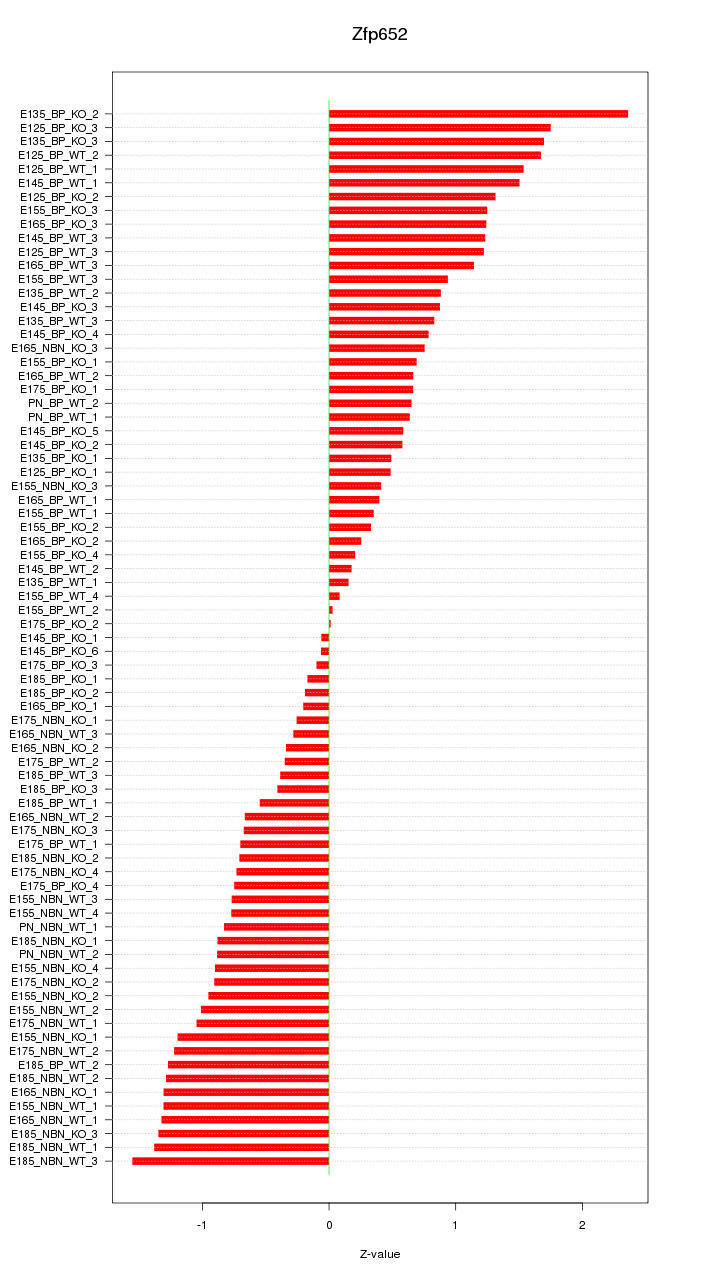
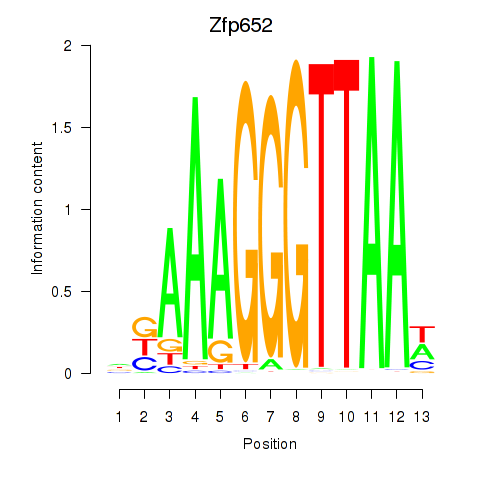
Motif ID: Zfp652
Z-value: 0.927
Transcription factors associated with Zfp652:
| Gene Symbol | Entrez ID | Gene Name |
|---|---|---|
| Zfp652 | ENSMUSG00000075595.3 | Zfp652 |
Activity-expression correlation:
| Gene Symbol | Promoter | Pearson corr. coef. | P-value | Plot |
|---|---|---|---|---|
| Zfp652 | mm10_v2_chr11_+_95749067_95749067 | 0.06 | 5.8e-01 | Click! |
Top targets:
Gene overrepresentation in biological_process category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 7.7 | 23.0 | GO:0090274 | positive regulation of somatostatin secretion(GO:0090274) |
| 3.2 | 16.0 | GO:0035262 | gonad morphogenesis(GO:0035262) |
| 2.2 | 30.9 | GO:0007213 | G-protein coupled acetylcholine receptor signaling pathway(GO:0007213) |
| 2.0 | 8.1 | GO:0060032 | notochord regression(GO:0060032) |
| 1.9 | 7.5 | GO:0046121 | deoxyribonucleoside catabolic process(GO:0046121) |
| 1.8 | 7.3 | GO:0060729 | intestinal epithelial structure maintenance(GO:0060729) |
| 1.4 | 7.0 | GO:0060266 | negative regulation of respiratory burst involved in inflammatory response(GO:0060266) |
| 1.2 | 4.9 | GO:0021775 | smoothened signaling pathway involved in ventral spinal cord interneuron specification(GO:0021775) smoothened signaling pathway involved in spinal cord motor neuron cell fate specification(GO:0021776) |
| 1.1 | 3.3 | GO:0060010 | Sertoli cell fate commitment(GO:0060010) |
| 1.1 | 8.4 | GO:0015074 | DNA integration(GO:0015074) |
| 1.0 | 3.0 | GO:0097350 | neutrophil clearance(GO:0097350) |
| 1.0 | 3.8 | GO:0021502 | neural fold elevation formation(GO:0021502) |
| 0.9 | 4.3 | GO:1903224 | regulation of endodermal cell differentiation(GO:1903224) |
| 0.8 | 2.4 | GO:0010579 | regulation of adenylate cyclase activity involved in G-protein coupled receptor signaling pathway(GO:0010578) positive regulation of adenylate cyclase activity involved in G-protein coupled receptor signaling pathway(GO:0010579) |
| 0.7 | 2.9 | GO:0010510 | regulation of acetyl-CoA biosynthetic process from pyruvate(GO:0010510) |
| 0.7 | 2.2 | GO:0086053 | AV node cell to bundle of His cell communication by electrical coupling(GO:0086053) |
| 0.7 | 6.8 | GO:0031119 | tRNA pseudouridine synthesis(GO:0031119) |
| 0.6 | 6.5 | GO:0034983 | peptidyl-lysine deacetylation(GO:0034983) regulation of skeletal muscle fiber development(GO:0048742) |
| 0.5 | 2.2 | GO:0006751 | glutathione catabolic process(GO:0006751) |
| 0.5 | 9.8 | GO:0050774 | negative regulation of dendrite morphogenesis(GO:0050774) |
