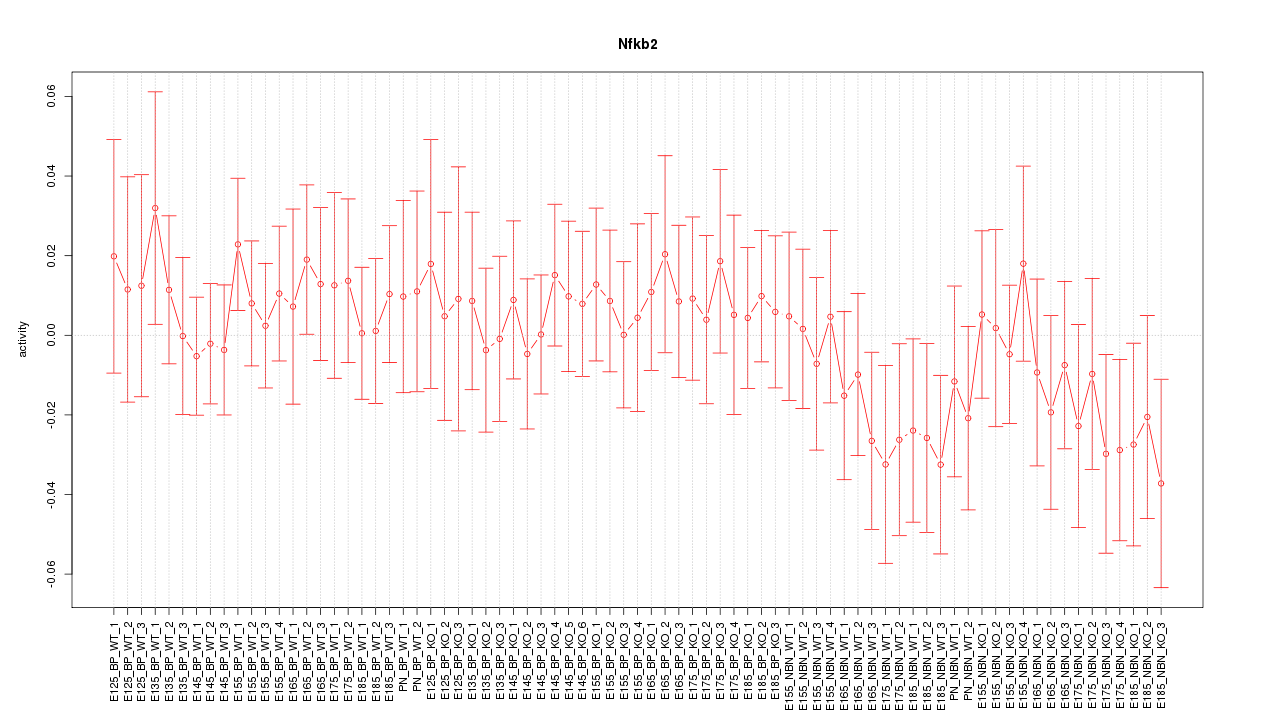
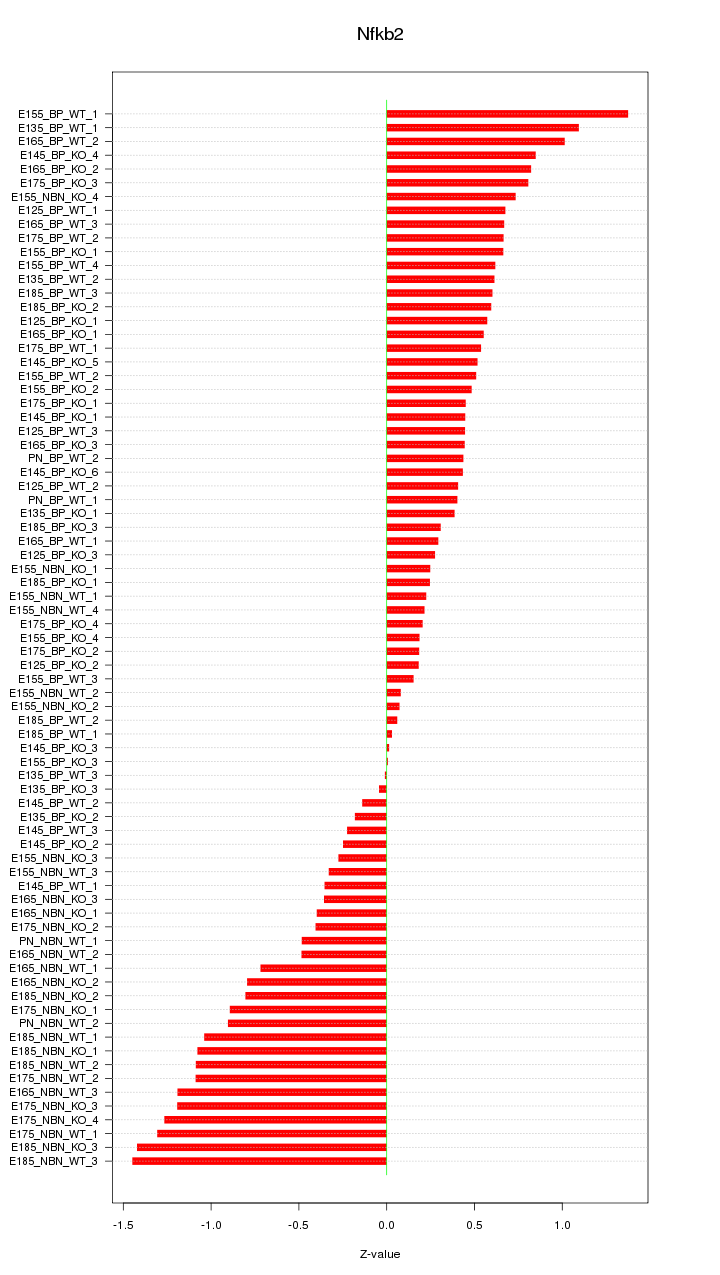
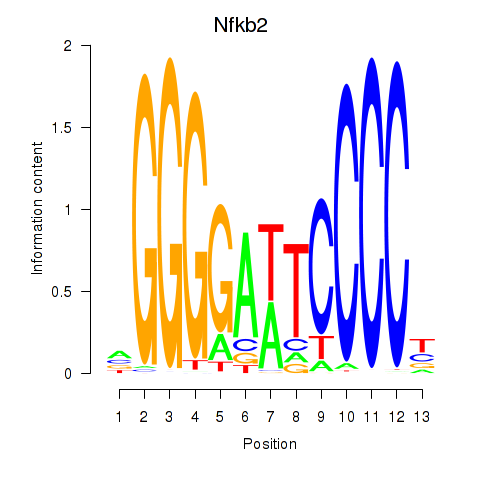
Motif ID: Nfkb2
Z-value: 0.661
Transcription factors associated with Nfkb2:
| Gene Symbol | Entrez ID | Gene Name |
|---|---|---|
| Nfkb2 | ENSMUSG00000025225.8 | Nfkb2 |
Activity-expression correlation:
| Gene Symbol | Promoter | Pearson corr. coef. | P-value | Plot |
|---|---|---|---|---|
| Nfkb2 | mm10_v2_chr19_+_46305682_46305741 | 0.54 | 4.4e-07 | Click! |
Top targets:
Gene overrepresentation in biological_process category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 8.0 | 24.0 | GO:0060598 | dichotomous subdivision of terminal units involved in mammary gland duct morphogenesis(GO:0060598) |
| 1.4 | 15.3 | GO:0042487 | regulation of odontogenesis of dentin-containing tooth(GO:0042487) |
| 1.1 | 10.8 | GO:0051256 | mitotic spindle midzone assembly(GO:0051256) |
| 0.8 | 2.5 | GO:1904059 | regulation of locomotor rhythm(GO:1904059) |
| 0.8 | 3.3 | GO:0002175 | protein localization to paranode region of axon(GO:0002175) |
| 0.8 | 4.1 | GO:0006680 | glucosylceramide catabolic process(GO:0006680) |
| 0.6 | 1.9 | GO:0050713 | negative regulation of interleukin-1 beta secretion(GO:0050713) negative regulation of interferon-gamma secretion(GO:1902714) response to chemokine(GO:1990868) cellular response to chemokine(GO:1990869) |
| 0.5 | 2.7 | GO:1990839 | response to endothelin(GO:1990839) |
| 0.4 | 2.6 | GO:0042535 | positive regulation of tumor necrosis factor biosynthetic process(GO:0042535) |
| 0.4 | 1.2 | GO:0070944 | neutrophil mediated killing of bacterium(GO:0070944) |
| 0.3 | 1.7 | GO:0033140 | negative regulation of peptidyl-serine phosphorylation of STAT protein(GO:0033140) |
| 0.3 | 2.2 | GO:1903003 | positive regulation of protein deubiquitination(GO:1903003) |
| 0.3 | 1.4 | GO:0090038 | negative regulation of protein kinase C signaling(GO:0090038) |
| 0.2 | 2.2 | GO:0042760 | very long-chain fatty acid catabolic process(GO:0042760) |
| 0.2 | 2.2 | GO:0061470 | T follicular helper cell differentiation(GO:0061470) |
| 0.2 | 2.9 | GO:0002467 | germinal center formation(GO:0002467) |
