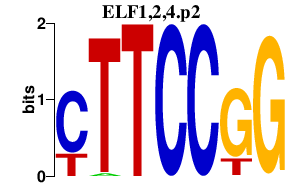
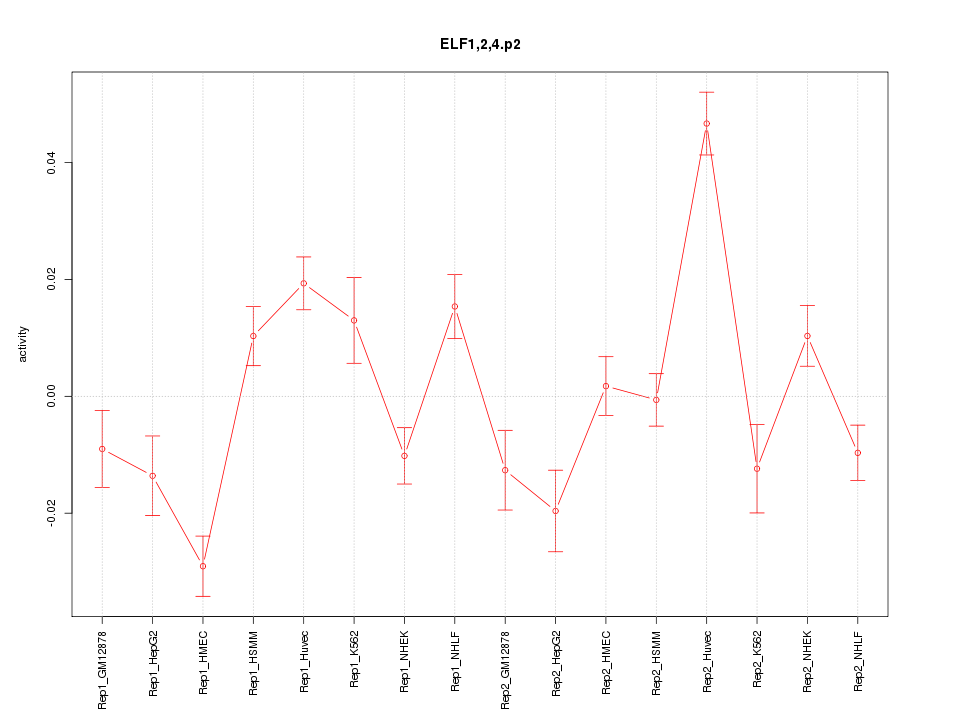
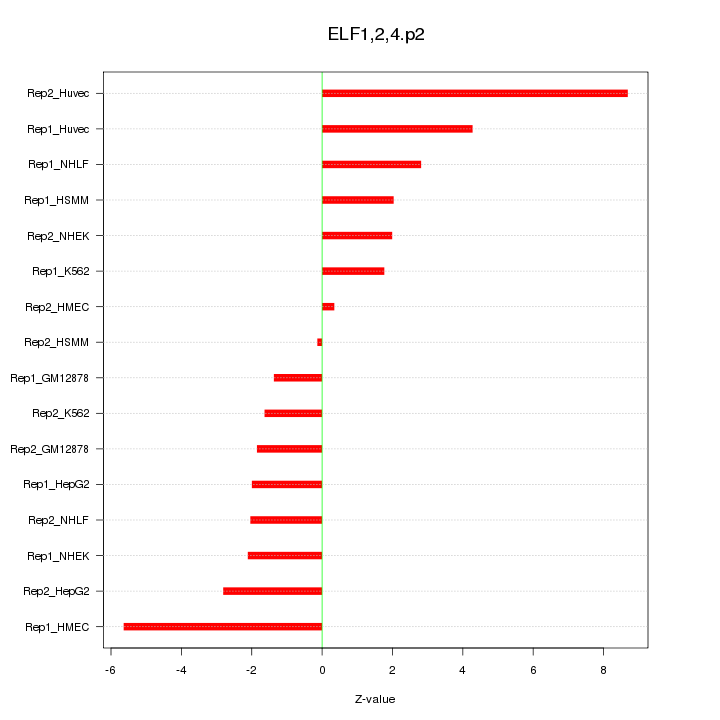
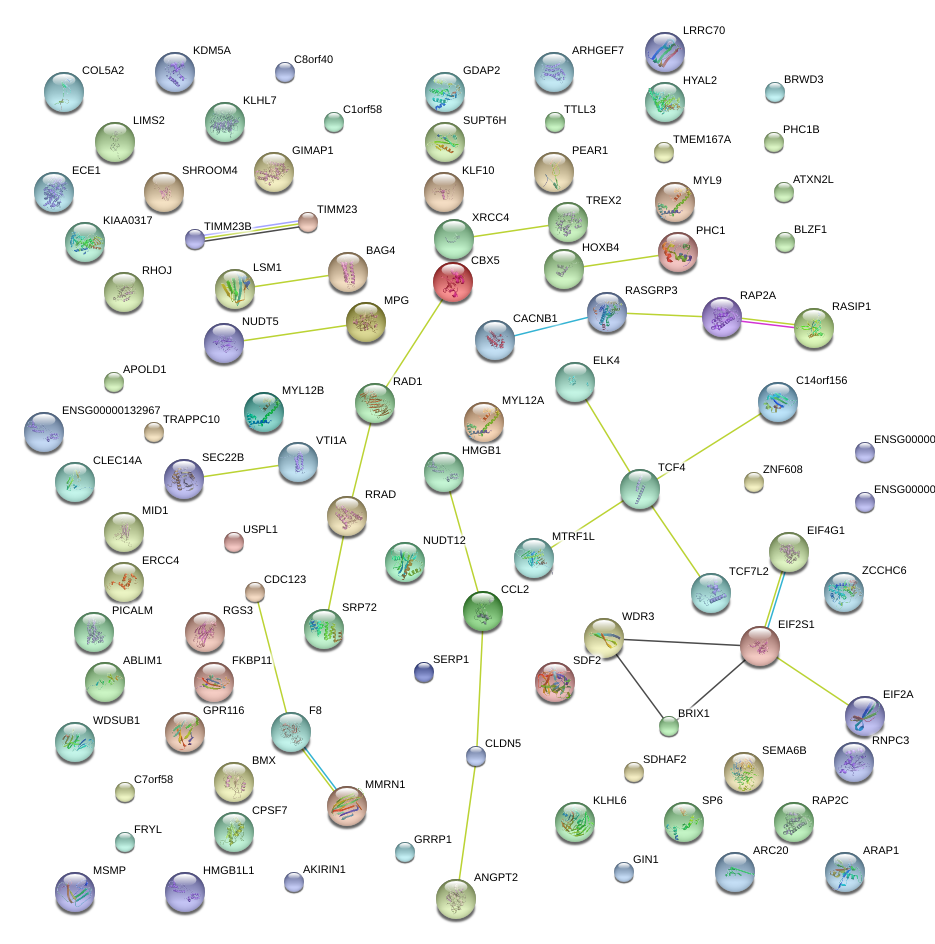
|
chr13_+_110653407
|
3.252
|
|
ARHGEF7
|
Rho guanine nucleotide exchange factor (GEF) 7
|
|
chr1_+_26358097
|
2.875
|
NM_024869
|
GRRP1
|
glycine/arginine rich protein 1
|
|
chr5_+_61910318
|
2.849
|
NM_181506
|
LRRC70
|
leucine rich repeat containing 70
|
|
chr13_-_30089509
|
2.399
|
|
HMGB1
|
high-mobility group box 1
|
|
chr18_+_3252184
|
2.315
|
|
MYL12B
|
myosin, light chain 12B, regulatory
|
|
chr19_-_53935648
|
2.252
|
NM_017805
|
RASIP1
|
Ras interacting protein 1
|
|
chr18_+_3252110
|
2.155
|
NM_033546
|
MYL12B
|
myosin, light chain 12B, regulatory
|
|
chr8_+_38153514
|
1.983
|
|
BAG4
|
BCL2-associated athanogene 4
|
|
chr9_+_115303527
|
1.982
|
NM_130795
|
RGS3
|
regulator of G-protein signaling 3
|
|
chr14_-_37795290
|
1.911
|
NM_175060
|
CLEC14A
|
C-type lectin domain family 14, member A
|
|
chr12_-_52939636
|
1.877
|
NM_001127321
|
CBX5
|
chromobox homolog 5
|
|
chr12_+_12829807
|
1.867
|
NM_030817
|
APOLD1
|
apolipoprotein L domain containing 1
|
|
chr1_-_21478571
|
1.859
|
NM_001113347
|
ECE1
|
endothelin converting enzyme 1
|
|
chr17_-_44010543
|
1.849
|
NM_024015
|
HOXB4
|
homeobox B4
|
|
chr3_-_184756101
|
1.840
|
NM_130446
|
KLHL6
|
kelch-like 6 (Drosophila)
|
|
chr1_+_155130146
|
1.809
|
NM_001080471
|
PEAR1
|
platelet endothelial aggregation receptor 1
|
|
chr8_+_38153453
|
1.809
|
|
BAG4
|
BCL2-associated athanogene 4
|
|
chr8_+_38153223
|
1.793
|
NM_004874
|
BAG4
|
BCL2-associated athanogene 4
|
|
chr2_+_33514896
|
1.785
|
NM_170672
|
RASGRP3
|
RAS guanyl releasing protein 3 (calcium and DAG-regulated)
|
|
chr22_-_17892859
|
1.753
|
NM_001130861
NM_003277
|
CLDN5
|
claudin 5
|
|
chr14_-_37795001
|
1.727
|
|
CLEC14A
|
C-type lectin domain family 14, member A
|
|
chr6_-_46997579
|
1.721
|
NM_001098518
|
GPR116
|
G protein-coupled receptor 116
|
|
chr10_-_116434364
|
1.679
|
NM_001003407
NM_001003408
|
ABLIM1
|
actin binding LIM protein 1
|
|
chr22_-_34566210
|
1.654
|
|
RBFOX2
|
RNA binding protein, fox-1 homolog (C. elegans) 2
|
|
chr7_+_150044530
|
1.594
