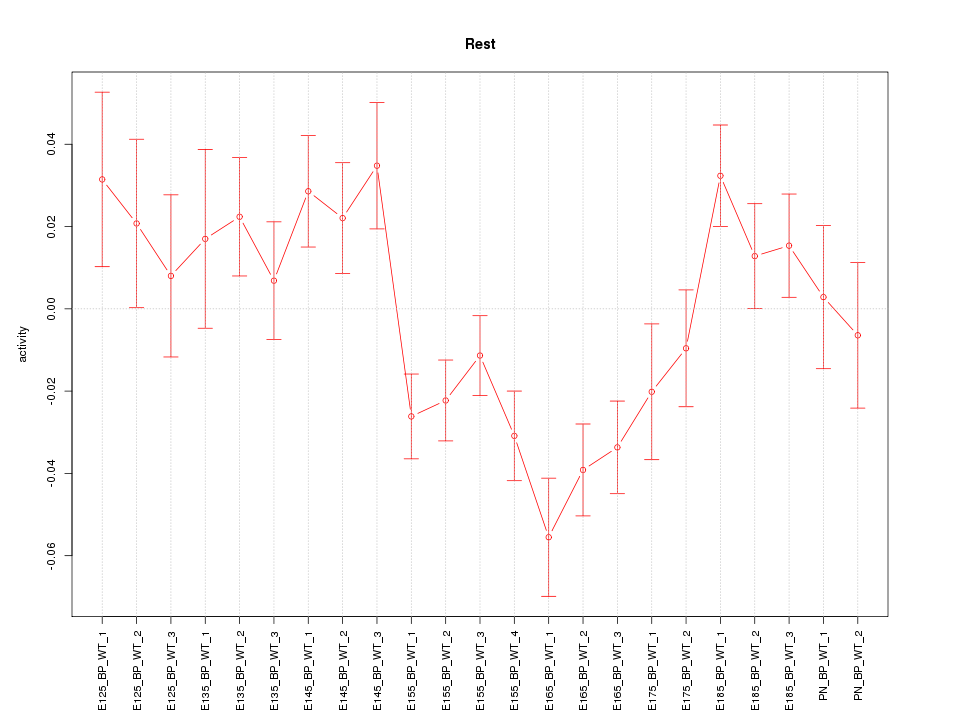
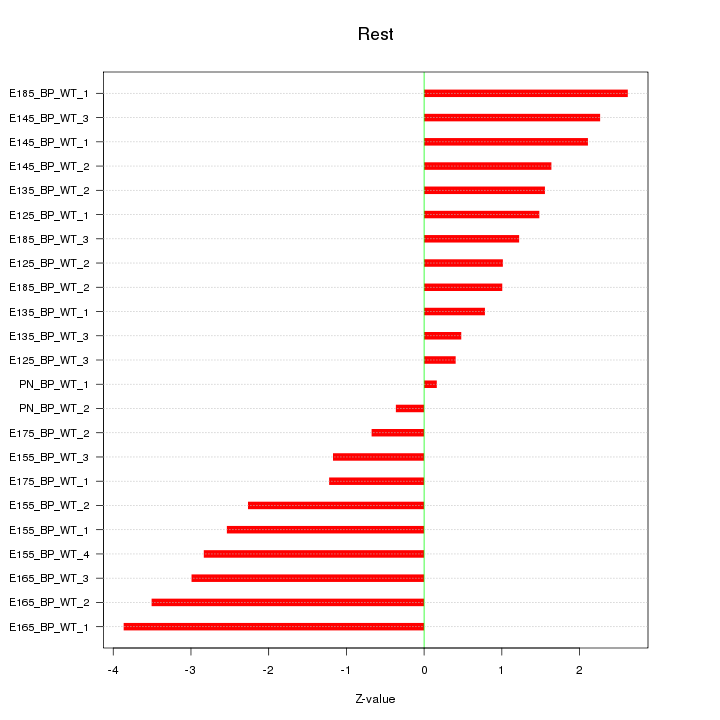
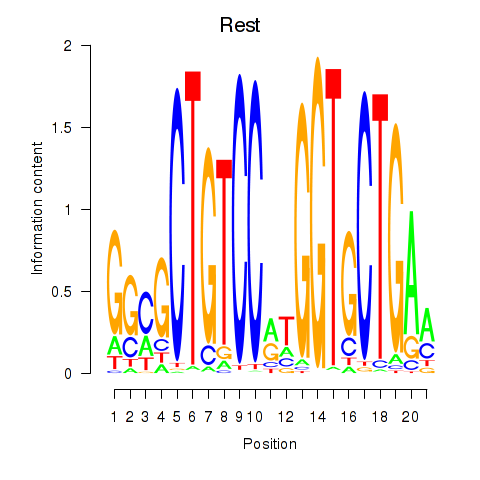
Motif ID: Rest
Z-value: 1.951
Transcription factors associated with Rest:
| Gene Symbol | Entrez ID | Gene Name |
|---|---|---|
| Rest | ENSMUSG00000029249.9 | Rest |
Activity-expression correlation:
| Gene Symbol | Promoter | Pearson corr. coef. | P-value | Plot |
|---|---|---|---|---|
| Rest | mm10_v2_chr5_+_77265454_77265549 | 0.42 | 4.7e-02 | Click! |
Top targets:
Gene overrepresentation in biological_process category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 1.2 | 3.7 | GO:1900673 | olefin metabolic process(GO:1900673) |
| 0.7 | 2.0 | GO:0070634 | transepithelial ammonium transport(GO:0070634) |
| 0.7 | 2.0 | GO:0035021 | negative regulation of Rac protein signal transduction(GO:0035021) |
| 0.6 | 3.1 | GO:0018364 | peptidyl-glutamine methylation(GO:0018364) |
| 0.5 | 5.9 | GO:0007220 | Notch receptor processing(GO:0007220) |
| 0.5 | 1.6 | GO:0070093 | negative regulation of glucagon secretion(GO:0070093) |
| 0.5 | 4.6 | GO:0060373 | regulation of atrial cardiac muscle cell membrane depolarization(GO:0060371) regulation of ventricular cardiac muscle cell membrane depolarization(GO:0060373) |
| 0.5 | 2.3 | GO:0097116 | gephyrin clustering involved in postsynaptic density assembly(GO:0097116) |
| 0.4 | 2.1 | GO:0071947 | protein deubiquitination involved in ubiquitin-dependent protein catabolic process(GO:0071947) |
| 0.4 | 10.0 | GO:0032011 | ARF protein signal transduction(GO:0032011) regulation of ARF protein signal transduction(GO:0032012) |
| 0.3 | 1.0 | GO:0038003 | opioid receptor signaling pathway(GO:0038003) regulation of opioid receptor signaling pathway(GO:2000474) |
| 0.3 | 10.4 | GO:0048791 | calcium ion-regulated exocytosis of neurotransmitter(GO:0048791) |
| 0.3 | 1.9 | GO:0045162 | clustering of voltage-gated sodium channels(GO:0045162) |
| 0.3 | 0.9 | GO:0046032 | ADP catabolic process(GO:0046032) |
| 0.3 | 1.7 | GO:0072318 | clathrin coat disassembly(GO:0072318) |
