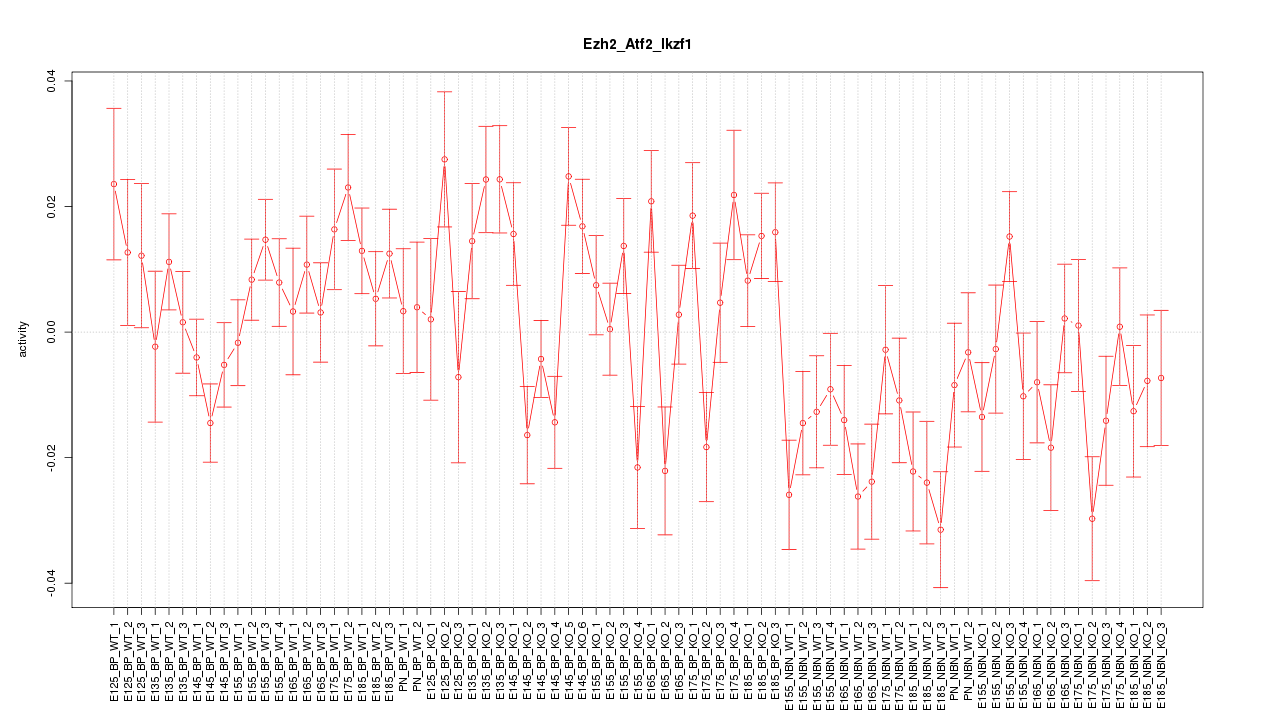
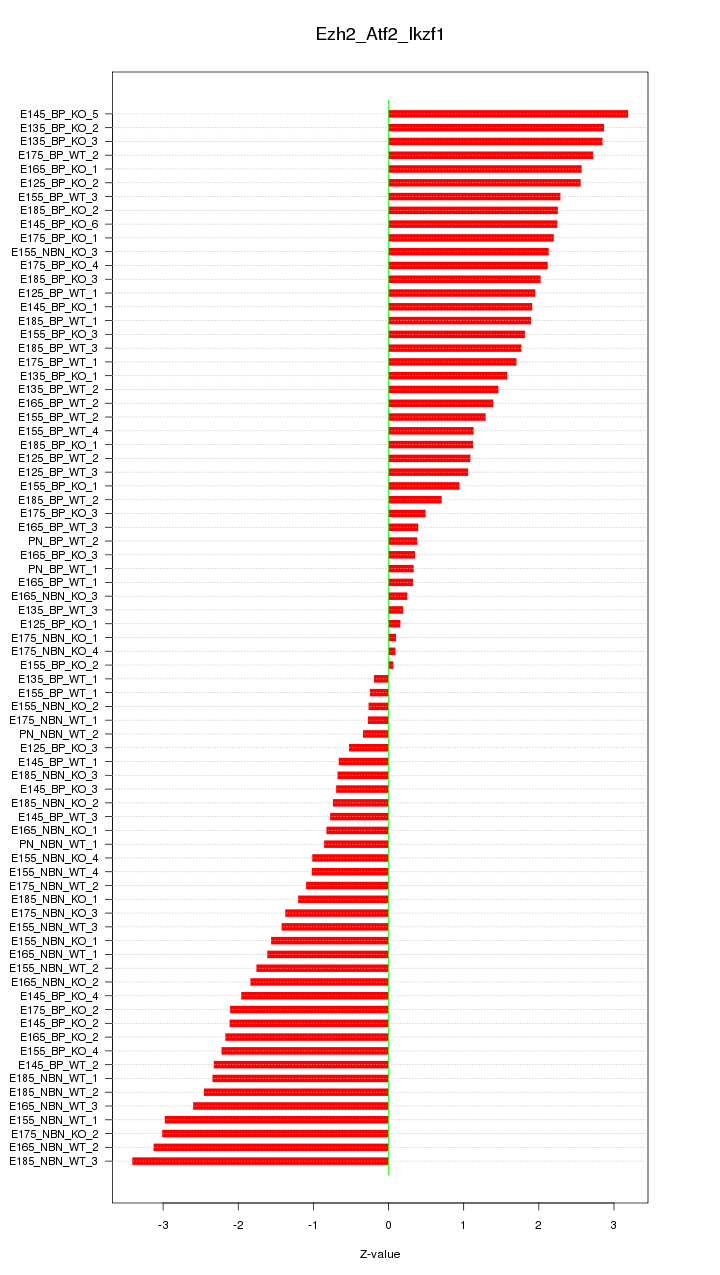
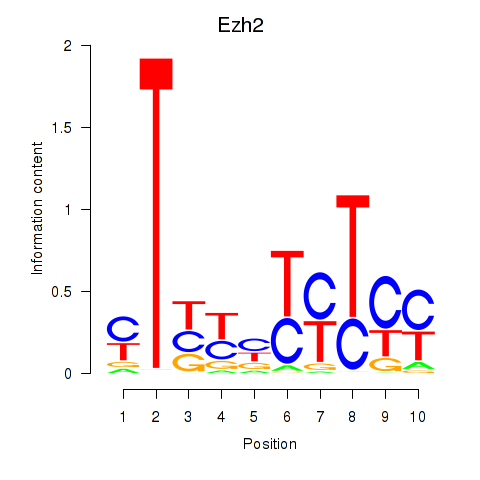
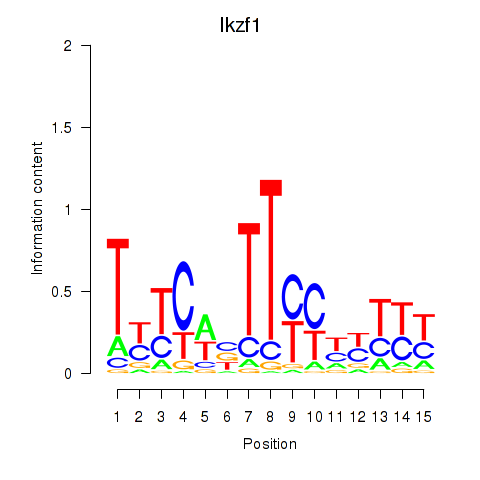
Motif ID: Ezh2_Atf2_Ikzf1
Z-value: 1.715
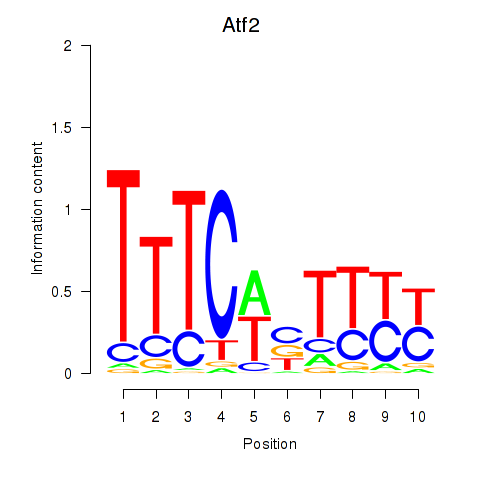
Transcription factors associated with Ezh2_Atf2_Ikzf1:
| Gene Symbol | Entrez ID | Gene Name |
|---|---|---|
| Atf2 | ENSMUSG00000027104.12 | Atf2 |
| Ezh2 | ENSMUSG00000029687.10 | Ezh2 |
| Ikzf1 | ENSMUSG00000018654.11 | Ikzf1 |
Activity-expression correlation:
| Gene Symbol | Promoter | Pearson corr. coef. | P-value | Plot |
|---|---|---|---|---|
| Ezh2 | mm10_v2_chr6_-_47594967_47595047 | 0.68 | 7.6e-12 | Click! |
| Atf2 | mm10_v2_chr2_-_73892588_73892616 | 0.55 | 1.9e-07 | Click! |
Top targets:
Gene overrepresentation in biological_process category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 20.6 | 61.8 | GO:0090425 | hepatocyte cell migration(GO:0002194) branching involved in pancreas morphogenesis(GO:0061114) acinar cell differentiation(GO:0090425) positive regulation of forebrain neuron differentiation(GO:2000979) |
| 9.5 | 28.6 | GO:0060244 | negative regulation of cell proliferation involved in contact inhibition(GO:0060244) |
| 8.8 | 26.3 | GO:0060067 | cervix development(GO:0060067) |
| 4.8 | 19.1 | GO:2000795 | negative regulation of epithelial cell proliferation involved in lung morphogenesis(GO:2000795) |
| 4.6 | 13.8 | GO:0002302 | CD8-positive, alpha-beta T cell differentiation involved in immune response(GO:0002302) mesodermal to mesenchymal transition involved in gastrulation(GO:0060809) |
| 4.0 | 12.0 | GO:0003221 | right ventricular cardiac muscle tissue morphogenesis(GO:0003221) |
| 3.6 | 21.6 | GO:0021797 | forebrain anterior/posterior pattern specification(GO:0021797) |
| 3.4 | 13.6 | GO:0021698 | cerebellar cortex structural organization(GO:0021698) |
| 3.2 | 19.5 | GO:0021965 | spinal cord ventral commissure morphogenesis(GO:0021965) |
| 2.6 | 5.2 | GO:0003404 | optic vesicle morphogenesis(GO:0003404) |
| 2.1 | 18.7 | GO:0042760 | very long-chain fatty acid catabolic process(GO:0042760) |
| 2.0 | 10.0 | GO:0032901 | positive regulation of neurotrophin production(GO:0032901) |
| 1.9 | 1.9 | GO:0070094 | positive regulation of glucagon secretion(GO:0070094) |
| 1.9 | 5.6 | GO:1900275 | negative regulation of phospholipase C activity(GO:1900275) |
| 1.9 | 7.5 | GO:0055099 | response to high density lipoprotein particle(GO:0055099) platelet dense granule organization(GO:0060155) |
| 1.8 | 11.0 | GO:0071883 | activation of MAPK activity by adrenergic receptor signaling pathway(GO:0071883) |
| 1.8 | 7.3 | GO:1903984 | positive regulation of TRAIL-activated apoptotic signaling pathway(GO:1903984) |
| 1.8 | 10.7 | GO:0048852 | diencephalon morphogenesis(GO:0048852) |
| 1.7 | 5.1 | GO:0090274 | positive regulation of somatostatin secretion(GO:0090274) |
| 1.7 | 21.5 | GO:0021796 | cerebral cortex regionalization(GO:0021796) |
| 1.6 | 19.3 | GO:0061302 | smooth muscle cell-matrix adhesion(GO:0061302) |
| 1.6 | 3.2 | GO:2000790 | regulation of mesenchymal cell proliferation involved in lung development(GO:2000790) negative regulation of mesenchymal cell proliferation involved in lung development(GO:2000791) |
| 1.6 | 11.0 | GO:0060613 | fat pad development(GO:0060613) |
| 1.5 | 6.1 | GO:0090126 | protein complex assembly involved in synapse maturation(GO:0090126) |
| 1.4 | 4.3 | GO:0007525 | somatic muscle development(GO:0007525) |
| 1.4 | 4.3 | GO:0023016 | signal transduction by trans-phosphorylation(GO:0023016) |
| 1.4 | 17.1 | GO:2001269 | positive regulation of cysteine-type endopeptidase activity involved in apoptotic signaling pathway(GO:2001269) |
| 1.4 | 22.0 | GO:0048387 | negative regulation of retinoic acid receptor signaling pathway(GO:0048387) |
| 1.4 | 9.6 | GO:0044806 | G-quadruplex DNA unwinding(GO:0044806) |
| 1.3 | 9.1 | GO:0021780 | oligodendrocyte cell fate specification(GO:0021778) oligodendrocyte cell fate commitment(GO:0021779) glial cell fate specification(GO:0021780) |
| 1.3 | 5.0 | GO:0060316 | positive regulation of ryanodine-sensitive calcium-release channel activity(GO:0060316) |
| 1.3 | 3.8 | GO:1903225 | negative regulation of endodermal cell differentiation(GO:1903225) |
| 1.2 | 11.0 | GO:0021979 | hypothalamus cell differentiation(GO:0021979) |
| 1.2 | 8.5 | GO:0097105 | presynaptic membrane assembly(GO:0097105) |
| 1.2 | 13.1 | GO:0060539 | diaphragm development(GO:0060539) |
| 1.2 | 3.5 | GO:0021759 | globus pallidus development(GO:0021759) |
| 1.1 | 6.8 | GO:0032196 | regulation of transposition(GO:0010528) negative regulation of transposition(GO:0010529) transposition(GO:0032196) |
| 1.1 | 8.7 | GO:0048625 | myoblast fate commitment(GO:0048625) |
| 1.1 | 4.3 | GO:0097476 | spinal cord motor neuron migration(GO:0097476) |
| 1.1 | 25.9 | GO:0010172 | embryonic body morphogenesis(GO:0010172) |
| 1.0 | 7.3 | GO:0032596 | protein transport into membrane raft(GO:0032596) |
| 1.0 | 5.1 | GO:0061073 | ciliary body morphogenesis(GO:0061073) |
| 1.0 | 1.0 | GO:0032747 | positive regulation of interleukin-23 production(GO:0032747) |
| 1.0 | 3.9 | GO:2001045 | negative regulation of integrin-mediated signaling pathway(GO:2001045) |
| 1.0 | 4.8 | GO:0090310 | negative regulation of methylation-dependent chromatin silencing(GO:0090310) |
| 1.0 | 3.8 | GO:0097393 | post-embryonic appendage morphogenesis(GO:0035120) post-embryonic limb morphogenesis(GO:0035127) post-embryonic forelimb morphogenesis(GO:0035128) telomeric repeat-containing RNA transcription(GO:0097393) telomeric repeat-containing RNA transcription from RNA pol II promoter(GO:0097394) regulation of telomeric RNA transcription from RNA pol II promoter(GO:1901580) negative regulation of telomeric RNA transcription from RNA pol II promoter(GO:1901581) positive regulation of telomeric RNA transcription from RNA pol II promoter(GO:1901582) |
| 0.9 | 9.4 | GO:1901386 | negative regulation of voltage-gated calcium channel activity(GO:1901386) |
| 0.9 | 2.7 | GO:0021691 | cerebellar Purkinje cell layer maturation(GO:0021691) |
| 0.9 | 2.7 | GO:0000821 | regulation of arginine metabolic process(GO:0000821) |
| 0.9 | 3.6 | GO:0060282 | positive regulation of oocyte development(GO:0060282) |
| 0.9 | 53.8 | GO:0006940 | regulation of smooth muscle contraction(GO:0006940) |
| 0.9 | 2.6 | GO:1902528 | regulation of protein linear polyubiquitination(GO:1902528) positive regulation of protein linear polyubiquitination(GO:1902530) |
| 0.8 | 1.7 | GO:0098914 | membrane repolarization during atrial cardiac muscle cell action potential(GO:0098914) |
| 0.8 | 3.3 | GO:0055071 | cellular manganese ion homeostasis(GO:0030026) Golgi calcium ion homeostasis(GO:0032468) manganese ion homeostasis(GO:0055071) |
| 0.8 | 2.4 | GO:0019085 | early viral transcription(GO:0019085) |
| 0.8 | 4.1 | GO:1903566 | positive regulation of protein localization to cilium(GO:1903566) |
| 0.8 | 2.4 | GO:0032696 | negative regulation of interleukin-13 production(GO:0032696) trachea submucosa development(GO:0061152) trachea gland development(GO:0061153) odontoblast differentiation(GO:0071895) regulation of estrogen receptor binding(GO:0071898) negative regulation of estrogen receptor binding(GO:0071899) |
| 0.8 | 3.2 | GO:0050904 | diapedesis(GO:0050904) |
| 0.8 | 11.2 | GO:0031915 | positive regulation of synaptic plasticity(GO:0031915) |
| 0.8 | 2.3 | GO:0046121 | deoxyribonucleoside catabolic process(GO:0046121) |
| 0.8 | 31.7 | GO:0032728 | positive regulation of interferon-beta production(GO:0032728) |
| 0.8 | 3.0 | GO:0033326 | cerebrospinal fluid secretion(GO:0033326) |
| 0.7 | 9.6 | GO:0006309 | apoptotic DNA fragmentation(GO:0006309) |
| 0.7 | 2.2 | GO:0051653 | establishment of mitotic spindle orientation(GO:0000132) establishment of mitotic spindle localization(GO:0040001) establishment of spindle localization(GO:0051293) establishment of spindle orientation(GO:0051294) spindle localization(GO:0051653) |
| 0.7 | 1.4 | GO:0042320 | regulation of circadian sleep/wake cycle, REM sleep(GO:0042320) circadian sleep/wake cycle, REM sleep(GO:0042747) positive regulation of circadian sleep/wake cycle, sleep(GO:0045938) |
| 0.7 | 2.1 | GO:0070315 | G1 to G0 transition involved in cell differentiation(GO:0070315) |
| 0.7 | 2.1 | GO:0007056 | spindle assembly involved in female meiosis(GO:0007056) |
| 0.7 | 4.8 | GO:0016584 | nucleosome positioning(GO:0016584) |
| 0.7 | 37.0 | GO:0007628 | adult walking behavior(GO:0007628) |
| 0.7 | 4.0 | GO:0097368 | establishment of Sertoli cell barrier(GO:0097368) |
| 0.7 | 2.0 | GO:0015793 | glycerol transport(GO:0015793) |
| 0.6 | 1.3 | GO:2000686 | regulation of rubidium ion transmembrane transporter activity(GO:2000686) |
| 0.6 | 13.8 | GO:0035855 | megakaryocyte development(GO:0035855) |
| 0.6 | 1.9 | GO:0007171 | activation of transmembrane receptor protein tyrosine kinase activity(GO:0007171) |
| 0.6 | 1.8 | GO:0070634 | transepithelial ammonium transport(GO:0070634) |
| 0.6 | 1.8 | GO:0002457 | T cell antigen processing and presentation(GO:0002457) |
| 0.6 | 4.8 | GO:0016576 | histone dephosphorylation(GO:0016576) |
| 0.6 | 1.8 | GO:0002191 | cap-dependent translational initiation(GO:0002191) |
| 0.6 | 1.8 | GO:0090649 | response to oxygen-glucose deprivation(GO:0090649) cellular response to oxygen-glucose deprivation(GO:0090650) |
| 0.6 | 4.0 | GO:1903715 | regulation of aerobic respiration(GO:1903715) |
| 0.6 | 1.7 | GO:0021960 | anterior commissure morphogenesis(GO:0021960) |
| 0.6 | 1.7 | GO:0060489 | orthogonal dichotomous subdivision of terminal units involved in lung branching morphogenesis(GO:0060488) planar dichotomous subdivision of terminal units involved in lung branching morphogenesis(GO:0060489) lateral sprouting involved in lung morphogenesis(GO:0060490) non-canonical Wnt signaling pathway involved in heart development(GO:0061341) planar cell polarity pathway involved in heart morphogenesis(GO:0061346) |
| 0.6 | 3.4 | GO:0045743 | positive regulation of fibroblast growth factor receptor signaling pathway(GO:0045743) |
| 0.6 | 7.2 | GO:0071481 | cellular response to X-ray(GO:0071481) |
| 0.6 | 2.2 | GO:0070829 | heterochromatin maintenance(GO:0070829) |
| 0.6 | 2.8 | GO:1901300 | positive regulation of hydrogen peroxide-mediated programmed cell death(GO:1901300) |
| 0.5 | 2.7 | GO:0071699 | olfactory placode formation(GO:0030910) olfactory placode development(GO:0071698) olfactory placode morphogenesis(GO:0071699) |
| 0.5 | 0.5 | GO:0039519 | modulation by virus of host autophagy(GO:0039519) |
| 0.5 | 2.1 | GO:0003365 | establishment of cell polarity involved in ameboidal cell migration(GO:0003365) |
| 0.5 | 2.1 | GO:0014719 | skeletal muscle satellite cell activation(GO:0014719) |
| 0.5 | 1.0 | GO:0032417 | positive regulation of sodium:proton antiporter activity(GO:0032417) |
| 0.5 | 2.1 | GO:0070934 | CRD-mediated mRNA stabilization(GO:0070934) |
| 0.5 | 5.1 | GO:0043586 | tongue development(GO:0043586) |
| 0.5 | 2.5 | GO:0035616 | histone H2B conserved C-terminal lysine deubiquitination(GO:0035616) |
| 0.5 | 1.5 | GO:0008594 | photoreceptor cell morphogenesis(GO:0008594) |
| 0.5 | 3.5 | GO:0031508 | pericentric heterochromatin assembly(GO:0031508) |
| 0.5 | 1.5 | GO:0019858 | cytosine metabolic process(GO:0019858) |
| 0.5 | 2.5 | GO:0035063 | nuclear speck organization(GO:0035063) |
| 0.5 | 3.0 | GO:0000160 | phosphorelay signal transduction system(GO:0000160) |
| 0.5 | 3.5 | GO:0030718 | germ-line stem cell population maintenance(GO:0030718) |
| 0.5 | 2.0 | GO:0051088 | PMA-inducible membrane protein ectodomain proteolysis(GO:0051088) |
| 0.5 | 0.5 | GO:0045085 | negative regulation of interleukin-2 biosynthetic process(GO:0045085) |
| 0.5 | 1.0 | GO:1904058 | positive regulation of sensory perception of pain(GO:1904058) |