|
NM_130759
|
GIMAP1
|
GTPase, IMAP family member 1
|
|
chr14_+_62740878
|
1.556
|
NM_020663
|
RHOJ
|
ras homolog gene family, member J
|
|
chrX_-_50403412
|
1.551
|
|
SHROOM4
|
shroom family member 4
|
|
chr1_-_203867402
|
1.519
|
|
ELK4
|
ELK4, ETS-domain protein (SRF accessory protein 1)
|
|
chr3_-_50335113
|
1.508
|
NM_003773
NM_033158
|
HYAL2
|
hyaluronoglucosaminidase 2
|
|
chr22_-_34566276
|
1.501
|
|
RBFOX2
|
RNA binding protein, fox-1 homolog (C. elegans) 2
|
|
chr8_-_6407882
|
1.486
|
NM_001118887
NM_001118888
NM_001147
|
ANGPT2
|
angiopoietin 2
|
|
chr17_+_29606408
|
1.484
|
NM_002982
|
CCL2
|
chemokine (C-C motif) ligand 2
|
|
chr4_+_91035025
|
1.466
|
NM_007351
|
MMRN1
|
multimerin 1
|
|
chr3_+_151747204
|
1.465
|
NM_032025
|
EIF2A
|
eukaryotic translation initiation factor 2A, 65kDa
|
|
chr1_+_167604056
|
1.453
|
|
BLZF1
|
basic leucine zipper nuclear factor 1
|
|
chr14_-_56805118
|
1.437
|
|
|
|
|
chr8_-_38152985
|
1.408
|
NM_014462
|
LSM1
|
LSM1 homolog, U6 small nuclear RNA associated (S. cerevisiae)
|
|
chrX_-_131179598
|
1.392
|
NM_021183
|
RAP2C
|
RAP2C, member of RAS oncogene family
|
|
chr2_-_189752575
|
1.381
|
NM_000393
|
COL5A2
|
collagen, type V, alpha 2
|
|
chr17_+_24013320
|
1.378
|
|
SUPT6H
|
suppressor of Ty 6 homolog (S. cerevisiae)
|
|
chrX_+_15428820
|
1.372
|
NM_203281
|
BMX
|
BMX non-receptor tyrosine kinase
|
|
chr5_-_102483680
|
1.367
|
NM_017676
|
GIN1
|
gypsy retrotransposon integrase 1
|
|
chr5_-_82409009
|
1.361
|
NM_174909
|
TMEM167A
|
transmembrane protein 167A
|
|
chr10_-_12278026
|
1.359
|
NM_014142
|
NUDT5
|
nudix (nucleoside diphosphate linked moiety X)-type motif 5
|
|
chr10_+_12278185
|
1.357
|
|
CDC123
|
cell division cycle 123 homolog (S. cerevisiae)
|
|
chr11_-_60953848
|
1.357
|
NM_024811
|
CPSF7
|
cleavage and polyadenylation specific factor 7, 59kDa
|
|
chr8_-_103735194
|
1.350
|
NM_001032282
|
KLF10
|
Kruppel-like factor 10
|
|
chrX_-_10504852
|
1.333
|
NM_001193277
|
MID1
|
midline 1 (Opitz/BBB syndrome)
|
|
chr5_-_124108672
|
1.329
|
NM_020747
|
ZNF608
|
zinc finger protein 608
|
|
chr6_-_153365532
|
1.329
|
|
MTRF1L
|
mitochondrial translational release factor 1-like
|
|
