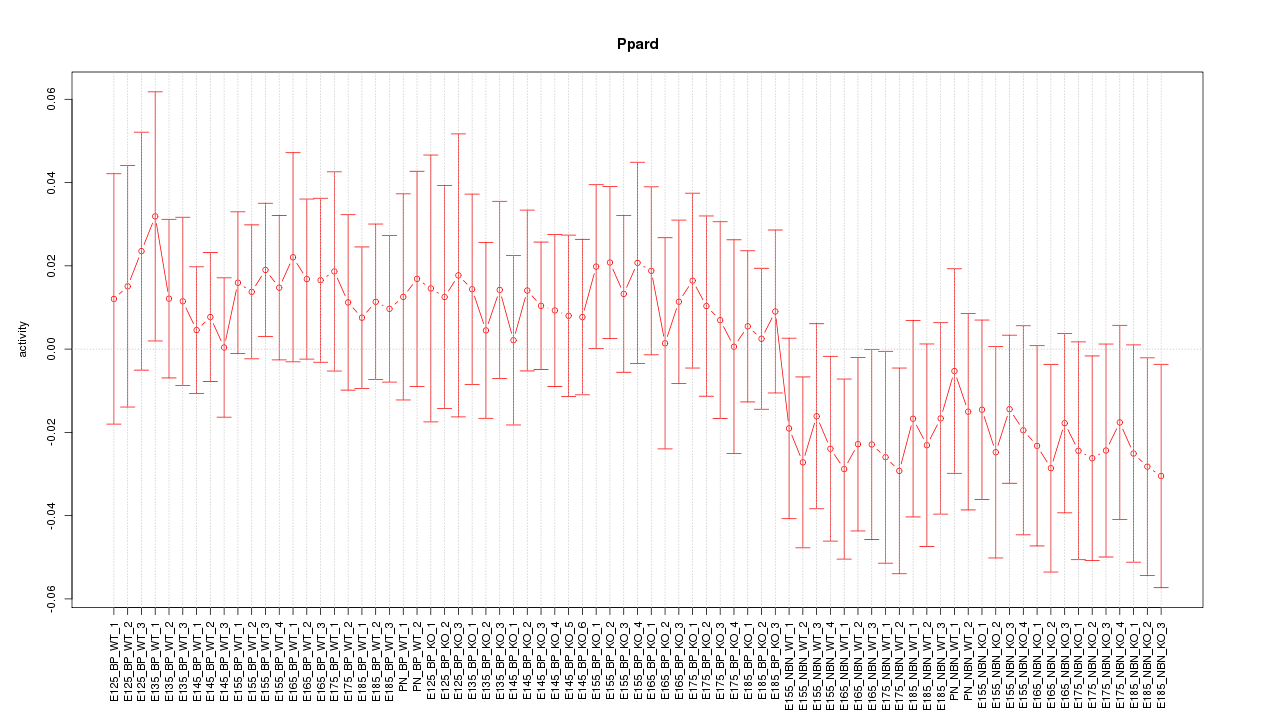
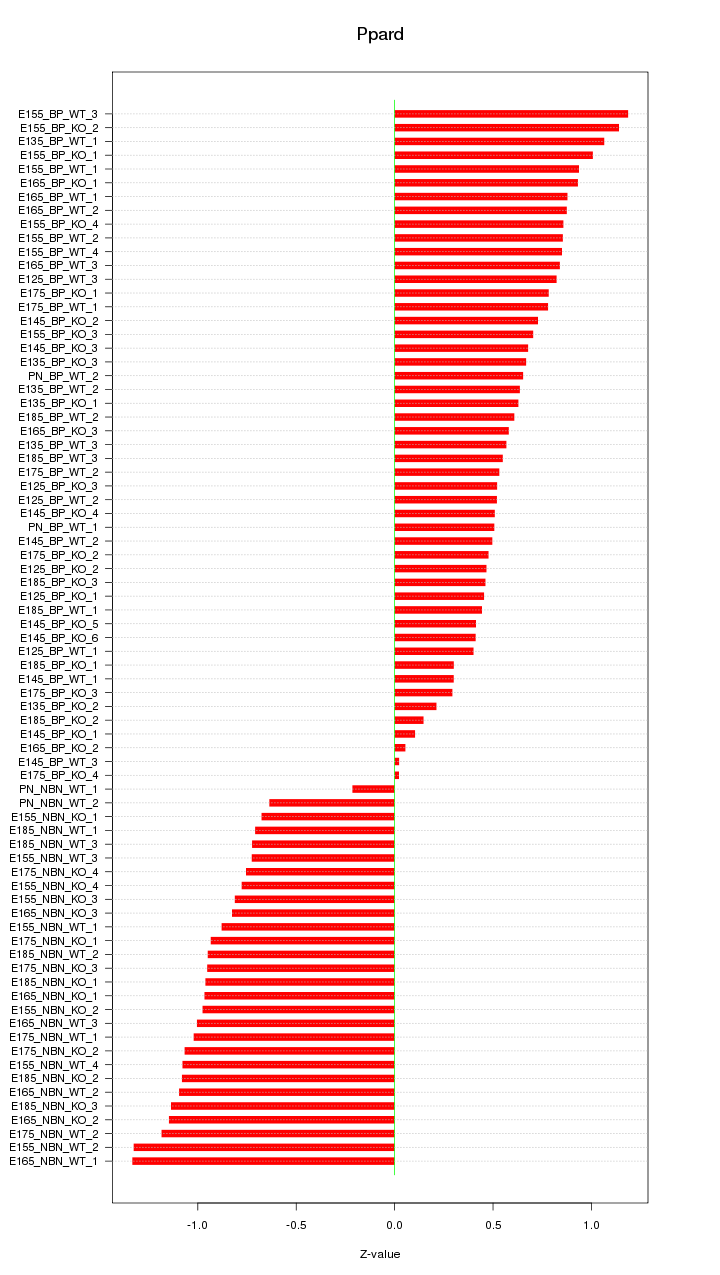
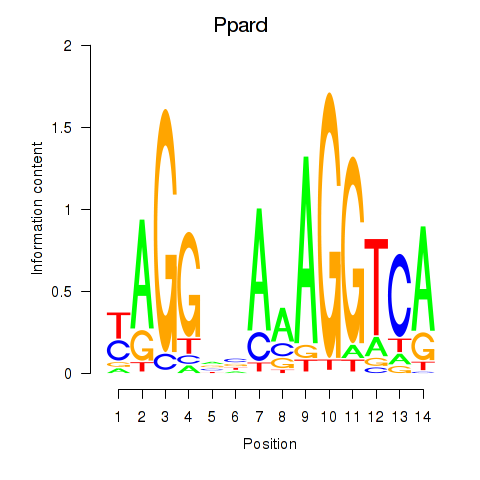
Motif ID: Ppard
Z-value: 0.776
Transcription factors associated with Ppard:
| Gene Symbol | Entrez ID | Gene Name |
|---|---|---|
| Ppard | ENSMUSG00000002250.9 | Ppard |
Activity-expression correlation:
| Gene Symbol | Promoter | Pearson corr. coef. | P-value | Plot |
|---|---|---|---|---|
| Ppard | mm10_v2_chr17_+_28232723_28232789 | 0.67 | 2.2e-11 | Click! |
Top targets:
Gene overrepresentation in biological_process category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 1.4 | 17.6 | GO:0042573 | retinoic acid metabolic process(GO:0042573) |
| 1.3 | 11.9 | GO:0046459 | short-chain fatty acid metabolic process(GO:0046459) |
| 1.0 | 7.2 | GO:0060372 | regulation of atrial cardiac muscle cell membrane repolarization(GO:0060372) |
| 0.9 | 4.7 | GO:0036151 | phosphatidylcholine acyl-chain remodeling(GO:0036151) |
| 0.9 | 2.8 | GO:2000620 | positive regulation of histone H4-K16 acetylation(GO:2000620) |
| 0.8 | 2.5 | GO:0045004 | DNA replication proofreading(GO:0045004) |
| 0.7 | 11.4 | GO:0033539 | fatty acid beta-oxidation using acyl-CoA dehydrogenase(GO:0033539) |
| 0.5 | 1.6 | GO:0019074 | viral genome packaging(GO:0019072) viral RNA genome packaging(GO:0019074) |
| 0.5 | 5.5 | GO:0060575 | intestinal epithelial cell differentiation(GO:0060575) |
| 0.4 | 4.8 | GO:0019368 | fatty acid elongation, saturated fatty acid(GO:0019367) fatty acid elongation, unsaturated fatty acid(GO:0019368) fatty acid elongation, monounsaturated fatty acid(GO:0034625) fatty acid elongation, polyunsaturated fatty acid(GO:0034626) |
| 0.4 | 2.0 | GO:0034196 | acylglycerol transport(GO:0034196) triglyceride transport(GO:0034197) |
| 0.4 | 2.1 | GO:0000098 | sulfur amino acid catabolic process(GO:0000098) |
| 0.2 | 1.0 | GO:0015793 | glycerol transport(GO:0015793) |
| 0.2 | 2.2 | GO:0048386 | positive regulation of retinoic acid receptor signaling pathway(GO:0048386) |
