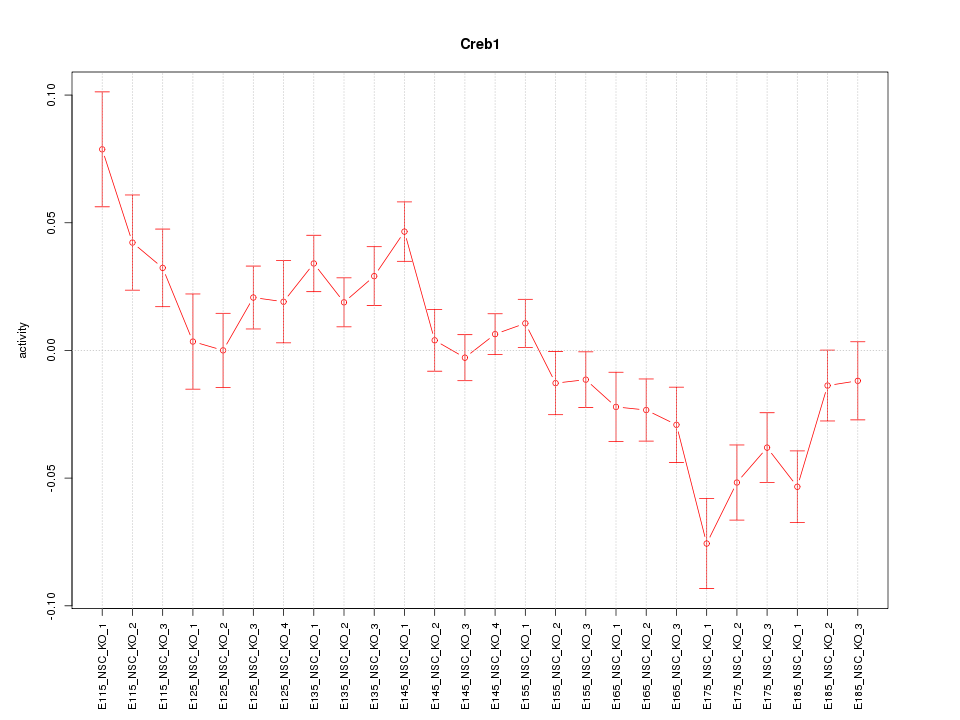
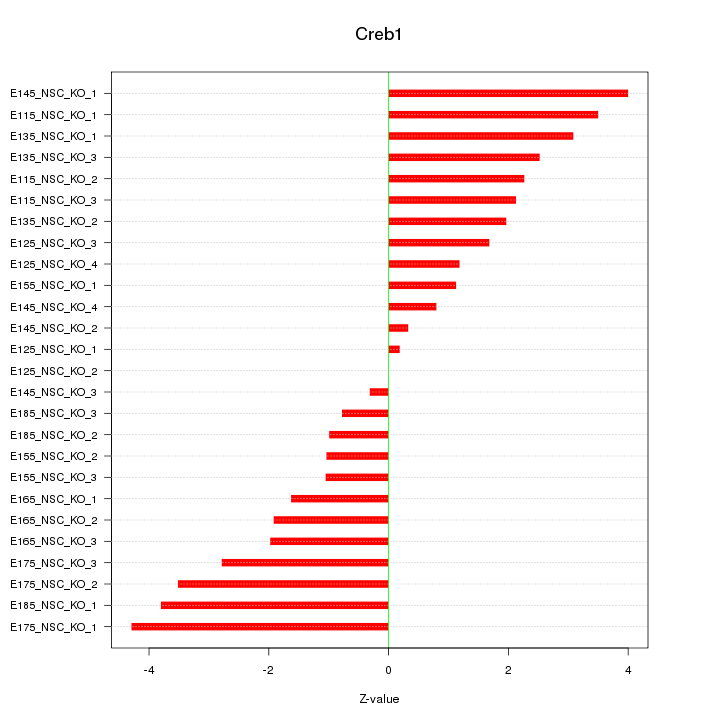
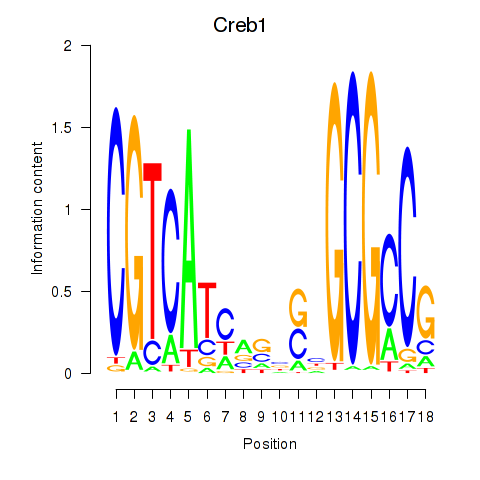
Motif ID: Creb1
Z-value: 2.245
Transcription factors associated with Creb1:
| Gene Symbol | Entrez ID | Gene Name |
|---|---|---|
| Creb1 | ENSMUSG00000025958.8 | Creb1 |
Activity-expression correlation:
| Gene Symbol | Promoter | Pearson corr. coef. | P-value | Plot |
|---|---|---|---|---|
| Creb1 | mm10_v2_chr1_+_64532790_64532815 | 0.33 | 1.0e-01 | Click! |
Top targets:
Gene overrepresentation in biological_process category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 2.4 | 7.3 | GO:0071921 | establishment of sister chromatid cohesion(GO:0034085) cohesin loading(GO:0071921) regulation of cohesin loading(GO:0071922) |
| 2.3 | 9.3 | GO:1904565 | response to 1-oleoyl-sn-glycerol 3-phosphate(GO:1904565) cellular response to 1-oleoyl-sn-glycerol 3-phosphate(GO:1904566) |
| 2.3 | 6.9 | GO:0032240 | negative regulation of nucleobase-containing compound transport(GO:0032240) negative regulation of RNA export from nucleus(GO:0046832) |
| 2.1 | 6.4 | GO:0021759 | globus pallidus development(GO:0021759) |
| 1.5 | 3.1 | GO:1904742 | regulation of telomeric DNA binding(GO:1904742) |
| 1.4 | 4.3 | GO:0034476 | U1 snRNA 3'-end processing(GO:0034473) U5 snRNA 3'-end processing(GO:0034476) |
| 1.4 | 6.8 | GO:0070345 | negative regulation of fat cell proliferation(GO:0070345) |
| 1.2 | 2.5 | GO:0010847 | regulation of chromatin assembly(GO:0010847) |
| 1.2 | 3.6 | GO:0042998 | positive regulation of Golgi to plasma membrane protein transport(GO:0042998) |
| 1.2 | 3.5 | GO:1902498 | regulation of protein autoubiquitination(GO:1902498) |
| 1.1 | 4.6 | GO:0051311 | meiotic metaphase plate congression(GO:0051311) |
| 1.1 | 6.8 | GO:0045875 | DNA strand renaturation(GO:0000733) negative regulation of sister chromatid cohesion(GO:0045875) |
| 0.9 | 2.8 | GO:0072554 | arterial endothelial cell fate commitment(GO:0060844) blood vessel endothelial cell fate commitment(GO:0060846) endothelial cell fate specification(GO:0060847) Notch signaling pathway involved in arterial endothelial cell fate commitment(GO:0060853) blood vessel lumenization(GO:0072554) blood vessel endothelial cell fate specification(GO:0097101) regulation of canonical Wnt signaling pathway involved in cardiac muscle cell fate commitment(GO:1901295) positive regulation of heart induction(GO:1901321) regulation of canonical Wnt signaling pathway involved in heart development(GO:1905066) |
| 0.9 | 6.3 | GO:0045842 | positive regulation of mitotic metaphase/anaphase transition(GO:0045842) positive regulation of metaphase/anaphase transition of cell cycle(GO:1902101) |
| 0.9 | 2.6 | GO:0021529 | spinal cord oligodendrocyte cell differentiation(GO:0021529) spinal cord oligodendrocyte cell fate specification(GO:0021530) musculoskeletal movement, spinal reflex action(GO:0050883) olfactory pit development(GO:0060166) |
| 0.9 | 3.5 | GO:0001306 | age-dependent response to oxidative stress(GO:0001306) age-dependent general metabolic decline(GO:0007571) |
| 0.8 | 2.4 | GO:1903225 | negative regulation of endodermal cell differentiation(GO:1903225) |
| 0.8 | 2.4 | GO:1903722 | endosome to lysosome transport via multivesicular body sorting pathway(GO:0032510) regulation of centriole elongation(GO:1903722) |
| 0.8 | 2.4 | GO:1901420 | negative regulation of response to alcohol(GO:1901420) |
| 0.8 | 2.3 | GO:1903896 | positive regulation of IRE1-mediated unfolded protein response(GO:1903896) |
| 0.7 | 2.9 | GO:1900864 | mitochondrial tRNA modification(GO:0070900) mitochondrial RNA modification(GO:1900864) |
| 0.7 | 2.1 | GO:0048296 | isotype switching to IgA isotypes(GO:0048290) regulation of isotype switching to IgA isotypes(GO:0048296) |
| 0.7 | 2.0 | GO:0035973 | aggrephagy(GO:0035973) |
| 0.6 | 3.0 | GO:0006189 | 'de novo' IMP biosynthetic process(GO:0006189) |
| 0.6 | 1.8 | GO:0036092 | phosphatidylinositol-3-phosphate biosynthetic process(GO:0036092) |