chr22_-_34566367
|
1.328
|
NM_001031695
NM_001082576
NM_001082577
NM_014309
|
RBFOX2
|
RNA binding protein, fox-1 homolog (C. elegans) 2
|
|
chr1_+_118273869
|
1.317
|
|
WDR3
|
WD repeat domain 3
|
|
chr13_+_30089829
|
1.297
|
NM_005800
|
USPL1
|
ubiquitin specific peptidase like 1
|
|
chr1_+_39229474
|
1.294
|
NM_001136275
NM_024595
|
AKIRIN1
|
akirin 1
|
|
chr10_+_12278213
|
1.293
|
|
CDC123
|
cell division cycle 123 homolog (S. cerevisiae)
|
|
chr6_-_153365542
|
1.288
|
|
MTRF1L
|
mitochondrial translational release factor 1-like
|
|
chr18_-_51219880
|
1.287
|
|
TCF4
|
transcription factor 4
|
|
chr17_+_24013359
|
1.287
|
NM_003170
|
SUPT6H
|
suppressor of Ty 6 homolog (S. cerevisiae)
|
|
chr1_-_118273701
|
1.274
|
|
GDAP2
|
ganglioside induced differentiation associated protein 2
|
|
chr17_-_43288137
|
1.259
|
NM_199262
|
SP6
|
Sp6 transcription factor
|
|
chr9_-_35744271
|
1.255
|
NM_001044264
|
MSMP
|
microseminoprotein, prostate associated
|
|
chr9_-_88159076
|
1.252
|
|
ZCCHC6
|
zinc finger, CCHC domain containing 6
|
|
chr5_-_82408917
|
1.251
|
|
TMEM167A
|
transmembrane protein 167A
|
|
chr1_+_220953278
|
1.251
|
|
BROX
|
BRO1 domain and CAAX motif containing
|
|
chr1_+_220953351
|
1.251
|
|
BROX
|
BRO1 domain and CAAX motif containing
|
|
chr2_-_128138435
|
1.250
|
NM_017980
|
LIMS2
|
LIM and senescent cell antigen-like domains 2
|
|
chr7_+_120416680
|
1.250
|
NM_001105533
|
C7orf58
|
chromosome 7 open reading frame 58
|
|
chr12_-_368517
|
1.242
|
|
KDM5A
|
lysine (K)-specific demethylase 5A
|
|
chr10_-_112668892
|
1.241
|
NM_001195304
NM_001195305
NM_001195307
|
BBIP1
|
BBSome interacting protein 1
|
|
chr12_-_52939584
|
1.237
|
NM_001127322
|
CBX5
|
chromobox homolog 5
|
|
chr1_+_118273887
|
1.232
|
NM_006784
|
WDR3
|
WD repeat domain 3
|
|
chr8_-_38153325
|
1.229
|
|
LSM1
|
LSM1 homolog, U6 small nuclear RNA associated (S. cerevisiae)
|
|
chr1_-_40496161
|
1.221
|
|
|
|
|
chr1_+_143807876
|
1.215
|
|
SEC22B
|
SEC22 vesicle trafficking protein homolog B (S. cerevisiae) (gene/pseudogene)
|
|
chr17_-_24013303
|
1.211
|
|
SDF2
|
stromal cell-derived factor 2
|
|
chr10_+_12278200
|
1.209
|
|
CDC123
|
cell division cycle 123 homolog (S. cerevisiae)