| 0.3 | 6.8 | GO:2000311 | regulation of alpha-amino-3-hydroxy-5-methyl-4-isoxazole propionate selective glutamate receptor activity(GO:2000311) |
| 0.3 | 14.6 | GO:0030901 | midbrain development(GO:0030901) |
| 0.2 | 3.3 | GO:0050910 | detection of mechanical stimulus involved in sensory perception of sound(GO:0050910) |
| 0.2 | 3.7 | GO:0007614 | short-term memory(GO:0007614) |
| 0.1 | 0.4 | GO:0051902 | negative regulation of mitochondrial depolarization(GO:0051902) negative regulation of membrane depolarization(GO:1904180) |
| 0.1 | 0.2 | GO:0021586 | pons maturation(GO:0021586) |
| 0.1 | 2.2 | GO:0071625 | vocalization behavior(GO:0071625) |
| 0.1 | 1.3 | GO:1900383 | regulation of synaptic plasticity by receptor localization to synapse(GO:1900383) |
| 0.1 | 1.0 | GO:0043084 | penile erection(GO:0043084) |
| 0.1 | 4.2 | GO:2001222 | regulation of neuron migration(GO:2001222) |
| 0.1 | 1.1 | GO:0060056 | mammary gland involution(GO:0060056) |
| 0.1 | 1.5 | GO:0007263 | nitric oxide mediated signal transduction(GO:0007263) |
| 0.1 | 2.8 | GO:0030514 | negative regulation of BMP signaling pathway(GO:0030514) |
| 0.1 | 1.3 | GO:0006828 | manganese ion transport(GO:0006828) |
| 0.0 | 0.7 | GO:0099517 | anterograde synaptic vesicle transport(GO:0048490) synaptic vesicle cytoskeletal transport(GO:0099514) synaptic vesicle transport along microtubule(GO:0099517) |
| 0.0 | 0.5 | GO:0015991 | energy coupled proton transmembrane transport, against electrochemical gradient(GO:0015988) ATP hydrolysis coupled proton transport(GO:0015991) ATP hydrolysis coupled transmembrane transport(GO:0090662) |
| 0.0 | 0.1 | GO:0090336 | positive regulation of brown fat cell differentiation(GO:0090336) |
| 0.0 | 0.9 | GO:0051965 | positive regulation of synapse assembly(GO:0051965) |
| 0.0 | 2.2 | GO:0060828 | regulation of canonical Wnt signaling pathway(GO:0060828) |
| 0.0 | 0.3 | GO:0006099 | tricarboxylic acid cycle(GO:0006099) |
| 0.0 | 0.7 | GO:0061077 | chaperone-mediated protein folding(GO:0061077) |
