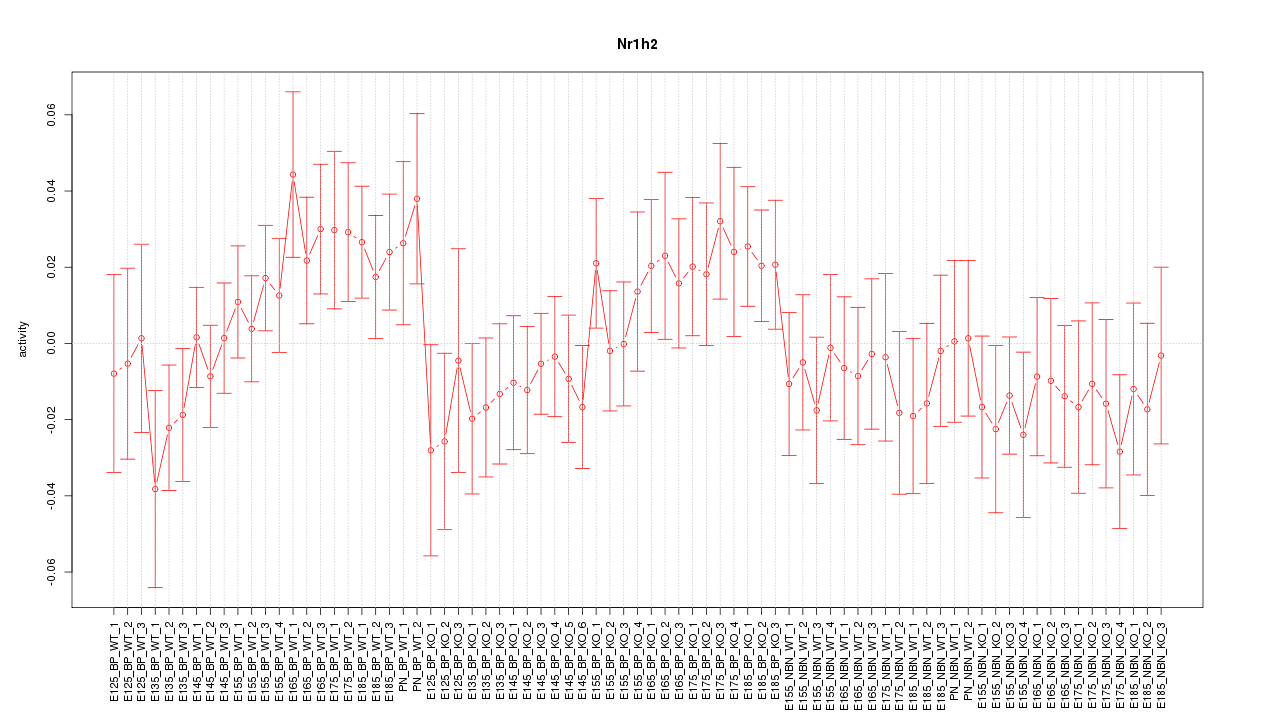
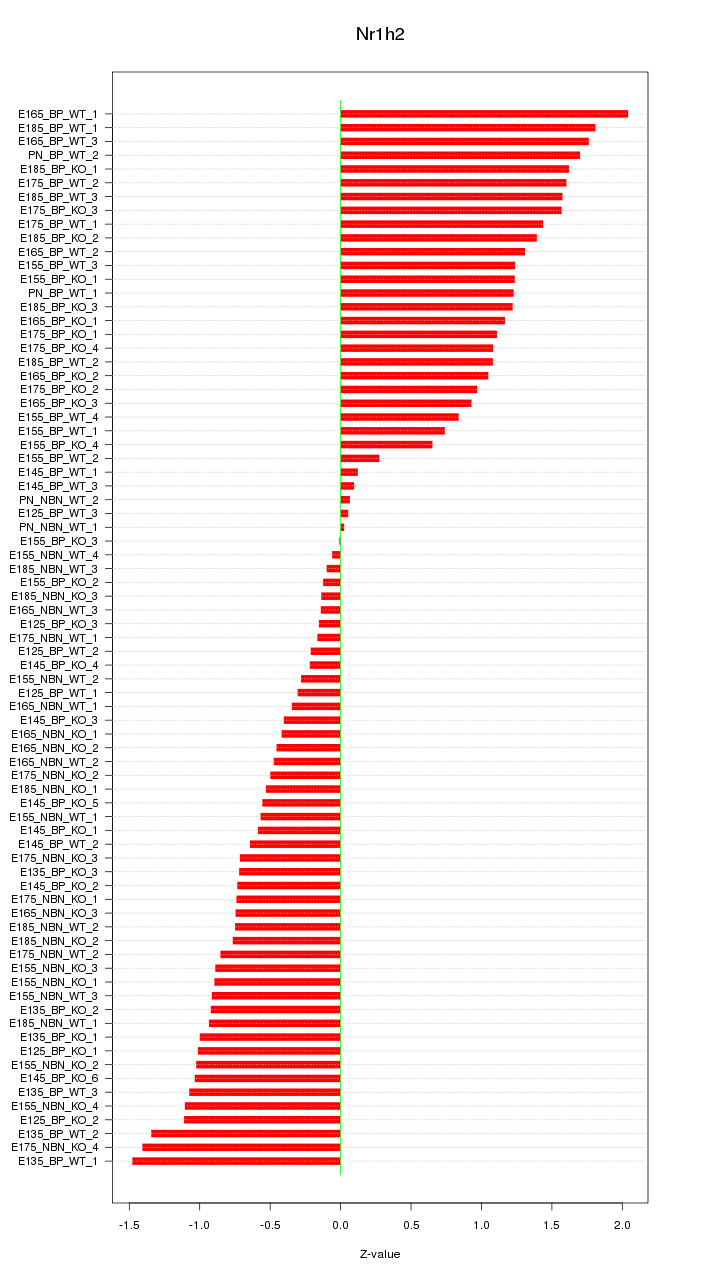
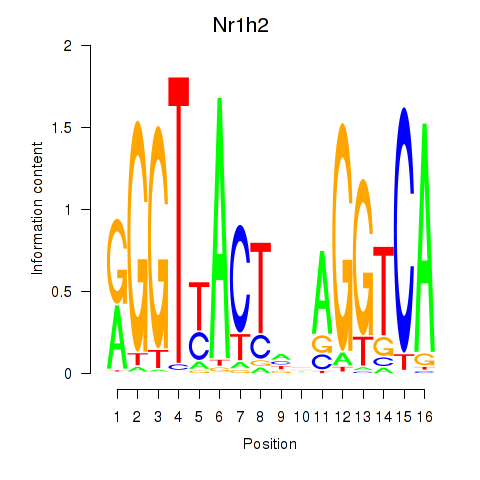
Motif ID: Nr1h2
Z-value: 0.958
Transcription factors associated with Nr1h2:
| Gene Symbol | Entrez ID | Gene Name |
|---|---|---|
| Nr1h2 | ENSMUSG00000060601.6 | Nr1h2 |
Activity-expression correlation:
| Gene Symbol | Promoter | Pearson corr. coef. | P-value | Plot |
|---|---|---|---|---|
| Nr1h2 | mm10_v2_chr7_-_44553901_44553955 | 0.85 | 9.5e-23 | Click! |
Top targets:
Gene overrepresentation in biological_process category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 4.8 | 19.1 | GO:0055099 | response to high density lipoprotein particle(GO:0055099) platelet dense granule organization(GO:0060155) |
| 2.0 | 15.7 | GO:0003062 | regulation of heart rate by chemical signal(GO:0003062) |
| 1.9 | 5.6 | GO:0014891 | striated muscle atrophy(GO:0014891) |
| 1.8 | 8.8 | GO:0046208 | spermine catabolic process(GO:0046208) |
| 1.5 | 5.8 | GO:0090238 | positive regulation of arachidonic acid secretion(GO:0090238) |
| 1.2 | 7.5 | GO:0032196 | regulation of transposition(GO:0010528) negative regulation of transposition(GO:0010529) transposition(GO:0032196) |
| 1.0 | 5.8 | GO:0070779 | D-aspartate transport(GO:0070777) D-aspartate import(GO:0070779) |
| 0.9 | 2.8 | GO:0071603 | endothelial cell-cell adhesion(GO:0071603) |
| 0.9 | 2.6 | GO:0050703 | interleukin-1 alpha secretion(GO:0050703) |
| 0.9 | 3.5 | GO:0072344 | rescue of stalled ribosome(GO:0072344) |
| 0.8 | 3.2 | GO:2000158 | positive regulation of ubiquitin-specific protease activity(GO:2000158) |
| 0.7 | 7.1 | GO:0051256 | mitotic spindle midzone assembly(GO:0051256) |
| 0.7 | 2.1 | GO:0046122 | purine deoxyribonucleoside metabolic process(GO:0046122) |
| 0.6 | 1.9 | GO:0015920 | lipopolysaccharide transport(GO:0015920) |
| 0.6 | 3.9 | GO:0002317 | plasma cell differentiation(GO:0002317) |
| 0.6 | 1.9 | GO:2000338 | toll-like receptor 5 signaling pathway(GO:0034146) positive regulation of interleukin-6 biosynthetic process(GO:0045410) chemokine (C-X-C motif) ligand 1 production(GO:0072566) regulation of chemokine (C-X-C motif) ligand 1 production(GO:2000338) |
| 0.6 | 14.0 | GO:0010866 | regulation of triglyceride biosynthetic process(GO:0010866) |
| 0.5 | 2.1 | GO:0016340 | calcium-dependent cell-matrix adhesion(GO:0016340) |
| 0.5 | 9.1 | GO:0002115 | store-operated calcium entry(GO:0002115) |