| 0.6 | 1.7 | GO:1901856 | negative regulation of cellular respiration(GO:1901856) |
| 0.6 | 2.9 | GO:0010792 | DNA double-strand break processing involved in repair via single-strand annealing(GO:0010792) |
| 0.6 | 4.0 | GO:0036265 | RNA (guanine-N7)-methylation(GO:0036265) |
| 0.6 | 6.6 | GO:0000244 | spliceosomal tri-snRNP complex assembly(GO:0000244) |
| 0.5 | 3.1 | GO:0045654 | positive regulation of megakaryocyte differentiation(GO:0045654) |
| 0.5 | 5.0 | GO:0035520 | monoubiquitinated protein deubiquitination(GO:0035520) |
| 0.5 | 1.5 | GO:0060743 | epithelial cell maturation involved in prostate gland development(GO:0060743) alveolar secondary septum development(GO:0061144) |
| 0.5 | 7.4 | GO:0034501 | protein localization to kinetochore(GO:0034501) |
| 0.5 | 1.4 | GO:0090526 | regulation of gluconeogenesis involved in cellular glucose homeostasis(GO:0090526) |
| 0.5 | 3.3 | GO:2000348 | regulation of CD40 signaling pathway(GO:2000348) |
| 0.5 | 1.9 | GO:0072675 | osteoclast fusion(GO:0072675) |
| 0.5 | 3.7 | GO:0043928 | exonucleolytic nuclear-transcribed mRNA catabolic process involved in deadenylation-dependent decay(GO:0043928) |
| 0.5 | 1.9 | GO:0002949 | tRNA threonylcarbamoyladenosine modification(GO:0002949) tRNA threonylcarbamoyladenosine metabolic process(GO:0070525) |
| 0.5 | 5.0 | GO:0035562 | negative regulation of chromatin binding(GO:0035562) |
| 0.4 | 11.6 | GO:0030488 | tRNA methylation(GO:0030488) |
| 0.4 | 2.2 | GO:0034196 | acylglycerol transport(GO:0034196) triglyceride transport(GO:0034197) |
| 0.4 | 3.5 | GO:0035845 | photoreceptor cell outer segment organization(GO:0035845) |
| 0.4 | 3.0 | GO:0006297 | nucleotide-excision repair, DNA gap filling(GO:0006297) |
| 0.4 | 2.1 | GO:0060011 | Sertoli cell proliferation(GO:0060011) |
| 0.4 | 5.1 | GO:0034773 | histone H4-K20 trimethylation(GO:0034773) |
| 0.4 | 4.5 | GO:0031468 | nuclear envelope reassembly(GO:0031468) |
| 0.4 | 2.5 | GO:0006344 | maintenance of chromatin silencing(GO:0006344) |
| 0.4 | 1.6 | GO:0035878 | nail development(GO:0035878) |
| 0.4 | 4.0 | GO:0051984 | positive regulation of chromosome segregation(GO:0051984) |
| 0.4 | 0.8 | GO:0000965 | mitochondrial RNA 3'-end processing(GO:0000965) |
| 0.4 | 4.9 | GO:0051382 | kinetochore assembly(GO:0051382) |
| 0.4 | 1.5 | GO:0015886 | heme transport(GO:0015886) |
| 0.4 | 2.2 | GO:1903755 | regulation of SUMO transferase activity(GO:1903182) positive regulation of SUMO transferase activity(GO:1903755) |
| 0.4 | 5.8 | GO:0015693 | magnesium ion transport(GO:0015693) |
| 0.4 | 1.1 | GO:0015014 | heparan sulfate proteoglycan biosynthetic process, polysaccharide chain biosynthetic process(GO:0015014) |
| 0.4 | 4.2 | GO:0021942 | radial glia guided migration of Purkinje cell(GO:0021942) |
| 0.4 | 1.1 | GO:0039530 | MDA-5 signaling pathway(GO:0039530) |
| 0.3 | 11.2 | GO:0006270 | DNA replication initiation(GO:0006270) |
| 0.3 | 1.0 | GO:0046836 | glycolipid transport(GO:0046836) |
| 0.3 | 1.0 | GO:0019085 | early viral transcription(GO:0019085) |
| 0.3 | 1.6 | GO:0042780 | tRNA 3'-end processing(GO:0042780) |
| 0.3 | 2.2 | GO:0051461 | regulation of corticotropin secretion(GO:0051459) positive regulation of corticotropin secretion(GO:0051461) |
| 0.3 | 1.6 | GO:0061084 | negative regulation of protein refolding(GO:0061084) |
| 0.3 | 1.9 | GO:2000210 | positive regulation of anoikis(GO:2000210) |
| 0.3 | 3.1 | GO:0016127 | cholesterol catabolic process(GO:0006707) sterol catabolic process(GO:0016127) |
| 0.3 | 0.9 | GO:0048312 | intracellular distribution of mitochondria(GO:0048312) |
| 0.3 | 2.3 | GO:0009125 | nucleoside monophosphate catabolic process(GO:0009125) |
| 0.3 | 2.6 | GO:0002318 | myeloid progenitor cell differentiation(GO:0002318) |
| 0.3 | 1.1 | GO:0072369 | regulation of lipid transport by positive regulation of transcription from RNA polymerase II promoter(GO:0072369) |
| 0.3 | 2.2 | GO:0070933 | DNA double-strand break processing(GO:0000729) histone H4 deacetylation(GO:0070933) |