Gene overrepresentation in cellular_component category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.9 | 3.7 | GO:0044307 | dendritic branch(GO:0044307) |
| 0.6 | 2.8 | GO:0032426 | stereocilium tip(GO:0032426) |
| 0.4 | 3.1 | GO:0031428 | box C/D snoRNP complex(GO:0031428) |
| 0.3 | 1.9 | GO:0008091 | spectrin(GO:0008091) |
| 0.3 | 4.6 | GO:0034706 | voltage-gated sodium channel complex(GO:0001518) sodium channel complex(GO:0034706) |
| 0.3 | 7.9 | GO:0060077 | inhibitory synapse(GO:0060077) |
| 0.2 | 0.5 | GO:0002141 | stereocilia ankle link(GO:0002141) |
| 0.2 | 6.8 | GO:0032281 | AMPA glutamate receptor complex(GO:0032281) |
| 0.1 | 1.0 | GO:0030285 | integral component of synaptic vesicle membrane(GO:0030285) intrinsic component of synaptic vesicle membrane(GO:0098563) |
| 0.1 | 4.0 | GO:0030672 | synaptic vesicle membrane(GO:0030672) exocytic vesicle membrane(GO:0099501) |
| 0.1 | 1.0 | GO:0097440 | apical dendrite(GO:0097440) |
| 0.1 | 0.2 | GO:0016533 | cyclin-dependent protein kinase 5 holoenzyme complex(GO:0016533) |
| 0.1 | 0.9 | GO:0031045 | dense core granule(GO:0031045) |
| 0.1 | 0.7 | GO:1904115 | axon cytoplasm(GO:1904115) |
| 0.1 | 0.5 | GO:0033180 | proton-transporting V-type ATPase, V1 domain(GO:0033180) |
| 0.0 | 0.1 | GO:0043202 | lysosomal lumen(GO:0043202) |
| 0.0 | 2.5 | GO:0043195 | terminal bouton(GO:0043195) |
| 0.0 | 0.3 | GO:0045252 | oxoglutarate dehydrogenase complex(GO:0045252) |
| 0.0 | 2.0 | GO:0009925 | basal plasma membrane(GO:0009925) |
| 0.0 | 1.8 | GO:0042734 | presynaptic membrane(GO:0042734) |
| 0.0 | 1.3 | GO:0005801 | cis-Golgi network(GO:0005801) |
| 0.0 | 1.0 | GO:0030667 | secretory granule membrane(GO:0030667) |
| 0.0 | 5.9 | GO:0005769 | early endosome(GO:0005769) |
| 0.0 | 7.9 | GO:0043025 | neuronal cell body(GO:0043025) |
| 0.0 | 2.0 | GO:0014069 | postsynaptic density(GO:0014069) postsynaptic specialization(GO:0099572) |