| 0.4 | 2.6 | GO:0002002 | regulation of angiotensin levels in blood(GO:0002002) |
| 0.4 | 4.7 | GO:0051574 | positive regulation of histone H3-K9 methylation(GO:0051574) |
| 0.3 | 1.0 | GO:0070543 | response to linoleic acid(GO:0070543) |
| 0.3 | 1.3 | GO:0006558 | L-phenylalanine metabolic process(GO:0006558) erythrose 4-phosphate/phosphoenolpyruvate family amino acid metabolic process(GO:1902221) |
| 0.3 | 1.9 | GO:0000707 | meiotic DNA recombinase assembly(GO:0000707) |
| 0.3 | 1.3 | GO:0002949 | tRNA threonylcarbamoyladenosine modification(GO:0002949) |
| 0.3 | 14.6 | GO:0046676 | negative regulation of insulin secretion(GO:0046676) |
| 0.3 | 6.9 | GO:0044381 | glucose import in response to insulin stimulus(GO:0044381) |
| 0.3 | 1.3 | GO:0018242 | protein O-linked glycosylation via serine(GO:0018242) |
| 0.3 | 4.7 | GO:0021684 | cerebellar granular layer formation(GO:0021684) cerebellar granule cell differentiation(GO:0021707) |
| 0.2 | 4.0 | GO:0016486 | peptide hormone processing(GO:0016486) |
| 0.2 | 1.2 | GO:0001575 | globoside metabolic process(GO:0001575) |
| 0.2 | 1.2 | GO:0006777 | Mo-molybdopterin cofactor biosynthetic process(GO:0006777) Mo-molybdopterin cofactor metabolic process(GO:0019720) |
| 0.2 | 1.9 | GO:0006265 | DNA topological change(GO:0006265) |
| 0.2 | 0.7 | GO:0009051 | pentose-phosphate shunt, oxidative branch(GO:0009051) |
| 0.2 | 1.4 | GO:0002934 | desmosome organization(GO:0002934) |
| 0.2 | 4.7 | GO:0001886 | endothelial cell morphogenesis(GO:0001886) |
| 0.2 | 1.6 | GO:0040016 | embryonic cleavage(GO:0040016) |
| 0.2 | 2.8 | GO:0045723 | positive regulation of fatty acid biosynthetic process(GO:0045723) |
| 0.2 | 1.2 | GO:0060050 | positive regulation of protein glycosylation(GO:0060050) |
| 0.1 | 2.0 | GO:0006654 | phosphatidic acid biosynthetic process(GO:0006654) |
| 0.1 | 1.4 | GO:2000671 | regulation of motor neuron apoptotic process(GO:2000671) |
| 0.1 | 2.5 | GO:0072112 | renal filtration cell differentiation(GO:0061318) glomerular visceral epithelial cell differentiation(GO:0072112) glomerular epithelial cell differentiation(GO:0072311) |
| 0.1 | 0.9 | GO:0090074 | negative regulation of protein homodimerization activity(GO:0090074) |
| 0.1 | 0.9 | GO:0030242 | pexophagy(GO:0030242) |
| 0.1 | 1.1 | GO:0006398 | mRNA 3'-end processing by stem-loop binding and cleavage(GO:0006398) |