| 0.5 | 1.4 | GO:0009174 | UMP biosynthetic process(GO:0006222) pyrimidine ribonucleoside monophosphate metabolic process(GO:0009173) pyrimidine ribonucleoside monophosphate biosynthetic process(GO:0009174) UMP metabolic process(GO:0046049) |
| 0.5 | 10.2 | GO:0051482 | positive regulation of cytosolic calcium ion concentration involved in phospholipase C-activating G-protein coupled signaling pathway(GO:0051482) |
| 0.4 | 2.3 | GO:0000160 | phosphorelay signal transduction system(GO:0000160) |
| 0.4 | 2.8 | GO:0048625 | myoblast fate commitment(GO:0048625) |
| 0.4 | 2.1 | GO:0030263 | apoptotic chromosome condensation(GO:0030263) |
| 0.3 | 1.0 | GO:0035672 | transepithelial chloride transport(GO:0030321) oligopeptide transmembrane transport(GO:0035672) |
| 0.3 | 1.8 | GO:2001054 | negative regulation of mesenchymal cell apoptotic process(GO:2001054) |
| 0.3 | 0.9 | GO:0061181 | regulation of chondrocyte development(GO:0061181) |
| 0.3 | 2.7 | GO:0048386 | positive regulation of retinoic acid receptor signaling pathway(GO:0048386) |
| 0.2 | 1.6 | GO:0010571 | positive regulation of nuclear cell cycle DNA replication(GO:0010571) |
| 0.2 | 7.3 | GO:0060445 | branching involved in salivary gland morphogenesis(GO:0060445) |
| 0.2 | 9.1 | GO:0048747 | muscle fiber development(GO:0048747) |
| 0.2 | 4.1 | GO:1902857 | positive regulation of nonmotile primary cilium assembly(GO:1902857) |
| 0.1 | 5.2 | GO:0043524 | negative regulation of neuron apoptotic process(GO:0043524) |
| 0.1 | 0.6 | GO:0001561 | fatty acid alpha-oxidation(GO:0001561) |
| 0.1 | 1.4 | GO:0031274 | regulation of pseudopodium assembly(GO:0031272) positive regulation of pseudopodium assembly(GO:0031274) |
| 0.1 | 4.5 | GO:0060325 | face morphogenesis(GO:0060325) |
| 0.1 | 0.4 | GO:0031581 | hemidesmosome assembly(GO:0031581) |
| 0.1 | 5.4 | GO:0000381 | regulation of alternative mRNA splicing, via spliceosome(GO:0000381) |
| 0.1 | 1.7 | GO:0003417 | growth plate cartilage development(GO:0003417) |
| 0.1 | 5.6 | GO:0061077 | chaperone-mediated protein folding(GO:0061077) |
| 0.1 | 2.8 | GO:0003170 | heart valve development(GO:0003170) |
| 0.1 | 0.2 | GO:0021698 | cerebellar cortex structural organization(GO:0021698) |
| 0.1 | 0.2 | GO:0090370 | negative regulation of cholesterol efflux(GO:0090370) |
| 0.1 | 11.4 | GO:0007266 | Rho protein signal transduction(GO:0007266) |
| 0.1 | 0.9 | GO:0022038 | corpus callosum development(GO:0022038) |
| 0.0 | 0.8 | GO:0002021 | response to dietary excess(GO:0002021) |
| 0.0 | 1.9 | GO:0031572 | G2 DNA damage checkpoint(GO:0031572) |
| 0.0 | 3.2 | GO:0042147 | retrograde transport, endosome to Golgi(GO:0042147) |
| 0.0 | 17.7 | GO:0045666 | positive regulation of neuron differentiation(GO:0045666) |