| 0.3 | 0.8 | GO:0035621 | ER to Golgi ceramide transport(GO:0035621) ceramide transport(GO:0035627) |
| 0.3 | 5.2 | GO:0038092 | nodal signaling pathway(GO:0038092) |
| 0.3 | 1.5 | GO:0098789 | pre-mRNA cleavage required for polyadenylation(GO:0098789) |
| 0.3 | 1.0 | GO:0070562 | regulation of vitamin D receptor signaling pathway(GO:0070562) |
| 0.3 | 3.8 | GO:0070816 | phosphorylation of RNA polymerase II C-terminal domain(GO:0070816) |
| 0.2 | 1.7 | GO:0008616 | queuosine biosynthetic process(GO:0008616) queuosine metabolic process(GO:0046116) |
| 0.2 | 0.9 | GO:1903288 | regulation of potassium ion import(GO:1903286) positive regulation of potassium ion import(GO:1903288) |
| 0.2 | 0.9 | GO:2000195 | negative regulation of female gonad development(GO:2000195) |
| 0.2 | 1.3 | GO:0006384 | transcription initiation from RNA polymerase III promoter(GO:0006384) |
| 0.2 | 2.4 | GO:0019368 | fatty acid elongation, saturated fatty acid(GO:0019367) fatty acid elongation, unsaturated fatty acid(GO:0019368) fatty acid elongation, monounsaturated fatty acid(GO:0034625) fatty acid elongation, polyunsaturated fatty acid(GO:0034626) |
| 0.2 | 0.9 | GO:0002326 | B cell lineage commitment(GO:0002326) ectopic germ cell programmed cell death(GO:0035234) |
| 0.2 | 0.9 | GO:0048319 | axial mesoderm morphogenesis(GO:0048319) |
| 0.2 | 1.2 | GO:0046602 | regulation of mitotic centrosome separation(GO:0046602) |
| 0.2 | 1.9 | GO:0042989 | sequestering of actin monomers(GO:0042989) |
| 0.2 | 2.0 | GO:0000054 | ribosomal subunit export from nucleus(GO:0000054) ribosome localization(GO:0033750) establishment of ribosome localization(GO:0033753) |
| 0.2 | 1.0 | GO:0043353 | enucleate erythrocyte differentiation(GO:0043353) |
| 0.2 | 2.6 | GO:0072600 | establishment of protein localization to Golgi(GO:0072600) |
| 0.2 | 1.0 | GO:0070475 | rRNA base methylation(GO:0070475) |
| 0.2 | 5.1 | GO:0006383 | transcription from RNA polymerase III promoter(GO:0006383) |
| 0.2 | 3.5 | GO:0070584 | mitochondrion morphogenesis(GO:0070584) |
| 0.2 | 2.5 | GO:0043162 | ubiquitin-dependent protein catabolic process via the multivesicular body sorting pathway(GO:0043162) |
| 0.1 | 0.7 | GO:1902035 | positive regulation of hematopoietic stem cell proliferation(GO:1902035) |
| 0.1 | 0.7 | GO:0040038 | polar body extrusion after meiotic divisions(GO:0040038) |
| 0.1 | 2.3 | GO:0006999 | nuclear pore organization(GO:0006999) |
| 0.1 | 1.1 | GO:0035871 | protein K11-linked deubiquitination(GO:0035871) |
| 0.1 | 2.3 | GO:0016973 | poly(A)+ mRNA export from nucleus(GO:0016973) |
| 0.1 | 0.5 | GO:0032494 | response to peptidoglycan(GO:0032494) |
| 0.1 | 1.5 | GO:0002227 | innate immune response in mucosa(GO:0002227) |
| 0.1 | 7.0 | GO:0045600 | positive regulation of fat cell differentiation(GO:0045600) |
| 0.1 | 1.4 | GO:2000001 | regulation of DNA damage checkpoint(GO:2000001) |
| 0.1 | 0.9 | GO:0046784 | viral mRNA export from host cell nucleus(GO:0046784) |
| 0.1 | 1.6 | GO:0043923 | positive regulation by host of viral transcription(GO:0043923) |
| 0.1 | 0.9 | GO:0051418 | interphase microtubule nucleation by interphase microtubule organizing center(GO:0051415) microtubule nucleation by microtubule organizing center(GO:0051418) |
| 0.1 | 1.6 | GO:0006654 | phosphatidic acid biosynthetic process(GO:0006654) phosphatidic acid metabolic process(GO:0046473) |
| 0.1 | 0.8 | GO:1902166 | negative regulation of intrinsic apoptotic signaling pathway in response to DNA damage by p53 class mediator(GO:1902166) |
| 0.1 | 1.1 | GO:0008063 | Toll signaling pathway(GO:0008063) |
| 0.1 | 0.2 | GO:0000491 | small nucleolar ribonucleoprotein complex assembly(GO:0000491) box C/D snoRNP assembly(GO:0000492) |
| 0.1 | 5.0 | GO:0010501 | RNA secondary structure unwinding(GO:0010501) |
| 0.1 | 1.3 | GO:0048280 | vesicle fusion with Golgi apparatus(GO:0048280) |
| 0.1 | 0.5 | GO:0045292 | mRNA cis splicing, via spliceosome(GO:0045292) |
| 0.1 | 0.6 | GO:0006450 | regulation of translational fidelity(GO:0006450) |
