Project
Epithelial-Mesenchymal Transition, human (Scheel, 2011)
Navigation
Downloads
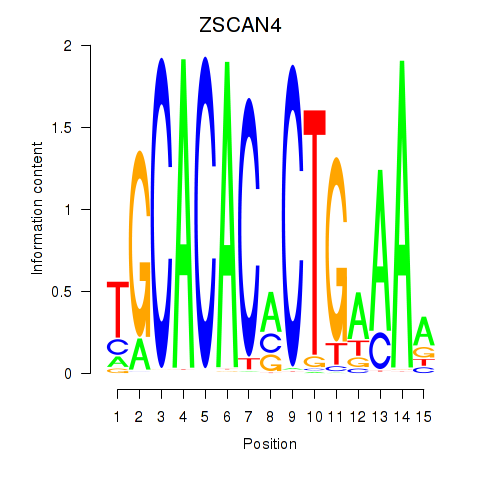
Results for ZSCAN4
Z-value: 0.64
Transcription factors associated with ZSCAN4
| Gene Symbol | Gene ID | Gene Info |
|---|---|---|
|
ZSCAN4
|
ENSG00000180532.6 | ZSCAN4 |
Activity-expression correlation:
| Gene | Promoter | Pearson corr. coef. | P-value | Plot |
|---|---|---|---|---|
| ZSCAN4 | hg19_v2_chr19_+_58180303_58180303 | -0.60 | 1.2e-01 | Click! |
Activity profile of ZSCAN4 motif
Sorted Z-values of ZSCAN4 motif
Network of associatons between targets according to the STRING database.
First level regulatory network of ZSCAN4
| Promoter | Score | Transcript | Gene | Gene Info |
|---|---|---|---|---|
| chr3_+_111393501 | 0.73 |
ENST00000393934.3 |
PLCXD2 |
phosphatidylinositol-specific phospholipase C, X domain containing 2 |
| chr21_+_25801041 | 0.58 |
ENST00000453784.2 ENST00000423581.1 |
AP000476.1 |
AP000476.1 |
| chr8_-_23712312 | 0.55 |
ENST00000290271.2 |
STC1 |
stanniocalcin 1 |
| chr1_+_159141397 | 0.55 |
ENST00000368124.4 ENST00000368125.4 ENST00000416746.1 |
CADM3 |
cell adhesion molecule 3 |
| chr13_-_114898016 | 0.35 |
ENST00000542651.1 ENST00000334062.7 |
RASA3 |
RAS p21 protein activator 3 |
| chr1_-_177133818 | 0.30 |
ENST00000424564.2 ENST00000361833.2 |
ASTN1 |
astrotactin 1 |
| chr4_-_89619386 | 0.30 |
ENST00000323061.5 |
NAP1L5 |
nucleosome assembly protein 1-like 5 |
| chr17_-_19648683 | 0.27 |
ENST00000573368.1 ENST00000457500.2 |
ALDH3A1 |
aldehyde dehydrogenase 3 family, member A1 |
| chr8_-_72987810 | 0.26 |
ENST00000262209.4 |
TRPA1 |
transient receptor potential cation channel, subfamily A, member 1 |
| chr17_-_19648916 | 0.26 |
ENST00000444455.1 ENST00000439102.2 |
ALDH3A1 |
aldehyde dehydrogenase 3 family, member A1 |
| chr11_+_45792967 | 0.19 |
ENST00000378779.2 |
CTD-2210P24.4 |
Uncharacterized protein |
| chr17_-_7080227 | 0.19 |
ENST00000574330.1 |
ASGR1 |
asialoglycoprotein receptor 1 |
| chr11_+_71938925 | 0.19 |
ENST00000538751.1 |
INPPL1 |
inositol polyphosphate phosphatase-like 1 |
| chr1_+_159409512 | 0.17 |
ENST00000423932.3 |
OR10J1 |
olfactory receptor, family 10, subfamily J, member 1 |
| chr12_+_129028500 | 0.16 |
ENST00000315208.8 |
TMEM132C |
transmembrane protein 132C |
| chr2_-_101034070 | 0.16 |
ENST00000264249.3 |
CHST10 |
carbohydrate sulfotransferase 10 |
| chr17_+_72462525 | 0.16 |
ENST00000360141.3 |
CD300A |
CD300a molecule |
| chr6_+_114178512 | 0.15 |
ENST00000368635.4 |
MARCKS |
myristoylated alanine-rich protein kinase C substrate |
| chr11_+_66624527 | 0.15 |
ENST00000393952.3 |
LRFN4 |
leucine rich repeat and fibronectin type III domain containing 4 |
| chr16_+_69140122 | 0.14 |
ENST00000219322.3 |
HAS3 |
hyaluronan synthase 3 |
| chr19_-_38714847 | 0.14 |
ENST00000420980.2 ENST00000355526.4 |
DPF1 |
D4, zinc and double PHD fingers family 1 |
| chrX_-_62571220 | 0.13 |
ENST00000374884.2 |
SPIN4 |
spindlin family, member 4 |
| chr2_-_175462934 | 0.13 |
ENST00000392546.2 ENST00000436221.1 |
WIPF1 |
WAS/WASL interacting protein family, member 1 |
| chr2_-_167232484 | 0.13 |
ENST00000375387.4 ENST00000303354.6 ENST00000409672.1 |
SCN9A |
sodium channel, voltage-gated, type IX, alpha subunit |
| chrX_-_62571187 | 0.13 |