| 0.1 | 6.2 | GO:0060487 | lung epithelial cell differentiation(GO:0060487) |
| 0.1 | 8.0 | GO:0070527 | platelet aggregation(GO:0070527) |
| 0.1 | 1.4 | GO:0015937 | coenzyme A biosynthetic process(GO:0015937) |
| 0.1 | 2.1 | GO:0060009 | Sertoli cell development(GO:0060009) |
| 0.1 | 1.9 | GO:0030575 | nuclear body organization(GO:0030575) |
| 0.1 | 1.1 | GO:0046548 | ovulation(GO:0030728) retinal rod cell development(GO:0046548) |
| 0.1 | 6.6 | GO:0006073 | glycogen metabolic process(GO:0005977) cellular glucan metabolic process(GO:0006073) glucan metabolic process(GO:0044042) |
| 0.1 | 0.5 | GO:0006013 | mannose metabolic process(GO:0006013) |
| 0.1 | 2.5 | GO:0090307 | mitotic spindle assembly(GO:0090307) |
| 0.0 | 1.7 | GO:0009988 | cell-cell recognition(GO:0009988) |
| 0.0 | 0.6 | GO:0080182 | histone H3-K4 trimethylation(GO:0080182) |
| 0.0 | 0.4 | GO:0048280 | vesicle fusion with Golgi apparatus(GO:0048280) |
| 0.0 | 0.1 | GO:1990022 | RNA polymerase II complex import to nucleus(GO:0044376) RNA polymerase III complex localization to nucleus(GO:1990022) |
| 0.0 | 1.9 | GO:0033045 | regulation of sister chromatid segregation(GO:0033045) |
| 0.0 | 1.2 | GO:0045600 | positive regulation of fat cell differentiation(GO:0045600) |
| 0.0 | 0.8 | GO:1901998 | toxin transport(GO:1901998) |
| 0.0 | 0.5 | GO:0006891 | intra-Golgi vesicle-mediated transport(GO:0006891) |
| 0.0 | 0.2 | GO:0071712 | ER-associated misfolded protein catabolic process(GO:0071712) |
| 0.0 | 3.7 | GO:0000398 | RNA splicing, via transesterification reactions with bulged adenosine as nucleophile(GO:0000377) mRNA splicing, via spliceosome(GO:0000398) |
Gene overrepresentation in cellular_component category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 1.1 | 19.1 | GO:0034364 | high-density lipoprotein particle(GO:0034364) |
| 0.9 | 6.6 | GO:0005964 | phosphorylase kinase complex(GO:0005964) |
| 0.9 | 5.6 | GO:0031265 | CD95 death-inducing signaling complex(GO:0031265) |
| 0.5 | 2.5 | GO:0031523 | Myb complex(GO:0031523) |
| 0.5 | 1.8 | GO:0045098 | type III intermediate filament(GO:0045098) |
| 0.4 | 6.9 | GO:0032593 | insulin-responsive compartment(GO:0032593) |
| 0.4 | 1.9 | GO:0033063 | Rad51B-Rad51C-Rad51D-XRCC2 complex(GO:0033063) |
| 0.3 | 1.2 | GO:0097447 | dendritic tree(GO:0097447) |
| 0.3 | 1.3 | GO:0000408 | EKC/KEOPS complex(GO:0000408) |
