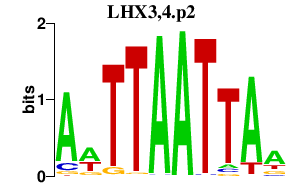
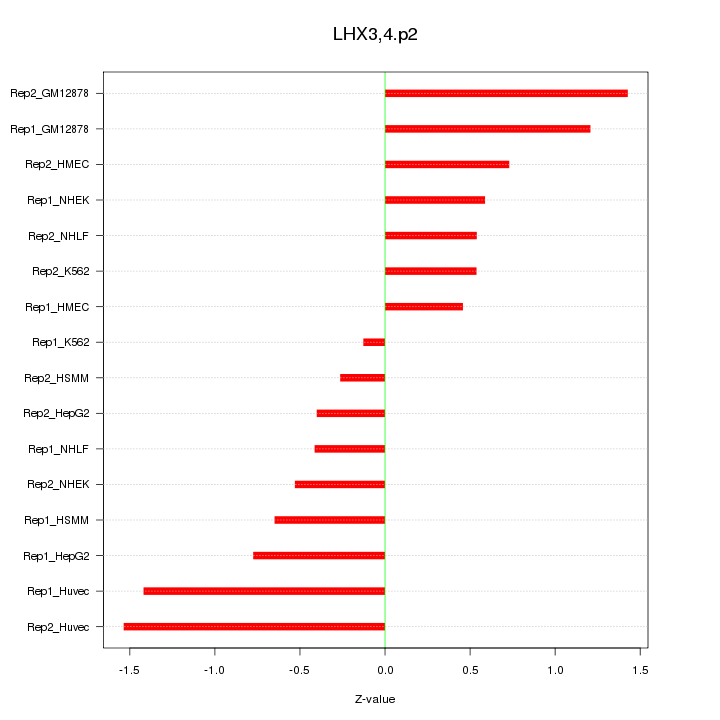
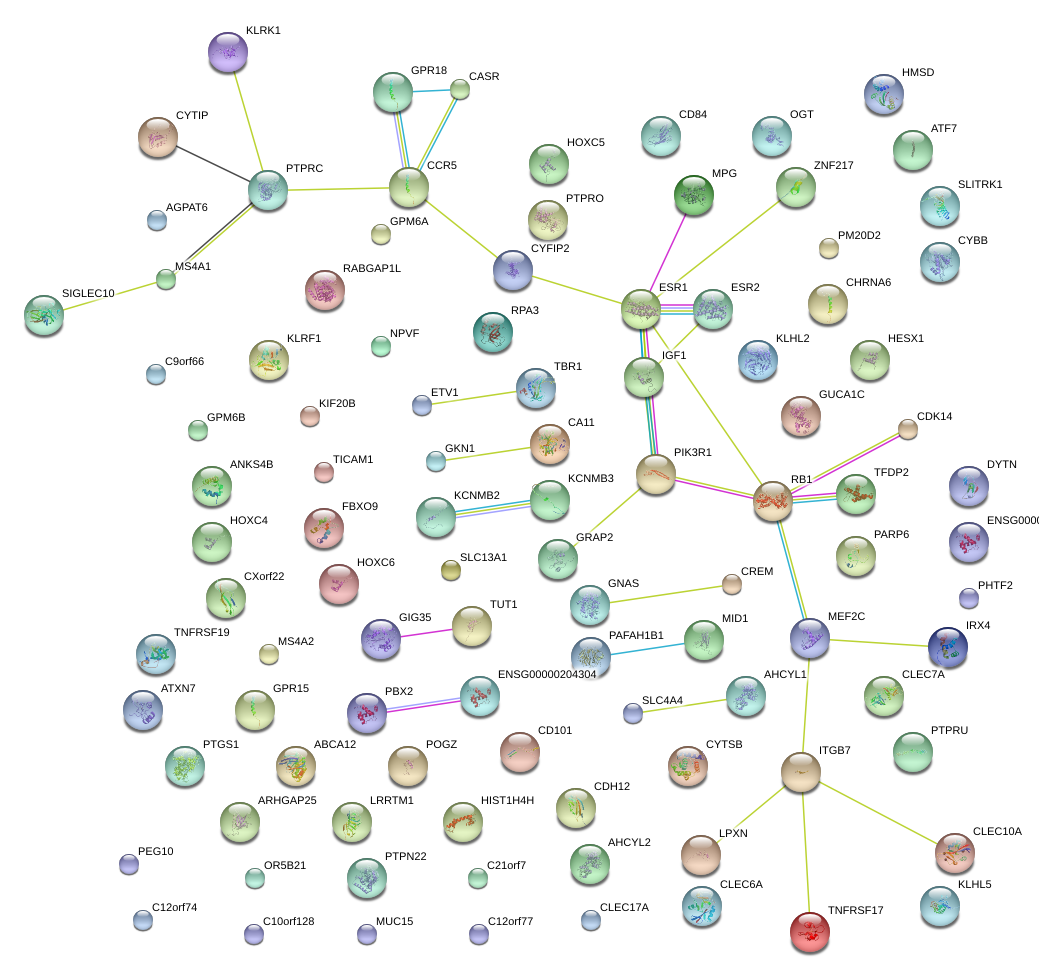
|
chr5_-_88155360
|
3.014
|
NM_001193348
NM_001193349
|
MEF2C
|
myocyte enhancer factor 2C
|
|
chr19_-_56612653
|
2.861
|
NM_001171156
NM_001171157
NM_001171158
NM_001171159
NM_001171160
NM_001171161
NM_033130
|
SIGLEC10
|
sialic acid binding Ig-like lectin 10
|
|
chr17_+_19917420
|
1.938
|
|
SPECC1
|
sperm antigen with calponin homology and coiled-coil domains 1
|
|
chr1_+_117345894
|
1.646
|
NM_004258
|
CD101
|
CD101 molecule
|
|
chr12_+_52696908
|
1.477
|
NM_014620
NM_153693
|
HOXC4
HOXC6
|
homeobox C4
homeobox C6
|
|
chr12_+_52697160
|
1.399
|
|
HOXC5
HOXC6
|
homeobox C5
homeobox C6
|
|
chr1_+_196874839
|
1.273
|
|
PTPRC
|
protein tyrosine phosphatase, receptor type, C
|
|
chr3_-_57209319
|
1.231
|
NM_003865
|
HESX1
|
HESX homeobox 1
|
|
chr12_-_101396459
|
1.210
|
NM_001111284
|
IGF1
|
insulin-like growth factor 1 (somatomedin C)
|
|
chr5_+_67622242
|
1.142
|
NM_181504
|
PIK3R1
|
phosphoinositide-3-kinase, regulatory subunit 1 (alpha)
|
|
chr13_-_83354425
|
1.054
|
|
SLITRK1
|
SLIT and NTRK-like family, member 1
|
|
chr5_+_156628929
|
1.048
|
NM_014376
|
CYFIP2
|
cytoplasmic FMR1 interacting protein 2
|
|
chr16_+_11966464
|
1.027
|
NM_001192
|
TNFRSF17
|
tumor necrosis factor receptor superfamily, member 17
|
|
chr10_-_50066386
|
1.008
|
NM_001010863
|
C10orf128
|
chromosome 10 open reading frame 128
|
|
chr7_+_90177082
|
0.976
|
|
CDK14
|
cyclin-dependent kinase 14
|
|
chr7_+_37689903
|
0.969
|
|
|
|
|
chr22_+_38627031
|
0.959
|
NM_004810
|
GRAP2
|
GRB2-related adaptor protein 2
|
|
chr10_+_35466715
|
0.957
|
NM_183011
NM_183012
|
CREM
|
cAMP responsive element modulator
|
|
chr12_+_52708460
|
0.944
|
NM_004503
|
HOXC6
|
homeobox C6
|
|
chr3_+_63928459
|
0.939
|
NM_001128149
|
ATXN7
|
ataxin 7
|
|
chr12_-_10173947
|
0.938
|
NM_022570
NM_197947
NM_197948
NM_197949
NM_197950
NM_197954
|
CLEC7A
|
C-type lectin domain family 7, member A
|
|
chr22_+_38627129
|
0.931
|
|
GRAP2
|
GRB2-related adaptor protein 2
|
|
chr10_+_91488704
|
0.926
|
|
KIF20B
|
kinesin family member 20B
|
|
chr1_-_114215768
|
0.921
|
NM_001193431