| 0.5 | 12.6 | GO:0010842 | retina layer formation(GO:0010842) |
| 0.5 | 1.9 | GO:0006272 | leading strand elongation(GO:0006272) |
| 0.5 | 3.7 | GO:0018344 | protein geranylgeranylation(GO:0018344) |
| 0.5 | 2.3 | GO:2000325 | regulation of ligand-dependent nuclear receptor transcription coactivator activity(GO:2000325) positive regulation of ligand-dependent nuclear receptor transcription coactivator activity(GO:2000327) |
| 0.5 | 7.0 | GO:0046697 | decidualization(GO:0046697) |
| 0.5 | 21.3 | GO:2001222 | regulation of neuron migration(GO:2001222) |
| 0.5 | 4.6 | GO:0007213 | G-protein coupled acetylcholine receptor signaling pathway(GO:0007213) |
| 0.5 | 1.4 | GO:1901629 | regulation of presynaptic membrane organization(GO:1901629) |
| 0.5 | 1.4 | GO:0070213 | protein auto-ADP-ribosylation(GO:0070213) |
| 0.4 | 1.3 | GO:0089700 | protein kinase D signaling(GO:0089700) |
| 0.4 | 6.7 | GO:0007288 | sperm axoneme assembly(GO:0007288) |
| 0.4 | 1.3 | GO:0048105 | establishment of body hair or bristle planar orientation(GO:0048104) establishment of body hair planar orientation(GO:0048105) |
| 0.4 | 3.6 | GO:2000781 | positive regulation of double-strand break repair(GO:2000781) |
| 0.4 | 4.0 | GO:0070236 | negative regulation of activation-induced cell death of T cells(GO:0070236) |
| 0.4 | 1.7 | GO:0070345 | negative regulation of fat cell proliferation(GO:0070345) |
| 0.4 | 1.3 | GO:0032650 | regulation of interleukin-1 alpha production(GO:0032650) positive regulation of interleukin-1 alpha production(GO:0032730) |
| 0.4 | 0.8 | GO:0002578 | negative regulation of antigen processing and presentation(GO:0002578) |
| 0.4 | 2.1 | GO:0048682 | axon extension involved in regeneration(GO:0048677) sprouting of injured axon(GO:0048682) |
| 0.4 | 2.0 | GO:2001168 | regulation of histone H2B ubiquitination(GO:2001166) positive regulation of histone H2B ubiquitination(GO:2001168) |
| 0.4 | 0.8 | GO:1901203 | positive regulation of extracellular matrix assembly(GO:1901203) |
| 0.4 | 0.8 | GO:1905244 | regulation of modification of synaptic structure(GO:1905244) |
| 0.4 | 4.0 | GO:0070423 | nucleotide-binding oligomerization domain containing signaling pathway(GO:0070423) nucleotide-binding oligomerization domain containing 2 signaling pathway(GO:0070431) |
| 0.4 | 1.6 | GO:0003360 | brainstem development(GO:0003360) |
| 0.4 | 3.9 | GO:0046602 | regulation of mitotic centrosome separation(GO:0046602) |
| 0.4 | 1.2 | GO:0030200 | heparan sulfate proteoglycan catabolic process(GO:0030200) |
| 0.4 | 3.5 | GO:0098700 | neurotransmitter loading into synaptic vesicle(GO:0098700) |
| 0.4 | 35.4 | GO:0043488 | regulation of mRNA stability(GO:0043488) |
| 0.4 | 1.6 | GO:1900147 | Schwann cell migration(GO:0036135) regulation of Schwann cell migration(GO:1900147) |
| 0.4 | 3.5 | GO:0010587 | miRNA catabolic process(GO:0010587) |
| 0.4 | 2.7 | GO:0060923 | cardiac muscle cell fate commitment(GO:0060923) |
| 0.4 | 1.2 | GO:0048633 | positive regulation of skeletal muscle tissue growth(GO:0048633) |
| 0.4 | 1.9 | GO:0046643 | regulation of gamma-delta T cell differentiation(GO:0045586) regulation of gamma-delta T cell activation(GO:0046643) |
| 0.4 | 3.0 | GO:0048664 | neuron fate determination(GO:0048664) |
| 0.4 | 1.5 | GO:0045162 | clustering of voltage-gated sodium channels(GO:0045162) |
| 0.4 | 3.6 | GO:0032876 | negative regulation of DNA endoreduplication(GO:0032876) |
| 0.4 | 2.9 | GO:0042095 | interferon-gamma biosynthetic process(GO:0042095) |
| 0.3 | 4.2 | GO:0003334 | keratinocyte development(GO:0003334) |
| 0.3 | 3.3 | GO:0042501 | serine phosphorylation of STAT protein(GO:0042501) |
| 0.3 | 1.6 | GO:0090074 | negative regulation of protein homodimerization activity(GO:0090074) |
| 0.3 | 1.6 | GO:1900169 | regulation of glucocorticoid mediated signaling pathway(GO:1900169) |
| 0.3 | 2.6 | GO:0071557 | histone H3-K27 demethylation(GO:0071557) |
| 0.3 | 6.1 | GO:0006353 | DNA-templated transcription, termination(GO:0006353) |
| 0.3 | 2.9 | GO:0071493 | cellular response to UV-B(GO:0071493) |
| 0.3 | 2.2 | GO:0007296 | vitellogenesis(GO:0007296) |
| 0.3 | 1.6 | GO:0044793 | negative regulation by host of viral process(GO:0044793) |
| 0.3 | 0.3 | GO:0006436 | tryptophanyl-tRNA aminoacylation(GO:0006436) |
| 0.3 | 0.9 | GO:0009814 | defense response, incompatible interaction(GO:0009814) |
| 0.3 | 1.2 | GO:0033603 | positive regulation of dopamine secretion(GO:0033603) positive regulation of catecholamine secretion(GO:0033605) |
| 0.3 | 4.2 | GO:0050774 | negative regulation of dendrite morphogenesis(GO:0050774) |
| 0.3 | 8.6 | GO:0001754 | eye photoreceptor cell differentiation(GO:0001754) |
| 0.3 | 1.2 | GO:0045654 | positive regulation of megakaryocyte differentiation(GO:0045654) |
| 0.3 | 2.0 | GO:0051660 | establishment of centrosome localization(GO:0051660) |
| 0.3 | 2.3 | GO:0035871 | protein K11-linked deubiquitination(GO:0035871) |
| 0.3 | 3.1 | GO:0060213 | regulation of nuclear-transcribed mRNA poly(A) tail shortening(GO:0060211) positive regulation of nuclear-transcribed mRNA poly(A) tail shortening(GO:0060213) |
| 0.3 | 2.8 | GO:0048934 | peripheral nervous system neuron differentiation(GO:0048934) peripheral nervous system neuron development(GO:0048935) |
| 0.3 | 1.4 | GO:0010792 | DNA double-strand break processing involved in repair via single-strand annealing(GO:0010792) |
| 0.3 | 2.2 | GO:0048102 | autophagic cell death(GO:0048102) |
| 0.3 | 1.9 | GO:0098535 | de novo centriole assembly(GO:0098535) |
| 0.3 | 1.1 | GO:0044849 | estrous cycle(GO:0044849) regulation of hepatocyte apoptotic process(GO:1903943) negative regulation of hepatocyte apoptotic process(GO:1903944) |
| 0.3 | 2.2 | GO:0090129 | positive regulation of synapse maturation(GO:0090129) |
| 0.3 | 1.6 | GO:0015074 | DNA integration(GO:0015074) |
| 0.3 | 7.0 | GO:0032011 | ARF protein signal transduction(GO:0032011) regulation of ARF protein signal transduction(GO:0032012) |
| 0.3 | 2.4 | GO:0031339 | negative regulation of vesicle fusion(GO:0031339) |
| 0.3 | 1.6 | GO:0021869 | forebrain ventricular zone progenitor cell division(GO:0021869) |
| 0.3 | 1.6 | GO:0007016 | cytoskeletal anchoring at plasma membrane(GO:0007016) |
| 0.3 | 1.0 | GO:0090238 | positive regulation of arachidonic acid secretion(GO:0090238) |
| 0.3 | 1.0 | GO:0014842 | regulation of skeletal muscle satellite cell proliferation(GO:0014842) |
| 0.3 | 2.3 | GO:0060009 | Sertoli cell development(GO:0060009) |
| 0.3 | 3.5 | GO:0045579 | positive regulation of B cell differentiation(GO:0045579) |
| 0.2 | 3.2 | GO:0042759 | long-chain fatty acid biosynthetic process(GO:0042759) |
| 0.2 | 0.7 | GO:1904879 | positive regulation of calcium ion transmembrane transport via high voltage-gated calcium channel(GO:1904879) |
| 0.2 | 0.5 | GO:0061370 | negative regulation of chronic inflammatory response(GO:0002677) testosterone biosynthetic process(GO:0061370) |
| 0.2 | 1.2 | GO:0030263 | apoptotic chromosome condensation(GO:0030263) |
| 0.2 | 1.0 | GO:0051890 | regulation of cardioblast differentiation(GO:0051890) positive regulation of secondary heart field cardioblast proliferation(GO:0072513) |
| 0.2 | 1.2 | GO:0015867 | ATP transport(GO:0015867) |
| 0.2 | 2.4 | GO:0090261 | positive regulation of inclusion body assembly(GO:0090261) |
| 0.2 | 1.2 | GO:0003406 | retinal pigment epithelium development(GO:0003406) |
| 0.2 | 0.5 | GO:0070827 | chromatin maintenance(GO:0070827) |
| 0.2 | 0.9 | GO:0060903 | positive regulation of meiosis I(GO:0060903) |
| 0.2 | 2.6 | GO:0015937 | coenzyme A biosynthetic process(GO:0015937) |
| 0.2 | 0.7 | GO:0021553 | olfactory nerve development(GO:0021553) |
| 0.2 | 0.4 | GO:0071442 | positive regulation of histone H3-K14 acetylation(GO:0071442) |
| 0.2 | 1.5 | GO:1903608 | protein localization to cytoplasmic stress granule(GO:1903608) |
| 0.2 | 0.7 | GO:0060708 | spongiotrophoblast differentiation(GO:0060708) |
| 0.2 | 1.1 | GO:0043383 | negative T cell selection(GO:0043383) negative thymic T cell selection(GO:0045060) |
| 0.2 | 16.0 | GO:0001578 | microtubule bundle formation(GO:0001578) |
| 0.2 | 1.5 | GO:0036444 | calcium ion transmembrane import into mitochondrion(GO:0036444) |
| 0.2 | 1.7 | GO:0001672 | regulation of chromatin assembly or disassembly(GO:0001672) |
| 0.2 | 5.0 | GO:0045736 | negative regulation of cyclin-dependent protein serine/threonine kinase activity(GO:0045736) negative regulation of cyclin-dependent protein kinase activity(GO:1904030) |
| 0.2 | 1.1 | GO:0048733 | sebaceous gland development(GO:0048733) |
| 0.2 | 2.7 | GO:0035563 | positive regulation of chromatin binding(GO:0035563) |
| 0.2 | 1.2 | GO:0035735 | intraciliary transport involved in cilium morphogenesis(GO:0035735) |
| 0.2 | 3.3 | GO:0070816 | phosphorylation of RNA polymerase II C-terminal domain(GO:0070816) |
| 0.2 | 0.6 | GO:0097274 | urea homeostasis(GO:0097274) |
| 0.2 | 0.6 | GO:1990839 | response to endothelin(GO:1990839) |
| 0.2 | 4.7 | GO:0043486 | histone exchange(GO:0043486) |
| 0.2 | 2.7 | GO:0045671 | negative regulation of osteoclast differentiation(GO:0045671) |
| 0.2 | 0.6 | GO:0034334 | adherens junction maintenance(GO:0034334) |
| 0.2 | 3.6 | GO:0015693 | magnesium ion transport(GO:0015693) |
| 0.2 | 1.2 | GO:0098964 | dendritic transport of ribonucleoprotein complex(GO:0098961) dendritic transport of messenger ribonucleoprotein complex(GO:0098963) anterograde dendritic transport of messenger ribonucleoprotein complex(GO:0098964) |
| 0.2 | 0.6 | GO:0070318 | positive regulation of G0 to G1 transition(GO:0070318) |
| 0.2 | 19.3 | GO:0045727 | positive regulation of translation(GO:0045727) |
| 0.2 | 1.4 | GO:0006691 | leukotriene metabolic process(GO:0006691) |
| 0.2 | 0.2 | GO:0010616 | negative regulation of cardiac muscle adaptation(GO:0010616) |
| 0.2 | 1.3 | GO:0010992 | ubiquitin homeostasis(GO:0010992) |
| 0.2 | 0.6 | GO:0050748 | negative regulation of lipoprotein metabolic process(GO:0050748) |
| 0.2 | 7.2 | GO:0014037 | Schwann cell differentiation(GO:0014037) |
| 0.2 | 1.9 | GO:0009404 | toxin metabolic process(GO:0009404) |
| 0.2 | 0.9 | GO:0061009 | common bile duct development(GO:0061009) |
| 0.2 | 0.9 | GO:0033148 | positive regulation of intracellular estrogen receptor signaling pathway(GO:0033148) |
| 0.2 | 1.1 | GO:0000289 | nuclear-transcribed mRNA poly(A) tail shortening(GO:0000289) |
| 0.2 | 0.4 | GO:0070164 | negative regulation of adiponectin secretion(GO:0070164) |
| 0.2 | 0.2 | GO:0036112 | medium-chain fatty-acyl-CoA metabolic process(GO:0036112) |
| 0.2 | 0.9 | GO:0015671 | oxygen transport(GO:0015671) |
| 0.2 | 0.5 | GO:0016598 | protein arginylation(GO:0016598) |
| 0.2 | 0.5 | GO:0060385 | axonogenesis involved in innervation(GO:0060385) |
| 0.2 | 2.8 | GO:0030308 | negative regulation of cell growth(GO:0030308) |
| 0.2 | 0.7 | GO:1901098 | positive regulation of autophagosome maturation(GO:1901098) |
| 0.2 | 1.4 | GO:2000096 | positive regulation of Wnt signaling pathway, planar cell polarity pathway(GO:2000096) |
| 0.2 | 0.5 | GO:1903966 | monounsaturated fatty acid metabolic process(GO:1903964) monounsaturated fatty acid biosynthetic process(GO:1903966) |
| 0.2 | 0.7 | GO:1901162 | serotonin biosynthetic process(GO:0042427) primary amino compound biosynthetic process(GO:1901162) |
| 0.2 | 1.7 | GO:0006620 | posttranslational protein targeting to membrane(GO:0006620) |
| 0.2 | 2.6 | GO:0014912 | negative regulation of smooth muscle cell migration(GO:0014912) |
| 0.2 | 6.1 | GO:0000381 | regulation of alternative mRNA splicing, via spliceosome(GO:0000381) |
| 0.2 | 3.5 | GO:0043029 | T cell homeostasis(GO:0043029) |
| 0.2 | 1.2 | GO:0035520 | monoubiquitinated protein deubiquitination(GO:0035520) |
| 0.2 | 1.3 | GO:0060330 | regulation of response to interferon-gamma(GO:0060330) |