|
|
chr11_-_85457469
|
1.209
|
|
PICALM
|
phosphatidylinositol binding clathrin assembly protein
|
|
chr6_-_153365564
|
1.198
|
NM_001114184
NM_019041
|
MTRF1L
|
mitochondrial translational release factor 1-like
|
|
chr5_+_34951538
|
1.198
|
NM_018321
|
BRIX1
|
BRX1, biogenesis of ribosomes, homolog (S. cerevisiae)
|
|
chr6_-_153365480
|
1.191
|
|
MTRF1L
|
mitochondrial translational release factor 1-like
|
|
chr5_+_82409072
|
1.181
|
NM_003401
NM_022406
NM_022550
|
XRCC4
|
X-ray repair complementing defective repair in Chinese hamster cells 4
|
|
chr4_-_48477038
|
1.168
|
NM_015030
|
FRYL
|
FRY-like
|
|
chr8_+_42515873
|
1.162
|
NM_138436
NM_001135674
NM_001135675
|
C8orf40
|
chromosome 8 open reading frame 40
|
|
chr4_-_48477021
|
1.160
|
|
FRYL
|
FRY-like
|
|
chr11_+_60954096
|
1.148
|
|
SDHAF2
|
succinate dehydrogenase complex assembly factor 2
|
|
chr14_-_74249492
|
1.147
|
NM_001039479
|
KIAA0317
|
KIAA0317
|
|
chr5_-_34951346
|
1.138
|
|
RAD1
|
RAD1 homolog (S. pombe)
|
|
chr5_-_102926349
|
1.127
|
NM_031438
|
NUDT12
|
nudix (nucleoside diphosphate linked moiety X)-type motif 12
|
|
chr11_-_60954031
|
1.122
|
NM_001136040
NM_001142565
|
CPSF7
|
cleavage and polyadenylation specific factor 7, 59kDa
|
|
chr14_+_77244177
|
1.110
|
NM_031210
|
C14orf156
|
chromosome 14 open reading frame 156
|
|
chr6_-_47030492
|
1.108
|
NM_015234
|
GPR116
|
G protein-coupled receptor 116
|
|
chr21_+_44256630
|
1.105
|
NM_003274
|
TRAPPC10
|
trafficking protein particle complex 10
|
|
chrX_-_153908408
|
1.104
|
|
F8
|
coagulation factor VIII, procoagulant component
|
|
chr19_-_4510713
|
1.098
|
|
SEMA6B
|
sema domain, transmembrane domain (TM), and cytoplasmic domain, (semaphorin) 6B
|
|
chr3_+_9809776
|
1.097
|
NM_001198780
|
ARPC4
|
actin related protein 2/3 complex, subunit 4, 20kDa
|
|
chr3_-_151746821
|
1.096
|
|
SERP1
|
stress-associated endoplasmic reticulum protein 1
|
|
chr17_-_24013259
|
1.095
|
|
SDF2
|
stromal cell-derived factor 2
|
|
chr3_+_185515543
|
1.093
|
NM_182917
|
EIF4G1
|
eukaryotic translation initiation factor 4 gamma, 1
|
|
chr10_-_51293220
|
1.077
|
NM_006327
|
TIMM23
TIMM23B
|
translocase of inner mitochondrial membrane 23 homolog (yeast)