ENST00000335144.3 |
SPIN4 |
spindlin family, member 4 |
| chr2_+_102759199 | 0.12 |
ENST00000409288.1 ENST00000410023.1 |
IL1R1 |
interleukin 1 receptor, type I |
| chr11_+_125462690 | 0.12 |
ENST00000392708.4 ENST00000529196.1 ENST00000531491.1 |
STT3A |
STT3A, subunit of the oligosaccharyltransferase complex (catalytic) |
| chr1_-_161519682 | 0.12 |
ENST00000367969.3 ENST00000443193.1 |
FCGR3A |
Fc fragment of IgG, low affinity IIIa, receptor (CD16a) |
| chr3_-_121740969 | 0.12 |
ENST00000393631.1 ENST00000273691.3 ENST00000344209.5 |
ILDR1 |
immunoglobulin-like domain containing receptor 1 |
| chr19_+_17516624 | 0.12 |
ENST00000596322.1 ENST00000600008.1 ENST00000601885.1 |
CTD-2521M24.9 |
CTD-2521M24.9 |
| chr12_-_96390108 | 0.12 |
ENST00000538703.1 ENST00000261208.3 |
HAL |
histidine ammonia-lyase |
| chr16_+_4674814 | 0.11 |
ENST00000415496.1 ENST00000587747.1 ENST00000399577.5 ENST00000588994.1 ENST00000586183.1 |
MGRN1 |
mahogunin ring finger 1, E3 ubiquitin protein ligase |
| chr6_-_153304697 | 0.11 |
ENST00000367241.3 |
FBXO5 |
F-box protein 5 |
| chr16_+_4674787 | 0.11 |
ENST00000262370.7 |
MGRN1 |
mahogunin ring finger 1, E3 ubiquitin protein ligase |
| chr17_-_33759509 | 0.11 |
ENST00000304905.5 |
SLFN12 |
schlafen family member 12 |
| chrX_+_49160148 | 0.11 |
ENST00000407599.3 |
GAGE10 |
G antigen 10 |
| chr14_-_55658252 | 0.11 |
ENST00000395425.2 |
DLGAP5 |
discs, large (Drosophila) homolog-associated protein 5 |
| chr14_-_23904861 | 0.10 |
ENST00000355349.3 |
MYH7 |
myosin, heavy chain 7, cardiac muscle, beta |
| chr15_+_78632666 | 0.10 |
ENST00000299529.6 |
CRABP1 |
cellular retinoic acid binding protein 1 |
| chr8_+_85095497 | 0.10 |
ENST00000522455.1 ENST00000521695.1 |
RALYL |
RALY RNA binding protein-like |
| chr9_+_125137565 | 0.10 |
ENST00000373698.5 |
PTGS1 |
prostaglandin-endoperoxide synthase 1 (prostaglandin G/H synthase and cyclooxygenase) |
| chr1_-_161519579 | 0.10 |
ENST00000426740.1 |
FCGR3A |
Fc fragment of IgG, low affinity IIIa, receptor (CD16a) |
| chr15_+_38544476 | 0.10 |
ENST00000299084.4 |
SPRED1 |
sprouty-related, EVH1 domain containing 1 |
| chr19_+_17516494 | 0.10 |
ENST00000534306.1 |
CTD-2521M24.9 |
CTD-2521M24.9 |
| chr1_+_32645645 | 0.10 |
ENST00000373609.1 |
TXLNA |
taxilin alpha |
| chr15_+_72947079 | 0.09 |
ENST00000421285.3 |
GOLGA6B |
golgin A6 family, member B |
| chr12_-_96390063 | 0.09 |
ENST00000541929.1 |
HAL |
histidine ammonia-lyase |
| chr8_+_85095769 | 0.09 |
ENST00000518566.1 |
RALYL |
RALY RNA binding protein-like |
| chr17_+_73089382 | 0.09 |
ENST00000538213.2 ENST00000584118.1 |
SLC16A5 |
solute carrier family 16 (monocarboxylate transporter), member 5 |
| chr20_-_16554078 | 0.09 |
ENST00000354981.2 ENST00000355755.3 ENST00000378003.2 ENST00000408042.1 |
KIF16B |
kinesin family member 16B |
| chr16_+_56385290 | 0.09 |
ENST00000564727.1 |
GNAO1 |
guanine nucleotide binding protein (G protein), alpha activating activity polypeptide O |
| chr7_-_155326532 | 0.09 |
ENST00000406197.1 ENST00000321736.5 |
CNPY1 |
canopy FGF signaling regulator 1 |
| chr15_-_37392703 | 0.09 |
ENST00000382766.2 ENST00000444725.1 |
MEIS2 |
Meis homeobox 2 |
| chr2_+_204732666 | 0.08 |
ENST00000295854.6 ENST00000472206.1 |
CTLA4 |