| 0.2 | 1.9 | GO:0042382 | paraspeckles(GO:0042382) |
| 0.2 | 4.7 | GO:0035098 | ESC/E(Z) complex(GO:0035098) |
| 0.1 | 5.8 | GO:0030673 | axolemma(GO:0030673) |
| 0.1 | 2.1 | GO:0043196 | varicosity(GO:0043196) |
| 0.1 | 0.9 | GO:1990316 | ATG1/ULK1 kinase complex(GO:1990316) |
| 0.1 | 7.1 | GO:0072686 | mitotic spindle(GO:0072686) |
| 0.1 | 3.5 | GO:0000315 | organellar large ribosomal subunit(GO:0000315) mitochondrial large ribosomal subunit(GO:0005762) |
| 0.1 | 3.2 | GO:0008023 | transcription elongation factor complex(GO:0008023) |
| 0.1 | 1.1 | GO:0005847 | mRNA cleavage and polyadenylation specificity factor complex(GO:0005847) |
| 0.1 | 5.9 | GO:0005814 | centriole(GO:0005814) |
| 0.0 | 9.1 | GO:0030426 | growth cone(GO:0030426) |
| 0.0 | 1.3 | GO:0005788 | endoplasmic reticulum lumen(GO:0005788) |
| 0.0 | 0.2 | GO:0065010 | extracellular membrane-bounded organelle(GO:0065010) |
| 0.0 | 2.3 | GO:0017053 | transcriptional repressor complex(GO:0017053) |
| 0.0 | 0.8 | GO:0030131 | clathrin adaptor complex(GO:0030131) |
| 0.0 | 1.9 | GO:0032587 | ruffle membrane(GO:0032587) |
| 0.0 | 0.4 | GO:0012507 | ER to Golgi transport vesicle membrane(GO:0012507) |
| 0.0 | 5.0 | GO:0030141 | secretory granule(GO:0030141) |
| 0.0 | 8.0 | GO:0005925 | focal adhesion(GO:0005925) |
| 0.0 | 4.0 | GO:0031965 | nuclear membrane(GO:0031965) |
| 0.0 | 1.0 | GO:0031526 | brush border membrane(GO:0031526) |
| 0.0 | 13.8 | GO:0031410 | cytoplasmic vesicle(GO:0031410) |
| 0.0 | 1.9 | GO:0000793 | condensed chromosome(GO:0000793) |
| 0.0 | 7.4 | GO:0005815 | microtubule organizing center(GO:0005815) |
| 0.0 | 0.5 | GO:0030173 | integral component of Golgi membrane(GO:0030173) |
| 0.0 | 2.3 | GO:0005938 | cell cortex(GO:0005938) |
| 0.0 | 0.6 | GO:0035097 | histone methyltransferase complex(GO:0035097) |
Gene overrepresentation in molecular_function category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 6.4 | 19.1 | GO:0030226 | apolipoprotein receptor activity(GO:0030226) apolipoprotein A-I receptor activity(GO:0034188) phosphatidylserine-translocating ATPase activity(GO:0090556) |
| 1.9 | 7.5 | GO:0047844 | deoxycytidine deaminase activity(GO:0047844) |
| 1.7 | 6.9 | GO:0055056 | D-glucose transmembrane transporter activity(GO:0055056) |
| 1.6 | 4.7 | GO:0004145 | diamine N-acetyltransferase activity(GO:0004145) |