| 0.2 | 2.0 | GO:0019886 | antigen processing and presentation of exogenous peptide antigen via MHC class II(GO:0019886) |
| 0.1 | 1.1 | GO:0033564 | anterior/posterior axon guidance(GO:0033564) |
| 0.1 | 8.0 | GO:0035690 | cellular response to drug(GO:0035690) |
| 0.1 | 3.9 | GO:0002474 | antigen processing and presentation of peptide antigen via MHC class I(GO:0002474) |
| 0.1 | 0.4 | GO:2000286 | receptor internalization involved in canonical Wnt signaling pathway(GO:2000286) |
| 0.1 | 5.8 | GO:0006096 | glycolytic process(GO:0006096) ATP generation from ADP(GO:0006757) |
| 0.1 | 6.8 | GO:0001942 | hair follicle development(GO:0001942) |
| 0.0 | 1.1 | GO:0060384 | innervation(GO:0060384) |
| 0.0 | 1.9 | GO:0060612 | adipose tissue development(GO:0060612) |
| 0.0 | 6.7 | GO:0043547 | positive regulation of GTPase activity(GO:0043547) |
| 0.0 | 5.0 | GO:0006869 | lipid transport(GO:0006869) |
Gene overrepresentation in cellular_component category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.6 | 5.8 | GO:0000015 | phosphopyruvate hydratase complex(GO:0000015) |
| 0.5 | 1.5 | GO:1990682 | CSF1-CSF1R complex(GO:1990682) |
| 0.4 | 2.6 | GO:0097512 | cardiac myofibril(GO:0097512) |
| 0.3 | 3.9 | GO:0042612 | MHC class I protein complex(GO:0042612) |
| 0.3 | 1.9 | GO:0042406 | extrinsic component of endoplasmic reticulum membrane(GO:0042406) |
| 0.3 | 2.0 | GO:0042613 | MHC class II protein complex(GO:0042613) |
| 0.2 | 1.6 | GO:0033256 | I-kappaB/NF-kappaB complex(GO:0033256) |
| 0.2 | 2.8 | GO:0042788 | polysomal ribosome(GO:0042788) |
| 0.1 | 3.3 | GO:0033270 | paranode region of axon(GO:0033270) |
| 0.1 | 10.8 | GO:0000922 | spindle pole(GO:0000922) |
| 0.1 | 8.5 | GO:0017053 | transcriptional repressor complex(GO:0017053) |
| 0.0 | 14.9 | GO:0005578 | proteinaceous extracellular matrix(GO:0005578) |
| 0.0 | 2.2 | GO:0042645 | nucleoid(GO:0009295) mitochondrial nucleoid(GO:0042645) |
| 0.0 | 3.6 | GO:0016363 | nuclear matrix(GO:0016363) |
| 0.0 | 8.1 | GO:0045121 | membrane raft(GO:0045121) membrane microdomain(GO:0098857) |
| 0.0 | 1.9 | GO:0005905 | clathrin-coated pit(GO:0005905) |
| 0.0 | 2.7 | GO:0010008 | endosome membrane(GO:0010008) |
| 0.0 | 3.6 | GO:0005802 | trans-Golgi network(GO:0005802) |