cytotoxic T-lymphocyte-associated protein 4 |
| chr8_-_134115118 | 0.08 |
ENST00000395352.3 ENST00000338087.5 |
SLA |
Src-like-adaptor |
| chr1_-_110283138 | 0.08 |
ENST00000256594.3 |
GSTM3 |
glutathione S-transferase mu 3 (brain) |
| chr16_-_80838195 | 0.08 |
ENST00000570137.2 |
CDYL2 |
chromodomain protein, Y-like 2 |
| chr19_-_40562063 | 0.08 |
ENST00000598845.1 ENST00000593605.1 ENST00000221355.6 ENST00000434248.1 |
ZNF780B |
zinc finger protein 780B |
| chr15_+_63414017 | 0.08 |
ENST00000413507.2 |
LACTB |
lactamase, beta |
| chr1_+_32645269 | 0.08 |
ENST00000373610.3 |
TXLNA |
taxilin alpha |
| chr14_-_94254821 | 0.08 |
ENST00000393140.1 |
PRIMA1 |
proline rich membrane anchor 1 |
| chr18_+_32173276 | 0.08 |
ENST00000591816.1 ENST00000588125.1 ENST00000598334.1 ENST00000588684.1 ENST00000554864.3 ENST00000399121.5 ENST00000595022.1 ENST00000269190.7 ENST00000399097.3 |
DTNA |
dystrobrevin, alpha |
| chr4_+_15779901 | 0.08 |
ENST00000226279.3 |
CD38 |
CD38 molecule |
| chr20_+_23168759 | 0.08 |
ENST00000411595.1 |
RP4-737E23.2 |
RP4-737E23.2 |
| chr12_+_66218212 | 0.08 |
ENST00000393578.3 ENST00000425208.2 ENST00000536545.1 ENST00000354636.3 |
HMGA2 |
high mobility group AT-hook 2 |
| chr5_+_149887672 | 0.08 |
ENST00000261797.6 |
NDST1 |
N-deacetylase/N-sulfotransferase (heparan glucosaminyl) 1 |
| chr19_-_11266471 | 0.07 |
ENST00000592540.1 |
SPC24 |
SPC24, NDC80 kinetochore complex component |
| chr7_+_107224364 | 0.07 |
ENST00000491150.1 |
BCAP29 |
B-cell receptor-associated protein 29 |
| chr12_-_57634475 | 0.07 |
ENST00000393825.1 |
NDUFA4L2 |
NADH dehydrogenase (ubiquinone) 1 alpha subcomplex, 4-like 2 |
| chr12_-_11214893 | 0.07 |
ENST00000533467.1 |
TAS2R46 |
taste receptor, type 2, member 46 |
| chr17_+_28256874 | 0.07 |
ENST00000541045.1 ENST00000536908.2 |
EFCAB5 |
EF-hand calcium binding domain 5 |
| chr1_+_157963063 | 0.07 |
ENST00000360089.4 ENST00000368173.3 ENST00000392272.2 |
KIRREL |
kin of IRRE like (Drosophila) |
| chr13_-_99404875 | 0.07 |
ENST00000376503.5 |
SLC15A1 |
solute carrier family 15 (oligopeptide transporter), member 1 |
| chr16_+_16472912 | 0.07 |
ENST00000530217.2 |
NPIPA7 |
nuclear pore complex interacting protein family, member A7 |
| chr8_+_38831683 | 0.07 |
ENST00000302495.4 |
HTRA4 |
HtrA serine peptidase 4 |
| chr21_-_46221684 | 0.07 |
ENST00000330942.5 |
UBE2G2 |
ubiquitin-conjugating enzyme E2G 2 |
| chr3_-_195538760 | 0.07 |
ENST00000475231.1 |
MUC4 |
mucin 4, cell surface associated |
| chr19_-_42636543 | 0.07 |
ENST00000528894.4 ENST00000560804.2 ENST00000560558.1 ENST00000560398.1 ENST00000526816.2 |
POU2F2 |
POU class 2 homeobox 2 |
| chr2_+_237994519 | 0.07 |
ENST00000392008.2 ENST00000409334.1 ENST00000409629.1 |
COPS8 |
COP9 signalosome subunit 8 |
| chr1_+_15764931 | 0.07 |
ENST00000375949.4 ENST00000375943.2 |
CTRC |
chymotrypsin C (caldecrin) |
| chr15_-_53082178 | 0.07 |
ENST00000305901.5 |
ONECUT1 |
one cut homeobox 1 |
| chr11_+_111807863 | 0.07 |
ENST00000440460.2 |
DIXDC1 |
DIX domain containing 1 |
| chr1_+_89829610 | 0.07 |
ENST00000370456.4 ENST00000535065.1 |
GBP6 |
guanylate binding protein family, member 6 |
| chr9_-_14910420 | 0.06 |
ENST00000380880.3 |
FREM1 |
FRAS1 related extracellular matrix 1 |
| chr21_+_46066331 | 0.06 |
ENST00000334670.8 |