| 0.2 | 1.4 | GO:0000160 | phosphorelay signal transduction system(GO:0000160) |
| 0.2 | 0.7 | GO:0070973 | COPI-coated vesicle budding(GO:0035964) protein localization to endoplasmic reticulum exit site(GO:0070973) |
| 0.2 | 1.0 | GO:1901668 | regulation of superoxide dismutase activity(GO:1901668) |
| 0.2 | 2.0 | GO:1903351 | response to dopamine(GO:1903350) cellular response to dopamine(GO:1903351) |
| 0.2 | 1.4 | GO:0006703 | estrogen biosynthetic process(GO:0006703) |
| 0.2 | 0.8 | GO:0006014 | D-ribose metabolic process(GO:0006014) |
| 0.2 | 1.7 | GO:0032286 | central nervous system myelin maintenance(GO:0032286) |
| 0.2 | 2.9 | GO:0000291 | nuclear-transcribed mRNA catabolic process, exonucleolytic(GO:0000291) |
| 0.1 | 2.5 | GO:0090344 | negative regulation of cell aging(GO:0090344) |
| 0.1 | 10.5 | GO:0051489 | regulation of filopodium assembly(GO:0051489) |
| 0.1 | 4.6 | GO:0048025 | negative regulation of mRNA splicing, via spliceosome(GO:0048025) |
| 0.1 | 6.5 | GO:0015909 | long-chain fatty acid transport(GO:0015909) |
| 0.1 | 4.0 | GO:0060351 | cartilage development involved in endochondral bone morphogenesis(GO:0060351) |
| 0.1 | 2.9 | GO:1900078 | positive regulation of cellular response to insulin stimulus(GO:1900078) |
| 0.1 | 4.1 | GO:0043001 | Golgi to plasma membrane protein transport(GO:0043001) |
| 0.1 | 1.3 | GO:1902043 | positive regulation of extrinsic apoptotic signaling pathway via death domain receptors(GO:1902043) |
| 0.0 | 4.7 | GO:0006096 | glycolytic process(GO:0006096) ATP generation from ADP(GO:0006757) |
| 0.0 | 0.3 | GO:0015879 | carnitine transport(GO:0015879) |
| 0.0 | 1.1 | GO:0015985 | energy coupled proton transport, down electrochemical gradient(GO:0015985) ATP synthesis coupled proton transport(GO:0015986) |
| 0.0 | 2.4 | GO:0045454 | cell redox homeostasis(GO:0045454) |
| 0.0 | 4.7 | GO:0046488 | phosphatidylinositol metabolic process(GO:0046488) |
| 0.0 | 14.3 | GO:0055114 | oxidation-reduction process(GO:0055114) |
| 0.0 | 2.0 | GO:0006665 | sphingolipid metabolic process(GO:0006665) |