| 0.1 | 2.4 | GO:0045722 | positive regulation of gluconeogenesis(GO:0045722) |
| 0.1 | 0.4 | GO:0070535 | histone H2A K63-linked ubiquitination(GO:0070535) |
| 0.1 | 0.4 | GO:1905098 | negative regulation of guanyl-nucleotide exchange factor activity(GO:1905098) response to amino acid starvation(GO:1990928) |
| 0.1 | 2.5 | GO:0045879 | negative regulation of smoothened signaling pathway(GO:0045879) |
| 0.1 | 2.2 | GO:0030261 | chromosome condensation(GO:0030261) |
| 0.1 | 2.0 | GO:0042059 | negative regulation of epidermal growth factor receptor signaling pathway(GO:0042059) |
| 0.1 | 2.5 | GO:0006890 | retrograde vesicle-mediated transport, Golgi to ER(GO:0006890) |
| 0.1 | 2.3 | GO:0031146 | SCF-dependent proteasomal ubiquitin-dependent protein catabolic process(GO:0031146) |
| 0.1 | 2.5 | GO:0006378 | mRNA polyadenylation(GO:0006378) |
| 0.1 | 1.5 | GO:0042026 | protein refolding(GO:0042026) |
| 0.1 | 0.5 | GO:0007296 | vitellogenesis(GO:0007296) |
| 0.1 | 3.2 | GO:0006418 | tRNA aminoacylation for protein translation(GO:0006418) |
| 0.1 | 1.1 | GO:0002021 | response to dietary excess(GO:0002021) |
| 0.1 | 0.3 | GO:0000447 | endonucleolytic cleavage in ITS1 to separate SSU-rRNA from 5.8S rRNA and LSU-rRNA from tricistronic rRNA transcript (SSU-rRNA, 5.8S rRNA, LSU-rRNA)(GO:0000447) endonucleolytic cleavage in 5'-ETS of tricistronic rRNA transcript (SSU-rRNA, 5.8S rRNA, LSU-rRNA)(GO:0000480) |
| 0.1 | 0.5 | GO:0006071 | glycerol metabolic process(GO:0006071) |
| 0.1 | 0.8 | GO:0006614 | SRP-dependent cotranslational protein targeting to membrane(GO:0006614) |
| 0.1 | 0.2 | GO:0045041 | protein import into mitochondrial intermembrane space(GO:0045041) |
| 0.1 | 9.3 | GO:0006364 | rRNA processing(GO:0006364) |
| 0.1 | 1.2 | GO:0045948 | positive regulation of translational initiation(GO:0045948) |
| 0.1 | 3.5 | GO:0000245 | spliceosomal complex assembly(GO:0000245) |
| 0.1 | 0.6 | GO:0080182 | histone H3-K4 trimethylation(GO:0080182) |
| 0.1 | 3.1 | GO:0006888 | ER to Golgi vesicle-mediated transport(GO:0006888) |
| 0.1 | 3.9 | GO:0051028 | mRNA transport(GO:0051028) |
| 0.1 | 5.3 | GO:0051297 | centrosome organization(GO:0051297) |
| 0.1 | 0.7 | GO:0006515 | misfolded or incompletely synthesized protein catabolic process(GO:0006515) chaperone-mediated protein complex assembly(GO:0051131) |
| 0.0 | 4.9 | GO:0098792 | xenophagy(GO:0098792) |
| 0.0 | 0.6 | GO:0060712 | spongiotrophoblast layer development(GO:0060712) |
| 0.0 | 0.7 | GO:0008209 | androgen metabolic process(GO:0008209) |
| 0.0 | 0.3 | GO:0008054 | negative regulation of cyclin-dependent protein serine/threonine kinase by cyclin degradation(GO:0008054) |
| 0.0 | 2.7 | GO:0043966 | histone H3 acetylation(GO:0043966) |
| 0.0 | 0.3 | GO:0018026 | peptidyl-lysine monomethylation(GO:0018026) |
| 0.0 | 1.7 | GO:0032543 | mitochondrial translation(GO:0032543) |
| 0.0 | 0.2 | GO:0036506 | maintenance of unfolded protein(GO:0036506) protein insertion into ER membrane(GO:0045048) tail-anchored membrane protein insertion into ER membrane(GO:0071816) maintenance of unfolded protein involved in ERAD pathway(GO:1904378) |
| 0.0 | 0.5 | GO:0003333 | amino acid transmembrane transport(GO:0003333) |
| 0.0 | 0.1 | GO:0015817 | histidine transport(GO:0015817) |
| 0.0 | 4.2 | GO:0006260 | DNA replication(GO:0006260) |
| 0.0 | 1.2 | GO:0006096 | glycolytic process(GO:0006096) ATP generation from ADP(GO:0006757) |
| 0.0 | 0.8 | GO:0007585 | respiratory gaseous exchange(GO:0007585) |
| 0.0 | 2.6 | GO:0030217 | T cell differentiation(GO:0030217) |
| 0.0 | 2.1 | GO:0042384 | cilium assembly(GO:0042384) |
| 0.0 | 2.7 | GO:0007018 | microtubule-based movement(GO:0007018) |
| 0.0 | 0.3 | GO:0000413 | protein peptidyl-prolyl isomerization(GO:0000413) |
| 0.0 | 1.1 | GO:0006633 | fatty acid biosynthetic process(GO:0006633) |
Gene overrepresentation in cellular_component category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 1.4 | 4.2 | GO:0043224 | nuclear SCF ubiquitin ligase complex(GO:0043224) |