Gene overrepresentation in molecular_function category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 1.0 | 9.5 | GO:0042301 | phosphate ion binding(GO:0042301) |
| 0.7 | 4.6 | GO:0086006 | voltage-gated sodium channel activity involved in cardiac muscle cell action potential(GO:0086006) |
| 0.3 | 3.7 | GO:0004972 | NMDA glutamate receptor activity(GO:0004972) |
| 0.3 | 10.0 | GO:0005086 | ARF guanyl-nucleotide exchange factor activity(GO:0005086) |
| 0.2 | 13.6 | GO:0005179 | hormone activity(GO:0005179) |
| 0.2 | 5.9 | GO:0005112 | Notch binding(GO:0005112) |
| 0.2 | 2.3 | GO:0097109 | neuroligin family protein binding(GO:0097109) |
| 0.2 | 3.9 | GO:0030506 | ankyrin binding(GO:0030506) |
| 0.2 | 6.8 | GO:0005245 | voltage-gated calcium channel activity(GO:0005245) |
| 0.1 | 0.9 | GO:0047631 | adenosine-diphosphatase activity(GO:0043262) ADP-ribose diphosphatase activity(GO:0047631) |
| 0.1 | 1.3 | GO:0015279 | store-operated calcium channel activity(GO:0015279) |
| 0.1 | 2.1 | GO:0070530 | K63-linked polyubiquitin binding(GO:0070530) |
| 0.1 | 1.0 | GO:0001786 | phosphatidylserine binding(GO:0001786) |
| 0.1 | 5.7 | GO:0043178 | alcohol binding(GO:0043178) |
| 0.1 | 7.0 | GO:0051219 | phosphoprotein binding(GO:0051219) |
| 0.1 | 3.5 | GO:0005546 | phosphatidylinositol-4,5-bisphosphate binding(GO:0005546) |
| 0.1 | 16.4 | GO:0001077 | transcriptional activator activity, RNA polymerase II core promoter proximal region sequence-specific binding(GO:0001077) |
| 0.0 | 3.1 | GO:0042054 | histone methyltransferase activity(GO:0042054) |
| 0.0 | 0.3 | GO:0004591 | oxoglutarate dehydrogenase (succinyl-transferring) activity(GO:0004591) |
| 0.0 | 1.5 | GO:0046875 | ephrin receptor binding(GO:0046875) |
| 0.0 | 0.7 | GO:0070888 | E-box binding(GO:0070888) |
| 0.0 | 0.7 | GO:0005528 | macrolide binding(GO:0005527) FK506 binding(GO:0005528) |
| 0.0 | 2.8 | GO:0046332 | SMAD binding(GO:0046332) |
| 0.0 | 0.5 | GO:0003841 | 1-acylglycerol-3-phosphate O-acyltransferase activity(GO:0003841) |
| 0.0 | 1.6 | GO:0004725 | protein tyrosine phosphatase activity(GO:0004725) |