| 1.3 | 4.0 | GO:0046592 | polyamine oxidase activity(GO:0046592) spermine:oxygen oxidoreductase (spermidine-forming) activity(GO:0052901) |
| 1.0 | 5.8 | GO:0015501 | glutamate:sodium symporter activity(GO:0015501) |
| 0.9 | 6.6 | GO:0004689 | phosphorylase kinase activity(GO:0004689) |
| 0.9 | 3.5 | GO:0004045 | aminoacyl-tRNA hydrolase activity(GO:0004045) |
| 0.8 | 3.2 | GO:0035800 | deubiquitinase activator activity(GO:0035800) |
| 0.8 | 9.1 | GO:0015279 | store-operated calcium channel activity(GO:0015279) |
| 0.7 | 2.1 | GO:0004731 | purine-nucleoside phosphorylase activity(GO:0004731) |
| 0.6 | 1.9 | GO:0071723 | lipopeptide binding(GO:0071723) |
| 0.6 | 5.6 | GO:0097200 | cysteine-type endopeptidase activity involved in execution phase of apoptosis(GO:0097200) |
| 0.6 | 5.8 | GO:0047499 | calcium-independent phospholipase A2 activity(GO:0047499) |
| 0.6 | 2.8 | GO:0070330 | aromatase activity(GO:0070330) |
| 0.4 | 1.3 | GO:0030158 | protein xylosyltransferase activity(GO:0030158) |
| 0.4 | 2.6 | GO:0050786 | RAGE receptor binding(GO:0050786) |
| 0.4 | 1.3 | GO:0061711 | N(6)-L-threonylcarbamoyladenine synthase(GO:0061711) |
| 0.4 | 1.9 | GO:0003917 | DNA topoisomerase type I activity(GO:0003917) |
| 0.4 | 1.9 | GO:0035662 | Toll-like receptor 4 binding(GO:0035662) |
| 0.4 | 2.6 | GO:0015016 | [heparan sulfate]-glucosamine N-sulfotransferase activity(GO:0015016) |
| 0.3 | 1.0 | GO:0070892 | lipoteichoic acid receptor activity(GO:0070892) |
| 0.3 | 2.5 | GO:0001135 | transcription factor activity, RNA polymerase II transcription factor recruiting(GO:0001135) |
| 0.3 | 1.4 | GO:0008545 | JUN kinase kinase activity(GO:0008545) |
| 0.3 | 1.9 | GO:0000150 | recombinase activity(GO:0000150) |
| 0.3 | 15.7 | GO:0004879 | RNA polymerase II transcription factor activity, ligand-activated sequence-specific DNA binding(GO:0004879) transcription factor activity, direct ligand regulated sequence-specific DNA binding(GO:0098531) |
| 0.2 | 0.7 | GO:0017057 | 6-phosphogluconolactonase activity(GO:0017057) |
| 0.2 | 1.4 | GO:0004169 | dolichyl-phosphate-mannose-protein mannosyltransferase activity(GO:0004169) |
| 0.2 | 1.1 | GO:0004095 | carnitine O-palmitoyltransferase activity(GO:0004095) O-palmitoyltransferase activity(GO:0016416) |
| 0.2 | 1.1 | GO:0047184 | 1-acylglycerophosphocholine O-acyltransferase activity(GO:0047184) |
| 0.1 | 7.1 | GO:0008138 | protein tyrosine/serine/threonine phosphatase activity(GO:0008138) |