| 0.2 | 0.7 | GO:0032375 | negative regulation of sterol transport(GO:0032372) negative regulation of cholesterol transport(GO:0032375) |
| 0.2 | 2.9 | GO:0007175 | negative regulation of epidermal growth factor-activated receptor activity(GO:0007175) |
| 0.2 | 2.9 | GO:0048368 | lateral mesoderm development(GO:0048368) |
| 0.2 | 0.6 | GO:1903756 | regulation of transcription from RNA polymerase II promoter by histone modification(GO:1903756) negative regulation of transcription from RNA polymerase II promoter by histone modification(GO:1903758) |
| 0.2 | 1.4 | GO:0030259 | lipid glycosylation(GO:0030259) |
| 0.2 | 0.5 | GO:1990928 | response to amino acid starvation(GO:1990928) |
| 0.2 | 1.1 | GO:0090557 | establishment of endothelial intestinal barrier(GO:0090557) |
| 0.2 | 1.6 | GO:0019368 | fatty acid elongation, saturated fatty acid(GO:0019367) fatty acid elongation, unsaturated fatty acid(GO:0019368) fatty acid elongation, monounsaturated fatty acid(GO:0034625) fatty acid elongation, polyunsaturated fatty acid(GO:0034626) |
| 0.2 | 1.1 | GO:0097428 | protein maturation by iron-sulfur cluster transfer(GO:0097428) |
| 0.1 | 0.7 | GO:0042769 | DNA damage response, detection of DNA damage(GO:0042769) |
| 0.1 | 1.0 | GO:0045742 | positive regulation of epidermal growth factor receptor signaling pathway(GO:0045742) positive regulation of ERBB signaling pathway(GO:1901186) |
| 0.1 | 0.6 | GO:0006547 | histidine metabolic process(GO:0006547) |
| 0.1 | 1.0 | GO:0048706 | embryonic skeletal system development(GO:0048706) |
| 0.1 | 0.1 | GO:0050883 | musculoskeletal movement, spinal reflex action(GO:0050883) |
| 0.1 | 0.9 | GO:0007196 | adenylate cyclase-inhibiting G-protein coupled glutamate receptor signaling pathway(GO:0007196) |
| 0.1 | 0.7 | GO:0090383 | phagosome acidification(GO:0090383) |
| 0.1 | 0.6 | GO:0010593 | negative regulation of lamellipodium assembly(GO:0010593) |
| 0.1 | 6.1 | GO:0030901 | midbrain development(GO:0030901) |
| 0.1 | 0.7 | GO:0060753 | regulation of mast cell chemotaxis(GO:0060753) |
| 0.1 | 1.2 | GO:1903298 | regulation of hypoxia-induced intrinsic apoptotic signaling pathway(GO:1903297) negative regulation of hypoxia-induced intrinsic apoptotic signaling pathway(GO:1903298) |
| 0.1 | 0.4 | GO:0045059 | positive thymic T cell selection(GO:0045059) |
| 0.1 | 2.6 | GO:0060049 | regulation of protein glycosylation(GO:0060049) |
| 0.1 | 0.3 | GO:0007158 | neuron cell-cell adhesion(GO:0007158) |
| 0.1 | 2.0 | GO:0045911 | positive regulation of isotype switching(GO:0045830) positive regulation of DNA recombination(GO:0045911) |
| 0.1 | 0.4 | GO:0036292 | DNA rewinding(GO:0036292) |
| 0.1 | 0.7 | GO:1903361 | protein localization to basolateral plasma membrane(GO:1903361) |
| 0.1 | 1.1 | GO:0018026 | peptidyl-lysine monomethylation(GO:0018026) |
| 0.1 | 0.7 | GO:0038018 | Wnt receptor catabolic process(GO:0038018) |
| 0.1 | 1.2 | GO:1904738 | vascular associated smooth muscle cell migration(GO:1904738) regulation of vascular associated smooth muscle cell migration(GO:1904752) positive regulation of vascular associated smooth muscle cell migration(GO:1904754) |
| 0.1 | 0.7 | GO:0043568 | positive regulation of insulin-like growth factor receptor signaling pathway(GO:0043568) |
| 0.1 | 0.7 | GO:0043117 | positive regulation of vascular permeability(GO:0043117) |
| 0.1 | 0.3 | GO:0060040 | retinal bipolar neuron differentiation(GO:0060040) |
| 0.1 | 0.3 | GO:0046462 | monoacylglycerol metabolic process(GO:0046462) |
| 0.1 | 0.9 | GO:0051611 | negative regulation of neurotransmitter uptake(GO:0051581) serotonin uptake(GO:0051610) regulation of serotonin uptake(GO:0051611) negative regulation of serotonin uptake(GO:0051612) |
| 0.1 | 0.8 | GO:0010642 | negative regulation of platelet-derived growth factor receptor signaling pathway(GO:0010642) |
| 0.1 | 1.4 | GO:0036120 | response to platelet-derived growth factor(GO:0036119) cellular response to platelet-derived growth factor stimulus(GO:0036120) |
| 0.1 | 0.4 | GO:1902083 | negative regulation of peptidyl-cysteine S-nitrosylation(GO:1902083) |
| 0.1 | 0.4 | GO:0014835 | myoblast differentiation involved in skeletal muscle regeneration(GO:0014835) |
| 0.1 | 3.6 | GO:0007588 | excretion(GO:0007588) |
| 0.1 | 2.6 | GO:0016180 | snRNA processing(GO:0016180) |
| 0.1 | 0.7 | GO:0035095 | behavioral response to nicotine(GO:0035095) |
| 0.1 | 1.2 | GO:0001675 | acrosome assembly(GO:0001675) |
| 0.1 | 0.3 | GO:0060912 | cardiac cell fate specification(GO:0060912) |
| 0.1 | 0.3 | GO:0030473 | modulation of microtubule cytoskeleton involved in cerebral cortex radial glia guided migration(GO:0021815) nucleokinesis involved in cell motility in cerebral cortex radial glia guided migration(GO:0021817) nuclear migration along microtubule(GO:0030473) nuclear migration along microfilament(GO:0031022) |
| 0.1 | 0.3 | GO:0002281 | macrophage activation involved in immune response(GO:0002281) |
| 0.1 | 0.7 | GO:0016266 | O-glycan processing(GO:0016266) |
| 0.1 | 0.7 | GO:0042633 | molting cycle(GO:0042303) hair cycle(GO:0042633) |
| 0.1 | 0.1 | GO:0060681 | branch elongation involved in ureteric bud branching(GO:0060681) |
| 0.1 | 0.6 | GO:0090527 | actin filament reorganization(GO:0090527) |
| 0.1 | 2.6 | GO:0046827 | positive regulation of protein export from nucleus(GO:0046827) |
| 0.1 | 0.6 | GO:0046684 | response to pyrethroid(GO:0046684) |
| 0.1 | 1.4 | GO:0000083 | regulation of transcription involved in G1/S transition of mitotic cell cycle(GO:0000083) |
| 0.1 | 0.9 | GO:0048742 | regulation of skeletal muscle fiber development(GO:0048742) |
| 0.1 | 5.4 | GO:0050684 | regulation of mRNA processing(GO:0050684) |
| 0.1 | 0.6 | GO:0033564 | anterior/posterior axon guidance(GO:0033564) |
| 0.1 | 0.5 | GO:0048840 | otolith development(GO:0048840) |
| 0.1 | 0.2 | GO:0072719 | cellular response to cisplatin(GO:0072719) |
| 0.1 | 2.5 | GO:0045494 | photoreceptor cell maintenance(GO:0045494) |
| 0.1 | 2.1 | GO:0014003 | oligodendrocyte development(GO:0014003) |
| 0.1 | 0.3 | GO:0050926 | regulation of positive chemotaxis(GO:0050926) positive regulation of positive chemotaxis(GO:0050927) |
| 0.1 | 2.0 | GO:0051443 | positive regulation of ubiquitin-protein transferase activity(GO:0051443) |
| 0.1 | 0.9 | GO:0032793 | positive regulation of CREB transcription factor activity(GO:0032793) |
| 0.1 | 3.7 | GO:0006270 | DNA replication initiation(GO:0006270) |
| 0.1 | 2.9 | GO:0070979 | protein K11-linked ubiquitination(GO:0070979) |
| 0.1 | 0.8 | GO:1902414 | protein localization to cell junction(GO:1902414) |
| 0.1 | 1.0 | GO:0043374 | CD8-positive, alpha-beta T cell differentiation(GO:0043374) |
| 0.1 | 0.5 | GO:0090292 | nuclear matrix organization(GO:0043578) nuclear matrix anchoring at nuclear membrane(GO:0090292) |
| 0.1 | 0.4 | GO:0046501 | protoporphyrinogen IX metabolic process(GO:0046501) |
| 0.1 | 1.0 | GO:0060117 | auditory receptor cell development(GO:0060117) |
| 0.1 | 0.4 | GO:0051096 | positive regulation of helicase activity(GO:0051096) |
| 0.1 | 1.2 | GO:0070914 | UV-damage excision repair(GO:0070914) |
| 0.1 | 4.5 | GO:0030512 | negative regulation of transforming growth factor beta receptor signaling pathway(GO:0030512) |
| 0.1 | 0.9 | GO:0051450 | myoblast proliferation(GO:0051450) |
| 0.1 | 8.5 | GO:0044782 | cilium organization(GO:0044782) |
| 0.1 | 1.5 | GO:0070536 | protein K63-linked deubiquitination(GO:0070536) |
| 0.1 | 0.4 | GO:0060056 | mammary gland involution(GO:0060056) |
| 0.1 | 1.5 | GO:0034453 | microtubule anchoring(GO:0034453) |
| 0.1 | 1.9 | GO:0002066 | columnar/cuboidal epithelial cell development(GO:0002066) |
| 0.1 | 0.8 | GO:0032463 | negative regulation of protein homooligomerization(GO:0032463) |
| 0.1 | 1.0 | GO:0042573 | retinoic acid metabolic process(GO:0042573) |
| 0.1 | 1.1 | GO:0016973 | poly(A)+ mRNA export from nucleus(GO:0016973) |
| 0.1 | 0.8 | GO:1990440 | positive regulation of transcription from RNA polymerase II promoter in response to endoplasmic reticulum stress(GO:1990440) |
| 0.1 | 0.5 | GO:0035331 | negative regulation of hippo signaling(GO:0035331) |
| 0.1 | 0.5 | GO:0061368 | behavioral response to chemical pain(GO:0061366) behavioral response to formalin induced pain(GO:0061368) |
| 0.1 | 0.5 | GO:0090160 | Golgi to lysosome transport(GO:0090160) |
| 0.1 | 2.6 | GO:0003333 | amino acid transmembrane transport(GO:0003333) |
| 0.1 | 0.9 | GO:0090435 | protein localization to nuclear envelope(GO:0090435) |
| 0.1 | 0.2 | GO:0060075 | regulation of resting membrane potential(GO:0060075) |
| 0.1 | 0.2 | GO:0090400 | stress-induced premature senescence(GO:0090400) |
| 0.1 | 0.3 | GO:0031547 | brain-derived neurotrophic factor receptor signaling pathway(GO:0031547) |
| 0.1 | 0.4 | GO:1902031 | regulation of NADP metabolic process(GO:1902031) |
| 0.1 | 0.1 | GO:0046110 | xanthine metabolic process(GO:0046110) |
| 0.1 | 0.6 | GO:0070072 | vacuolar proton-transporting V-type ATPase complex assembly(GO:0070072) |
| 0.1 | 0.8 | GO:0050667 | homocysteine metabolic process(GO:0050667) |
| 0.1 | 1.6 | GO:0045931 | positive regulation of mitotic cell cycle(GO:0045931) |
| 0.1 | 0.2 | GO:0019336 | phenol-containing compound catabolic process(GO:0019336) |
| 0.1 | 0.1 | GO:0072592 | oxygen metabolic process(GO:0072592) |
| 0.1 | 0.8 | GO:0006614 | SRP-dependent cotranslational protein targeting to membrane(GO:0006614) |
| 0.1 | 1.0 | GO:0051156 | glucose 6-phosphate metabolic process(GO:0051156) |
| 0.1 | 1.1 | GO:0031297 | replication fork processing(GO:0031297) |
| 0.1 | 0.4 | GO:2000253 | positive regulation of feeding behavior(GO:2000253) |
| 0.1 | 1.1 | GO:1902187 | negative regulation of viral release from host cell(GO:1902187) |
| 0.1 | 0.3 | GO:1903055 | positive regulation of extracellular matrix organization(GO:1903055) |
| 0.1 | 0.9 | GO:0021859 | pyramidal neuron differentiation(GO:0021859) pyramidal neuron development(GO:0021860) |
| 0.1 | 1.1 | GO:0000038 | very long-chain fatty acid metabolic process(GO:0000038) |
| 0.1 | 1.0 | GO:0048194 | Golgi vesicle budding(GO:0048194) |
| 0.1 | 2.0 | GO:0000460 | maturation of 5.8S rRNA(GO:0000460) |
| 0.1 | 2.8 | GO:0051310 | metaphase plate congression(GO:0051310) |
| 0.1 | 1.8 | GO:0003301 | physiological muscle hypertrophy(GO:0003298) physiological cardiac muscle hypertrophy(GO:0003301) cell growth involved in cardiac muscle cell development(GO:0061049) |
| 0.1 | 0.2 | GO:0035672 | oligopeptide transmembrane transport(GO:0035672) xenobiotic transport(GO:0042908) |
| 0.1 | 0.3 | GO:0007253 | cytoplasmic sequestering of NF-kappaB(GO:0007253) |
| 0.1 | 1.2 | GO:0002335 | mature B cell differentiation(GO:0002335) |
| 0.1 | 0.7 | GO:0060349 | bone morphogenesis(GO:0060349) |
| 0.1 | 0.5 | GO:0031274 | positive regulation of pseudopodium assembly(GO:0031274) |
| 0.1 | 0.4 | GO:0072189 | ureter development(GO:0072189) |
| 0.1 | 0.3 | GO:0019919 | peptidyl-arginine methylation, to asymmetrical-dimethyl arginine(GO:0019919) |
| 0.1 | 0.1 | GO:0072553 | terminal button organization(GO:0072553) |
| 0.1 | 0.4 | GO:0032836 | glomerular basement membrane development(GO:0032836) |
| 0.1 | 0.4 | GO:0034587 | piRNA metabolic process(GO:0034587) |
| 0.1 | 0.5 | GO:0015838 | amino-acid betaine transport(GO:0015838) |
| 0.1 | 0.5 | GO:0006020 | inositol metabolic process(GO:0006020) |
| 0.1 | 0.6 | GO:1904262 | negative regulation of TORC1 signaling(GO:1904262) |
| 0.1 | 0.3 | GO:0046677 | response to antibiotic(GO:0046677) |
| 0.1 | 1.1 | GO:0030520 | intracellular estrogen receptor signaling pathway(GO:0030520) |
| 0.1 | 2.7 | GO:0006367 | transcription initiation from RNA polymerase II promoter(GO:0006367) |
| 0.1 | 0.7 | GO:0032526 | response to retinoic acid(GO:0032526) |