| 0.0 | 1.8 | GO:0042098 | T cell proliferation(GO:0042098) |
| 0.0 | 0.4 | GO:0090162 | establishment of epithelial cell polarity(GO:0090162) |
Gene overrepresentation in cellular_component category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 1.9 | 7.4 | GO:0045251 | mitochondrial electron transfer flavoprotein complex(GO:0017133) electron transfer flavoprotein complex(GO:0045251) |
| 1.0 | 2.9 | GO:0031251 | PAN complex(GO:0031251) |
| 0.5 | 4.7 | GO:0000015 | phosphopyruvate hydratase complex(GO:0000015) |
| 0.5 | 2.5 | GO:0008622 | epsilon DNA polymerase complex(GO:0008622) |
| 0.2 | 1.0 | GO:1990130 | Iml1 complex(GO:1990130) |
| 0.2 | 4.7 | GO:0097038 | perinuclear endoplasmic reticulum(GO:0097038) |
| 0.2 | 10.5 | GO:0016459 | myosin complex(GO:0016459) |
| 0.1 | 2.0 | GO:0031528 | microvillus membrane(GO:0031528) |
| 0.1 | 4.0 | GO:0042645 | nucleoid(GO:0009295) mitochondrial nucleoid(GO:0042645) |
| 0.1 | 1.1 | GO:0000276 | mitochondrial proton-transporting ATP synthase complex, coupling factor F(o)(GO:0000276) |
| 0.1 | 13.6 | GO:0005777 | peroxisome(GO:0005777) microbody(GO:0042579) |
| 0.1 | 2.4 | GO:0030173 | integral component of Golgi membrane(GO:0030173) |
| 0.1 | 8.4 | GO:0005741 | mitochondrial outer membrane(GO:0005741) |
| 0.1 | 0.3 | GO:0034715 | U7 snRNP(GO:0005683) pICln-Sm protein complex(GO:0034715) |
| 0.0 | 4.8 | GO:0030176 | integral component of endoplasmic reticulum membrane(GO:0030176) |
| 0.0 | 1.3 | GO:0016592 | mediator complex(GO:0016592) |
| 0.0 | 2.2 | GO:0016605 | PML body(GO:0016605) |
| 0.0 | 1.8 | GO:0005604 | basement membrane(GO:0005604) |
| 0.0 | 1.0 | GO:0031526 | brush border membrane(GO:0031526) |
| 0.0 | 2.6 | GO:0005759 | mitochondrial matrix(GO:0005759) |
| 0.0 | 18.3 | GO:0005783 | endoplasmic reticulum(GO:0005783) |
| 0.0 | 3.6 | GO:0016607 | nuclear speck(GO:0016607) |
Gene overrepresentation in molecular_function category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 2.9 | 17.6 | GO:0019841 | retinol binding(GO:0019841) |