NM_012411
NM_015967
|
PTPN22
|
protein tyrosine phosphatase, non-receptor type 22 (lymphoid)
|
|
chr9_-_205892
|
0.906
|
NM_152569
|
C9orf66
|
chromosome 9 open reading frame 66
|
|
chr20_-_51633042
|
0.905
|
NM_006526
|
ZNF217
|
zinc finger protein 217
|
|
chr13_-_98708549
|
0.898
|
NM_001098200
NM_005292
|
GPR18
|
G protein-coupled receptor 18
|
|
chr7_+_128802719
|
0.886
|
NM_001130722
|
AHCYL2
|
adenosylhomocysteinase-like 2
|
|
chr8_-_42742729
|
0.878
|
NM_004198
|
CHRNA6
|
cholinergic receptor, nicotinic, alpha 6
|
|
chr2_-_215711313
|
0.860
|
NM_173076
|
ABCA12
|
ATP-binding cassette, sub-family A (ABC1), member 12
|
|
chr6_+_152053323
|
0.833
|
NM_001122742
|
ESR1
|
estrogen receptor 1
|
|
chr2_+_161981138
|
0.804
|
|
TBR1
|
T-box, brain, 1
|
|
chr19_-_53841064
|
0.796
|
NM_001217
|
CA11
|
carbonic anhydrase XI
|
|
chr3_-_143230124
|
0.775
|
NM_001178141
NM_001178142
NM_006286
|
TFDP2
|
transcription factor Dp-2 (E2F dimerization partner 2)
|
|
chr5_-_22889192
|
0.747
|
|
CDH12
|
cadherin 12, type 2 (N-cadherin 2)
|
|
chrX_+_70682204
|
0.742
|
|
OGT
|
O-linked N-acetylglucosamine (GlcNAc) transferase (UDP-N-acetylglucosamine:polypeptide-N-acetylglucosaminyl transferase)
|
|
chr3_+_46386636
|
0.738
|
NM_000579
NM_001100168
|
CCR5
|
chemokine (C-C motif) receptor 5
|
|
chr3_+_123385219
|
0.736
|
NM_000388
|
CASR
|
calcium-sensing receptor
|
|
chr1_-_158815895
|
0.707
|
NM_001184879
NM_001184881
NM_001184882
NM_003874
|
CD84
|
CD84 molecule
|
|
chr18_+_59767567
|
0.702
|
NM_001123366
|
HMSD
|
histocompatibility (minor) serpin domain containing
|
|
chr19_+_14554895
|
0.702
|
NM_207390
|
CLEC17A
|
C-type lectin domain family 17, member A
|
|
chr2_+_161981046
|
0.683
|
|
TBR1
|
T-box, brain, 1
|
|
chr3_+_179759181
|
0.681
|
NM_005832
|
KCNMB2
|
potassium large conductance calcium-activated channel, subfamily M, beta member 2
|
|
chr20_+_56919233
|
0.678
|
|
GNAS
|
GNAS complex locus
|
|
chr7_-_13992351
|
0.678
|
NM_001163150
NM_001163151
NM_001163152
|
ETV1
|
ets variant 1
|
|
chr13_+_23042722
|
0.677
|
NM_148957
|
TNFRSF19
|
tumor necrosis factor receptor superfamily, member 19
|
|
chr4_+_166350620
|
0.676
|
NM_001161521
NM_001161522
|
KLHL2