| 0.0 | 5.7 | GO:0007283 | spermatogenesis(GO:0007283) |
Gene overrepresentation in cellular_component category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 1.8 | 7.3 | GO:0043259 | laminin-10 complex(GO:0043259) |
| 1.0 | 2.9 | GO:0005967 | mitochondrial pyruvate dehydrogenase complex(GO:0005967) |
| 0.6 | 1.9 | GO:0005588 | collagen type V trimer(GO:0005588) |
| 0.6 | 8.1 | GO:0097542 | ciliary tip(GO:0097542) |
| 0.6 | 2.3 | GO:1902937 | inward rectifier potassium channel complex(GO:1902937) |
| 0.5 | 19.6 | GO:0032809 | neuronal cell body membrane(GO:0032809) |
| 0.4 | 1.7 | GO:0005594 | collagen type IX trimer(GO:0005594) |
| 0.3 | 2.8 | GO:0070369 | beta-catenin-TCF7L2 complex(GO:0070369) |
| 0.3 | 5.0 | GO:0035102 | PRC1 complex(GO:0035102) |
| 0.3 | 4.1 | GO:0031105 | septin complex(GO:0031105) |
| 0.3 | 2.4 | GO:0032591 | dendritic spine membrane(GO:0032591) |
| 0.2 | 2.1 | GO:0061574 | ASAP complex(GO:0061574) |
| 0.2 | 10.2 | GO:0097610 | cleavage furrow(GO:0032154) cell surface furrow(GO:0097610) |
| 0.2 | 23.0 | GO:0043195 | terminal bouton(GO:0043195) |
| 0.2 | 2.2 | GO:0005922 | connexon complex(GO:0005922) |
| 0.2 | 25.0 | GO:0090575 | RNA polymerase II transcription factor complex(GO:0090575) |
| 0.1 | 3.0 | GO:0043657 | host(GO:0018995) host cell part(GO:0033643) host cell(GO:0043657) |
| 0.1 | 18.6 | GO:0001650 | fibrillar center(GO:0001650) |
| 0.1 | 5.2 | GO:0005882 | intermediate filament(GO:0005882) |
| 0.1 | 6.5 | GO:0035097 | histone methyltransferase complex(GO:0035097) |
| 0.1 | 0.4 | GO:0030056 | hemidesmosome(GO:0030056) |
| 0.1 | 1.0 | GO:0097225 | sperm midpiece(GO:0097225) |
| 0.0 | 0.6 | GO:0034098 | VCP-NPL4-UFD1 AAA ATPase complex(GO:0034098) |
| 0.0 | 0.7 | GO:0001673 | male germ cell nucleus(GO:0001673) |
| 0.0 | 1.9 | GO:0016235 | aggresome(GO:0016235) |
| 0.0 | 0.9 | GO:0046658 | anchored component of plasma membrane(GO:0046658) |
| 0.0 | 0.2 | GO:0032937 | SREBP-SCAP-Insig complex(GO:0032937) |
| 0.0 | 0.6 | GO:0005779 | integral component of peroxisomal membrane(GO:0005779) |
Gene overrepresentation in molecular_function category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 1.5 | 10.2 | GO:0070915 | lysophosphatidic acid receptor activity(GO:0070915) |
| 1.4 | 11.4 | GO:0005095 | GTPase inhibitor activity(GO:0005095) |
| 1.0 | 2.9 | GO:0019966 | interleukin-1 binding(GO:0019966) |
| 0.8 | 15.6 | GO:0035497 | cAMP response element binding(GO:0035497) |
| 0.8 | 30.9 | GO:0001965 | G-protein alpha-subunit binding(GO:0001965) |
| 0.8 | 2.4 | GO:0048019 | receptor antagonist activity(GO:0048019) |