| 0.1 | 14.6 | GO:0019905 | syntaxin binding(GO:0019905) |
| 0.1 | 1.8 | GO:0019215 | intermediate filament binding(GO:0019215) |
| 0.1 | 1.2 | GO:0009922 | fatty acid elongase activity(GO:0009922) 3-oxo-arachidoyl-CoA synthase activity(GO:0102336) 3-oxo-cerotoyl-CoA synthase activity(GO:0102337) 3-oxo-lignoceronyl-CoA synthase activity(GO:0102338) |
| 0.1 | 4.7 | GO:0032452 | histone demethylase activity(GO:0032452) |
| 0.1 | 1.2 | GO:0015643 | toxic substance binding(GO:0015643) |
| 0.1 | 0.8 | GO:0030306 | ADP-ribosylation factor binding(GO:0030306) |
| 0.1 | 1.1 | GO:0051393 | alpha-actinin binding(GO:0051393) |
| 0.1 | 0.9 | GO:0048273 | mitogen-activated protein kinase p38 binding(GO:0048273) |
| 0.1 | 1.1 | GO:0008409 | 5'-3' exonuclease activity(GO:0008409) |
| 0.1 | 1.4 | GO:0016881 | acid-amino acid ligase activity(GO:0016881) |
| 0.0 | 1.0 | GO:0003841 | 1-acylglycerol-3-phosphate O-acyltransferase activity(GO:0003841) |
| 0.0 | 0.5 | GO:0015924 | mannosyl-oligosaccharide mannosidase activity(GO:0015924) |
| 0.0 | 0.2 | GO:0004035 | alkaline phosphatase activity(GO:0004035) |
| 0.0 | 0.7 | GO:0004198 | calcium-dependent cysteine-type endopeptidase activity(GO:0004198) |
| 0.0 | 1.3 | GO:0004364 | glutathione transferase activity(GO:0004364) |
| 0.0 | 3.8 | GO:0005178 | integrin binding(GO:0005178) |
| 0.0 | 1.2 | GO:0005109 | frizzled binding(GO:0005109) |
| 0.0 | 1.2 | GO:0051539 | 4 iron, 4 sulfur cluster binding(GO:0051539) |
| 0.0 | 2.6 | GO:0051082 | unfolded protein binding(GO:0051082) |
| 0.0 | 1.9 | GO:0005262 | calcium channel activity(GO:0005262) |
| 0.0 | 2.3 | GO:0003697 | single-stranded DNA binding(GO:0003697) |
| 0.0 | 0.4 | GO:0043236 | laminin binding(GO:0043236) |
| 0.0 | 1.4 | GO:0008186 | RNA helicase activity(GO:0003724) ATP-dependent RNA helicase activity(GO:0004004) RNA-dependent ATPase activity(GO:0008186) |
| 0.0 | 1.6 | GO:0008022 | protein C-terminus binding(GO:0008022) |
| 0.0 | 0.6 | GO:0000979 | RNA polymerase II core promoter sequence-specific DNA binding(GO:0000979) |
| 0.0 | 0.4 | GO:0005484 | SNAP receptor activity(GO:0005484) |
| 0.0 | 5.9 | GO:0001228 | transcriptional activator activity, RNA polymerase II transcription regulatory region sequence-specific binding(GO:0001228) |
| 0.0 | 11.6 | GO:0042803 | protein homodimerization activity(GO:0042803) |
| 0.0 | 0.9 | GO:0032947 | protein complex scaffold(GO:0032947) |