| 0.1 | 1.5 | GO:0034723 | DNA replication-dependent nucleosome assembly(GO:0006335) DNA replication-dependent nucleosome organization(GO:0034723) |
| 0.1 | 1.3 | GO:0035335 | peptidyl-tyrosine dephosphorylation(GO:0035335) |
| 0.0 | 0.1 | GO:0048014 | Tie signaling pathway(GO:0048014) |
| 0.0 | 0.7 | GO:0033539 | fatty acid beta-oxidation using acyl-CoA dehydrogenase(GO:0033539) |
| 0.0 | 1.1 | GO:0035411 | catenin import into nucleus(GO:0035411) |
| 0.0 | 0.8 | GO:0035315 | hair cell differentiation(GO:0035315) auditory receptor cell differentiation(GO:0042491) |
| 0.0 | 1.9 | GO:0007338 | single fertilization(GO:0007338) |
| 0.0 | 1.5 | GO:0060612 | adipose tissue development(GO:0060612) |
| 0.0 | 0.6 | GO:0000380 | alternative mRNA splicing, via spliceosome(GO:0000380) |
| 0.0 | 0.5 | GO:0070168 | negative regulation of biomineral tissue development(GO:0070168) |
| 0.0 | 0.3 | GO:0035265 | organ growth(GO:0035265) |
| 0.0 | 0.2 | GO:1990504 | dense core granule exocytosis(GO:1990504) |
| 0.0 | 0.4 | GO:0071802 | negative regulation of podosome assembly(GO:0071802) |
| 0.0 | 0.7 | GO:0099500 | synaptic vesicle fusion to presynaptic active zone membrane(GO:0031629) vesicle fusion to plasma membrane(GO:0099500) |
| 0.0 | 0.4 | GO:0071763 | nuclear membrane organization(GO:0071763) |
| 0.0 | 0.7 | GO:0030033 | microvillus assembly(GO:0030033) |
| 0.0 | 1.0 | GO:0033235 | positive regulation of protein sumoylation(GO:0033235) |
| 0.0 | 0.7 | GO:0006120 | mitochondrial electron transport, NADH to ubiquinone(GO:0006120) |
| 0.0 | 0.6 | GO:0000288 | nuclear-transcribed mRNA catabolic process, deadenylation-dependent decay(GO:0000288) |
| 0.0 | 1.8 | GO:0015914 | phospholipid transport(GO:0015914) |
| 0.0 | 0.9 | GO:0010390 | histone monoubiquitination(GO:0010390) |
| 0.0 | 1.0 | GO:0046329 | negative regulation of JNK cascade(GO:0046329) |
| 0.0 | 0.3 | GO:0035235 | ionotropic glutamate receptor signaling pathway(GO:0035235) |
| 0.0 | 0.2 | GO:0021855 | hypothalamus cell migration(GO:0021855) |
| 0.0 | 0.9 | GO:0035036 | binding of sperm to zona pellucida(GO:0007339) sperm-egg recognition(GO:0035036) |
| 0.0 | 0.3 | GO:0030382 | sperm mitochondrion organization(GO:0030382) |
| 0.0 | 0.4 | GO:1900047 | negative regulation of blood coagulation(GO:0030195) negative regulation of hemostasis(GO:1900047) |
| 0.0 | 0.6 | GO:0051482 | positive regulation of cytosolic calcium ion concentration involved in phospholipase C-activating G-protein coupled signaling pathway(GO:0051482) |
| 0.0 | 0.3 | GO:0009409 | response to cold(GO:0009409) |
| 0.0 | 0.0 | GO:2000426 | interleukin-4-mediated signaling pathway(GO:0035771) negative regulation of apoptotic cell clearance(GO:2000426) |
| 0.0 | 2.3 | GO:0051289 | protein homotetramerization(GO:0051289) |
| 0.0 | 0.9 | GO:0006376 | mRNA splice site selection(GO:0006376) |
| 0.0 | 0.5 | GO:0010506 | regulation of autophagy(GO:0010506) |
| 0.0 | 4.3 | GO:0010466 | negative regulation of peptidase activity(GO:0010466) |
| 0.0 | 0.1 | GO:0072176 | nephric duct development(GO:0072176) nephric duct morphogenesis(GO:0072178) |
| 0.0 | 0.4 | GO:0007097 | nuclear migration(GO:0007097) establishment of nucleus localization(GO:0040023) |
| 0.0 | 0.4 | GO:0001833 | inner cell mass cell proliferation(GO:0001833) |
| 0.0 | 0.5 | GO:0051894 | positive regulation of focal adhesion assembly(GO:0051894) positive regulation of cell junction assembly(GO:1901890) |
| 0.0 | 0.7 | GO:0007094 | mitotic spindle assembly checkpoint(GO:0007094) spindle assembly checkpoint(GO:0071173) |
| 0.0 | 0.3 | GO:0051602 | response to electrical stimulus(GO:0051602) |
| 0.0 | 0.5 | GO:0007064 | mitotic sister chromatid cohesion(GO:0007064) |
| 0.0 | 0.1 | GO:1901341 | positive regulation of store-operated calcium channel activity(GO:1901341) |
| 0.0 | 1.9 | GO:0000725 | double-strand break repair via homologous recombination(GO:0000724) recombinational repair(GO:0000725) |
| 0.0 | 0.2 | GO:0048149 | behavioral response to ethanol(GO:0048149) |
| 0.0 | 0.1 | GO:0003431 | growth plate cartilage chondrocyte development(GO:0003431) |
| 0.0 | 0.5 | GO:0050873 | brown fat cell differentiation(GO:0050873) |
| 0.0 | 0.7 | GO:0018345 | protein palmitoylation(GO:0018345) |
| 0.0 | 0.2 | GO:0090070 | positive regulation of ribosome biogenesis(GO:0090070) positive regulation of rRNA processing(GO:2000234) |
| 0.0 | 0.8 | GO:0008333 | endosome to lysosome transport(GO:0008333) |
| 0.0 | 0.0 | GO:1902608 | regulation of large conductance calcium-activated potassium channel activity(GO:1902606) positive regulation of large conductance calcium-activated potassium channel activity(GO:1902608) |
| 0.0 | 0.1 | GO:0002484 | antigen processing and presentation of endogenous peptide antigen via MHC class I via ER pathway(GO:0002484) antigen processing and presentation of endogenous peptide antigen via MHC class I via ER pathway, TAP-dependent(GO:0002485) |
| 0.0 | 0.1 | GO:0097067 | cellular response to thyroid hormone stimulus(GO:0097067) |
| 0.0 | 0.7 | GO:0006829 | zinc II ion transport(GO:0006829) |
| 0.0 | 0.1 | GO:0030828 | positive regulation of cGMP biosynthetic process(GO:0030828) |
| 0.0 | 0.2 | GO:0010650 | positive regulation of cell communication by electrical coupling(GO:0010650) positive regulation of membrane depolarization during cardiac muscle cell action potential(GO:1900827) |
| 0.0 | 0.1 | GO:0006903 | vesicle targeting(GO:0006903) |
| 0.0 | 0.5 | GO:0002244 | hematopoietic progenitor cell differentiation(GO:0002244) |
| 0.0 | 0.2 | GO:0046085 | adenosine metabolic process(GO:0046085) |
| 0.0 | 0.7 | GO:0035904 | aorta development(GO:0035904) |
| 0.0 | 0.2 | GO:0001921 | positive regulation of receptor recycling(GO:0001921) |
| 0.0 | 0.1 | GO:0016560 | protein import into peroxisome matrix, docking(GO:0016560) |
| 0.0 | 0.6 | GO:0000387 | spliceosomal snRNP assembly(GO:0000387) |
| 0.0 | 0.2 | GO:0032331 | negative regulation of chondrocyte differentiation(GO:0032331) |
| 0.0 | 1.7 | GO:0000398 | RNA splicing, via transesterification reactions with bulged adenosine as nucleophile(GO:0000377) mRNA splicing, via spliceosome(GO:0000398) |
| 0.0 | 0.2 | GO:0006895 | Golgi to endosome transport(GO:0006895) |
| 0.0 | 0.3 | GO:0032024 | positive regulation of insulin secretion(GO:0032024) |
| 0.0 | 0.2 | GO:0042407 | cristae formation(GO:0042407) |
| 0.0 | 0.4 | GO:0097320 | membrane tubulation(GO:0097320) |
| 0.0 | 0.4 | GO:0034644 | cellular response to UV(GO:0034644) |
| 0.0 | 0.1 | GO:0043653 | mitochondrial fragmentation involved in apoptotic process(GO:0043653) |
| 0.0 | 0.8 | GO:0016579 | protein deubiquitination(GO:0016579) |
| 0.0 | 0.0 | GO:0010820 | positive regulation of T cell chemotaxis(GO:0010820) |
| 0.0 | 0.1 | GO:0017144 | drug metabolic process(GO:0017144) |
| 0.0 | 0.3 | GO:1900087 | positive regulation of G1/S transition of mitotic cell cycle(GO:1900087) |
| 0.0 | 0.0 | GO:0001180 | transcription initiation from RNA polymerase I promoter for nuclear large rRNA transcript(GO:0001180) |
| 0.0 | 0.9 | GO:0007030 | Golgi organization(GO:0007030) |
| 0.0 | 0.3 | GO:0006826 | iron ion transport(GO:0006826) |
Gene overrepresentation in cellular_component category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 8.9 | 80.2 | GO:0030478 | actin cap(GO:0030478) |
| 2.6 | 38.5 | GO:0042788 | polysomal ribosome(GO:0042788) |
| 2.0 | 8.0 | GO:0070033 | synaptobrevin 2-SNAP-25-syntaxin-1a-complexin II complex(GO:0070033) |
| 1.7 | 15.2 | GO:0097442 | CA3 pyramidal cell dendrite(GO:0097442) |
| 1.6 | 3.3 | GO:0000015 | phosphopyruvate hydratase complex(GO:0000015) |
| 1.5 | 6.2 | GO:0097129 | cyclin D2-CDK4 complex(GO:0097129) |
| 1.5 | 16.1 | GO:0032584 | growth cone membrane(GO:0032584) |
| 1.3 | 4.0 | GO:0038045 | large latent transforming growth factor-beta complex(GO:0038045) |
| 1.3 | 3.8 | GO:0005588 | collagen type V trimer(GO:0005588) |
| 1.0 | 4.1 | GO:0042105 | alpha-beta T cell receptor complex(GO:0042105) |
| 1.0 | 11.1 | GO:1990907 | beta-catenin-TCF complex(GO:1990907) |
| 1.0 | 2.9 | GO:0070557 | PCNA-p21 complex(GO:0070557) |
| 1.0 | 2.9 | GO:0071149 | TEAD-2-YAP complex(GO:0071149) |
| 1.0 | 3.8 | GO:1990421 | subtelomeric heterochromatin(GO:1990421) nuclear subtelomeric heterochromatin(GO:1990707) |
| 0.9 | 2.8 | GO:0002944 | cyclin K-CDK12 complex(GO:0002944) |
| 0.8 | 6.9 | GO:0002177 | manchette(GO:0002177) |
| 0.8 | 3.0 | GO:0060187 | cell pole(GO:0060187) |
| 0.8 | 16.0 | GO:0044300 | cerebellar mossy fiber(GO:0044300) |
| 0.7 | 3.7 | GO:0005968 | Rab-protein geranylgeranyltransferase complex(GO:0005968) |
| 0.6 | 5.1 | GO:0033648 | host intracellular organelle(GO:0033647) host intracellular membrane-bounded organelle(GO:0033648) |
| 0.6 | 1.9 | GO:0005899 | insulin receptor complex(GO:0005899) |
| 0.5 | 1.6 | GO:0097059 | CNTFR-CLCF1 complex(GO:0097059) |
| 0.5 | 6.5 | GO:0097542 | ciliary tip(GO:0097542) |
| 0.5 | 5.4 | GO:0005677 | chromatin silencing complex(GO:0005677) |
| 0.5 | 3.3 | GO:0044352 | pinosome(GO:0044352) macropinosome(GO:0044354) |
| 0.5 | 1.9 | GO:0030981 | cortical microtubule cytoskeleton(GO:0030981) |
| 0.4 | 14.7 | GO:0051233 | spindle midzone(GO:0051233) |
| 0.4 | 6.1 | GO:0000974 | Prp19 complex(GO:0000974) |
| 0.4 | 2.1 | GO:0070937 | CRD-mediated mRNA stability complex(GO:0070937) |
| 0.4 | 7.3 | GO:0034364 | high-density lipoprotein particle(GO:0034364) |
| 0.4 | 2.4 | GO:0043256 | laminin complex(GO:0043256) |
| 0.4 | 2.8 | GO:0005964 | phosphorylase kinase complex(GO:0005964) |
| 0.4 | 1.6 | GO:0045098 | type III intermediate filament(GO:0045098) |
| 0.4 | 4.1 | GO:0034464 | BBSome(GO:0034464) |
| 0.4 | 1.8 | GO:0005658 | alpha DNA polymerase:primase complex(GO:0005658) |
| 0.4 | 8.1 | GO:0030285 | integral component of synaptic vesicle membrane(GO:0030285) intrinsic component of synaptic vesicle membrane(GO:0098563) |
| 0.4 | 5.1 | GO:0005922 | connexon complex(GO:0005922) |
| 0.3 | 18.4 | GO:0097610 | cleavage furrow(GO:0032154) cell surface furrow(GO:0097610) |
| 0.3 | 6.4 | GO:0071004 | U2-type prespliceosome(GO:0071004) |
| 0.3 | 4.3 | GO:0070938 | contractile ring(GO:0070938) |
| 0.3 | 1.9 | GO:0098536 | deuterosome(GO:0098536) |
| 0.3 | 1.5 | GO:1990246 | uniplex complex(GO:1990246) |
| 0.3 | 2.7 | GO:0036056 | filtration diaphragm(GO:0036056) slit diaphragm(GO:0036057) |
| 0.3 | 2.0 | GO:0033503 | HULC complex(GO:0033503) |
| 0.3 | 3.5 | GO:0031597 | cytosolic proteasome complex(GO:0031597) |
| 0.3 | 1.1 | GO:0044307 | dendritic branch(GO:0044307) |
| 0.3 | 1.1 | GO:0031673 | H zone(GO:0031673) |
| 0.3 | 2.1 | GO:0072687 | meiotic spindle(GO:0072687) |
| 0.3 | 6.6 | GO:0005680 | anaphase-promoting complex(GO:0005680) |
| 0.3 | 3.6 | GO:0031616 | spindle pole centrosome(GO:0031616) |
| 0.3 | 4.6 | GO:0005640 | nuclear outer membrane(GO:0005640) |
| 0.3 | 3.8 | GO:0001518 | voltage-gated sodium channel complex(GO:0001518) |
| 0.2 | 1.2 | GO:0005672 | transcription factor TFIIA complex(GO:0005672) |
| 0.2 | 1.4 | GO:0097504 | Gemini of coiled bodies(GO:0097504) |
| 0.2 | 2.1 | GO:0008024 | cyclin/CDK positive transcription elongation factor complex(GO:0008024) |
| 0.2 | 8.3 | GO:0032809 | neuronal cell body membrane(GO:0032809) |
| 0.2 | 2.0 | GO:0097197 | tetraspanin-enriched microdomain(GO:0097197) |
| 0.2 | 21.6 | GO:0005814 | centriole(GO:0005814) |
| 0.2 | 0.6 | GO:1990047 | spindle matrix(GO:1990047) |
| 0.2 | 1.0 | GO:0070876 | SOSS complex(GO:0070876) |
| 0.2 | 0.8 | GO:0098837 | postsynaptic recycling endosome(GO:0098837) |
| 0.2 | 1.8 | GO:0036157 | outer dynein arm(GO:0036157) |
| 0.2 | 1.5 | GO:0097165 | nuclear stress granule(GO:0097165) |
| 0.2 | 2.9 | GO:0032593 | insulin-responsive compartment(GO:0032593) |
| 0.2 | 1.0 | GO:0005587 | collagen type IV trimer(GO:0005587) network-forming collagen trimer(GO:0098642) collagen network(GO:0098645) basement membrane collagen trimer(GO:0098651) |