translocase of inner mitochondrial membrane 23 homolog B (yeast)
|
|
chr12_+_8958397
|
1.075
|
NM_004426
|
PHC1
|
polyhomeotic homolog 1 (Drosophila)
|
|
chr10_+_114196698
|
1.073
|
NM_145206
|
VTI1A
|
vesicle transport through interaction with t-SNAREs homolog 1A (yeast)
|
|
chr1_+_202647097
|
1.071
|
|
|
|
|
chrX_-_79951838
|
1.069
|
NM_153252
|
BRWD3
|
bromodomain and WD repeat domain containing 3
|
|
chr11_-_72111027
|
1.064
|
NM_001135190
NM_015242
|
ARAP1
|
ArfGAP with RhoGAP domain, ankyrin repeat and PH domain 1
|
|
chr5_-_34951470
|
1.064
|
NM_002853
|
RAD1
|
RAD1 homolog (S. pombe)
|
|
chr1_+_167603817
|
1.064
|
NM_003666
|
BLZF1
|
basic leucine zipper nuclear factor 1
|
|
chr4_+_57028536
|
1.063
|
|
SRP72
|
signal recognition particle 72kDa
|
|
chr10_-_116434102
|
1.062
|
|
ABLIM1
|
actin binding LIM protein 1
|
|
chrX_+_114734074
|
1.061
|
NM_001136025
NM_001172335
|
PLS3
|
plastin 3
|
|
chr3_+_44665220
|
1.060
|
NM_003420
|
ZNF35
|
zinc finger protein 35
|
|
chr3_-_71262405
|
1.058
|
|
FOXP1
|
forkhead box P1
|
|
chr6_+_147567200
|
1.056
|
NM_001127715
NM_139244
|
STXBP5
|
syntaxin binding protein 5 (tomosyn)
|
|
chr5_+_102483856
|
1.049
|
|
PPIP5K2
|
diphosphoinositol pentakisphosphate kinase 2
|
|
chrX_+_153908426
|
1.047
|
|
FUNDC2
|
FUN14 domain containing 2
|
|
chr3_-_32519240
|
1.045
|
|
CMTM6
|
CKLF-like MARVEL transmembrane domain containing 6
|
|
chr3_-_71262426
|
1.039
|
|
FOXP1
|
forkhead box P1
|
|
chr10_-_27483144
|
1.039
|
|
YME1L1
|
YME1-like 1 (S. cerevisiae)
|
|
chr3_-_130362563
|
1.038
|
NM_020701
|
ISY1
|
ISY1 splicing factor homolog (S. cerevisiae)
|
|
chr3_-_193928038
|
1.034
|
NM_004113
|
FGF12
|
fibroblast growth factor 12
|
|
chr12_-_6103945
|
1.030
|
NM_000552
|
VWF
|
von Willebrand factor
|
|
chr15_+_21361901
|
1.029
|
|
MKRN3
|
makorin ring finger protein 3
|
|
chr1_+_143807748
|
1.026
|
NM_004892
|
SEC22B
|
SEC22 vesicle trafficking protein homolog B (S. cerevisiae) (gene/pseudogene)
|
|
chr1_+_84382562
|
1.025
|
|
PRKACB
|
protein kinase, cAMP-dependent, catalytic, beta
|
|
chr14_+_74249599
|
1.025
|
NM_015962
|
FCF1
|
FCF1 small subunit (SSU) processome component homolog (S. cerevisiae)