KRTAP10-11 |
keratin associated protein 10-11 |
| chr1_+_169337172 | 0.06 |
ENST00000367807.3 ENST00000367808.3 ENST00000329281.2 ENST00000420531.1 |
BLZF1 |
basic leucine zipper nuclear factor 1 |
| chr2_+_189156389 | 0.06 |
ENST00000409843.1 |
GULP1 |
GULP, engulfment adaptor PTB domain containing 1 |
| chr17_+_9066252 | 0.06 |
ENST00000436734.1 |
NTN1 |
netrin 1 |
| chr1_-_161600822 | 0.06 |
ENST00000534776.1 ENST00000540048.1 |
FCGR3B FCGR3A |
Fc fragment of IgG, low affinity IIIb, receptor (CD16b) Fc fragment of IgG, low affinity IIIa, receptor (CD16a) |
| chr12_-_81331460 | 0.06 |
ENST00000549417.1 |
LIN7A |
lin-7 homolog A (C. elegans) |
| chr9_+_103790991 | 0.06 |
ENST00000374874.3 |
LPPR1 |
Lipid phosphate phosphatase-related protein type 1 |
| chr13_+_60971080 | 0.06 |
ENST00000377894.2 |
TDRD3 |
tudor domain containing 3 |
| chr19_-_9649253 | 0.06 |
ENST00000593003.1 |
ZNF426 |
zinc finger protein 426 |
| chr16_-_55909255 | 0.05 |
ENST00000290567.9 |
CES5A |
carboxylesterase 5A |
| chr16_-_55909272 | 0.05 |
ENST00000319165.9 |
CES5A |
carboxylesterase 5A |
| chr2_-_219696511 | 0.05 |
ENST00000439262.2 |
PRKAG3 |
protein kinase, AMP-activated, gamma 3 non-catalytic subunit |
| chr16_-_55909211 | 0.05 |
ENST00000520435.1 |
CES5A |
carboxylesterase 5A |
| chr3_-_195538728 | 0.05 |
ENST00000349607.4 ENST00000346145.4 |
MUC4 |
mucin 4, cell surface associated |
| chr19_+_51630287 | 0.05 |
ENST00000599948.1 |
SIGLEC9 |
sialic acid binding Ig-like lectin 9 |
| chr3_+_9944492 | 0.05 |
ENST00000383814.3 ENST00000454190.2 ENST00000454992.1 |
IL17RE |
interleukin 17 receptor E |
| chr12_+_122242597 | 0.05 |
ENST00000267197.5 |
SETD1B |
SET domain containing 1B |
| chr1_-_161600990 | 0.05 |
ENST00000531221.1 |
FCGR3B |
Fc fragment of IgG, low affinity IIIb, receptor (CD16b) |
| chr17_-_73761222 | 0.05 |
ENST00000437911.1 ENST00000225614.2 |
GALK1 |
galactokinase 1 |
| chr17_-_58499698 | 0.05 |
ENST00000590297.1 |
USP32 |
ubiquitin specific peptidase 32 |
| chr3_+_9944303 | 0.05 |
ENST00000421412.1 ENST00000295980.3 |
IL17RE |
interleukin 17 receptor E |
| chr10_+_90354503 | 0.05 |
ENST00000531458.1 |
LIPJ |
lipase, family member J |
| chr1_-_161600942 | 0.04 |
ENST00000421702.2 |
FCGR3B |
Fc fragment of IgG, low affinity IIIb, receptor (CD16b) |
| chr14_+_53173910 | 0.04 |
ENST00000606149.1 ENST00000555339.1 ENST00000556813.1 |
PSMC6 |
proteasome (prosome, macropain) 26S subunit, ATPase, 6 |
| chr14_+_53173890 | 0.04 |
ENST00000445930.2 |
PSMC6 |
proteasome (prosome, macropain) 26S subunit, ATPase, 6 |
| chr13_+_60971427 | 0.04 |
ENST00000535286.1 ENST00000377881.2 |
TDRD3 |
tudor domain containing 3 |
| chr15_+_75575176 | 0.04 |
ENST00000434739.3 |
GOLGA6D |
golgin A6 family, member D |
| chr11_-_62783276 | 0.04 |
ENST00000535878.1 ENST00000545207.1 |
SLC22A8 |
solute carrier family 22 (organic anion transporter), member 8 |
| chr11_-_4414880 | 0.04 |
ENST00000254436.7 ENST00000543625.1 |
TRIM21 |
tripartite motif containing 21 |
| chrX_+_2984874 | 0.04 |
ENST00000359361.2 |
ARSF |
arylsulfatase F |
| chr19_-_46999755 | 0.04 |
ENST00000599531.1 |
PNMAL2 |
paraneoplastic Ma antigen family-like 2 |
| chr12_+_133613878 | 0.04 |
ENST00000392319.2 ENST00000543758.1 |
ZNF84 |
zinc finger protein 84 |