| 0.2 | 1.5 | GO:0031313 | extrinsic component of endosome membrane(GO:0031313) |
| 0.2 | 0.5 | GO:0020018 | ciliary pocket(GO:0020016) ciliary pocket membrane(GO:0020018) |
| 0.2 | 1.2 | GO:0031465 | Cul4B-RING E3 ubiquitin ligase complex(GO:0031465) |
| 0.2 | 2.6 | GO:0032039 | integrator complex(GO:0032039) |
| 0.2 | 11.6 | GO:0005657 | replication fork(GO:0005657) |
| 0.2 | 3.9 | GO:0005942 | phosphatidylinositol 3-kinase complex(GO:0005942) |
| 0.2 | 1.0 | GO:0070776 | H3 histone acetyltransferase complex(GO:0070775) MOZ/MORF histone acetyltransferase complex(GO:0070776) |
| 0.2 | 1.5 | GO:0005850 | eukaryotic translation initiation factor 2 complex(GO:0005850) |
| 0.2 | 1.2 | GO:0000940 | condensed chromosome outer kinetochore(GO:0000940) |
| 0.2 | 13.1 | GO:0005778 | peroxisomal membrane(GO:0005778) microbody membrane(GO:0031903) |
| 0.2 | 1.1 | GO:0035985 | senescence-associated heterochromatin focus(GO:0035985) |
| 0.2 | 11.5 | GO:0036064 | ciliary basal body(GO:0036064) |
| 0.2 | 3.2 | GO:0097228 | sperm principal piece(GO:0097228) |
| 0.2 | 1.7 | GO:0016342 | catenin complex(GO:0016342) |
| 0.2 | 3.2 | GO:0005721 | pericentric heterochromatin(GO:0005721) |
| 0.2 | 20.9 | GO:0032993 | protein-DNA complex(GO:0032993) |
| 0.1 | 0.4 | GO:0000235 | astral microtubule(GO:0000235) |
| 0.1 | 1.2 | GO:0061574 | ASAP complex(GO:0061574) |
| 0.1 | 1.8 | GO:0072546 | ER membrane protein complex(GO:0072546) |
| 0.1 | 2.4 | GO:0016580 | Sin3 complex(GO:0016580) |
| 0.1 | 1.5 | GO:0000242 | pericentriolar material(GO:0000242) |
| 0.1 | 3.9 | GO:0005876 | spindle microtubule(GO:0005876) |
| 0.1 | 1.3 | GO:0080008 | Cul4-RING E3 ubiquitin ligase complex(GO:0080008) |
| 0.1 | 0.7 | GO:1990454 | L-type voltage-gated calcium channel complex(GO:1990454) |
| 0.1 | 5.0 | GO:0005902 | microvillus(GO:0005902) |
| 0.1 | 1.2 | GO:0035253 | ciliary rootlet(GO:0035253) |
| 0.1 | 0.6 | GO:0032797 | SMN complex(GO:0032797) |
| 0.1 | 0.1 | GO:0016939 | kinesin II complex(GO:0016939) |
| 0.1 | 0.9 | GO:0005833 | hemoglobin complex(GO:0005833) |
| 0.1 | 1.1 | GO:0070545 | PeBoW complex(GO:0070545) |
| 0.1 | 0.4 | GO:0030891 | VCB complex(GO:0030891) |
| 0.1 | 0.8 | GO:0070652 | HAUS complex(GO:0070652) |
| 0.1 | 1.2 | GO:0002080 | acrosomal membrane(GO:0002080) |
| 0.1 | 7.7 | GO:0030864 | cortical actin cytoskeleton(GO:0030864) |
| 0.1 | 1.9 | GO:0030014 | CCR4-NOT complex(GO:0030014) |
| 0.1 | 0.6 | GO:0005664 | origin recognition complex(GO:0000808) nuclear origin of replication recognition complex(GO:0005664) |
| 0.1 | 0.8 | GO:0005786 | signal recognition particle, endoplasmic reticulum targeting(GO:0005786) |
| 0.1 | 0.5 | GO:0034993 | microtubule organizing center attachment site(GO:0034992) LINC complex(GO:0034993) |
| 0.1 | 2.7 | GO:0005815 | microtubule organizing center(GO:0005815) |
| 0.1 | 0.7 | GO:0046581 | intercellular canaliculus(GO:0046581) |
| 0.1 | 0.3 | GO:0071001 | U4/U6 snRNP(GO:0071001) |
| 0.1 | 0.3 | GO:0043083 | synaptic cleft(GO:0043083) |
| 0.1 | 3.5 | GO:0002102 | podosome(GO:0002102) |
| 0.1 | 1.2 | GO:0042405 | nuclear inclusion body(GO:0042405) |
| 0.1 | 1.1 | GO:0097470 | ribbon synapse(GO:0097470) |
| 0.1 | 11.5 | GO:0005813 | centrosome(GO:0005813) |
| 0.1 | 0.9 | GO:0031229 | integral component of nuclear inner membrane(GO:0005639) intrinsic component of nuclear inner membrane(GO:0031229) nuclear membrane part(GO:0044453) |
| 0.1 | 0.6 | GO:0072669 | tRNA-splicing ligase complex(GO:0072669) |
| 0.1 | 0.8 | GO:0005671 | Ada2/Gcn5/Ada3 transcription activator complex(GO:0005671) |
| 0.1 | 2.9 | GO:0016235 | aggresome(GO:0016235) |
| 0.1 | 1.0 | GO:0016010 | dystrophin-associated glycoprotein complex(GO:0016010) glycoprotein complex(GO:0090665) |
| 0.1 | 0.2 | GO:0090498 | extrinsic component of Golgi membrane(GO:0090498) |
| 0.1 | 0.7 | GO:1990316 | ATG1/ULK1 kinase complex(GO:1990316) |
| 0.1 | 1.4 | GO:0046658 | anchored component of plasma membrane(GO:0046658) |
| 0.1 | 8.5 | GO:0090575 | RNA polymerase II transcription factor complex(GO:0090575) |
| 0.1 | 1.1 | GO:0097014 | axoneme(GO:0005930) ciliary plasm(GO:0097014) |
| 0.1 | 0.6 | GO:0005883 | neurofilament(GO:0005883) |
| 0.1 | 1.0 | GO:0000777 | condensed chromosome kinetochore(GO:0000777) |
| 0.1 | 0.3 | GO:0035867 | alphav-beta3 integrin-IGF-1-IGF1R complex(GO:0035867) |
| 0.1 | 2.0 | GO:1990391 | DNA repair complex(GO:1990391) |
| 0.1 | 0.5 | GO:0000127 | transcription factor TFIIIC complex(GO:0000127) |
| 0.1 | 3.0 | GO:0005801 | cis-Golgi network(GO:0005801) |
| 0.1 | 0.7 | GO:0005892 | acetylcholine-gated channel complex(GO:0005892) |
| 0.1 | 0.7 | GO:0017119 | Golgi transport complex(GO:0017119) |
| 0.1 | 2.2 | GO:0005913 | cell-cell adherens junction(GO:0005913) |
| 0.1 | 2.3 | GO:0001669 | acrosomal vesicle(GO:0001669) |
| 0.1 | 0.5 | GO:0031209 | SCAR complex(GO:0031209) |
| 0.1 | 0.3 | GO:0097227 | sperm annulus(GO:0097227) |
| 0.1 | 2.5 | GO:0008180 | COP9 signalosome(GO:0008180) |
| 0.1 | 2.4 | GO:0031463 | Cul3-RING ubiquitin ligase complex(GO:0031463) |
| 0.1 | 0.7 | GO:0000164 | protein phosphatase type 1 complex(GO:0000164) |
| 0.1 | 0.6 | GO:0061700 | GATOR2 complex(GO:0061700) |
| 0.1 | 1.5 | GO:0055038 | recycling endosome membrane(GO:0055038) |
| 0.1 | 2.4 | GO:0000784 | nuclear chromosome, telomeric region(GO:0000784) |
| 0.1 | 2.0 | GO:0005811 | lipid particle(GO:0005811) |
| 0.1 | 0.4 | GO:0042629 | mast cell granule(GO:0042629) |
| 0.0 | 1.4 | GO:0035145 | exon-exon junction complex(GO:0035145) |
| 0.0 | 7.8 | GO:0016323 | basolateral plasma membrane(GO:0016323) |
| 0.0 | 0.8 | GO:0016327 | apicolateral plasma membrane(GO:0016327) |
| 0.0 | 0.6 | GO:0000346 | transcription export complex(GO:0000346) |
| 0.0 | 0.6 | GO:0001891 | phagocytic cup(GO:0001891) |
| 0.0 | 2.7 | GO:0010494 | cytoplasmic stress granule(GO:0010494) |
| 0.0 | 0.2 | GO:0071438 | invadopodium membrane(GO:0071438) |
| 0.0 | 270.7 | GO:0005634 | nucleus(GO:0005634) |
| 0.0 | 0.2 | GO:0061617 | MICOS complex(GO:0061617) |
| 0.0 | 2.1 | GO:0043195 | terminal bouton(GO:0043195) |
| 0.0 | 0.7 | GO:0031307 | integral component of mitochondrial outer membrane(GO:0031307) |
| 0.0 | 0.3 | GO:0008305 | integrin complex(GO:0008305) protein complex involved in cell adhesion(GO:0098636) |
| 0.0 | 0.4 | GO:0014704 | intercalated disc(GO:0014704) |
| 0.0 | 0.6 | GO:0035770 | ribonucleoprotein granule(GO:0035770) |
| 0.0 | 0.7 | GO:0045335 | phagocytic vesicle(GO:0045335) |
| 0.0 | 0.7 | GO:0005788 | endoplasmic reticulum lumen(GO:0005788) |
| 0.0 | 0.7 | GO:0031201 | SNARE complex(GO:0031201) |
| 0.0 | 0.2 | GO:0032839 | dendrite cytoplasm(GO:0032839) |
| 0.0 | 0.7 | GO:0005770 | late endosome(GO:0005770) |
| 0.0 | 0.2 | GO:0005844 | polysome(GO:0005844) |
| 0.0 | 0.0 | GO:0042612 | MHC class I protein complex(GO:0042612) |
| 0.0 | 2.8 | GO:0015630 | microtubule cytoskeleton(GO:0015630) |
Gene overrepresentation in molecular_function category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 5.2 | 61.8 | GO:0050693 | LBD domain binding(GO:0050693) |
| 4.8 | 14.5 | GO:0005110 | frizzled-2 binding(GO:0005110) |
| 3.8 | 30.3 | GO:0001075 | transcription factor activity, RNA polymerase II core promoter sequence-specific binding involved in preinitiation complex assembly(GO:0001075) |
| 3.4 | 37.8 | GO:0044323 | retinoic acid-responsive element binding(GO:0044323) |
| 3.2 | 9.6 | GO:0033680 | ATP-dependent DNA/RNA helicase activity(GO:0033680) |
| 2.5 | 7.6 | GO:0004531 | deoxyribonuclease II activity(GO:0004531) |
| 2.5 | 7.5 | GO:0030226 | apolipoprotein receptor activity(GO:0030226) apolipoprotein A-I receptor activity(GO:0034188) phosphatidylserine-translocating ATPase activity(GO:0090556) |
| 2.4 | 21.2 | GO:0038062 | protein tyrosine kinase collagen receptor activity(GO:0038062) |
| 2.2 | 13.0 | GO:0005042 | netrin receptor activity(GO:0005042) |
| 1.8 | 14.3 | GO:0005519 | cytoskeletal regulatory protein binding(GO:0005519) |
| 1.7 | 6.8 | GO:0047844 | deoxycytidine deaminase activity(GO:0047844) |
| 1.6 | 4.9 | GO:0031871 | proteinase activated receptor binding(GO:0031871) |
| 1.5 | 18.4 | GO:0070742 | C2H2 zinc finger domain binding(GO:0070742) |
| 1.5 | 5.9 | GO:0004921 | interleukin-11 receptor activity(GO:0004921) interleukin-11 binding(GO:0019970) |
| 1.1 | 11.0 | GO:0004691 | cAMP-dependent protein kinase activity(GO:0004691) |
| 1.1 | 40.5 | GO:0017091 | AU-rich element binding(GO:0017091) |
| 1.0 | 6.3 | GO:0070579 | methylcytosine dioxygenase activity(GO:0070579) |
| 1.0 | 4.0 | GO:0050436 | microfibril binding(GO:0050436) |
| 1.0 | 35.4 | GO:0001106 | RNA polymerase II transcription corepressor activity(GO:0001106) |
| 1.0 | 2.9 | GO:0019912 | cyclin-dependent protein kinase activating kinase activity(GO:0019912) |
| 0.9 | 7.4 | GO:0048403 | brain-derived neurotrophic factor binding(GO:0048403) |
| 0.8 | 3.3 | GO:0015410 | manganese-transporting ATPase activity(GO:0015410) |
| 0.8 | 6.1 | GO:0019869 | chloride channel inhibitor activity(GO:0019869) |
| 0.7 | 2.2 | GO:0016174 | NAD(P)H oxidase activity(GO:0016174) |
| 0.7 | 8.4 | GO:0008191 | metalloendopeptidase inhibitor activity(GO:0008191) |
| 0.7 | 3.5 | GO:0005095 | GTPase inhibitor activity(GO:0005095) |
| 0.6 | 11.0 | GO:0003906 | DNA-(apurinic or apyrimidinic site) lyase activity(GO:0003906) |
| 0.6 | 3.2 | GO:0019960 | C-X3-C chemokine binding(GO:0019960) |
| 0.6 | 6.1 | GO:0001055 | RNA polymerase II activity(GO:0001055) |
| 0.6 | 1.8 | GO:0030350 | iron-responsive element binding(GO:0030350) |
| 0.6 | 1.7 | GO:0004994 | somatostatin receptor activity(GO:0004994) |
| 0.6 | 10.8 | GO:0045295 | gamma-catenin binding(GO:0045295) |
| 0.6 | 13.8 | GO:0043395 | heparan sulfate proteoglycan binding(GO:0043395) |
| 0.6 | 3.9 | GO:0046935 | 1-phosphatidylinositol-3-kinase regulator activity(GO:0046935) |
| 0.5 | 17.6 | GO:1990841 | promoter-specific chromatin binding(GO:1990841) |
| 0.5 | 3.8 | GO:0070087 | DNA translocase activity(GO:0015616) chromo shadow domain binding(GO:0070087) |
| 0.5 | 3.2 | GO:0010997 | anaphase-promoting complex binding(GO:0010997) |
| 0.5 | 2.6 | GO:0005113 | patched binding(GO:0005113) |
| 0.5 | 2.6 | GO:0004594 | pantothenate kinase activity(GO:0004594) |
| 0.5 | 1.0 | GO:0010484 | H3 histone acetyltransferase activity(GO:0010484) |
| 0.5 | 2.5 | GO:0005324 | long-chain fatty acid transporter activity(GO:0005324) |
| 0.5 | 1.5 | GO:0048019 | receptor antagonist activity(GO:0048019) |
| 0.5 | 3.0 | GO:0000155 | phosphorelay sensor kinase activity(GO:0000155) |
| 0.5 | 0.5 | GO:0042289 | MHC class II protein binding(GO:0042289) |
| 0.5 | 2.4 | GO:0097322 | 7SK snRNA binding(GO:0097322) |
| 0.5 | 1.9 | GO:0008142 | oxysterol binding(GO:0008142) |
| 0.5 | 1.4 | GO:0017099 | very-long-chain-acyl-CoA dehydrogenase activity(GO:0017099) |
| 0.5 | 9.1 | GO:0001222 | transcription corepressor binding(GO:0001222) |
| 0.5 | 8.6 | GO:0070410 | co-SMAD binding(GO:0070410) |
| 0.4 | 6.7 | GO:0017017 | MAP kinase tyrosine/serine/threonine phosphatase activity(GO:0017017) |
| 0.4 | 1.3 | GO:0043533 | inositol 1,3,4,5 tetrakisphosphate binding(GO:0043533) |
| 0.4 | 5.0 | GO:0004861 | cyclin-dependent protein serine/threonine kinase inhibitor activity(GO:0004861) |
| 0.4 | 3.2 | GO:0019911 | structural constituent of myelin sheath(GO:0019911) |
| 0.4 | 2.8 | GO:0004689 | phosphorylase kinase activity(GO:0004689) |
| 0.4 | 1.6 | GO:0038049 | glucocorticoid receptor activity(GO:0004883) transcription factor activity, ligand-activated RNA polymerase II transcription factor binding(GO:0038049) glucocorticoid-activated RNA polymerase II transcription factor binding transcription factor activity(GO:0038051) |
| 0.4 | 1.2 | GO:0034190 | apolipoprotein receptor binding(GO:0034190) |