|
|
chr7_+_106293112
|
1.020
|
NM_002649
|
PIK3CG
|
phosphoinositide-3-kinase, catalytic, gamma polypeptide
|
|
chr12_+_99185334
|
1.019
|
NM_017988
|
SCYL2
|
SCY1-like 2 (S. cerevisiae)
|
|
chr1_-_159369041
|
1.011
|
|
DEDD
|
death effector domain containing
|
|
chr2_-_144994349
|
1.008
|
NM_001171653
NM_014795
|
ZEB2
|
zinc finger E-box binding homeobox 2
|
|
chr9_+_76893232
|
1.005
|
|
OSTF1
|
osteoclast stimulating factor 1
|
|
chr10_+_12277935
|
1.004
|
NM_006023
|
CDC123
|
cell division cycle 123 homolog (S. cerevisiae)
|
|
chr7_+_149842850
|
1.003
|
NM_153236
|
GIMAP7
|
GTPase, IMAP family member 7
|
|
chr7_+_44207131
|
1.000
|
|
YKT6
|
YKT6 v-SNARE homolog (S. cerevisiae)
|
|
chr12_-_47774837
|
0.999
|
NM_021044
|
DHH
|
desert hedgehog
|
|
chr1_+_84382575
|
0.997
|
|
PRKACB
|
protein kinase, cAMP-dependent, catalytic, beta
|
|
chr12_+_11694172
|
0.996
|
|
ETV6
|
ets variant 6
|
|
chr14_+_77244191
|
0.992
|
|
C14orf156
|
chromosome 14 open reading frame 156
|
|
chr15_-_40352765
|
0.986
|
NM_001110503
NM_015497
|
TMEM87A
|
transmembrane protein 87A
|
|
chr3_-_120878910
|
0.984
|
NM_005694
|
COX17
|
COX17 cytochrome c oxidase assembly homolog (S. cerevisiae)
|
|
chr10_+_114196977
|
0.980
|
|
VTI1A
|
vesicle transport through interaction with t-SNAREs homolog 1A (yeast)
|
|
chr8_+_125620523
|
0.979
|
NM_005005
|
NDUFB9
|
NADH dehydrogenase (ubiquinone) 1 beta subcomplex, 9, 22kDa
|
|
chr12_+_120840829
|
0.974
|
NM_001178003
NM_144668
|
WDR66
|
WD repeat domain 66
|
|
chr1_+_84382529
|
0.972
|
NM_182948
|
PRKACB
|
protein kinase, cAMP-dependent, catalytic, beta
|
|
chr5_-_60493985
|
0.969
|
NM_001048249
|
C5orf43
|
chromosome 5 open reading frame 43
|
|
chr8_-_42515774
|
0.969
|
|
SLC20A2
|
solute carrier family 20 (phosphate transporter), member 2
|
|
chr8_-_38153369
|
0.968
|
|
LSM1
|
LSM1 homolog, U6 small nuclear RNA associated (S. cerevisiae)
|
|
chr1_-_159369081
|
0.968
|
NM_001039711
NM_032998
|
DEDD
|
death effector domain containing
|
|
chr7_-_91713044
|
0.967
|
NM_004912
NM_001013406
NM_194454
|
KRIT1
|
KRIT1, ankyrin repeat containing
|
|
chr5_+_78944151
|
0.963
|
|
PAPD4
|
PAP associated domain containing 4