| chr10_+_18549645 | 0.04 |
ENST00000396576.2 |
CACNB2 |
calcium channel, voltage-dependent, beta 2 subunit |
| chr11_-_33774944 | 0.04 |
ENST00000532057.1 ENST00000531080.1 |
FBXO3 |
F-box protein 3 |
| chr9_+_124048864 | 0.04 |
ENST00000545652.1 |
GSN |
gelsolin |
| chr20_-_21494654 | 0.04 |
ENST00000377142.4 |
NKX2-2 |
NK2 homeobox 2 |
| chr15_+_28623784 | 0.04 |
ENST00000526619.2 ENST00000337838.7 ENST00000532622.2 |
GOLGA8F |
golgin A8 family, member F |
| chr11_-_77791156 | 0.04 |
ENST00000281031.4 |
NDUFC2 |
NADH dehydrogenase (ubiquinone) 1, subcomplex unknown, 2, 14.5kDa |
| chr5_-_172036436 | 0.04 |
ENST00000601856.1 |
AC027309.1 |
AC027309.1 |
| chr19_-_35626104 | 0.04 |
ENST00000310123.3 ENST00000392225.3 |
LGI4 |
leucine-rich repeat LGI family, member 4 |
| chr1_-_193075180 | 0.03 |
ENST00000367440.3 |
GLRX2 |
glutaredoxin 2 |
| chr6_-_31620149 | 0.03 |
ENST00000435080.1 ENST00000375976.4 ENST00000441054.1 |
BAG6 |
BCL2-associated athanogene 6 |
| chr9_-_114090713 | 0.03 |
ENST00000302681.1 ENST00000374428.1 |
OR2K2 |
olfactory receptor, family 2, subfamily K, member 2 |
| chr1_+_31971828 | 0.03 |
ENST00000418618.1 |
RP11-439L8.3 |
RP11-439L8.3 |
| chr7_-_135194822 | 0.03 |
ENST00000428680.2 ENST00000315544.5 ENST00000423368.2 ENST00000451834.1 ENST00000361528.4 ENST00000356162.4 ENST00000541284.1 |
CNOT4 |
CCR4-NOT transcription complex, subunit 4 |
| chr7_+_27282319 | 0.03 |
ENST00000222761.3 |
EVX1 |
even-skipped homeobox 1 |
| chr17_+_30348024 | 0.03 |
ENST00000327564.7 ENST00000584368.1 ENST00000394713.3 ENST00000341671.7 |
LRRC37B |
leucine rich repeat containing 37B |
| chr1_-_228603694 | 0.03 |
ENST00000366697.2 |
TRIM17 |
tripartite motif containing 17 |
| chr6_-_76203454 | 0.03 |
ENST00000237172.7 |
FILIP1 |
filamin A interacting protein 1 |
| chr10_+_80828774 | 0.03 |
ENST00000334512.5 |
ZMIZ1 |
zinc finger, MIZ-type containing 1 |
| chrX_+_1387721 | 0.03 |
ENST00000419094.1 ENST00000381509.3 ENST00000494969.2 ENST00000355805.2 ENST00000355432.3 |
CSF2RA |
colony stimulating factor 2 receptor, alpha, low-affinity (granulocyte-macrophage) |
| chr2_+_183580954 | 0.03 |
ENST00000264065.7 |
DNAJC10 |
DnaJ (Hsp40) homolog, subfamily C, member 10 |
| chr19_-_10530784 | 0.03 |
ENST00000593124.1 |
CDC37 |
cell division cycle 37 |
| chr10_+_88414338 | 0.02 |
ENST00000241891.5 ENST00000443292.1 |
OPN4 |
opsin 4 |
| chrX_+_119495934 | 0.02 |
ENST00000218008.3 ENST00000361319.3 ENST00000539306.1 |
ATP1B4 |
ATPase, Na+/K+ transporting, beta 4 polypeptide |
| chr13_-_52733980 | 0.02 |
ENST00000339406.3 |
NEK3 |
NIMA-related kinase 3 |
| chr5_+_152870287 | 0.02 |
ENST00000340592.5 |
GRIA1 |
glutamate receptor, ionotropic, AMPA 1 |
| chr11_+_61717842 | 0.02 |
ENST00000449131.2 |
BEST1 |
bestrophin 1 |
| chr1_-_154580616 | 0.02 |
ENST00000368474.4 |
ADAR |
adenosine deaminase, RNA-specific |
| chr10_+_88414298 | 0.02 |
ENST00000372071.2 |
OPN4 |
opsin 4 |
| chr17_+_2699697 | 0.02 |
ENST00000254695.8 ENST00000366401.4 ENST00000542807.1 |
RAP1GAP2 |
RAP1 GTPase activating protein 2 |
| chr20_+_43343517 | 0.02 |
ENST00000372865.4 |
WISP2 |
WNT1 inducible signaling pathway protein 2 |
| chr2_-_99952769 | 0.02 |
ENST00000409434.1 ENST00000434323.1 ENST00000264255.3 |
TXNDC9 |