| 0.0 | 1.6 | GO:0030674 | protein binding, bridging(GO:0030674) |
| 0.0 | 1.1 | GO:0043130 | ubiquitin binding(GO:0043130) |
Gene overrepresentation in C2:CP category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.9 | 34.8 | PID_RXR_VDR_PATHWAY | RXR and RAR heterodimerization with other nuclear receptor |
| 0.7 | 5.6 | SA_FAS_SIGNALING | The TNF-type receptor Fas induces apoptosis on ligand binding. |
| 0.2 | 6.9 | PID_INSULIN_GLUCOSE_PATHWAY | Insulin-mediated glucose transport |
| 0.2 | 4.7 | PID_INTEGRIN_A9B1_PATHWAY | Alpha9 beta1 integrin signaling events |
| 0.2 | 9.1 | PID_CD8_TCR_PATHWAY | TCR signaling in naïve CD8+ T cells |
| 0.1 | 1.9 | PID_TOLL_ENDOGENOUS_PATHWAY | Endogenous TLR signaling |
| 0.1 | 1.4 | PID_ANTHRAX_PATHWAY | Cellular roles of Anthrax toxin |
| 0.0 | 1.9 | PID_FANCONI_PATHWAY | Fanconi anemia pathway |
| 0.0 | 2.2 | PID_HES_HEY_PATHWAY | Notch-mediated HES/HEY network |
| 0.0 | 1.9 | PID_IL6_7_PATHWAY | IL6-mediated signaling events |
| 0.0 | 2.5 | PID_E2F_PATHWAY | E2F transcription factor network |
| 0.0 | 3.9 | NABA_ECM_AFFILIATED | Genes encoding proteins affiliated structurally or functionally to extracellular matrix proteins |
| 0.0 | 1.4 | PID_TAP63_PATHWAY | Validated transcriptional targets of TAp63 isoforms |
| 0.0 | 1.1 | PID_HNF3B_PATHWAY | FOXA2 and FOXA3 transcription factor networks |
| 0.0 | 3.8 | NABA_SECRETED_FACTORS | Genes encoding secreted soluble factors |
| 0.0 | 0.4 | PID_ARF_3PATHWAY | Arf1 pathway |
| 0.0 | 0.9 | PID_ARF6_TRAFFICKING_PATHWAY | Arf6 trafficking events |
Gene overrepresentation in C2:CP:REACTOME category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 1.5 | 9.1 | REACTOME_ELEVATION_OF_CYTOSOLIC_CA2_LEVELS | Genes involved in Elevation of cytosolic Ca2+ levels |
| 1.5 | 19.1 | REACTOME_HDL_MEDIATED_LIPID_TRANSPORT | Genes involved in HDL-mediated lipid transport |
| 0.6 | 6.9 | REACTOME_FACILITATIVE_NA_INDEPENDENT_GLUCOSE_TRANSPORTERS | Genes involved in Facilitative Na+-independent glucose transporters |
| 0.5 | 8.8 | REACTOME_METABOLISM_OF_POLYAMINES | Genes involved in Metabolism of polyamines |
| 0.5 | 15.7 | REACTOME_RORA_ACTIVATES_CIRCADIAN_EXPRESSION | Genes involved in RORA Activates Circadian Expression |
| 0.5 | 6.9 | REACTOME_ACYL_CHAIN_REMODELLING_OF_PC | Genes involved in Acyl chain remodelling of PC |