|
|
chr6_+_43592812
|
0.959
|
|
POLR1C
|
polymerase (RNA) I polypeptide C, 30kDa
|
|
chr10_-_27483285
|
0.959
|
|
YME1L1
|
YME1-like 1 (S. cerevisiae)
|
|
chr7_+_44207094
|
0.954
|
NM_006555
|
YKT6
|
YKT6 v-SNARE homolog (S. cerevisiae)
|
|
chr16_+_66131247
|
0.953
|
|
FAM65A
|
family with sequence similarity 65, member A
|
|
chr5_+_87600485
|
0.953
|
|
LOC100505894
|
hypothetical LOC100505894
|
|
chr4_+_2783743
|
0.951
|
NM_001145855
|
SH3BP2
|
SH3-domain binding protein 2
|
|
chr1_+_9926072
|
0.948
|
NM_022787
|
NMNAT1
|
nicotinamide nucleotide adenylyltransferase 1
|
|
chr3_-_113763049
|
0.942
|
NM_022488
|
ATG3
|
ATG3 autophagy related 3 homolog (S. cerevisiae)
|
|
chr3_-_151746924
|
0.939
|
|
SERP1
|
stress-associated endoplasmic reticulum protein 1
|
|
chr6_-_76050837
|
0.937
|
|
|
|
|
chr3_-_32519323
|
0.937
|
|
CMTM6
|
CKLF-like MARVEL transmembrane domain containing 6
|
|
chr12_-_3732614
|
0.935
|
NM_001144958
NM_001144959
NM_032680
|
EFCAB4B
|
EF-hand calcium binding domain 4B
|
|
chr1_+_168030798
|
0.934
|
|
C1orf112
|
chromosome 1 open reading frame 112
|
|
chr9_+_123003549
|
0.934
|
|
GSN
|
gelsolin
|
|
chr7_-_27158726
|
0.933
|
|
|
|
|
chr8_+_56848552
|
0.931
|
|
TGS1
|
trimethylguanosine synthase 1
|
|
chr10_-_27483315
|
0.929
|
NM_014263
NM_139312
|
YME1L1
|
YME1-like 1 (S. cerevisiae)
|
|
chr1_+_40279036
|
0.928
|
|
CAP1
|
CAP, adenylate cyclase-associated protein 1 (yeast)
|
|
chr14_-_77244073
|
0.923
|
NM_006020
|
ALKBH1
|
alkB, alkylation repair homolog 1 (E. coli)
|
|
chr13_-_51878235
|
0.922
|
|
THSD1
|
thrombospondin, type I, domain containing 1
|
|
chr11_+_10786137
|
0.918
|
|
LOC144017
|
hypothetical protein LOC144017
|
|
chr7_+_134114701
|
0.917
|
NM_004342
NM_033138
NM_033157
|
CALD1
|
caldesmon 1
|
|
chr9_+_99435900
|
0.912
|
|
NCBP1
|
nuclear cap binding protein subunit 1, 80kDa
|
|
chr3_+_29297806
|
0.912
|
NM_001003792
NM_001003793
NM_001177711
NM_001177712
NM_014483
|
RBMS3
|
RNA binding motif, single stranded interacting protein 3
|
|
chr5_-_87600371
|
0.905
|
NM_153354
|
TMEM161B
|
transmembrane protein 161B
|
|
chr18_+_64616296
|
0.904
|
NM_024781
|
CCDC102B
|
coiled-coil domain containing 102B