| 1.6 | 4.7 | GO:0030156 | benzodiazepine receptor binding(GO:0030156) |
| 1.5 | 6.0 | GO:0004174 | electron-transferring-flavoprotein dehydrogenase activity(GO:0004174) oxidoreductase activity, acting on the CH-NH group of donors, quinone or similar compound as acceptor(GO:0016649) |
| 1.3 | 4.0 | GO:0017099 | very-long-chain-acyl-CoA dehydrogenase activity(GO:0017099) |
| 1.3 | 6.5 | GO:0004095 | carnitine O-palmitoyltransferase activity(GO:0004095) O-palmitoyltransferase activity(GO:0016416) |
| 0.9 | 7.2 | GO:0086008 | voltage-gated potassium channel activity involved in cardiac muscle cell action potential repolarization(GO:0086008) |
| 0.8 | 2.5 | GO:0008310 | single-stranded DNA 3'-5' exodeoxyribonuclease activity(GO:0008310) |
| 0.8 | 11.9 | GO:0003995 | acyl-CoA dehydrogenase activity(GO:0003995) |
| 0.6 | 2.4 | GO:0016971 | flavin-linked sulfhydryl oxidase activity(GO:0016971) |
| 0.5 | 4.7 | GO:0004634 | phosphopyruvate hydratase activity(GO:0004634) |
| 0.4 | 4.8 | GO:0009922 | fatty acid elongase activity(GO:0009922) 3-oxo-arachidoyl-CoA synthase activity(GO:0102336) 3-oxo-cerotoyl-CoA synthase activity(GO:0102337) 3-oxo-lignoceronyl-CoA synthase activity(GO:0102338) |
| 0.4 | 1.4 | GO:0009055 | electron carrier activity(GO:0009055) |
| 0.3 | 1.4 | GO:0000155 | phosphorelay sensor kinase activity(GO:0000155) |
| 0.3 | 2.0 | GO:0017040 | ceramidase activity(GO:0017040) |
| 0.3 | 4.6 | GO:1990825 | sequence-specific mRNA binding(GO:1990825) |
| 0.3 | 1.4 | GO:0070404 | NADH binding(GO:0070404) |
| 0.3 | 1.6 | GO:0009374 | biotin binding(GO:0009374) |
| 0.2 | 4.7 | GO:0004435 | phosphatidylinositol phospholipase C activity(GO:0004435) |
| 0.2 | 1.0 | GO:0015254 | glycerol channel activity(GO:0015254) |
| 0.2 | 10.5 | GO:0030507 | spectrin binding(GO:0030507) |
| 0.2 | 2.9 | GO:0004535 | poly(A)-specific ribonuclease activity(GO:0004535) |
| 0.1 | 0.3 | GO:0015226 | amino-acid betaine transmembrane transporter activity(GO:0015199) carnitine transmembrane transporter activity(GO:0015226) |