|
kelch-like 2, Mayven (Drosophila)
|
|
chrX_-_10761729
|
0.666
|
NM_033290
|
MID1
|
midline 1 (Opitz/BBB syndrome)
|
|
chrX_-_13744933
|
0.644
|
|
GPM6B
|
glycoprotein M6B
|
|
chr11_+_59979857
|
0.640
|
NM_021950
NM_152866
|
MS4A1
|
membrane-spanning 4-domains, subfamily A, member 1
|
|
chr7_-_25234582
|
0.624
|
NM_022150
|
NPVF
|
neuropeptide VF precursor
|
|
chr12_-_51887266
|
0.605
|
NM_000889
|
ITGB7
|
integrin, beta 7
|
|
chr12_+_9871343
|
0.603
|
NM_016523
|
KLRF1
|
killer cell lectin-like receptor subfamily F, member 1
|
|
chr13_+_47948586
|
0.599
|
|
RB1
|
retinoblastoma 1
|
|
chr2_-_158008763
|
0.598
|
NM_004288
|
CYTIP
|
cytohesin 1 interacting protein
|
|
chr12_+_15590811
|
0.586
|
|
PTPRO
|
protein tyrosine phosphatase, receptor type, O
|
|
chr3_+_99733432
|
0.578
|
NM_005290
|
GPR15
|
G protein-coupled receptor 15
|
|
chr1_+_173111306
|
0.571
|
NM_001035230
|
RABGAP1L
|
RAB GTPase activating protein 1-like
|
|
chr9_+_124173143
|
0.569
|
|
PTGS1
|
prostaglandin-endoperoxide synthase 1 (prostaglandin G/H synthase and cyclooxygenase)
|
|
chr2_-_207291364
|
0.565
|
NM_001093730
|
DYTN
|
dystrotelin
|
|
chr1_+_196874867
|
0.563
|
|
PTPRC
|
protein tyrosine phosphatase, receptor type, C
|
|
chr4_-_176970372
|
0.558
|
|
GPM6A
|
glycoprotein M6A
|
|
chr9_+_124173049
|
0.554
|
NM_000962
NM_080591
|
PTGS1
|
prostaglandin-endoperoxide synthase 1 (prostaglandin G/H synthase and cyclooxygenase)
|
|
chr1_-_149681265
|
0.540
|
NM_207171
|
POGZ
|
pogo transposable element with ZNF domain
|
|
chr7_-_132371208
|
0.539
|
|
EEF1G
|
eukaryotic translation elongation factor 1 gamma
|
|
chr4_+_72423633
|
0.538
|
NM_003759
|
SLC4A4
|
solute carrier family 4, sodium bicarbonate cotransporter, member 4
|
|
chr12_-_51887318
|
0.535
|
|
ITGB7
|
integrin, beta 7
|
|
chr7_+_77307382
|
0.529
|
NM_001127357
NM_001127359
NM_020432
|
PHTF2
|
putative homeodomain transcription factor 2
|
|
chr2_+_68815416
|
0.529
|
NM_001007231
NM_001166277
|
ARHGAP25
|
Rho GTPase activating protein 25
|
|
chr12_-_52281050
|
0.518
|
NM_001130059
|
ATF7
|
activating transcription factor 7
|
|
chrX_+_37524190
|
0.514
|
NM_000397
|
CYBB
|
cytochrome b-245, beta polypeptide
|
|