| 1.4 | 4.2 | GO:0043527 | tRNA methyltransferase complex(GO:0043527) |
| 1.4 | 8.4 | GO:0005663 | DNA replication factor C complex(GO:0005663) |
| 1.4 | 6.9 | GO:0044615 | nuclear pore nuclear basket(GO:0044615) |
| 0.8 | 2.5 | GO:0005658 | alpha DNA polymerase:primase complex(GO:0005658) |
| 0.8 | 2.4 | GO:0005588 | collagen type V trimer(GO:0005588) |
| 0.8 | 7.3 | GO:0008278 | cohesin complex(GO:0008278) |
| 0.7 | 4.0 | GO:1990726 | Lsm1-7-Pat1 complex(GO:1990726) |
| 0.6 | 1.9 | GO:0000408 | EKC/KEOPS complex(GO:0000408) |
| 0.6 | 6.7 | GO:0042405 | nuclear inclusion body(GO:0042405) |
| 0.6 | 2.8 | GO:0002193 | MAML1-RBP-Jkappa- ICN1 complex(GO:0002193) |
| 0.5 | 3.1 | GO:0030127 | COPII vesicle coat(GO:0030127) |
| 0.5 | 5.1 | GO:0090576 | RNA polymerase III transcription factor complex(GO:0090576) |
| 0.5 | 3.9 | GO:0070652 | HAUS complex(GO:0070652) |
| 0.5 | 3.2 | GO:0097452 | GAIT complex(GO:0097452) |
| 0.4 | 1.3 | GO:0034274 | Atg12-Atg5-Atg16 complex(GO:0034274) |
| 0.4 | 4.3 | GO:0000778 | condensed nuclear chromosome kinetochore(GO:0000778) |
| 0.4 | 1.2 | GO:0002944 | cyclin K-CDK12 complex(GO:0002944) |
| 0.4 | 2.4 | GO:0030289 | protein phosphatase 4 complex(GO:0030289) |
| 0.3 | 2.4 | GO:0090543 | Flemming body(GO:0090543) |
| 0.3 | 3.0 | GO:0000347 | THO complex(GO:0000347) THO complex part of transcription export complex(GO:0000445) |
| 0.3 | 5.0 | GO:0000177 | cytoplasmic exosome (RNase complex)(GO:0000177) |
| 0.3 | 2.0 | GO:0071141 | SMAD protein complex(GO:0071141) |
| 0.3 | 1.4 | GO:0030689 | Noc complex(GO:0030689) |
| 0.3 | 2.4 | GO:0000439 | core TFIIH complex(GO:0000439) |
| 0.3 | 0.8 | GO:0005785 | signal recognition particle receptor complex(GO:0005785) |
| 0.3 | 2.0 | GO:0030915 | Smc5-Smc6 complex(GO:0030915) |
| 0.2 | 1.0 | GO:0031084 | BLOC-2 complex(GO:0031084) |
| 0.2 | 2.0 | GO:0042382 | paraspeckles(GO:0042382) |
| 0.2 | 3.9 | GO:0001650 | fibrillar center(GO:0001650) |
| 0.2 | 2.6 | GO:0005687 | U4 snRNP(GO:0005687) |
| 0.2 | 0.9 | GO:0000931 | gamma-tubulin large complex(GO:0000931) gamma-tubulin ring complex(GO:0008274) |
| 0.2 | 13.0 | GO:0005871 | kinesin complex(GO:0005871) |
| 0.2 | 0.8 | GO:0008537 | proteasome activator complex(GO:0008537) |
| 0.2 | 0.7 | GO:0016602 | CCAAT-binding factor complex(GO:0016602) |
| 0.2 | 2.2 | GO:0051286 | cell tip(GO:0051286) |
| 0.2 | 8.0 | GO:0000307 | cyclin-dependent protein kinase holoenzyme complex(GO:0000307) |
| 0.2 | 3.5 | GO:0030014 | CCR4-NOT complex(GO:0030014) |
| 0.2 | 1.9 | GO:0072546 | ER membrane protein complex(GO:0072546) |
| 0.1 | 2.2 | GO:0031528 | microvillus membrane(GO:0031528) |
| 0.1 | 0.6 | GO:0042827 | platelet dense granule(GO:0042827) |
| 0.1 | 5.6 | GO:0035869 | ciliary transition zone(GO:0035869) |
| 0.1 | 2.7 | GO:0005671 | Ada2/Gcn5/Ada3 transcription activator complex(GO:0005671) |
| 0.1 | 0.9 | GO:0005958 | DNA-dependent protein kinase-DNA ligase 4 complex(GO:0005958) |
| 0.1 | 2.0 | GO:0016600 | flotillin complex(GO:0016600) |
| 0.1 | 1.0 | GO:0000940 | condensed chromosome outer kinetochore(GO:0000940) |
| 0.1 | 8.5 | GO:0005643 | nuclear pore(GO:0005643) |
| 0.1 | 1.6 | GO:0030056 | hemidesmosome(GO:0030056) |
| 0.1 | 0.9 | GO:0005890 | sodium:potassium-exchanging ATPase complex(GO:0005890) |
| 0.1 | 2.9 | GO:0070822 | Sin3-type complex(GO:0070822) |
| 0.1 | 2.0 | GO:0005852 | eukaryotic translation initiation factor 3 complex(GO:0005852) |
| 0.1 | 15.4 | GO:0000776 | kinetochore(GO:0000776) |
| 0.1 | 1.2 | GO:1904115 | axon cytoplasm(GO:1904115) |
| 0.1 | 1.7 | GO:0042555 | MCM complex(GO:0042555) |
| 0.1 | 0.5 | GO:0071006 | U2-type catalytic step 1 spliceosome(GO:0071006) |
| 0.1 | 1.1 | GO:0044232 | organelle membrane contact site(GO:0044232) |
| 0.1 | 5.6 | GO:0005801 | cis-Golgi network(GO:0005801) |
| 0.1 | 1.5 | GO:0034518 | mRNA cap binding complex(GO:0005845) RNA cap binding complex(GO:0034518) |
| 0.1 | 2.0 | GO:0071564 | npBAF complex(GO:0071564) |