| 0.8 | 4.6 | GO:0008239 | dipeptidyl-peptidase activity(GO:0008239) |
| 0.8 | 9.1 | GO:0008191 | metalloendopeptidase inhibitor activity(GO:0008191) |
| 0.7 | 2.9 | GO:0004740 | pyruvate dehydrogenase (acetyl-transferring) kinase activity(GO:0004740) |
| 0.7 | 2.2 | GO:0086077 | gap junction channel activity involved in AV node cell-bundle of His cell electrical coupling(GO:0086077) |
| 0.6 | 5.2 | GO:0031730 | CCR5 chemokine receptor binding(GO:0031730) |
| 0.6 | 7.5 | GO:0016832 | aldehyde-lyase activity(GO:0016832) |
| 0.5 | 7.0 | GO:0017017 | MAP kinase tyrosine/serine/threonine phosphatase activity(GO:0017017) mitogen-activated protein kinase p38 binding(GO:0048273) |
| 0.5 | 2.3 | GO:1902282 | voltage-gated potassium channel activity involved in ventricular cardiac muscle cell action potential repolarization(GO:1902282) |
| 0.4 | 23.0 | GO:0008528 | G-protein coupled peptide receptor activity(GO:0008528) |
| 0.4 | 6.8 | GO:0009982 | pseudouridine synthase activity(GO:0009982) |
| 0.4 | 2.4 | GO:0030284 | estrogen receptor activity(GO:0030284) |
| 0.4 | 3.8 | GO:0070700 | BMP receptor binding(GO:0070700) |
| 0.3 | 2.2 | GO:0008242 | omega peptidase activity(GO:0008242) |
| 0.3 | 6.5 | GO:0031078 | histone deacetylase activity (H3-K14 specific)(GO:0031078) NAD-dependent histone deacetylase activity (H3-K14 specific)(GO:0032041) |
| 0.3 | 1.4 | GO:0004849 | uridine kinase activity(GO:0004849) |
| 0.3 | 5.6 | GO:0005528 | macrolide binding(GO:0005527) FK506 binding(GO:0005528) |
| 0.3 | 3.0 | GO:0032036 | myosin heavy chain binding(GO:0032036) |
| 0.3 | 3.3 | GO:0035014 | retinoic acid binding(GO:0001972) phosphatidylinositol 3-kinase regulator activity(GO:0035014) |
| 0.3 | 1.0 | GO:0015563 | uptake transmembrane transporter activity(GO:0015563) |
| 0.2 | 8.1 | GO:1990841 | promoter-specific chromatin binding(GO:1990841) |
| 0.2 | 4.5 | GO:0070182 | DNA polymerase binding(GO:0070182) |
| 0.1 | 2.8 | GO:0070016 | gamma-catenin binding(GO:0045295) armadillo repeat domain binding(GO:0070016) |
| 0.1 | 6.2 | GO:0070888 | E-box binding(GO:0070888) |
| 0.1 | 1.8 | GO:0003680 | AT DNA binding(GO:0003680) |
| 0.1 | 1.9 | GO:0050321 | tau-protein kinase activity(GO:0050321) |
| 0.1 | 0.8 | GO:0015217 | ATP transmembrane transporter activity(GO:0005347) ADP transmembrane transporter activity(GO:0015217) |
| 0.1 | 6.4 | GO:0097110 | scaffold protein binding(GO:0097110) |
| 0.1 | 1.4 | GO:0017049 | GTP-Rho binding(GO:0017049) |
| 0.1 | 0.5 | GO:0008440 | inositol-1,4,5-trisphosphate 3-kinase activity(GO:0008440) |
| 0.1 | 3.2 | GO:0017112 | Rab guanyl-nucleotide exchange factor activity(GO:0017112) |