|
|
chr19_-_20011013
|
0.903
|
NM_001077349
|
ZNF682
|
zinc finger protein 682
|
|
chrX_+_119622570
|
0.901
|
NM_001137554
|
MCTS1
|
malignant T cell amplified sequence 1
|
|
chr19_-_55008172
|
0.901
|
NM_001171937
NM_025129
|
FUZ
|
fuzzy homolog (Drosophila)
|
|
chr1_-_112963247
|
0.896
|
NM_017744
NM_138727
NM_138728
NM_138729
|
ST7L
|
suppression of tumorigenicity 7 like
|
|
chr5_-_60493903
|
0.895
|
|
C5orf43
|
chromosome 5 open reading frame 43
|
|
chr7_+_44207080
|
0.892
|
|
YKT6
|
YKT6 v-SNARE homolog (S. cerevisiae)
|
|
chr6_+_151815298
|
0.890
|
|
C6orf211
|
chromosome 6 open reading frame 211
|
|
chr1_+_9926113
|
0.889
|
|
NMNAT1
|
nicotinamide nucleotide adenylyltransferase 1
|
|
chr8_-_68136881
|
0.886
|
|
COPS5
|
COP9 constitutive photomorphogenic homolog subunit 5 (Arabidopsis)
|
|
chr15_+_46271158
|
0.885
|
NM_001145668
|
CTXN2
SLC12A1
|
cortexin 2
solute carrier family 12 (sodium/potassium/chloride transporters), member 1
|
|
chr7_+_106292975
|
0.884
|
|
PIK3CG
|
phosphoinositide-3-kinase, catalytic, gamma polypeptide
|
|
chr15_+_75500284
|
0.884
|
NM_018200
|
HMG20A
|
high-mobility group 20A
|
|
chr9_-_26937165
|
0.884
|
|
PLAA
|
phospholipase A2-activating protein
|
|
chr8_-_68136700
|
0.884
|
|
COPS5
|
COP9 constitutive photomorphogenic homolog subunit 5 (Arabidopsis)
|
|
chrX_+_102518764
|
0.884
|
NM_014380
|
NGFRAP1
|
nerve growth factor receptor (TNFRSF16) associated protein 1
|
|
chr8_+_56848733
|
0.882
|
|
|
|
|
chr15_+_40353076
|
0.875
|
|
GANC
|
glucosidase, alpha; neutral C
|
|
chr12_-_70343656
|
0.875
|
|
|
|
|
chr1_-_158579583
|
0.873
|
|
|
|
|
chr1_-_118273728
|
0.866
|
NM_001135589
NM_017686
|
GDAP2
|
ganglioside induced differentiation associated protein 2
|
|
chr8_-_95634845
|
0.866
|
NM_015496
NM_183009
|
KIAA1429
|
KIAA1429
|
|
chr14_+_73423260
|
0.864
|
NM_021188
|
ZNF410
|
zinc finger protein 410
|
|
chr7_+_44207163
|
0.860
|
|
YKT6
|
YKT6 v-SNARE homolog (S. cerevisiae)
|
|
chr3_-_151746970
|
0.860
|
NM_014445
|
SERP1
|
stress-associated endoplasmic reticulum protein 1
|
|
chr10_-_69504956
|
0.858
|
|
HERC4
|
hect domain and RLD 4
|
|
chr12_+_99185289
|
0.857
|
|
SCYL2
|
SCY1-like 2 (S. cerevisiae)
|