
| 0.0 | 2.1 | REACTOME CELL JUNCTION ORGANIZATION | Genes involved in Cell junction organization |