thioredoxin domain containing 9 |
| chr2_-_224810070 | 0.02 |
ENST00000429915.1 ENST00000233055.4 |
WDFY1 |
WD repeat and FYVE domain containing 1 |
| chr2_-_36779411 | 0.02 |
ENST00000406220.1 |
AC007401.2 |
Uncharacterized protein |
| chr15_+_63413990 | 0.02 |
ENST00000261893.4 |
LACTB |
lactamase, beta |
| chr8_+_68864330 | 0.02 |
ENST00000288368.4 |
PREX2 |
phosphatidylinositol-3,4,5-trisphosphate-dependent Rac exchange factor 2 |
| chr1_+_209929377 | 0.02 |
ENST00000400959.3 ENST00000367025.3 |
TRAF3IP3 |
TRAF3 interacting protein 3 |
| chr10_+_82009466 | 0.02 |
ENST00000356374.4 |
AL359195.1 |
Uncharacterized protein; cDNA FLJ46261 fis, clone TESTI4025062 |
| chr1_+_209929494 | 0.02 |
ENST00000367026.3 |
TRAF3IP3 |
TRAF3 interacting protein 3 |
| chr13_-_79980315 | 0.02 |
ENST00000438737.2 |
RBM26 |
RNA binding motif protein 26 |
| chr3_+_130569429 | 0.02 |
ENST00000505330.1 ENST00000504381.1 ENST00000507488.2 ENST00000393221.4 |
ATP2C1 |
ATPase, Ca++ transporting, type 2C, member 1 |
| chr2_-_214016314 | 0.02 |
ENST00000434687.1 ENST00000374319.4 |
IKZF2 |
IKAROS family zinc finger 2 (Helios) |
| chr17_-_46799872 | 0.02 |
ENST00000290294.3 |
PRAC1 |
prostate cancer susceptibility candidate 1 |
| chr12_+_122459757 | 0.01 |
ENST00000261822.4 |
BCL7A |
B-cell CLL/lymphoma 7A |
| chr16_-_86542652 | 0.01 |
ENST00000599749.1 |
FENDRR |
FOXF1 adjacent non-coding developmental regulatory RNA |
| chr15_+_30375158 | 0.01 |
ENST00000341650.6 ENST00000567927.1 |
GOLGA8J |
golgin A8 family, member J |
| chr15_+_76135622 | 0.01 |
ENST00000338677.4 ENST00000267938.4 ENST00000569423.1 |
UBE2Q2 |
ubiquitin-conjugating enzyme E2Q family member 2 |
| chr7_-_99381798 | 0.01 |
ENST00000415003.1 ENST00000354593.2 |
CYP3A4 |
cytochrome P450, family 3, subfamily A, polypeptide 4 |
| chr3_-_53290016 | 0.01 |
ENST00000423525.2 ENST00000423516.1 ENST00000296289.6 ENST00000462138.1 |
TKT |
transketolase |
| chr10_-_62332357 | 0.01 |
ENST00000503366.1 |
ANK3 |
ankyrin 3, node of Ranvier (ankyrin G) |
| chr8_-_16050142 | 0.01 |
ENST00000536385.1 ENST00000381998.4 |
MSR1 |
macrophage scavenger receptor 1 |
| chr3_+_130569592 | 0.01 |
ENST00000533801.2 |
ATP2C1 |
ATPase, Ca++ transporting, type 2C, member 1 |
| chr11_+_61717535 | 0.01 |
ENST00000534553.1 ENST00000301774.9 |
BEST1 |
bestrophin 1 |
| chr6_-_76203345 | 0.01 |
ENST00000393004.2 |
FILIP1 |
filamin A interacting protein 1 |
| chrX_+_17393543 | 0.01 |
ENST00000380060.3 |
NHS |
Nance-Horan syndrome (congenital cataracts and dental anomalies) |
| chr4_-_168155730 | 0.01 |
ENST00000502330.1 ENST00000357154.3 |
SPOCK3 |
sparc/osteonectin, cwcv and kazal-like domains proteoglycan (testican) 3 |
| chr3_+_52009110 | 0.01 |
ENST00000491470.1 |
ABHD14A |
abhydrolase domain containing 14A |
| chr2_+_240323439 | 0.01 |
ENST00000428471.1 ENST00000413029.1 |
AC062017.1 |
Uncharacterized protein |
| chr16_+_23847267 | 0.01 |
ENST00000321728.7 |
PRKCB |
protein kinase C, beta |
| chr4_-_152682129 | 0.01 |
ENST00000512306.1 ENST00000508611.1 ENST00000515812.1 ENST00000263985.6 |
PET112 |
PET112 homolog (yeast) |
| chr1_+_231376941 | 0.01 |
ENST00000436239.1 ENST00000366647.4 ENST00000366646.3 ENST00000416000.1 |
GNPAT |
glyceronephosphate O-acyltransferase |
| chr13_+_114238997 | 0.00 |