| 0.4 | 1.6 | GO:0004897 | ciliary neurotrophic factor receptor activity(GO:0004897) |
| 0.4 | 66.1 | GO:0001078 | transcriptional repressor activity, RNA polymerase II core promoter proximal region sequence-specific binding(GO:0001078) |
| 0.4 | 1.6 | GO:0046790 | virion binding(GO:0046790) |
| 0.4 | 1.6 | GO:1990254 | keratin filament binding(GO:1990254) |
| 0.4 | 1.5 | GO:0001011 | transcription factor activity, sequence-specific DNA binding, RNA polymerase recruiting(GO:0001011) transcription factor activity, TFIIB-class binding(GO:0001087) |
| 0.4 | 1.9 | GO:0043515 | kinetochore binding(GO:0043515) |
| 0.4 | 7.0 | GO:0030215 | semaphorin receptor binding(GO:0030215) |
| 0.4 | 2.6 | GO:0071558 | histone demethylase activity (H3-K27 specific)(GO:0071558) |
| 0.4 | 1.8 | GO:0050265 | RNA uridylyltransferase activity(GO:0050265) |
| 0.4 | 1.5 | GO:0097157 | pre-mRNA intronic binding(GO:0097157) |
| 0.4 | 1.8 | GO:0072542 | protein phosphatase activator activity(GO:0072542) |
| 0.4 | 2.9 | GO:0001134 | transcription factor activity, transcription factor recruiting(GO:0001134) |
| 0.4 | 1.5 | GO:0048408 | epidermal growth factor binding(GO:0048408) |
| 0.4 | 2.2 | GO:0071074 | eukaryotic initiation factor eIF2 binding(GO:0071074) |
| 0.4 | 18.3 | GO:0001102 | RNA polymerase II activating transcription factor binding(GO:0001102) |
| 0.4 | 5.0 | GO:0043495 | protein anchor(GO:0043495) |
| 0.4 | 1.8 | GO:0046974 | histone methyltransferase activity (H3-K9 specific)(GO:0046974) |
| 0.4 | 2.1 | GO:0043559 | insulin binding(GO:0043559) |
| 0.4 | 1.4 | GO:0015254 | glycerol channel activity(GO:0015254) |
| 0.3 | 0.7 | GO:0017116 | single-stranded DNA-dependent ATP-dependent DNA helicase activity(GO:0017116) |
| 0.3 | 3.5 | GO:0004386 | helicase activity(GO:0004386) |
| 0.3 | 2.4 | GO:0036402 | proteasome-activating ATPase activity(GO:0036402) |
| 0.3 | 1.0 | GO:0030368 | interleukin-17 receptor activity(GO:0030368) |
| 0.3 | 1.0 | GO:0004936 | alpha-adrenergic receptor activity(GO:0004936) |
| 0.3 | 0.7 | GO:0019958 | C-X-C chemokine binding(GO:0019958) |
| 0.3 | 1.0 | GO:0035497 | cAMP response element binding(GO:0035497) |
| 0.3 | 21.1 | GO:0003730 | mRNA 3'-UTR binding(GO:0003730) |
| 0.3 | 5.4 | GO:0001103 | RNA polymerase II repressing transcription factor binding(GO:0001103) |
| 0.3 | 0.6 | GO:0097493 | structural molecule activity conferring elasticity(GO:0097493) |
| 0.3 | 2.0 | GO:0001206 | transcriptional repressor activity, RNA polymerase II distal enhancer sequence-specific binding(GO:0001206) |
| 0.3 | 4.0 | GO:0070888 | E-box binding(GO:0070888) |
| 0.3 | 1.4 | GO:0004032 | alditol:NADP+ 1-oxidoreductase activity(GO:0004032) |
| 0.3 | 4.0 | GO:0005251 | delayed rectifier potassium channel activity(GO:0005251) |
| 0.3 | 2.0 | GO:0070915 | lysophosphatidic acid receptor activity(GO:0070915) |
| 0.3 | 1.7 | GO:0086008 | voltage-gated potassium channel activity involved in cardiac muscle cell action potential repolarization(GO:0086008) |
| 0.3 | 0.6 | GO:0061665 | SUMO ligase activity(GO:0061665) |
| 0.3 | 0.8 | GO:0004677 | DNA-dependent protein kinase activity(GO:0004677) |
| 0.3 | 7.4 | GO:0005086 | ARF guanyl-nucleotide exchange factor activity(GO:0005086) |
| 0.3 | 1.1 | GO:0046975 | histone methyltransferase activity (H3-K36 specific)(GO:0046975) |
| 0.3 | 2.1 | GO:0000983 | transcription factor activity, RNA polymerase II core promoter sequence-specific(GO:0000983) |
| 0.3 | 1.1 | GO:0042809 | vitamin D receptor binding(GO:0042809) |
| 0.3 | 1.6 | GO:0004984 | olfactory receptor activity(GO:0004984) |
| 0.3 | 72.4 | GO:0001077 | transcriptional activator activity, RNA polymerase II core promoter proximal region sequence-specific binding(GO:0001077) |
| 0.3 | 1.8 | GO:0004614 | phosphoglucomutase activity(GO:0004614) |
| 0.3 | 0.8 | GO:0016501 | prostacyclin receptor activity(GO:0016501) |
| 0.3 | 1.3 | GO:0042134 | rRNA primary transcript binding(GO:0042134) |
| 0.3 | 3.0 | GO:0097109 | neuroligin family protein binding(GO:0097109) |
| 0.2 | 1.5 | GO:0005544 | calcium-dependent phospholipid binding(GO:0005544) |
| 0.2 | 5.0 | GO:0043422 | protein kinase B binding(GO:0043422) |
| 0.2 | 14.4 | GO:0001158 | enhancer sequence-specific DNA binding(GO:0001158) |
| 0.2 | 1.4 | GO:0047276 | N-acetyllactosaminide 3-alpha-galactosyltransferase activity(GO:0047276) |
| 0.2 | 1.2 | GO:0045545 | syndecan binding(GO:0045545) |
| 0.2 | 1.4 | GO:0035381 | extracellular ATP-gated cation channel activity(GO:0004931) ATP-gated ion channel activity(GO:0035381) |
| 0.2 | 0.7 | GO:0004510 | tryptophan 5-monooxygenase activity(GO:0004510) |
| 0.2 | 1.8 | GO:0015087 | cobalt ion transmembrane transporter activity(GO:0015087) |
| 0.2 | 0.9 | GO:0031721 | hemoglobin alpha binding(GO:0031721) |
| 0.2 | 0.7 | GO:0009383 | rRNA (cytosine-C5-)-methyltransferase activity(GO:0009383) |
| 0.2 | 1.1 | GO:0008172 | S-methyltransferase activity(GO:0008172) |
| 0.2 | 0.9 | GO:0001639 | PLC activating G-protein coupled glutamate receptor activity(GO:0001639) G-protein coupled receptor activity involved in regulation of postsynaptic membrane potential(GO:0099530) |
| 0.2 | 1.1 | GO:0004971 | AMPA glutamate receptor activity(GO:0004971) |
| 0.2 | 1.8 | GO:0008525 | phosphatidylcholine transporter activity(GO:0008525) |
| 0.2 | 3.6 | GO:0004535 | poly(A)-specific ribonuclease activity(GO:0004535) |
| 0.2 | 0.8 | GO:0008131 | primary amine oxidase activity(GO:0008131) |
| 0.2 | 1.0 | GO:0043532 | angiostatin binding(GO:0043532) |
| 0.2 | 2.0 | GO:0003964 | telomerase activity(GO:0003720) RNA-directed DNA polymerase activity(GO:0003964) |
| 0.2 | 6.3 | GO:0008574 | ATP-dependent microtubule motor activity, plus-end-directed(GO:0008574) |
| 0.2 | 1.4 | GO:0032453 | histone demethylase activity (H3-K4 specific)(GO:0032453) |
| 0.2 | 18.7 | GO:0001047 | core promoter binding(GO:0001047) |
| 0.2 | 0.4 | GO:0032794 | GTPase activating protein binding(GO:0032794) |
| 0.2 | 0.4 | GO:0004844 | uracil DNA N-glycosylase activity(GO:0004844) deaminated base DNA N-glycosylase activity(GO:0097506) |
| 0.2 | 2.7 | GO:0005242 | inward rectifier potassium channel activity(GO:0005242) |
| 0.2 | 2.2 | GO:0016805 | dipeptidase activity(GO:0016805) |
| 0.2 | 2.5 | GO:0015174 | basic amino acid transmembrane transporter activity(GO:0015174) |
| 0.2 | 0.7 | GO:0032450 | alpha-1,4-glucosidase activity(GO:0004558) maltose alpha-glucosidase activity(GO:0032450) |
| 0.2 | 2.3 | GO:0032454 | histone demethylase activity (H3-K9 specific)(GO:0032454) |
| 0.2 | 2.1 | GO:0035259 | glucocorticoid receptor binding(GO:0035259) |
| 0.2 | 0.7 | GO:0002162 | dystroglycan binding(GO:0002162) |
| 0.2 | 5.7 | GO:0005112 | Notch binding(GO:0005112) |
| 0.2 | 2.0 | GO:0048273 | mitogen-activated protein kinase p38 binding(GO:0048273) |
| 0.2 | 1.2 | GO:0052629 | phosphatidylinositol-3,5-bisphosphate 3-phosphatase activity(GO:0052629) |
| 0.2 | 3.6 | GO:0070006 | metalloaminopeptidase activity(GO:0070006) |
| 0.2 | 2.8 | GO:0034236 | protein kinase A catalytic subunit binding(GO:0034236) |
| 0.2 | 3.2 | GO:0031402 | sodium ion binding(GO:0031402) |
| 0.2 | 1.1 | GO:0016618 | hydroxypyruvate reductase activity(GO:0016618) glyoxylate reductase (NADP) activity(GO:0030267) |
| 0.2 | 0.8 | GO:0030942 | endoplasmic reticulum signal peptide binding(GO:0030942) |
| 0.2 | 1.4 | GO:0070097 | delta-catenin binding(GO:0070097) |
| 0.2 | 1.5 | GO:0016274 | arginine N-methyltransferase activity(GO:0016273) protein-arginine N-methyltransferase activity(GO:0016274) |
| 0.2 | 2.0 | GO:0016832 | aldehyde-lyase activity(GO:0016832) |
| 0.2 | 2.1 | GO:0008349 | MAP kinase kinase kinase kinase activity(GO:0008349) |
| 0.1 | 0.6 | GO:0051022 | Rho GDP-dissociation inhibitor binding(GO:0051022) |
| 0.1 | 6.3 | GO:0042169 | SH2 domain binding(GO:0042169) |
| 0.1 | 2.4 | GO:0035035 | histone acetyltransferase binding(GO:0035035) |
| 0.1 | 1.0 | GO:1990247 | N6-methyladenosine-containing RNA binding(GO:1990247) |
| 0.1 | 0.6 | GO:0042834 | peptidoglycan binding(GO:0042834) |
| 0.1 | 1.6 | GO:0009922 | fatty acid elongase activity(GO:0009922) 3-oxo-arachidoyl-CoA synthase activity(GO:0102336) 3-oxo-cerotoyl-CoA synthase activity(GO:0102337) 3-oxo-lignoceronyl-CoA synthase activity(GO:0102338) |
| 0.1 | 2.4 | GO:0045505 | dynein intermediate chain binding(GO:0045505) |
| 0.1 | 5.5 | GO:0031624 | ubiquitin conjugating enzyme binding(GO:0031624) |
| 0.1 | 0.8 | GO:0031419 | cobalamin binding(GO:0031419) |
| 0.1 | 2.8 | GO:0008353 | RNA polymerase II carboxy-terminal domain kinase activity(GO:0008353) |
| 0.1 | 0.4 | GO:0005105 | type 1 fibroblast growth factor receptor binding(GO:0005105) |
| 0.1 | 2.1 | GO:0030553 | cGMP binding(GO:0030553) |
| 0.1 | 6.2 | GO:0031491 | nucleosome binding(GO:0031491) |
| 0.1 | 0.5 | GO:0032896 | palmitoyl-CoA 9-desaturase activity(GO:0032896) |
| 0.1 | 1.9 | GO:0004559 | alpha-mannosidase activity(GO:0004559) |
| 0.1 | 3.7 | GO:0003785 | actin monomer binding(GO:0003785) |
| 0.1 | 0.5 | GO:0000832 | inositol hexakisphosphate kinase activity(GO:0000828) inositol hexakisphosphate 5-kinase activity(GO:0000832) inositol hexakisphosphate 1-kinase activity(GO:0052723) inositol hexakisphosphate 3-kinase activity(GO:0052724) |
| 0.1 | 5.6 | GO:0042393 | histone binding(GO:0042393) |
| 0.1 | 0.4 | GO:0004473 | malic enzyme activity(GO:0004470) malate dehydrogenase (decarboxylating) (NAD+) activity(GO:0004471) malate dehydrogenase (decarboxylating) (NADP+) activity(GO:0004473) |
| 0.1 | 0.7 | GO:0005005 | transmembrane-ephrin receptor activity(GO:0005005) |
| 0.1 | 1.4 | GO:0016701 | oxidoreductase activity, acting on single donors with incorporation of molecular oxygen(GO:0016701) oxidoreductase activity, acting on single donors with incorporation of molecular oxygen, incorporation of two atoms of oxygen(GO:0016702) |
| 0.1 | 6.0 | GO:0017022 | myosin binding(GO:0017022) |
| 0.1 | 0.5 | GO:0050733 | RS domain binding(GO:0050733) |
| 0.1 | 3.1 | GO:0001085 | RNA polymerase II transcription factor binding(GO:0001085) |
| 0.1 | 0.6 | GO:0016841 | ammonia-lyase activity(GO:0016841) |
| 0.1 | 0.8 | GO:0035612 | AP-2 adaptor complex binding(GO:0035612) |
| 0.1 | 1.0 | GO:0045125 | bioactive lipid receptor activity(GO:0045125) |
| 0.1 | 1.8 | GO:0015095 | magnesium ion transmembrane transporter activity(GO:0015095) |
| 0.1 | 2.5 | GO:0080025 | phosphatidylinositol-3,5-bisphosphate binding(GO:0080025) |
| 0.1 | 0.8 | GO:0005004 | GPI-linked ephrin receptor activity(GO:0005004) |
| 0.1 | 0.3 | GO:0004830 | tryptophan-tRNA ligase activity(GO:0004830) |
| 0.1 | 4.1 | GO:0043027 | cysteine-type endopeptidase inhibitor activity involved in apoptotic process(GO:0043027) |
| 0.1 | 1.7 | GO:0046527 | glucosyltransferase activity(GO:0046527) |
| 0.1 | 0.4 | GO:0002151 | G-quadruplex RNA binding(GO:0002151) |
| 0.1 | 0.2 | GO:0005375 | copper ion transmembrane transporter activity(GO:0005375) |
| 0.1 | 4.1 | GO:0030544 | Hsp70 protein binding(GO:0030544) |
| 0.1 | 43.0 | GO:0003779 | actin binding(GO:0003779) |
| 0.1 | 1.0 | GO:0047499 | calcium-independent phospholipase A2 activity(GO:0047499) |
| 0.1 | 1.8 | GO:0008519 | ammonium transmembrane transporter activity(GO:0008519) |
| 0.1 | 0.6 | GO:0016712 | oxidoreductase activity, acting on paired donors, with incorporation or reduction of molecular oxygen, reduced flavin or flavoprotein as one donor, and incorporation of one atom of oxygen(GO:0016712) |
| 0.1 | 0.6 | GO:0070728 | leucine binding(GO:0070728) |
| 0.1 | 4.2 | GO:0004867 | serine-type endopeptidase inhibitor activity(GO:0004867) |
| 0.1 | 1.1 | GO:0034237 | protein kinase A regulatory subunit binding(GO:0034237) |
| 0.1 | 2.0 | GO:0003950 | NAD+ ADP-ribosyltransferase activity(GO:0003950) |
| 0.1 | 14.7 | GO:0044822 | mRNA binding(GO:0003729) poly(A) RNA binding(GO:0044822) |