| 0.0 | 1.2 | GO:0000123 | histone acetyltransferase complex(GO:0000123) |
Gene overrepresentation in molecular_function category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 1.0 | 4.1 | GO:0070976 | calcium-independent protein kinase C activity(GO:0004699) TIR domain binding(GO:0070976) |
| 0.7 | 2.2 | GO:0017099 | very-long-chain-acyl-CoA dehydrogenase activity(GO:0017099) |
| 0.7 | 2.7 | GO:0048408 | epidermal growth factor binding(GO:0048408) |
| 0.6 | 5.8 | GO:0004634 | phosphopyruvate hydratase activity(GO:0004634) |
| 0.3 | 13.8 | GO:0005109 | frizzled binding(GO:0005109) |
| 0.3 | 3.9 | GO:0030881 | beta-2-microglobulin binding(GO:0030881) |
| 0.3 | 5.0 | GO:0051861 | glycolipid binding(GO:0051861) |
| 0.2 | 24.0 | GO:0000980 | RNA polymerase II distal enhancer sequence-specific DNA binding(GO:0000980) |
| 0.2 | 10.8 | GO:0008138 | protein tyrosine/serine/threonine phosphatase activity(GO:0008138) |
| 0.2 | 7.7 | GO:0004864 | protein phosphatase inhibitor activity(GO:0004864) |
| 0.1 | 1.9 | GO:0035925 | mRNA 3'-UTR AU-rich region binding(GO:0035925) |
| 0.1 | 2.0 | GO:0042605 | peptide antigen binding(GO:0042605) |
| 0.1 | 2.6 | GO:0044183 | protein binding involved in protein folding(GO:0044183) |
| 0.0 | 0.5 | GO:0019956 | chemokine binding(GO:0019956) |
| 0.0 | 0.3 | GO:0046976 | histone methyltransferase activity (H3-K27 specific)(GO:0046976) |
| 0.0 | 2.2 | GO:0051019 | mitogen-activated protein kinase binding(GO:0051019) |
| 0.0 | 8.5 | GO:0003714 | transcription corepressor activity(GO:0003714) |
| 0.0 | 2.5 | GO:0001046 | core promoter sequence-specific DNA binding(GO:0001046) |
| 0.0 | 10.3 | GO:0005096 | GTPase activator activity(GO:0005096) |
| 0.0 | 8.4 | GO:0001077 | transcriptional activator activity, RNA polymerase II core promoter proximal region sequence-specific binding(GO:0001077) |
| 0.0 | 1.8 | GO:0004725 | protein tyrosine phosphatase activity(GO:0004725) |
| 0.0 | 3.3 | GO:0017124 | SH3 domain binding(GO:0017124) |
Gene overrepresentation in C2:CP category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.4 | 23.8 | PID_TAP63_PATHWAY | Validated transcriptional targets of TAp63 isoforms |
| 0.2 | 1.8 | PID_TCR_RAS_PATHWAY | Ras signaling in the CD4+ TCR pathway |