chr7_+_90176647
|
0.511
|
NM_012395
|
CDK14
|
cyclin-dependent kinase 14
|
|
chr12_+_8499788
|
0.504
|
NM_001007033
|
CLEC6A
|
C-type lectin domain family 6, member A
|
|
chr11_-_26550243
|
0.504
|
NM_001135091
NM_001135092
NM_145650
|
MUC15
|
mucin 15, cell surface associated
|
|
chr6_-_32265489
|
0.502
|
NM_002586
|
PBX2
|
pre-B-cell leukemia homeobox 2
|
|
chr7_+_94135384
|
0.498
|
|
PEG10
|
paternally expressed 10
|
|
chr7_-_7724762
|
0.498
|
NM_002947
|
RPA3
|
replication protein A3, 14kDa
|
|
chr13_-_83354474
|
0.494
|
NM_052910
|
SLITRK1
|
SLIT and NTRK-like family, member 1
|
|
chr12_+_91620749
|
0.493
|
NM_001037671
NM_001178097
|
C12orf74
|
chromosome 12 open reading frame 74
|
|
chr3_-_110155237
|
0.470
|
NM_005459
|
GUCA1C
|
guanylate cyclase activator 1C
|
|
chr11_-_58032153
|
0.462
|
NM_001005218
|
OR5B21
|
olfactory receptor, family 5, subfamily B, member 21
|
|
chr21_+_29374743
|
0.454
|
NM_020152
|
C21orf7
|
chromosome 21 open reading frame 7
|
|
chr12_-_101398429
|
0.451
|
|
IGF1
|
insulin-like growth factor 1 (somatomedin C)
|
|
chr2_-_80384454
|
0.451
|
|
LRRTM1
|
leucine rich repeat transmembrane neuronal 1
|
|
chr6_+_53043729
|
0.445
|
NM_012347
|
FBXO9
|
F-box protein 9
|
|
chr7_-_122627235
|
0.441
|
NM_022444
|
SLC13A1
|
solute carrier family 13 (sodium/sulfate symporters), member 1
|
|
chr2_+_161980865
|
0.441
|
NM_006593
|
TBR1
|
T-box, brain, 1
|
|
chr1_+_196874759
|
0.440
|
NM_002838
NM_080921
NM_080923
|
PTPRC
|
protein tyrosine phosphatase, receptor type, C
|
|
chr5_-_1935764
|
0.439
|
NM_016358
|
IRX4
|
iroquois homeobox 4
|
|
chr12_-_25041639
|
0.434
|
NM_001101339
|
C12orf77
|
chromosome 12 open reading frame 77
|
|
chr17_-_1195096
|
0.433
|
|
PAFAH1B1
|
platelet-activating factor acetylhydrolase 1b, regulatory subunit 1 (45kDa)
|
|
chr15_-_70350609
|
0.432
|
NM_020214
|
PARP6
|
poly (ADP-ribose) polymerase family, member 6
|
|
chr4_-_176970895
|
0.431
|
|
GPM6A
|
glycoprotein M6A
|
|
chr4_+_38740246
|
0.430
|
NM_015990
NM_199039
|
KLHL5
|
kelch-like 5 (Drosophila)
|
|
chr9_+_124172644
|
0.427
|
|
PTGS1
|
prostaglandin-endoperoxide synthase 1 (prostaglandin G/H synthase and cyclooxygenase)
|
|
chrX_+_37524250
|
0.426