| 0.1 | 3.8 | GO:0000315 | organellar large ribosomal subunit(GO:0000315) mitochondrial large ribosomal subunit(GO:0005762) |
| 0.1 | 0.9 | GO:1990909 | Wnt signalosome(GO:1990909) |
| 0.1 | 1.8 | GO:0005942 | phosphatidylinositol 3-kinase complex(GO:0005942) |
| 0.1 | 6.9 | GO:0000793 | condensed chromosome(GO:0000793) |
| 0.1 | 1.4 | GO:0000242 | pericentriolar material(GO:0000242) |
| 0.1 | 2.6 | GO:0035861 | site of double-strand break(GO:0035861) |
| 0.1 | 5.5 | GO:0005811 | lipid particle(GO:0005811) |
| 0.1 | 1.4 | GO:0000930 | gamma-tubulin complex(GO:0000930) |
| 0.1 | 0.4 | GO:0005850 | eukaryotic translation initiation factor 2 complex(GO:0005850) |
| 0.1 | 1.6 | GO:0005669 | transcription factor TFIID complex(GO:0005669) |
| 0.1 | 1.9 | GO:0002102 | podosome(GO:0002102) |
| 0.1 | 1.1 | GO:0035631 | CD40 receptor complex(GO:0035631) |
| 0.1 | 9.3 | GO:0030139 | endocytic vesicle(GO:0030139) |
| 0.1 | 2.1 | GO:0030687 | preribosome, large subunit precursor(GO:0030687) |
| 0.1 | 0.3 | GO:0032021 | NELF complex(GO:0032021) |
| 0.1 | 1.1 | GO:0005736 | DNA-directed RNA polymerase I complex(GO:0005736) |
| 0.1 | 0.7 | GO:0005686 | U2 snRNP(GO:0005686) |
| 0.1 | 0.2 | GO:0005726 | perichromatin fibrils(GO:0005726) |
| 0.1 | 0.7 | GO:0071203 | WASH complex(GO:0071203) |
| 0.1 | 1.9 | GO:0035145 | exon-exon junction complex(GO:0035145) |
| 0.0 | 2.4 | GO:0022627 | cytosolic small ribosomal subunit(GO:0022627) |
| 0.0 | 3.6 | GO:0017053 | transcriptional repressor complex(GO:0017053) |
| 0.0 | 3.5 | GO:0090575 | RNA polymerase II transcription factor complex(GO:0090575) |
| 0.0 | 1.4 | GO:0008023 | transcription elongation factor complex(GO:0008023) |
| 0.0 | 12.3 | GO:0005667 | transcription factor complex(GO:0005667) |
| 0.0 | 4.8 | GO:0000784 | nuclear chromosome, telomeric region(GO:0000784) |
| 0.0 | 0.2 | GO:0005745 | m-AAA complex(GO:0005745) |
| 0.0 | 0.5 | GO:0044292 | dendrite terminus(GO:0044292) |
| 0.0 | 1.7 | GO:0042645 | nucleoid(GO:0009295) mitochondrial nucleoid(GO:0042645) |
| 0.0 | 2.2 | GO:0019005 | SCF ubiquitin ligase complex(GO:0019005) |
| 0.0 | 2.5 | GO:0005758 | mitochondrial intermembrane space(GO:0005758) |
| 0.0 | 24.5 | GO:0005730 | nucleolus(GO:0005730) |
| 0.0 | 4.0 | GO:0030176 | integral component of endoplasmic reticulum membrane(GO:0030176) |
| 0.0 | 0.6 | GO:0016592 | mediator complex(GO:0016592) |
| 0.0 | 1.7 | GO:0031519 | PcG protein complex(GO:0031519) |
| 0.0 | 11.2 | GO:0005694 | chromosome(GO:0005694) |
| 0.0 | 2.6 | GO:0031513 | nonmotile primary cilium(GO:0031513) |
| 0.0 | 1.0 | GO:0016605 | PML body(GO:0016605) |
| 0.0 | 1.6 | GO:0032587 | ruffle membrane(GO:0032587) |
| 0.0 | 0.2 | GO:0071818 | BAT3 complex(GO:0071818) ER membrane insertion complex(GO:0072379) |
| 0.0 | 0.9 | GO:0031901 | early endosome membrane(GO:0031901) |
| 0.0 | 0.2 | GO:0032580 | Golgi cisterna membrane(GO:0032580) |
| 0.0 | 2.4 | GO:0005759 | mitochondrial matrix(GO:0005759) |
| 0.0 | 0.4 | GO:0005839 | proteasome core complex(GO:0005839) |
| 0.0 | 0.6 | GO:0030964 | mitochondrial respiratory chain complex I(GO:0005747) NADH dehydrogenase complex(GO:0030964) respiratory chain complex I(GO:0045271) |
Gene overrepresentation in molecular_function category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 1.5 | 9.3 | GO:0035727 | lysophosphatidic acid binding(GO:0035727) lysophosphatidic acid receptor activity(GO:0070915) |
| 1.5 | 4.6 | GO:0010484 | H3 histone acetyltransferase activity(GO:0010484) |
| 1.4 | 6.8 | GO:0000405 | bubble DNA binding(GO:0000405) |
| 1.3 | 4.0 | GO:0008176 | tRNA (guanine-N7-)-methyltransferase activity(GO:0008176) |
| 1.1 | 4.3 | GO:0043515 | kinetochore binding(GO:0043515) |
| 1.0 | 4.0 | GO:0000403 | Y-form DNA binding(GO:0000403) |
| 0.8 | 8.4 | GO:0003689 | DNA clamp loader activity(GO:0003689) protein-DNA loading ATPase activity(GO:0033170) |
| 0.7 | 2.2 | GO:0070137 | ubiquitin-like protein-specific endopeptidase activity(GO:0070137) SUMO-specific endopeptidase activity(GO:0070139) |
| 0.7 | 3.4 | GO:0070840 | dynein complex binding(GO:0070840) |