ENST00000538138.1 ENST00000375370.5 |
TFDP1 |
transcription factor Dp-1 |
| chrX_-_20134713 | 0.00 |
ENST00000452324.3 |
MAP7D2 |
MAP7 domain containing 2 |
| chr3_-_52569023 | 0.00 |
ENST00000307076.4 |
NT5DC2 |
5'-nucleotidase domain containing 2 |
| chr4_-_168155417 | 0.00 |
ENST00000511269.1 ENST00000506697.1 ENST00000512042.1 |
SPOCK3 |
sparc/osteonectin, cwcv and kazal-like domains proteoglycan (testican) 3 |
| chr1_+_94884023 | 0.00 |
ENST00000315713.5 |
ABCD3 |
ATP-binding cassette, sub-family D (ALD), member 3 |
| chr21_-_33651324 | 0.00 |
ENST00000290130.3 |
MIS18A |
MIS18 kinetochore protein A |
| chr5_+_31193847 | 0.00 |
ENST00000514738.1 ENST00000265071.2 |
CDH6 |
cadherin 6, type 2, K-cadherin (fetal kidney) |
| chr19_-_40596767 | 0.00 |
ENST00000599972.1 ENST00000450241.2 ENST00000595687.2 |
ZNF780A |
zinc finger protein 780A |
| chr19_-_42636617 | 0.00 |
ENST00000529067.1 ENST00000529952.1 ENST00000533720.1 ENST00000389341.5 ENST00000342301.4 |
POU2F2 |
POU class 2 homeobox 2 |
| chr22_+_17082732 | 0.00 |
ENST00000558085.2 ENST00000592918.1 ENST00000400593.2 ENST00000592107.1 ENST00000426585.1 ENST00000591299.1 |
TPTEP1 |
transmembrane phosphatase with tensin homology pseudogene 1 |
Gene Ontology Analysis
Gene overrepresentation in biological_process category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.1 | 0.3 | GO:0050968 | thermoception(GO:0050955) detection of chemical stimulus involved in sensory perception of pain(GO:0050968) |
| 0.1 | 0.2 | GO:1905154 | negative regulation of MyD88-dependent toll-like receptor signaling pathway(GO:0034125) negative regulation of eosinophil activation(GO:1902567) negative regulation of membrane invagination(GO:1905154) negative regulation of eosinophil migration(GO:2000417) |
| 0.0 | 0.5 | GO:0001886 | endothelial cell morphogenesis(GO:0001886) |
| 0.0 | 0.1 | GO:0045226 | extracellular polysaccharide biosynthetic process(GO:0045226) extracellular polysaccharide metabolic process(GO:0046379) |
| 0.0 | 0.2 | GO:0019557 | histidine catabolic process to glutamate and formamide(GO:0019556) histidine catabolic process to glutamate and formate(GO:0019557) formamide metabolic process(GO:0043606) |
| 0.0 | 0.1 | GO:2000661 | positive regulation of interleukin-1-mediated signaling pathway(GO:2000661) |
| 0.0 | 0.1 | GO:0043686 | co-translational protein modification(GO:0043686) |
| 0.0 | 0.1 | GO:0007057 | spindle assembly involved in female meiosis I(GO:0007057) |
| 0.0 | 0.1 | GO:0014724 | regulation of twitch skeletal muscle contraction(GO:0014724) |
| 0.0 | 0.1 | GO:0016103 | diterpenoid catabolic process(GO:0016103) retinoic acid catabolic process(GO:0034653) |
| 0.0 | 0.1 | GO:0042938 | dipeptide transport(GO:0042938) |
| 0.0 | 0.1 | GO:0003131 | mesodermal-endodermal cell signaling(GO:0003131) programmed DNA elimination(GO:0031049) chromosome breakage(GO:0031052) histone H2A-S139 phosphorylation(GO:0035978) positive regulation of cellular response to X-ray(GO:2000685) |
| 0.0 | 0.2 | GO:0043951 | negative regulation of cAMP-mediated signaling(GO:0043951) |
| 0.0 | 0.1 | GO:0007079 | mitotic chromosome movement towards spindle pole(GO:0007079) |
| 0.0 | 0.1 | GO:0010387 | COP9 signalosome assembly(GO:0010387) |