|
|
CYBB
|
cytochrome b-245, beta polypeptide
|
|
chr6_+_31538934
|
0.426
|
NM_006674
|
HCP5
|
HLA complex P5
|
|
chr11_-_58099884
|
0.421
|
NM_004811
|
LPXN
|
leupaxin
|
|
chr3_+_188222337
|
0.418
|
NM_003032
|
ST6GAL1
|
ST6 beta-galactosamide alpha-2,6-sialyltranferase 1
|
|
chr3_-_55489973
|
0.415
|
|
WNT5A
|
wingless-type MMTV integration site family, member 5A
|
|
chr3_-_168674502
|
0.409
|
NM_006217
|
SERPINI2
|
serpin peptidase inhibitor, clade I (pancpin), member 2
|
|
chr1_+_246682699
|
0.406
|
NM_001004136
|
OR2T2
|
olfactory receptor, family 2, subfamily T, member 2
|
|
chr9_+_124172606
|
0.396
|
|
PTGS1
|
prostaglandin-endoperoxide synthase 1 (prostaglandin G/H synthase and cyclooxygenase)
|
|
chr1_+_166516901
|
0.394
|
NM_005149
|
TBX19
|
T-box 19
|
|
chr16_-_66075216
|
0.393
|
NM_001138
|
AGRP
|
agouti related protein homolog (mouse)
|
|
chrX_-_55074125
|
0.389
|
NM_000032
NM_001037967
NM_001037968
|
ALAS2
|
aminolevulinate, delta-, synthase 2
|
|
chr7_-_86433139
|
0.386
|
NM_152748
|
KIAA1324L
|
KIAA1324-like
|
|
chr7_-_36730525
|
0.385
|
NM_001177506
NM_001177507
NM_001637
|
AOAH
|
acyloxyacyl hydrolase (neutrophil)
|
|
chr8_+_98772537
|
0.385
|
|
MTDH
|
metadherin
|
|
chr11_-_35235325
|
0.385
|
|
SLC1A2
|
solute carrier family 1 (glial high affinity glutamate transporter), member 2
|
|
chr12_-_101036407
|
0.385
|
NM_024057
|
NUP37
|
nucleoporin 37kDa
|
|
chr4_-_176970407
|
0.376
|
|
GPM6A
|
glycoprotein M6A
|
|
chr8_-_7207899
|
0.376
|
NM_001164457
|
ZNF705G
|
zinc finger protein 705G
|
|
chr9_-_116305273
|
0.373
|
NM_001083885
|
DFNB31
|
deafness, autosomal recessive 31
|
|
chr2_-_175170674
|
0.372
|
|
WIPF1
|
WAS/WASL interacting protein family, member 1
|
|
chr6_+_31661934
|
0.372
|
NM_205838
NM_205839
|
LST1
|
leukocyte specific transcript 1
|
|
chr17_-_38087366
|
0.371
|
NM_016602
|
CCR10
|
chemokine (C-C motif) receptor 10
|
|
chrX_+_99726445
|
0.366
|
NM_022144
|
TNMD
|
tenomodulin
|
|
chr3_-_11598829
|
0.364
|
NM_001128220
|
VGLL4
|
vestigial like 4 (Drosophila)
|
|
chr9_+_92603832
|
0.363
|
NM_001135052
NM_003177
|
SYK
|
spleen tyrosine kinase
|
|
chr5_+_60969392
|
0.361
|
NM_173667
|
FLJ37543
|
hypothetical protein FLJ37543