| 0.1 | 1.9 | GO:0005092 | GDP-dissociation inhibitor activity(GO:0005092) |
| 0.1 | 0.4 | GO:0017162 | aryl hydrocarbon receptor binding(GO:0017162) |
| 0.1 | 0.3 | GO:0003846 | 2-acylglycerol O-acyltransferase activity(GO:0003846) |
| 0.1 | 0.9 | GO:0015217 | ATP transmembrane transporter activity(GO:0005347) ADP transmembrane transporter activity(GO:0015217) |
| 0.1 | 3.3 | GO:0051539 | 4 iron, 4 sulfur cluster binding(GO:0051539) |
| 0.1 | 6.0 | GO:0004197 | cysteine-type endopeptidase activity(GO:0004197) |
| 0.1 | 0.6 | GO:0061578 | Lys63-specific deubiquitinase activity(GO:0061578) |
| 0.1 | 1.6 | GO:0071855 | neuropeptide receptor binding(GO:0071855) |
| 0.1 | 0.8 | GO:0015279 | store-operated calcium channel activity(GO:0015279) |
| 0.1 | 2.8 | GO:0003682 | chromatin binding(GO:0003682) |
| 0.1 | 0.4 | GO:0008408 | 3'-5' exonuclease activity(GO:0008408) |
| 0.1 | 0.7 | GO:0022848 | acetylcholine-gated cation channel activity(GO:0022848) |
| 0.1 | 0.3 | GO:0030621 | U4 snRNA binding(GO:0030621) |
| 0.1 | 7.5 | GO:0003714 | transcription corepressor activity(GO:0003714) |
| 0.1 | 0.7 | GO:0031957 | very long-chain fatty acid-CoA ligase activity(GO:0031957) |
| 0.1 | 1.8 | GO:0019707 | protein-cysteine S-palmitoyltransferase activity(GO:0019706) protein-cysteine S-acyltransferase activity(GO:0019707) |
| 0.1 | 3.3 | GO:0019003 | GDP binding(GO:0019003) |
| 0.1 | 1.5 | GO:0004697 | protein kinase C activity(GO:0004697) |
| 0.1 | 0.4 | GO:0003828 | alpha-N-acetylneuraminate alpha-2,8-sialyltransferase activity(GO:0003828) |
| 0.1 | 4.0 | GO:0008186 | RNA helicase activity(GO:0003724) ATP-dependent RNA helicase activity(GO:0004004) RNA-dependent ATPase activity(GO:0008186) |
| 0.1 | 0.4 | GO:0031995 | insulin-like growth factor II binding(GO:0031995) |
| 0.1 | 0.8 | GO:0005172 | vascular endothelial growth factor receptor binding(GO:0005172) |
| 0.1 | 0.2 | GO:0003996 | acyl-CoA ligase activity(GO:0003996) fatty-acyl-CoA synthase activity(GO:0004321) |
| 0.1 | 1.2 | GO:0071889 | 14-3-3 protein binding(GO:0071889) |
| 0.1 | 0.9 | GO:0005528 | macrolide binding(GO:0005527) FK506 binding(GO:0005528) |
| 0.1 | 0.8 | GO:0017025 | TBP-class protein binding(GO:0017025) |
| 0.1 | 0.4 | GO:0015016 | [heparan sulfate]-glucosamine N-sulfotransferase activity(GO:0015016) |
| 0.1 | 0.9 | GO:0008510 | sodium:bicarbonate symporter activity(GO:0008510) |
| 0.1 | 0.2 | GO:0005326 | neurotransmitter transporter activity(GO:0005326) |
| 0.1 | 0.8 | GO:0004190 | aspartic-type endopeptidase activity(GO:0004190) aspartic-type peptidase activity(GO:0070001) |
| 0.0 | 0.1 | GO:0001591 | dopamine neurotransmitter receptor activity, coupled via Gi/Go(GO:0001591) |
| 0.0 | 1.1 | GO:0042813 | Wnt-activated receptor activity(GO:0042813) |
| 0.0 | 3.7 | GO:0015171 | amino acid transmembrane transporter activity(GO:0015171) |
| 0.0 | 2.1 | GO:0097110 | scaffold protein binding(GO:0097110) |
| 0.0 | 0.4 | GO:0036310 | annealing helicase activity(GO:0036310) |
| 0.0 | 1.1 | GO:0018024 | histone-lysine N-methyltransferase activity(GO:0018024) |
| 0.0 | 0.4 | GO:0030296 | protein tyrosine kinase activator activity(GO:0030296) |
| 0.0 | 1.4 | GO:0005201 | extracellular matrix structural constituent(GO:0005201) |
| 0.0 | 0.8 | GO:0031435 | mitogen-activated protein kinase kinase kinase binding(GO:0031435) |
| 0.0 | 0.7 | GO:0070577 | lysine-acetylated histone binding(GO:0070577) |
| 0.0 | 0.2 | GO:0004563 | beta-N-acetylhexosaminidase activity(GO:0004563) |
| 0.0 | 0.3 | GO:0016634 | oxidoreductase activity, acting on the CH-CH group of donors, oxygen as acceptor(GO:0016634) |
| 0.0 | 0.8 | GO:0070273 | phosphatidylinositol-4-phosphate binding(GO:0070273) |
| 0.0 | 0.1 | GO:0043546 | molybdenum ion binding(GO:0030151) molybdopterin cofactor binding(GO:0043546) |
| 0.0 | 0.3 | GO:0071253 | connexin binding(GO:0071253) |
| 0.0 | 0.2 | GO:0001515 | opioid peptide activity(GO:0001515) |
| 0.0 | 0.4 | GO:1990226 | histone methyltransferase binding(GO:1990226) |
| 0.0 | 1.2 | GO:0004364 | glutathione transferase activity(GO:0004364) |
| 0.0 | 0.4 | GO:0017154 | semaphorin receptor activity(GO:0017154) |
| 0.0 | 0.5 | GO:0017049 | GTP-Rho binding(GO:0017049) |
| 0.0 | 0.5 | GO:0043014 | alpha-tubulin binding(GO:0043014) |
| 0.0 | 7.5 | GO:0061659 | ubiquitin protein ligase activity(GO:0061630) ubiquitin-like protein ligase activity(GO:0061659) |
| 0.0 | 1.0 | GO:0050699 | WW domain binding(GO:0050699) |
| 0.0 | 2.1 | GO:0036459 | thiol-dependent ubiquitinyl hydrolase activity(GO:0036459) ubiquitinyl hydrolase activity(GO:0101005) |
| 0.0 | 0.7 | GO:0005385 | zinc ion transmembrane transporter activity(GO:0005385) |
| 0.0 | 0.2 | GO:0000175 | 3'-5'-exoribonuclease activity(GO:0000175) |
| 0.0 | 0.4 | GO:0030374 | ligand-dependent nuclear receptor transcription coactivator activity(GO:0030374) |
| 0.0 | 0.1 | GO:0030171 | voltage-gated proton channel activity(GO:0030171) |
| 0.0 | 0.1 | GO:0004035 | alkaline phosphatase activity(GO:0004035) |
| 0.0 | 0.0 | GO:1990430 | extracellular matrix protein binding(GO:1990430) |
| 0.0 | 0.2 | GO:0022841 | potassium ion leak channel activity(GO:0022841) |
| 0.0 | 0.7 | GO:0001784 | phosphotyrosine binding(GO:0001784) |
| 0.0 | 0.4 | GO:0051019 | mitogen-activated protein kinase binding(GO:0051019) |
| 0.0 | 0.3 | GO:0070064 | proline-rich region binding(GO:0070064) |
| 0.0 | 0.8 | GO:0000978 | RNA polymerase II core promoter proximal region sequence-specific DNA binding(GO:0000978) |
| 0.0 | 0.7 | GO:0005484 | SNAP receptor activity(GO:0005484) |
| 0.0 | 0.1 | GO:0017034 | Rap guanyl-nucleotide exchange factor activity(GO:0017034) |
| 0.0 | 1.7 | GO:0042626 | ATPase activity, coupled to transmembrane movement of substances(GO:0042626) |
| 0.0 | 0.1 | GO:0005052 | peroxisome matrix targeting signal-1 binding(GO:0005052) |
| 0.0 | 0.9 | GO:0043130 | ubiquitin binding(GO:0043130) |
| 0.0 | 0.8 | GO:0020037 | heme binding(GO:0020037) tetrapyrrole binding(GO:0046906) |
| 0.0 | 2.7 | GO:0030246 | carbohydrate binding(GO:0030246) |
| 0.0 | 3.2 | GO:0008134 | transcription factor binding(GO:0008134) |
| 0.0 | 0.5 | GO:0032947 | protein complex scaffold(GO:0032947) |
| 0.0 | 0.3 | GO:0050136 | NADH dehydrogenase (ubiquinone) activity(GO:0008137) NADH dehydrogenase (quinone) activity(GO:0050136) |
| 0.0 | 0.2 | GO:0005234 | extracellular-glutamate-gated ion channel activity(GO:0005234) |
| 0.0 | 0.2 | GO:0030414 | peptidase inhibitor activity(GO:0030414) |
| 0.0 | 0.2 | GO:0031702 | angiotensin receptor binding(GO:0031701) type 1 angiotensin receptor binding(GO:0031702) |
| 0.0 | 0.0 | GO:0097001 | ceramide binding(GO:0097001) |
| 0.0 | 0.4 | GO:0070063 | RNA polymerase binding(GO:0070063) |
| 0.0 | 0.1 | GO:0005507 | copper ion binding(GO:0005507) |
| 0.0 | 0.1 | GO:0008269 | JAK pathway signal transduction adaptor activity(GO:0008269) |
| 0.0 | 0.1 | GO:0017160 | Ral GTPase binding(GO:0017160) |
| 0.0 | 0.3 | GO:0030676 | Rac guanyl-nucleotide exchange factor activity(GO:0030676) |
Gene overrepresentation in C2:CP category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.5 | 4.6 | SA_G2_AND_M_PHASES | Cdc25 activates the cdc2/cyclin B complex to induce the G2/M transition. |
| 0.5 | 15.2 | PID_WNT_SIGNALING_PATHWAY | Wnt signaling network |
| 0.4 | 20.5 | PID_HNF3A_PATHWAY | FOXA1 transcription factor network |
| 0.4 | 11.2 | PID_WNT_NONCANONICAL_PATHWAY | Noncanonical Wnt signaling pathway |
| 0.4 | 1.1 | PID_IGF1_PATHWAY | IGF1 pathway |
| 0.3 | 2.8 | PID_PI3K_PLC_TRK_PATHWAY | Trk receptor signaling mediated by PI3K and PLC-gamma |
| 0.3 | 2.4 | PID_INTEGRIN4_PATHWAY | Alpha6 beta4 integrin-ligand interactions |
| 0.3 | 8.8 | PID_HEDGEHOG_2PATHWAY | Signaling events mediated by the Hedgehog family |
| 0.3 | 14.9 | PID_IL12_2PATHWAY | IL12-mediated signaling events |
| 0.2 | 8.6 | PID_RET_PATHWAY | Signaling events regulated by Ret tyrosine kinase |
| 0.2 | 8.0 | PID_NETRIN_PATHWAY | Netrin-mediated signaling events |
| 0.2 | 3.6 | PID_P38_GAMMA_DELTA_PATHWAY | Signaling mediated by p38-gamma and p38-delta |
| 0.2 | 8.0 | PID_NEPHRIN_NEPH1_PATHWAY | Nephrin/Neph1 signaling in the kidney podocyte |
| 0.2 | 10.6 | PID_LKB1_PATHWAY | LKB1 signaling events |
| 0.2 | 9.9 | PID_THROMBIN_PAR1_PATHWAY | PAR1-mediated thrombin signaling events |
| 0.2 | 2.7 | PID_ERBB2_ERBB3_PATHWAY | ErbB2/ErbB3 signaling events |
| 0.2 | 9.1 | PID_TAP63_PATHWAY | Validated transcriptional targets of TAp63 isoforms |
| 0.2 | 20.5 | PID_BETA_CATENIN_NUC_PATHWAY | Regulation of nuclear beta catenin signaling and target gene transcription |
| 0.2 | 6.6 | PID_RXR_VDR_PATHWAY | RXR and RAR heterodimerization with other nuclear receptor |
| 0.2 | 1.4 | PID_ERBB1_RECEPTOR_PROXIMAL_PATHWAY | EGF receptor (ErbB1) signaling pathway |
| 0.2 | 4.5 | PID_FOXM1_PATHWAY | FOXM1 transcription factor network |
| 0.2 | 10.0 | PID_CASPASE_PATHWAY | Caspase cascade in apoptosis |
| 0.2 | 8.7 | PID_PLK1_PATHWAY | PLK1 signaling events |
| 0.1 | 2.7 | PID_HIF1A_PATHWAY | Hypoxic and oxygen homeostasis regulation of HIF-1-alpha |
| 0.1 | 8.4 | PID_TELOMERASE_PATHWAY | Regulation of Telomerase |
| 0.1 | 2.4 | PID_INSULIN_GLUCOSE_PATHWAY | Insulin-mediated glucose transport |
| 0.1 | 1.8 | ST_GRANULE_CELL_SURVIVAL_PATHWAY | Granule Cell Survival Pathway is a specific case of more general PAC1 Receptor Pathway. |
| 0.1 | 2.2 | PID_DELTA_NP63_PATHWAY | Validated transcriptional targets of deltaNp63 isoforms |
| 0.1 | 2.7 | PID_PS1_PATHWAY | Presenilin action in Notch and Wnt signaling |
| 0.1 | 2.5 | PID_ECADHERIN_NASCENT_AJ_PATHWAY | E-cadherin signaling in the nascent adherens junction |
| 0.1 | 2.7 | PID_AURORA_A_PATHWAY | Aurora A signaling |
| 0.1 | 4.8 | PID_ECADHERIN_STABILIZATION_PATHWAY | Stabilization and expansion of the E-cadherin adherens junction |
| 0.1 | 1.0 | PID_IL8_CXCR2_PATHWAY | IL8- and CXCR2-mediated signaling events |
| 0.1 | 2.9 | PID_FAK_PATHWAY | Signaling events mediated by focal adhesion kinase |
| 0.1 | 3.0 | PID_RHOA_PATHWAY | RhoA signaling pathway |
| 0.1 | 2.0 | ST_ADRENERGIC | Adrenergic Pathway |
| 0.1 | 2.0 | ST_WNT_BETA_CATENIN_PATHWAY | Wnt/beta-catenin Pathway |
| 0.1 | 1.3 | PID_WNT_CANONICAL_PATHWAY | Canonical Wnt signaling pathway |
| 0.1 | 2.2 | PID_CERAMIDE_PATHWAY | Ceramide signaling pathway |
| 0.1 | 1.7 | SA_B_CELL_RECEPTOR_COMPLEXES | Antigen binding to B cell receptors activates protein tyrosine kinases, such as the Src family, which ultimate activate MAP kinases. |
| 0.1 | 1.3 | PID_EPHA_FWDPATHWAY | EPHA forward signaling |
| 0.1 | 2.9 | PID_HNF3B_PATHWAY | FOXA2 and FOXA3 transcription factor networks |
| 0.1 | 1.3 | PID_IL4_2PATHWAY | IL4-mediated signaling events |
| 0.1 | 4.0 | PID_NOTCH_PATHWAY | Notch signaling pathway |
| 0.1 | 1.0 | PID_INTEGRIN5_PATHWAY | Beta5 beta6 beta7 and beta8 integrin cell surface interactions |
| 0.1 | 1.5 | PID_P38_ALPHA_BETA_DOWNSTREAM_PATHWAY | Signaling mediated by p38-alpha and p38-beta |
| 0.1 | 1.5 | PID_P38_ALPHA_BETA_PATHWAY | Regulation of p38-alpha and p38-beta |
| 0.1 | 5.7 | PID_MYC_ACTIV_PATHWAY | Validated targets of C-MYC transcriptional activation |
| 0.1 | 1.9 | PID_AURORA_B_PATHWAY | Aurora B signaling |
| 0.1 | 1.4 | PID_HES_HEY_PATHWAY | Notch-mediated HES/HEY network |
| 0.1 | 1.2 | PID_P38_MK2_PATHWAY | p38 signaling mediated by MAPKAP kinases |
| 0.1 | 1.0 | PID_LPA4_PATHWAY | LPA4-mediated signaling events |
| 0.1 | 0.5 | PID_ARF6_DOWNSTREAM_PATHWAY | Arf6 downstream pathway |
| 0.1 | 6.7 | NABA_ECM_REGULATORS | Genes encoding enzymes and their regulators involved in the remodeling of the extracellular matrix |
| 0.0 | 0.6 | ST_JAK_STAT_PATHWAY | Jak-STAT Pathway |
| 0.0 | 0.6 | PID_NFKAPPAB_CANONICAL_PATHWAY | Canonical NF-kappaB pathway |
| 0.0 | 0.4 | PID_IL1_PATHWAY | IL1-mediated signaling events |