| 0.1 | 0.6 | GO:0000268 | peroxisome targeting sequence binding(GO:0000268) |
| 0.1 | 5.8 | GO:0008013 | beta-catenin binding(GO:0008013) |
| 0.1 | 18.6 | GO:0001077 | transcriptional activator activity, RNA polymerase II core promoter proximal region sequence-specific binding(GO:0001077) |
| 0.1 | 1.9 | GO:0005201 | extracellular matrix structural constituent(GO:0005201) |
| 0.0 | 1.9 | GO:0004896 | cytokine receptor activity(GO:0004896) |
| 0.0 | 4.1 | GO:0045296 | cadherin binding(GO:0045296) |
| 0.0 | 0.2 | GO:0070410 | co-SMAD binding(GO:0070410) |
| 0.0 | 5.4 | GO:0044822 | mRNA binding(GO:0003729) poly(A) RNA binding(GO:0044822) |
| 0.0 | 2.8 | GO:0001078 | transcriptional repressor activity, RNA polymerase II core promoter proximal region sequence-specific binding(GO:0001078) |
| 0.0 | 0.7 | GO:0003684 | damaged DNA binding(GO:0003684) |
| 0.0 | 2.1 | GO:0003700 | nucleic acid binding transcription factor activity(GO:0001071) transcription factor activity, sequence-specific DNA binding(GO:0003700) |
Gene overrepresentation in C2:CP category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.9 | 7.3 | PID_INTEGRIN4_PATHWAY | Alpha6 beta4 integrin-ligand interactions |
| 0.3 | 8.1 | PID_HEDGEHOG_2PATHWAY | Signaling events mediated by the Hedgehog family |
| 0.2 | 7.3 | SA_PTEN_PATHWAY | PTEN is a tumor suppressor that dephosphorylates the lipid messenger phosphatidylinositol triphosphate. |
| 0.2 | 7.0 | ST_P38_MAPK_PATHWAY | p38 MAPK Pathway |
| 0.2 | 1.9 | ST_INTERFERON_GAMMA_PATHWAY | Interferon gamma pathway. |
| 0.2 | 9.8 | PID_RHOA_PATHWAY | RhoA signaling pathway |
| 0.1 | 6.5 | PID_HDAC_CLASSII_PATHWAY | Signaling events mediated by HDAC Class II |
| 0.1 | 3.3 | PID_RXR_VDR_PATHWAY | RXR and RAR heterodimerization with other nuclear receptor |
| 0.1 | 4.9 | PID_HEDGEHOG_GLI_PATHWAY | Hedgehog signaling events mediated by Gli proteins |
| 0.1 | 3.3 | ST_WNT_BETA_CATENIN_PATHWAY | Wnt/beta-catenin Pathway |
| 0.1 | 3.6 | PID_SYNDECAN_1_PATHWAY | Syndecan-1-mediated signaling events |
| 0.1 | 4.5 | ST_FAS_SIGNALING_PATHWAY | Fas Signaling Pathway |
| 0.0 | 1.7 | PID_DELTA_NP63_PATHWAY | Validated transcriptional targets of deltaNp63 isoforms |
| 0.0 | 1.9 | PID_LKB1_PATHWAY | LKB1 signaling events |
| 0.0 | 2.8 | PID_BETA_CATENIN_NUC_PATHWAY | Regulation of nuclear beta catenin signaling and target gene transcription |
| 0.0 | 0.7 | PID_ATM_PATHWAY | ATM pathway |
| 0.0 | 0.2 | SA_CASPASE_CASCADE | Apoptosis is mediated by caspases, cysteine proteases arranged in a proteolytic cascade. |
Gene overrepresentation in C2:CP:REACTOME category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 1.1 | 10.2 | REACTOME_P2Y_RECEPTORS | Genes involved in P2Y receptors |