| 0.6 | 1.9 | GO:0061711 | N(6)-L-threonylcarbamoyladenine synthase(GO:0061711) |
| 0.6 | 4.2 | GO:0019788 | NEDD8 transferase activity(GO:0019788) |
| 0.6 | 1.8 | GO:0019166 | trans-2-enoyl-CoA reductase (NADPH) activity(GO:0019166) |
| 0.6 | 7.0 | GO:0001161 | intronic transcription regulatory region sequence-specific DNA binding(GO:0001161) |
| 0.5 | 3.1 | GO:0050786 | RAGE receptor binding(GO:0050786) |
| 0.5 | 2.6 | GO:0030621 | U4 snRNA binding(GO:0030621) |
| 0.5 | 1.5 | GO:0000340 | RNA 7-methylguanosine cap binding(GO:0000340) |
| 0.4 | 1.3 | GO:0019776 | Atg8 ligase activity(GO:0019776) |
| 0.4 | 3.5 | GO:0043495 | protein anchor(GO:0043495) |
| 0.4 | 1.9 | GO:0016176 | superoxide-generating NADPH oxidase activator activity(GO:0016176) |
| 0.4 | 1.5 | GO:0015232 | heme transporter activity(GO:0015232) |
| 0.4 | 2.2 | GO:0061656 | SUMO conjugating enzyme activity(GO:0061656) |
| 0.4 | 5.0 | GO:0070182 | DNA polymerase binding(GO:0070182) |
| 0.4 | 2.5 | GO:0003896 | DNA primase activity(GO:0003896) |
| 0.4 | 3.5 | GO:0052872 | 3-(3-hydroxyphenyl)propionate hydroxylase activity(GO:0008688) 4-chlorobenzaldehyde oxidase activity(GO:0018471) 3,5-xylenol methylhydroxylase activity(GO:0018630) phenylacetate hydroxylase activity(GO:0018631) 4-nitrophenol 4-monooxygenase activity(GO:0018632) dimethyl sulfide monooxygenase activity(GO:0018633) alpha-pinene monooxygenase [NADH] activity(GO:0018634) phenanthrene 9,10-monooxygenase activity(GO:0018636) 1-hydroxy-2-naphthoate hydroxylase activity(GO:0018637) toluene 4-monooxygenase activity(GO:0018638) xylene monooxygenase activity(GO:0018639) dibenzothiophene monooxygenase activity(GO:0018640) 6-hydroxy-3-methyl-2-oxo-1,2-dihydroquinoline 6-monooxygenase activity(GO:0018641) chlorophenol 4-monooxygenase activity(GO:0018642) carbon disulfide oxygenase activity(GO:0018643) toluene 2-monooxygenase activity(GO:0018644) 1-hydroxy-2-oxolimonene 1,2-monooxygenase activity(GO:0018646) phenanthrene 1,2-monooxygenase activity(GO:0018647) tetrahydrofuran hydroxylase activity(GO:0018649) styrene monooxygenase activity(GO:0018650) toluene-4-sulfonate monooxygenase activity(GO:0018651) toluene-sulfonate methyl-monooxygenase activity(GO:0018652) 3-methyl-2-oxo-1,2-dihydroquinoline 6-monooxygenase activity(GO:0018653) 2-hydroxy-phenylacetate hydroxylase activity(GO:0018654) 2-oxo-delta3-4,5,5-trimethylcyclopentenylacetyl-CoA 1,2-monooxygenase activity(GO:0018655) phenanthrene 3,4-monooxygenase activity(GO:0018656) toluene 3-monooxygenase activity(GO:0018657) 4-hydroxyphenylacetate,NADH:oxygen oxidoreductase (3-hydroxylating) activity(GO:0018660) limonene monooxygenase activity(GO:0019113) 2-methylnaphthalene hydroxylase activity(GO:0034526) 1-methylnaphthalene hydroxylase activity(GO:0034534) bisphenol A hydroxylase A activity(GO:0034560) salicylate 5-hydroxylase activity(GO:0034785) isobutylamine N-hydroxylase activity(GO:0034791) branched-chain dodecylbenzene sulfonate monooxygenase activity(GO:0034802) 3-HSA hydroxylase activity(GO:0034819) 4-hydroxypyridine-3-hydroxylase activity(GO:0034894) 2-octaprenyl-3-methyl-6-methoxy-1,4-benzoquinol hydroxylase activity(GO:0043719) 6-hydroxynicotinate 3-monooxygenase activity(GO:0043731) tocotrienol omega-hydroxylase activity(GO:0052872) thalianol hydroxylase activity(GO:0080014) |
| 0.3 | 1.6 | GO:0038132 | neuregulin binding(GO:0038132) |
| 0.3 | 2.2 | GO:0030911 | TPR domain binding(GO:0030911) |
| 0.3 | 1.6 | GO:0032564 | dATP binding(GO:0032564) |
| 0.3 | 2.4 | GO:0015087 | cobalt ion transmembrane transporter activity(GO:0015087) |
| 0.3 | 3.6 | GO:0016884 | carbon-nitrogen ligase activity, with glutamine as amido-N-donor(GO:0016884) |
| 0.3 | 2.9 | GO:0000014 | single-stranded DNA endodeoxyribonuclease activity(GO:0000014) |
| 0.3 | 3.1 | GO:0010521 | telomerase inhibitor activity(GO:0010521) |
| 0.3 | 3.3 | GO:0000700 | mismatch base pair DNA N-glycosylase activity(GO:0000700) |
| 0.2 | 4.4 | GO:0005487 | nucleocytoplasmic transporter activity(GO:0005487) |