|
|
chr14_+_74968589
|
0.360
|
NM_001135049
|
JDP2
|
Jun dimerization protein 2
|
|
chr14_-_106249999
|
0.360
|
|
IGHA1
|
immunoglobulin heavy constant alpha 1
|
|
chr3_+_123779122
|
0.359
|
NM_001113523
|
PARP15
|
poly (ADP-ribose) polymerase family, member 15
|
|
chr5_+_140772691
|
0.358
|
NM_018913
NM_032090
|
PCDHGA10
|
protocadherin gamma subfamily A, 10
|
|
chr16_-_3567354
|
0.354
|
NM_178844
|
NLRC3
|
NLR family, CARD domain containing 3
|
|
chr8_+_32905027
|
0.351
|
|
|
|
|
chr6_-_76022565
|
0.348
|
|
TMEM30A
|
transmembrane protein 30A
|
|
chr3_+_161425946
|
0.342
|
NM_001168214
|
LOC401097
|
hypothetical protein LOC401097
|
|
chr16_+_2749934
|
0.341
|
|
|
|
|
chr4_-_185921384
|
0.338
|
|
ACSL1
|
acyl-CoA synthetase long-chain family member 1
|
|
chr17_-_9863929
|
0.337
|
|
GAS7
|
growth arrest-specific 7
|
|
chr12_-_16652202
|
0.337
|
NM_001001395
|
LMO3
|
LIM domain only 3 (rhombotin-like 2)
|
|
chr8_-_101384633
|
0.335
|
NM_015435
|
RNF19A
|
ring finger protein 19A
|
|
chr1_+_13903936
|
0.334
|
NM_012231
NM_015866
|
PRDM2
|
PR domain containing 2, with ZNF domain
|
|
chr21_-_30510113
|
0.333
|
NM_199328
|
CLDN8
|
claudin 8
|
|
chr18_+_27281729
|
0.332
|
NM_001944
|
DSG3
|
desmoglein 3
|
|
chr4_-_77163609
|
0.332
|
NM_001565
|
CXCL10
|
chemokine (C-X-C motif) ligand 10
|
|
chr1_+_157067729
|
0.330
|
NM_002432
|
MNDA
|
myeloid cell nuclear differentiation antigen
|
|
chr2_-_65013070
|
0.326
|
|
LOC400958
|
hypothetical LOC400958
|
|
chrX_+_115215985
|
0.323
|
NM_000686
|
AGTR2
|
angiotensin II receptor, type 2
|
|
chr7_+_28418668
|
0.322
|
NM_182898
|
CREB5
|
cAMP responsive element binding protein 5
|
|
chr1_-_152845399
|
0.321
|
NM_001193495
|
ADAR
|
adenosine deaminase, RNA-specific
|
|
chr8_-_72436519
|
0.320
|
|
EYA1
|
eyes absent homolog 1 (Drosophila)
|
|
chr6_+_167445284
|
0.318
|
NM_004367
|
CCR6
|
chemokine (C-C motif) receptor 6
|
|
chr3_-_159872960
|
0.317
|
NM_020169
|
LXN
|
latexin
|
|
chr1_+_242283129
|
0.314
|
NM_006352
|
ZNF238
|
zinc finger protein 238
|
|
chr8_-_82536312
|
0.312
|
NM_001080526
|
FABP9
|
fatty acid binding protein 9, testis
|
|
chr4_+_88790477
|
0.308
|
NM_001079911
NM_004407
|
DMP1
|