| 0.1 | 0.8 | GO:0048531 | beta-1,3-galactosyltransferase activity(GO:0048531) |
| 0.1 | 5.0 | GO:0003707 | steroid hormone receptor activity(GO:0003707) |
| 0.1 | 0.9 | GO:0046933 | proton-transporting ATP synthase activity, rotational mechanism(GO:0046933) |
| 0.0 | 2.1 | GO:0016831 | carboxy-lyase activity(GO:0016831) |
| 0.0 | 2.5 | GO:0050660 | flavin adenine dinucleotide binding(GO:0050660) |
| 0.0 | 2.0 | GO:0005548 | phospholipid transporter activity(GO:0005548) |
| 0.0 | 17.5 | GO:0016491 | oxidoreductase activity(GO:0016491) |
| 0.0 | 0.7 | GO:0080025 | phosphatidylinositol-3,5-bisphosphate binding(GO:0080025) |
| 0.0 | 2.0 | GO:0033613 | activating transcription factor binding(GO:0033613) |
| 0.0 | 4.7 | GO:0003714 | transcription corepressor activity(GO:0003714) |
| 0.0 | 1.3 | GO:0001104 | RNA polymerase II transcription cofactor activity(GO:0001104) |
| 0.0 | 0.8 | GO:0019200 | carbohydrate kinase activity(GO:0019200) |
| 0.0 | 4.0 | GO:0000287 | magnesium ion binding(GO:0000287) |
| 0.0 | 1.8 | GO:0005178 | integrin binding(GO:0005178) |
Gene overrepresentation in C2:CP category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.2 | 10.4 | PID_HNF3B_PATHWAY | FOXA2 and FOXA3 transcription factor networks |
| 0.2 | 10.5 | PID_NETRIN_PATHWAY | Netrin-mediated signaling events |
| 0.1 | 4.0 | PID_LKB1_PATHWAY | LKB1 signaling events |
| 0.0 | 2.2 | PID_AR_TF_PATHWAY | Regulation of Androgen receptor activity |
| 0.0 | 0.7 | PID_ARF_3PATHWAY | Arf1 pathway |
Gene overrepresentation in C2:CP:REACTOME category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.9 | 15.8 | REACTOME_MITOCHONDRIAL_FATTY_ACID_BETA_OXIDATION | Genes involved in Mitochondrial Fatty Acid Beta-Oxidation |
| 0.3 | 6.8 | REACTOME_ACTIVATED_AMPK_STIMULATES_FATTY_ACID_OXIDATION_IN_MUSCLE | Genes involved in Activated AMPK stimulates fatty-acid oxidation in muscle |
| 0.2 | 6.8 | REACTOME_TIGHT_JUNCTION_INTERACTIONS | Genes involved in Tight junction interactions |