| 0.0 | 0.3 | PID_LYSOPHOSPHOLIPID_PATHWAY | LPA receptor mediated events |
| 0.0 | 0.3 | PID_VEGFR1_PATHWAY | VEGFR1 specific signals |
| 0.0 | 1.5 | PID_SMAD2_3NUCLEAR_PATHWAY | Regulation of nuclear SMAD2/3 signaling |
| 0.0 | 2.3 | PID_P53_DOWNSTREAM_PATHWAY | Direct p53 effectors |
| 0.0 | 0.1 | ST_G_ALPHA_S_PATHWAY | G alpha s Pathway |
| 0.0 | 0.6 | PID_LIS1_PATHWAY | Lissencephaly gene (LIS1) in neuronal migration and development |
| 0.0 | 0.2 | PID_S1P_META_PATHWAY | Sphingosine 1-phosphate (S1P) pathway |
| 0.0 | 0.4 | PID_ARF6_PATHWAY | Arf6 signaling events |
| 0.0 | 0.4 | PID_TCR_PATHWAY | TCR signaling in naïve CD4+ T cells |
| 0.0 | 0.4 | PID_CDC42_PATHWAY | CDC42 signaling events |
| 0.0 | 0.1 | PID_RAC1_PATHWAY | RAC1 signaling pathway |
| 0.0 | 0.2 | PID_TCR_CALCIUM_PATHWAY | Calcium signaling in the CD4+ TCR pathway |
Gene overrepresentation in C2:CP:REACTOME category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 2.0 | 77.0 | REACTOME_SMOOTH_MUSCLE_CONTRACTION | Genes involved in Smooth Muscle Contraction |
| 1.3 | 14.2 | REACTOME_ROLE_OF_SECOND_MESSENGERS_IN_NETRIN1_SIGNALING | Genes involved in Role of second messengers in netrin-1 signaling |
| 0.6 | 16.6 | REACTOME_RNA_POL_III_TRANSCRIPTION_TERMINATION | Genes involved in RNA Polymerase III Transcription Termination |
| 0.6 | 12.0 | REACTOME_ABCA_TRANSPORTERS_IN_LIPID_HOMEOSTASIS | Genes involved in ABCA transporters in lipid homeostasis |
| 0.5 | 3.0 | REACTOME_TRANSLOCATION_OF_ZAP_70_TO_IMMUNOLOGICAL_SYNAPSE | Genes involved in Translocation of ZAP-70 to Immunological synapse |
| 0.5 | 7.1 | REACTOME_RECEPTOR_LIGAND_BINDING_INITIATES_THE_SECOND_PROTEOLYTIC_CLEAVAGE_OF_NOTCH_RECEPTOR | Genes involved in Receptor-ligand binding initiates the second proteolytic cleavage of Notch receptor |
| 0.5 | 5.7 | REACTOME_VIRAL_MESSENGER_RNA_SYNTHESIS | Genes involved in Viral Messenger RNA Synthesis |
| 0.5 | 1.8 | REACTOME_INHIBITION_OF_REPLICATION_INITIATION_OF_DAMAGED_DNA_BY_RB1_E2F1 | Genes involved in Inhibition of replication initiation of damaged DNA by RB1/E2F1 |
| 0.5 | 19.7 | REACTOME_G1_PHASE | Genes involved in G1 Phase |
| 0.4 | 9.6 | REACTOME_HS_GAG_DEGRADATION | Genes involved in HS-GAG degradation |
| 0.4 | 4.8 | REACTOME_GAP_JUNCTION_ASSEMBLY | Genes involved in Gap junction assembly |
| 0.3 | 6.3 | REACTOME_CONVERSION_FROM_APC_C_CDC20_TO_APC_C_CDH1_IN_LATE_ANAPHASE | Genes involved in Conversion from APC/C:Cdc20 to APC/C:Cdh1 in late anaphase |
| 0.3 | 11.2 | REACTOME_DOWNREGULATION_OF_SMAD2_3_SMAD4_TRANSCRIPTIONAL_ACTIVITY | Genes involved in Downregulation of SMAD2/3:SMAD4 transcriptional activity |
| 0.3 | 4.0 | REACTOME_HDL_MEDIATED_LIPID_TRANSPORT | Genes involved in HDL-mediated lipid transport |
| 0.3 | 12.1 | REACTOME_VOLTAGE_GATED_POTASSIUM_CHANNELS | Genes involved in Voltage gated Potassium channels |
| 0.3 | 19.3 | REACTOME_NUCLEAR_RECEPTOR_TRANSCRIPTION_PATHWAY | Genes involved in Nuclear Receptor transcription pathway |
| 0.3 | 2.7 | REACTOME_RNA_POL_III_TRANSCRIPTION_INITIATION_FROM_TYPE_3_PROMOTER | Genes involved in RNA Polymerase III Transcription Initiation From Type 3 Promoter |
| 0.3 | 4.3 | REACTOME_REGULATION_OF_INSULIN_LIKE_GROWTH_FACTOR_IGF_ACTIVITY_BY_INSULIN_LIKE_GROWTH_FACTOR_BINDING_PROTEINS_IGFBPS | Genes involved in Regulation of Insulin-like Growth Factor (IGF) Activity by Insulin-like Growth Factor Binding Proteins (IGFBPs) |
| 0.3 | 0.8 | REACTOME_EICOSANOID_LIGAND_BINDING_RECEPTORS | Genes involved in Eicosanoid ligand-binding receptors |
| 0.3 | 2.8 | REACTOME_SIGNALING_BY_NODAL | Genes involved in Signaling by NODAL |
| 0.2 | 2.0 | REACTOME_P2Y_RECEPTORS | Genes involved in P2Y receptors |
| 0.2 | 1.0 | REACTOME_INFLUENZA_LIFE_CYCLE | Genes involved in Influenza Life Cycle |
| 0.2 | 2.8 | REACTOME_IL_7_SIGNALING | Genes involved in Interleukin-7 signaling |
| 0.2 | 4.8 | REACTOME_OTHER_SEMAPHORIN_INTERACTIONS | Genes involved in Other semaphorin interactions |
| 0.2 | 2.4 | REACTOME_ELONGATION_ARREST_AND_RECOVERY | Genes involved in Elongation arrest and recovery |
| 0.2 | 5.5 | REACTOME_ACTIVATED_NOTCH1_TRANSMITS_SIGNAL_TO_THE_NUCLEUS | Genes involved in Activated NOTCH1 Transmits Signal to the Nucleus |
| 0.2 | 2.8 | REACTOME_SEMA3A_PLEXIN_REPULSION_SIGNALING_BY_INHIBITING_INTEGRIN_ADHESION | Genes involved in SEMA3A-Plexin repulsion signaling by inhibiting Integrin adhesion |
| 0.2 | 2.6 | REACTOME_VITAMIN_B5_PANTOTHENATE_METABOLISM | Genes involved in Vitamin B5 (pantothenate) metabolism |
| 0.2 | 2.2 | REACTOME_GENERATION_OF_SECOND_MESSENGER_MOLECULES | Genes involved in Generation of second messenger molecules |
| 0.2 | 4.2 | REACTOME_CASPASE_MEDIATED_CLEAVAGE_OF_CYTOSKELETAL_PROTEINS | Genes involved in Caspase-mediated cleavage of cytoskeletal proteins |
| 0.2 | 10.0 | REACTOME_DEPOSITION_OF_NEW_CENPA_CONTAINING_NUCLEOSOMES_AT_THE_CENTROMERE | Genes involved in Deposition of New CENPA-containing Nucleosomes at the Centromere |
| 0.2 | 1.4 | REACTOME_PASSIVE_TRANSPORT_BY_AQUAPORINS | Genes involved in Passive Transport by Aquaporins |
| 0.2 | 2.3 | REACTOME_OXYGEN_DEPENDENT_PROLINE_HYDROXYLATION_OF_HYPOXIA_INDUCIBLE_FACTOR_ALPHA | Genes involved in Oxygen-dependent Proline Hydroxylation of Hypoxia-inducible Factor Alpha |
| 0.2 | 4.7 | REACTOME_KINESINS | Genes involved in Kinesins |
| 0.2 | 6.4 | REACTOME_MEIOTIC_SYNAPSIS | Genes involved in Meiotic Synapsis |
| 0.2 | 4.4 | REACTOME_SIGNALING_BY_FGFR1_FUSION_MUTANTS | Genes involved in Signaling by FGFR1 fusion mutants |
| 0.1 | 3.8 | REACTOME_YAP1_AND_WWTR1_TAZ_STIMULATED_GENE_EXPRESSION | Genes involved in YAP1- and WWTR1 (TAZ)-stimulated gene expression |
| 0.1 | 0.9 | REACTOME_PROLACTIN_RECEPTOR_SIGNALING | Genes involved in Prolactin receptor signaling |
| 0.1 | 1.0 | REACTOME_G_ALPHA1213_SIGNALLING_EVENTS | Genes involved in G alpha (12/13) signalling events |
| 0.1 | 0.8 | REACTOME_REGULATION_OF_THE_FANCONI_ANEMIA_PATHWAY | Genes involved in Regulation of the Fanconi anemia pathway |
| 0.1 | 3.8 | REACTOME_DEADENYLATION_OF_MRNA | Genes involved in Deadenylation of mRNA |
| 0.1 | 1.2 | REACTOME_REGULATION_OF_IFNG_SIGNALING | Genes involved in Regulation of IFNG signaling |
| 0.1 | 2.1 | REACTOME_GLYCOGEN_BREAKDOWN_GLYCOGENOLYSIS | Genes involved in Glycogen breakdown (glycogenolysis) |
| 0.1 | 1.0 | REACTOME_E2F_ENABLED_INHIBITION_OF_PRE_REPLICATION_COMPLEX_FORMATION | Genes involved in E2F-enabled inhibition of pre-replication complex formation |
| 0.1 | 0.7 | REACTOME_APOPTOTIC_CLEAVAGE_OF_CELL_ADHESION_PROTEINS | Genes involved in Apoptotic cleavage of cell adhesion proteins |
| 0.1 | 1.7 | REACTOME_SYNTHESIS_SECRETION_AND_INACTIVATION_OF_GIP | Genes involved in Synthesis, Secretion, and Inactivation of Glucose-dependent Insulinotropic Polypeptide (GIP) |
| 0.1 | 1.6 | REACTOME_IL_RECEPTOR_SHC_SIGNALING | Genes involved in Interleukin receptor SHC signaling |
| 0.1 | 3.0 | REACTOME_CGMP_EFFECTS | Genes involved in cGMP effects |
| 0.1 | 1.6 | REACTOME_DEGRADATION_OF_THE_EXTRACELLULAR_MATRIX | Genes involved in Degradation of the extracellular matrix |
| 0.1 | 6.2 | REACTOME_TRANSCRIPTIONAL_REGULATION_OF_WHITE_ADIPOCYTE_DIFFERENTIATION | Genes involved in Transcriptional Regulation of White Adipocyte Differentiation |
| 0.1 | 1.5 | REACTOME_DOWNREGULATION_OF_ERBB2_ERBB3_SIGNALING | Genes involved in Downregulation of ERBB2:ERBB3 signaling |
| 0.1 | 8.1 | REACTOME_LOSS_OF_NLP_FROM_MITOTIC_CENTROSOMES | Genes involved in Loss of Nlp from mitotic centrosomes |
| 0.1 | 8.1 | REACTOME_NRAGE_SIGNALS_DEATH_THROUGH_JNK | Genes involved in NRAGE signals death through JNK |
| 0.1 | 4.0 | REACTOME_INTRINSIC_PATHWAY_FOR_APOPTOSIS | Genes involved in Intrinsic Pathway for Apoptosis |
| 0.1 | 1.0 | REACTOME_ACTIVATION_OF_CHAPERONE_GENES_BY_ATF6_ALPHA | Genes involved in Activation of Chaperone Genes by ATF6-alpha |
| 0.1 | 2.1 | REACTOME_SIGNALING_BY_HIPPO | Genes involved in Signaling by Hippo |
| 0.1 | 2.5 | REACTOME_PHASE1_FUNCTIONALIZATION_OF_COMPOUNDS | Genes involved in Phase 1 - Functionalization of compounds |
| 0.1 | 1.5 | REACTOME_MITOCHONDRIAL_FATTY_ACID_BETA_OXIDATION | Genes involved in Mitochondrial Fatty Acid Beta-Oxidation |
| 0.1 | 0.9 | REACTOME_AMINE_DERIVED_HORMONES | Genes involved in Amine-derived hormones |
| 0.1 | 1.2 | REACTOME_SYNTHESIS_OF_PIPS_AT_THE_EARLY_ENDOSOME_MEMBRANE | Genes involved in Synthesis of PIPs at the early endosome membrane |
| 0.1 | 1.8 | REACTOME_APOPTOTIC_CLEAVAGE_OF_CELLULAR_PROTEINS | Genes involved in Apoptotic cleavage of cellular proteins |
| 0.1 | 0.5 | REACTOME_RNA_POL_III_TRANSCRIPTION | Genes involved in RNA Polymerase III Transcription |
| 0.1 | 0.6 | REACTOME_REGULATION_OF_SIGNALING_BY_CBL | Genes involved in Regulation of signaling by CBL |
| 0.1 | 0.5 | REACTOME_FGFR4_LIGAND_BINDING_AND_ACTIVATION | Genes involved in FGFR4 ligand binding and activation |
| 0.1 | 1.4 | REACTOME_ACTIVATED_TAK1_MEDIATES_P38_MAPK_ACTIVATION | Genes involved in activated TAK1 mediates p38 MAPK activation |
| 0.1 | 0.4 | REACTOME_THE_NLRP3_INFLAMMASOME | Genes involved in The NLRP3 inflammasome |
| 0.1 | 1.1 | REACTOME_ENOS_ACTIVATION_AND_REGULATION | Genes involved in eNOS activation and regulation |
| 0.1 | 2.1 | REACTOME_MEIOSIS | Genes involved in Meiosis |
| 0.1 | 1.2 | REACTOME_AMINE_COMPOUND_SLC_TRANSPORTERS | Genes involved in Amine compound SLC transporters |
| 0.1 | 1.0 | REACTOME_TRAF6_MEDIATED_IRF7_ACTIVATION | Genes involved in TRAF6 mediated IRF7 activation |
| 0.1 | 0.2 | REACTOME_METAL_ION_SLC_TRANSPORTERS | Genes involved in Metal ion SLC transporters |
| 0.0 | 1.2 | REACTOME_RNA_POL_II_TRANSCRIPTION_PRE_INITIATION_AND_PROMOTER_OPENING | Genes involved in RNA Polymerase II Transcription Pre-Initiation And Promoter Opening |
| 0.0 | 1.2 | REACTOME_GLUTATHIONE_CONJUGATION | Genes involved in Glutathione conjugation |
| 0.0 | 1.2 | REACTOME_PI3K_AKT_ACTIVATION | Genes involved in PI3K/AKT activation |
| 0.0 | 0.6 | REACTOME_ACYL_CHAIN_REMODELLING_OF_PC | Genes involved in Acyl chain remodelling of PC |
| 0.0 | 1.9 | REACTOME_MRNA_3_END_PROCESSING | Genes involved in mRNA 3'-end processing |
| 0.0 | 0.7 | REACTOME_SEMA4D_INDUCED_CELL_MIGRATION_AND_GROWTH_CONE_COLLAPSE | Genes involved in Sema4D induced cell migration and growth-cone collapse |
| 0.0 | 1.1 | REACTOME_MYOGENESIS | Genes involved in Myogenesis |
| 0.0 | 0.1 | REACTOME_REGULATION_OF_HYPOXIA_INDUCIBLE_FACTOR_HIF_BY_OXYGEN | Genes involved in Regulation of Hypoxia-inducible Factor (HIF) by Oxygen |
| 0.0 | 0.2 | REACTOME_TANDEM_PORE_DOMAIN_POTASSIUM_CHANNELS | Genes involved in Tandem pore domain potassium channels |
| 0.0 | 0.3 | REACTOME_PURINE_CATABOLISM | Genes involved in Purine catabolism |
| 0.0 | 0.7 | REACTOME_ANTIVIRAL_MECHANISM_BY_IFN_STIMULATED_GENES | Genes involved in Antiviral mechanism by IFN-stimulated genes |
| 0.0 | 1.7 | REACTOME_CLASS_B_2_SECRETIN_FAMILY_RECEPTORS | Genes involved in Class B/2 (Secretin family receptors) |
| 0.0 | 0.7 | REACTOME_ION_TRANSPORT_BY_P_TYPE_ATPASES | Genes involved in Ion transport by P-type ATPases |
| 0.0 | 1.3 | REACTOME_FACTORS_INVOLVED_IN_MEGAKARYOCYTE_DEVELOPMENT_AND_PLATELET_PRODUCTION | Genes involved in Factors involved in megakaryocyte development and platelet production |
| 0.0 | 0.1 | REACTOME_NOD1_2_SIGNALING_PATHWAY | Genes involved in NOD1/2 Signaling Pathway |
| 0.0 | 0.1 | REACTOME_HORMONE_SENSITIVE_LIPASE_HSL_MEDIATED_TRIACYLGLYCEROL_HYDROLYSIS | Genes involved in Hormone-sensitive lipase (HSL)-mediated triacylglycerol hydrolysis |
| 0.0 | 0.2 | REACTOME_IONOTROPIC_ACTIVITY_OF_KAINATE_RECEPTORS | Genes involved in Ionotropic activity of Kainate Receptors |
| 0.0 | 0.1 | REACTOME_SYNTHESIS_SECRETION_AND_DEACYLATION_OF_GHRELIN | Genes involved in Synthesis, Secretion, and Deacylation of Ghrelin |