dentin matrix acidic phosphoprotein 1
|
|
chr7_+_129057152
|
0.307
|
NM_001040110
|
NRF1
|
nuclear respiratory factor 1
|
|
chr12_+_54672766
|
0.307
|
|
RAB5B
|
RAB5B, member RAS oncogene family
|
|
chr6_+_31691745
|
0.305
|
NM_032955
|
AIF1
|
allograft inflammatory factor 1
|
|
chr1_-_155336223
|
0.302
|
NM_001004341
|
ETV3L
|
ets variant 3-like
|
|
chr11_+_33306652
|
0.300
|
|
HIPK3
|
homeodomain interacting protein kinase 3
|
|
chr14_+_66901237
|
0.298
|
|
EIF2S1
|
eukaryotic translation initiation factor 2, subunit 1 alpha, 35kDa
|
|
chr6_+_158654890
|
0.298
|
|
|
|
|
chrX_+_107174855
|
0.297
|
NM_001170553
NM_182607
|
VSIG1
|
V-set and immunoglobulin domain containing 1
|
|
chr1_+_84540921
|
0.296
|
NM_001134664
|
SAMD13
|
sterile alpha motif domain containing 13
|
|
chr14_+_69307914
|
0.294
|
|
SRSF5
|
serine/arginine-rich splicing factor 5
|
|
chr18_+_70352779
|
0.293
|
|
CNDP1
|
carnosine dipeptidase 1 (metallopeptidase M20 family)
|
|
chr3_-_12561962
|
0.292
|
NM_001195279
|
LOC100129480
|
hypothetical protein LOC100129480
|
|
chr9_-_21375258
|
0.292
|
NM_000605
|
IFNA2
|
interferon, alpha 2
|
|
chr3_+_52212969
|
0.288
|
|
ALAS1
|
aminolevulinate, delta-, synthase 1
|
|
chr3_-_152403636
|
0.288
|
NM_013308
|
GPR171
|
G protein-coupled receptor 171
|
|
chr2_-_191580567
|
0.287
|
|
STAT1
|
signal transducer and activator of transcription 1, 91kDa
|
|
chr5_+_57823017
|
0.286
|
NM_152687
|
GAPT
|
GRB2-binding adaptor protein, transmembrane
|
|
chrX_+_153186602
|
0.285
|
NM_001145934
|
TKTL1
|
transketolase-like 1
|
|
chr15_+_97990705
|
0.281
|
NM_001130928
|
MEF2A
|
myocyte enhancer factor 2A
|
|
chr14_-_105610144
|
0.280
|
|
IGHA1
IGHG1
IGHG3
|
immunoglobulin heavy constant alpha 1
immunoglobulin heavy constant gamma 1 (G1m marker)
immunoglobulin heavy constant gamma 3 (G3m marker)
|
|
chr13_+_51933228
|
0.279
|
|
CKAP2
|
cytoskeleton associated protein 2
|
|
chr1_+_111215343
|
0.278
|
NM_001040033
|
CD53
|
CD53 molecule
|
|
chr1_-_151587645
|
0.278
|
NM_020393
|
PGLYRP4
|
peptidoglycan recognition protein 4
|
|
chr8_-_102872372
|
0.278
|
|
NCALD
|
neurocalcin delta
|
|
chr1_+_40635058
|
0.277
|
NM_001198980
|
SMAP2
|
small ArfGAP2
|
|