| 0.4 | 5.6 | REACTOME_EXTRINSIC_PATHWAY_FOR_APOPTOSIS | Genes involved in Extrinsic Pathway for Apoptosis |
| 0.4 | 2.8 | REACTOME_ENDOGENOUS_STEROLS | Genes involved in Endogenous sterols |
| 0.4 | 7.1 | REACTOME_CONVERSION_FROM_APC_C_CDC20_TO_APC_C_CDH1_IN_LATE_ANAPHASE | Genes involved in Conversion from APC/C:Cdc20 to APC/C:Cdh1 in late anaphase |
| 0.3 | 6.6 | REACTOME_GLYCOGEN_BREAKDOWN_GLYCOGENOLYSIS | Genes involved in Glycogen breakdown (glycogenolysis) |
| 0.2 | 3.2 | REACTOME_TRAF6_MEDIATED_IRF7_ACTIVATION | Genes involved in TRAF6 mediated IRF7 activation |
| 0.2 | 2.1 | REACTOME_PURINE_CATABOLISM | Genes involved in Purine catabolism |
| 0.1 | 5.8 | REACTOME_AMINO_ACID_AND_OLIGOPEPTIDE_SLC_TRANSPORTERS | Genes involved in Amino acid and oligopeptide SLC transporters |
| 0.1 | 1.4 | REACTOME_VITAMIN_B5_PANTOTHENATE_METABOLISM | Genes involved in Vitamin B5 (pantothenate) metabolism |
| 0.1 | 2.5 | REACTOME_G0_AND_EARLY_G1 | Genes involved in G0 and Early G1 |
| 0.1 | 2.6 | REACTOME_HS_GAG_BIOSYNTHESIS | Genes involved in HS-GAG biosynthesis |
| 0.1 | 1.4 | REACTOME_JNK_C_JUN_KINASES_PHOSPHORYLATION_AND_ACTIVATION_MEDIATED_BY_ACTIVATED_HUMAN_TAK1 | Genes involved in JNK (c-Jun kinases) phosphorylation and activation mediated by activated human TAK1 |
| 0.1 | 1.1 | REACTOME_PROCESSING_OF_INTRONLESS_PRE_MRNAS | Genes involved in Processing of Intronless Pre-mRNAs |
| 0.1 | 1.2 | REACTOME_ALPHA_LINOLENIC_ACID_ALA_METABOLISM | Genes involved in alpha-linolenic acid (ALA) metabolism |
| 0.1 | 1.1 | REACTOME_ACTIVATED_AMPK_STIMULATES_FATTY_ACID_OXIDATION_IN_MUSCLE | Genes involved in Activated AMPK stimulates fatty-acid oxidation in muscle |
| 0.0 | 1.0 | REACTOME_SYNTHESIS_OF_PA | Genes involved in Synthesis of PA |
| 0.0 | 1.3 | REACTOME_PRE_NOTCH_EXPRESSION_AND_PROCESSING | Genes involved in Pre-NOTCH Expression and Processing |
| 0.0 | 1.2 | REACTOME_METABOLISM_OF_VITAMINS_AND_COFACTORS | Genes involved in Metabolism of vitamins and cofactors |
| 0.0 | 1.9 | REACTOME_MYD88_MAL_CASCADE_INITIATED_ON_PLASMA_MEMBRANE | Genes involved in MyD88:Mal cascade initiated on plasma membrane |
| 0.0 | 1.0 | REACTOME_TRANSCRIPTIONAL_REGULATION_OF_WHITE_ADIPOCYTE_DIFFERENTIATION | Genes involved in Transcriptional Regulation of White Adipocyte Differentiation |
| 0.0 | 0.5 | REACTOME_SIGNALING_BY_FGFR1_FUSION_MUTANTS | Genes involved in Signaling by FGFR1 fusion mutants |