| 0.1 | 1.5 | SA_MMP_CYTOKINE_CONNECTION | Cytokines can induce activation of matrix metalloproteinases, which degrade extracellular matrix. |
| 0.1 | 13.8 | WNT_SIGNALING | Genes related to Wnt-mediated signal transduction |
| 0.1 | 7.7 | PID_ILK_PATHWAY | Integrin-linked kinase signaling |
| 0.1 | 6.2 | ST_G_ALPHA_I_PATHWAY | G alpha i Pathway |
| 0.1 | 2.9 | ST_GAQ_PATHWAY | G alpha q Pathway |
| 0.1 | 2.6 | PID_P38_MK2_PATHWAY | p38 signaling mediated by MAPKAP kinases |
| 0.1 | 2.2 | PID_HNF3B_PATHWAY | FOXA2 and FOXA3 transcription factor networks |
| 0.0 | 1.1 | NABA_BASEMENT_MEMBRANES | Genes encoding structural components of basement membranes |
| 0.0 | 2.5 | PID_CMYB_PATHWAY | C-MYB transcription factor network |
Gene overrepresentation in C2:CP:REACTOME category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.6 | 10.8 | REACTOME_CONVERSION_FROM_APC_C_CDC20_TO_APC_C_CDH1_IN_LATE_ANAPHASE | Genes involved in Conversion from APC/C:Cdc20 to APC/C:Cdh1 in late anaphase |
| 0.2 | 7.6 | REACTOME_G_ALPHA_Z_SIGNALLING_EVENTS | Genes involved in G alpha (z) signalling events |
| 0.1 | 12.3 | REACTOME_CLASS_B_2_SECRETIN_FAMILY_RECEPTORS | Genes involved in Class B/2 (Secretin family receptors) |
| 0.1 | 5.8 | REACTOME_GLUCONEOGENESIS | Genes involved in Gluconeogenesis |
| 0.1 | 1.9 | REACTOME_NEF_MEDIATED_DOWNREGULATION_OF_MHC_CLASS_I_COMPLEX_CELL_SURFACE_EXPRESSION | Genes involved in Nef mediated downregulation of MHC class I complex cell surface expression |
| 0.1 | 2.2 | REACTOME_MITOCHONDRIAL_FATTY_ACID_BETA_OXIDATION | Genes involved in Mitochondrial Fatty Acid Beta-Oxidation |
| 0.1 | 2.9 | REACTOME_RIP_MEDIATED_NFKB_ACTIVATION_VIA_DAI | Genes involved in RIP-mediated NFkB activation via DAI |
| 0.1 | 2.7 | REACTOME_SIGNAL_ATTENUATION | Genes involved in Signal attenuation |
| 0.1 | 1.1 | REACTOME_ROLE_OF_SECOND_MESSENGERS_IN_NETRIN1_SIGNALING | Genes involved in Role of second messengers in netrin-1 signaling |
| 0.0 | 2.5 | REACTOME_DESTABILIZATION_OF_MRNA_BY_AUF1_HNRNP_D0 | Genes involved in Destabilization of mRNA by AUF1 (hnRNP D0) |
| 0.0 | 3.3 | REACTOME_L1CAM_INTERACTIONS | Genes involved in L1CAM interactions |