| 0.2 | 4.8 | REACTOME_SYNTHESIS_OF_VERY_LONG_CHAIN_FATTY_ACYL_COAS | Genes involved in Synthesis of very long-chain fatty acyl-CoAs |
| 0.2 | 10.5 | REACTOME_NETRIN1_SIGNALING | Genes involved in Netrin-1 signaling |
| 0.2 | 2.5 | REACTOME_REPAIR_SYNTHESIS_FOR_GAP_FILLING_BY_DNA_POL_IN_TC_NER | Genes involved in Repair synthesis for gap-filling by DNA polymerase in TC-NER |
| 0.1 | 2.0 | REACTOME_CHYLOMICRON_MEDIATED_LIPID_TRANSPORT | Genes involved in Chylomicron-mediated lipid transport |
| 0.1 | 1.0 | REACTOME_PASSIVE_TRANSPORT_BY_AQUAPORINS | Genes involved in Passive Transport by Aquaporins |
| 0.1 | 4.7 | REACTOME_GLUCONEOGENESIS | Genes involved in Gluconeogenesis |
| 0.1 | 4.6 | REACTOME_MRNA_3_END_PROCESSING | Genes involved in mRNA 3'-end processing |
| 0.1 | 6.6 | REACTOME_RESPIRATORY_ELECTRON_TRANSPORT | Genes involved in Respiratory electron transport |
| 0.1 | 2.1 | REACTOME_SULFUR_AMINO_ACID_METABOLISM | Genes involved in Sulfur amino acid metabolism |
| 0.1 | 2.5 | REACTOME_PEROXISOMAL_LIPID_METABOLISM | Genes involved in Peroxisomal lipid metabolism |
| 0.1 | 2.0 | REACTOME_GLYCOSPHINGOLIPID_METABOLISM | Genes involved in Glycosphingolipid metabolism |
| 0.1 | 0.7 | REACTOME_COPI_MEDIATED_TRANSPORT | Genes involved in COPI Mediated Transport |
| 0.0 | 0.8 | REACTOME_METABOLISM_OF_STEROID_HORMONES_AND_VITAMINS_A_AND_D | Genes involved in Metabolism of steroid hormones and vitamins A and D |
| 0.0 | 2.9 | REACTOME_NUCLEAR_RECEPTOR_TRANSCRIPTION_PATHWAY | Genes involved in Nuclear Receptor transcription pathway |
| 0.0 | 1.4 | REACTOME_VOLTAGE_GATED_POTASSIUM_CHANNELS | Genes involved in Voltage gated Potassium channels |
| 0.0 | 1.8 | REACTOME_CELL_JUNCTION_ORGANIZATION | Genes involved in Cell junction organization |
| 0.0 | 0.3 | REACTOME_PROCESSING_OF_CAPPED_INTRONLESS_PRE_MRNA | Genes involved in Processing of Capped Intronless Pre-mRNA |
| 0.0 | 0.8 | REACTOME_O_LINKED_GLYCOSYLATION_OF_MUCINS | Genes involved in O-linked glycosylation of mucins |