| 0.5 | 17.7 | REACTOME_MYOGENESIS | Genes involved in Myogenesis |
| 0.3 | 30.9 | REACTOME_G_ALPHA_I_SIGNALLING_EVENTS | Genes involved in G alpha (i) signalling events |
| 0.3 | 7.3 | REACTOME_REGULATION_OF_GENE_EXPRESSION_IN_BETA_CELLS | Genes involved in Regulation of gene expression in beta cells |
| 0.3 | 23.0 | REACTOME_PEPTIDE_LIGAND_BINDING_RECEPTORS | Genes involved in Peptide ligand-binding receptors |
| 0.2 | 2.9 | REACTOME_REGULATION_OF_PYRUVATE_DEHYDROGENASE_PDH_COMPLEX | Genes involved in Regulation of pyruvate dehydrogenase (PDH) complex |
| 0.2 | 2.2 | REACTOME_GAP_JUNCTION_ASSEMBLY | Genes involved in Gap junction assembly |
| 0.2 | 1.9 | REACTOME_REGULATION_OF_IFNG_SIGNALING | Genes involved in Regulation of IFNG signaling |
| 0.1 | 5.6 | REACTOME_ACTIVATION_OF_CHAPERONE_GENES_BY_XBP1S | Genes involved in Activation of Chaperone Genes by XBP1(S) |
| 0.1 | 2.2 | REACTOME_GLUTATHIONE_CONJUGATION | Genes involved in Glutathione conjugation |
| 0.1 | 3.5 | REACTOME_VOLTAGE_GATED_POTASSIUM_CHANNELS | Genes involved in Voltage gated Potassium channels |
| 0.1 | 3.6 | REACTOME_NCAM1_INTERACTIONS | Genes involved in NCAM1 interactions |
| 0.1 | 5.1 | REACTOME_NOTCH1_INTRACELLULAR_DOMAIN_REGULATES_TRANSCRIPTION | Genes involved in NOTCH1 Intracellular Domain Regulates Transcription |
| 0.1 | 1.4 | REACTOME_PYRIMIDINE_METABOLISM | Genes involved in Pyrimidine metabolism |
| 0.1 | 3.8 | REACTOME_INTEGRIN_CELL_SURFACE_INTERACTIONS | Genes involved in Integrin cell surface interactions |
| 0.1 | 2.5 | REACTOME_APOPTOTIC_CLEAVAGE_OF_CELLULAR_PROTEINS | Genes involved in Apoptotic cleavage of cellular proteins |
| 0.0 | 3.3 | REACTOME_NUCLEAR_RECEPTOR_TRANSCRIPTION_PATHWAY | Genes involved in Nuclear Receptor transcription pathway |
| 0.0 | 0.7 | REACTOME_HOMOLOGOUS_RECOMBINATION_REPAIR_OF_REPLICATION_INDEPENDENT_DOUBLE_STRAND_BREAKS | Genes involved in Homologous recombination repair of replication-independent double-strand breaks |
| 0.0 | 7.9 | REACTOME_ANTIGEN_PROCESSING_UBIQUITINATION_PROTEASOME_DEGRADATION | Genes involved in Antigen processing: Ubiquitination & Proteasome degradation |
| 0.0 | 0.4 | REACTOME_STRIATED_MUSCLE_CONTRACTION | Genes involved in Striated Muscle Contraction |
| 0.0 | 0.2 | REACTOME_DOWNREGULATION_OF_SMAD2_3_SMAD4_TRANSCRIPTIONAL_ACTIVITY | Genes involved in Downregulation of SMAD2/3:SMAD4 transcriptional activity |
| 0.0 | 1.0 | REACTOME_TRANSPORT_OF_INORGANIC_CATIONS_ANIONS_AND_AMINO_ACIDS_OLIGOPEPTIDES | Genes involved in Transport of inorganic cations/anions and amino acids/oligopeptides |