| 0.0 | 0.1 | GO:0045590 | negative regulation of regulatory T cell differentiation(GO:0045590) |
| 0.0 | 0.1 | GO:0008065 | establishment of blood-nerve barrier(GO:0008065) |
| 0.0 | 0.1 | GO:0060979 | vasculogenesis involved in coronary vascular morphogenesis(GO:0060979) |
| 0.0 | 0.0 | GO:0006059 | hexitol metabolic process(GO:0006059) |
| 0.0 | 0.0 | GO:0021529 | spinal cord oligodendrocyte cell differentiation(GO:0021529) spinal cord oligodendrocyte cell fate specification(GO:0021530) |
| 0.0 | 0.0 | GO:0090086 | negative regulation of protein deubiquitination(GO:0090086) |
| 0.0 | 0.0 | GO:0021913 | regulation of transcription from RNA polymerase II promoter involved in ventral spinal cord interneuron specification(GO:0021913) |
Gene overrepresentation in cellular_component category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.1 | 0.2 | GO:0042585 | germinal vesicle(GO:0042585) |
| 0.0 | 0.1 | GO:0061689 | tricellular tight junction(GO:0061689) |
| 0.0 | 0.3 | GO:0031235 | intrinsic component of the cytoplasmic side of the plasma membrane(GO:0031235) |
| 0.0 | 0.1 | GO:0031262 | Ndc80 complex(GO:0031262) |
Gene overrepresentation in molecular_function category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.1 | 0.5 | GO:0018479 | benzaldehyde dehydrogenase (NAD+) activity(GO:0018479) |
| 0.1 | 0.3 | GO:0097604 | temperature-gated cation channel activity(GO:0097604) |
| 0.0 | 0.1 | GO:0070506 | high-density lipoprotein particle receptor activity(GO:0070506) |
| 0.0 | 0.1 | GO:0050501 | hyaluronan synthase activity(GO:0050501) |
| 0.0 | 0.1 | GO:0005173 | stem cell factor receptor binding(GO:0005173) |
| 0.0 | 0.1 | GO:0004909 | interleukin-1, Type I, activating receptor activity(GO:0004909) |
| 0.0 | 0.2 | GO:0016841 | ammonia-lyase activity(GO:0016841) |
| 0.0 | 0.2 | GO:0004873 | asialoglycoprotein receptor activity(GO:0004873) |
| 0.0 | 0.1 | GO:0050135 | NAD(P)+ nucleosidase activity(GO:0050135) |
| 0.0 | 0.1 | GO:0004666 | prostaglandin-endoperoxide synthase activity(GO:0004666) |
| 0.0 | 0.4 | GO:0019864 | IgG binding(GO:0019864) |
| 0.0 | 0.1 | GO:0030197 | extracellular matrix constituent, lubricant activity(GO:0030197) |
| 0.0 | 0.1 | GO:0035501 | MH1 domain binding(GO:0035501) |
| 0.0 | 0.2 | GO:0034485 | phosphatidylinositol-3,4,5-trisphosphate 5-phosphatase activity(GO:0034485) |
| 0.0 | 0.1 | GO:0050119 | N-acetylglucosamine deacetylase activity(GO:0050119) |
| 0.0 | 0.1 | GO:0015333 | peptide:proton symporter activity(GO:0015333) proton-dependent peptide secondary active transmembrane transporter activity(GO:0022897) oligopeptide transmembrane transporter activity(GO:0035673) |
| 0.0 | 0.3 | GO:0099604 | calcium-release channel activity(GO:0015278) ligand-gated calcium channel activity(GO:0099604) |
| 0.0 | 0.0 | GO:0004335 | galactokinase activity(GO:0004335) |
| 0.0 | 0.1 | GO:0055104 | ligase inhibitor activity(GO:0055104) ubiquitin ligase inhibitor activity(GO:1990948) |
| 0.0 | 0.1 | GO:0030368 | interleukin-17 receptor activity(GO:0030368) |
Gene overrepresentation in C2:CP:REACTOME category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.0 | 0.2 | REACTOME REGULATION OF INSULIN SECRETION BY ACETYLCHOLINE | Genes involved in Regulation of Insulin Secretion by Acetylcholine |