chr1_+_158636987
|
0.277
|
NM_020335
|
VANGL2
|
vang-like 2 (van gogh, Drosophila)
|
|
chr1_+_154120170
|
0.276
|
|
SYT11
|
synaptotagmin XI
|
|
chr3_-_166278969
|
0.276
|
NM_001041
|
SI
|
sucrase-isomaltase (alpha-glucosidase)
|
|
chr4_+_71806538
|
0.274
|
NM_001037442
NM_014961
|
RUFY3
|
RUN and FYVE domain containing 3
|
|
chr3_+_45902999
|
0.274
|
NM_006641
NM_031200
|
CCR9
|
chemokine (C-C motif) receptor 9
|
|
chr7_+_110518297
|
0.272
|
NM_001099658
NM_001099660
NM_018334
|
LRRN3
|
leucine rich repeat neuronal 3
|
|
chr12_-_101398503
|
0.271
|
NM_000618
NM_001111283
NM_001111285
|
IGF1
|
insulin-like growth factor 1 (somatomedin C)
|
|
chr6_-_56824371
|
0.269
|
|
DST
|
dystonin
|
|
chr11_-_58368555
|
0.268
|
NM_145016
|
GLYATL2
|
glycine-N-acyltransferase-like 2
|
|
chr19_-_60458915
|
0.264
|
|
PPP6R1
|
protein phosphatase 6, regulatory subunit 1
|
|
chr9_-_14076901
|
0.264
|
|
NFIB
|
nuclear factor I/B
|
|
chr4_-_54152385
|
0.264
|
|
LNX1
|
ligand of numb-protein X 1
|
|
chr7_+_18296569
|
0.264
|
|
HDAC9
|
histone deacetylase 9
|
|
chr6_-_29635680
|
0.262
|
NM_006398
|
UBD
|
ubiquitin D
|
|
chr11_-_102081654
|
0.262
|
NM_022122
|
MMP27
|
matrix metallopeptidase 27
|
|
chr12_-_101398288
|
0.261
|
|
IGF1
|
insulin-like growth factor 1 (somatomedin C)
|
|
chr2_+_161701711
|
0.258
|
NM_004180
NM_133484
|
TANK
|
TRAF family member-associated NFKB activator
|
|
chr1_+_156991133
|
0.257
|
NM_001005184
|
OR6K6
|
olfactory receptor, family 6, subfamily K, member 6
|
|
chr3_-_114047479
|
0.256
|
NM_001008784
|
CD200R1L
|
CD200 receptor 1-like
|
|
chr17_+_18027816
|
0.253
|
|
ALKBH5
|
alkB, alkylation repair homolog 5 (E. coli)
|
|
chrX_+_135558001
|
0.252
|
NM_000074
|
CD40LG
|
CD40 ligand
|
|
chr15_+_100175888
|
0.250
|
NM_001001674
|
OR4F15
|
olfactory receptor, family 4, subfamily F, member 15
|
|
chr13_+_48923688
|
0.249
|
NM_001160308
|
SETDB2
|
SET domain, bifurcated 2
|
|
chr12_-_56422120
|
0.249
|
NM_014770
|
AGAP2
|
ArfGAP with GTPase domain, ankyrin repeat and PH domain 2
|
|
chr5_-_22889487
|
0.249
|
NM_004061
|
CDH12
|
cadherin 12, type 2 (N-cadherin 2)
|
|
chr12_-_101398470
|
0.247
|
|
IGF1
|
insulin-like growth factor 1 (somatomedin C)
|