| 0.2 | 1.7 | GO:0008479 | queuine tRNA-ribosyltransferase activity(GO:0008479) |
| 0.2 | 2.4 | GO:0008474 | palmitoyl-(protein) hydrolase activity(GO:0008474) palmitoyl hydrolase activity(GO:0098599) |
| 0.2 | 4.6 | GO:0017056 | structural constituent of nuclear pore(GO:0017056) |
| 0.2 | 7.0 | GO:0017147 | Wnt-protein binding(GO:0017147) |
| 0.2 | 12.6 | GO:0000049 | tRNA binding(GO:0000049) |
| 0.2 | 2.4 | GO:0102338 | fatty acid elongase activity(GO:0009922) 3-oxo-arachidoyl-CoA synthase activity(GO:0102336) 3-oxo-cerotoyl-CoA synthase activity(GO:0102337) 3-oxo-lignoceronyl-CoA synthase activity(GO:0102338) |
| 0.2 | 3.3 | GO:0015095 | magnesium ion transmembrane transporter activity(GO:0015095) |
| 0.2 | 3.1 | GO:0016709 | oxidoreductase activity, acting on paired donors, with incorporation or reduction of molecular oxygen, NAD(P)H as one donor, and incorporation of one atom of oxygen(GO:0016709) |
| 0.2 | 1.0 | GO:0030618 | transforming growth factor beta receptor, pathway-specific cytoplasmic mediator activity(GO:0030618) |
| 0.2 | 13.0 | GO:0003777 | microtubule motor activity(GO:0003777) |
| 0.2 | 3.5 | GO:0004535 | poly(A)-specific ribonuclease activity(GO:0004535) |
| 0.2 | 3.6 | GO:0102391 | decanoate--CoA ligase activity(GO:0102391) |
| 0.2 | 3.6 | GO:0008353 | RNA polymerase II carboxy-terminal domain kinase activity(GO:0008353) |
| 0.2 | 1.1 | GO:1990829 | C-rich single-stranded DNA binding(GO:1990829) |
| 0.2 | 1.8 | GO:0035005 | 1-phosphatidylinositol-4-phosphate 3-kinase activity(GO:0035005) |
| 0.2 | 1.6 | GO:0000182 | rDNA binding(GO:0000182) |
| 0.2 | 3.2 | GO:0035613 | RNA stem-loop binding(GO:0035613) |
| 0.2 | 4.0 | GO:0017160 | Ral GTPase binding(GO:0017160) |
| 0.2 | 2.4 | GO:0048407 | platelet-derived growth factor binding(GO:0048407) |
| 0.2 | 2.6 | GO:0042809 | vitamin D receptor binding(GO:0042809) |
| 0.2 | 4.5 | GO:0033130 | acetylcholine receptor binding(GO:0033130) |
| 0.2 | 0.8 | GO:0097001 | sphingolipid transporter activity(GO:0046624) ceramide binding(GO:0097001) |
| 0.2 | 3.5 | GO:0019789 | SUMO transferase activity(GO:0019789) |
| 0.2 | 7.2 | GO:0016538 | cyclin-dependent protein serine/threonine kinase regulator activity(GO:0016538) |
| 0.2 | 0.8 | GO:0005047 | signal recognition particle binding(GO:0005047) |
| 0.2 | 2.0 | GO:0043024 | ribosomal small subunit binding(GO:0043024) |
| 0.2 | 2.1 | GO:0051400 | BH domain binding(GO:0051400) |
| 0.1 | 4.3 | GO:0017091 | AU-rich element binding(GO:0017091) |
| 0.1 | 1.1 | GO:0008429 | phosphatidylethanolamine binding(GO:0008429) |
| 0.1 | 8.5 | GO:0008186 | ATP-dependent RNA helicase activity(GO:0004004) RNA-dependent ATPase activity(GO:0008186) |
| 0.1 | 0.8 | GO:1990050 | phosphatidic acid transporter activity(GO:1990050) |
| 0.1 | 2.0 | GO:0036312 | phosphatidylinositol 3-kinase regulatory subunit binding(GO:0036312) |
| 0.1 | 1.4 | GO:0008758 | thiamine-pyrophosphatase activity(GO:0004787) UDP-2,3-diacylglucosamine hydrolase activity(GO:0008758) dATP pyrophosphohydrolase activity(GO:0008828) dihydroneopterin monophosphate phosphatase activity(GO:0019176) dihydroneopterin triphosphate pyrophosphohydrolase activity(GO:0019177) dTTP diphosphatase activity(GO:0036218) phosphocholine hydrolase activity(GO:0044606) |
| 0.1 | 0.4 | GO:0035851 | histone deacetylase activity (H4-K16 specific)(GO:0034739) Krueppel-associated box domain binding(GO:0035851) |
| 0.1 | 0.8 | GO:0061133 | endopeptidase activator activity(GO:0061133) |
| 0.1 | 0.3 | GO:0003963 | RNA-3'-phosphate cyclase activity(GO:0003963) |
| 0.1 | 2.2 | GO:0034185 | apolipoprotein binding(GO:0034185) |
| 0.1 | 0.4 | GO:0008240 | tripeptidyl-peptidase activity(GO:0008240) |
| 0.1 | 0.9 | GO:0051011 | microtubule minus-end binding(GO:0051011) |
| 0.1 | 2.8 | GO:0031435 | mitogen-activated protein kinase kinase kinase binding(GO:0031435) |
| 0.1 | 0.9 | GO:0008556 | sodium:potassium-exchanging ATPase activity(GO:0005391) potassium-transporting ATPase activity(GO:0008556) |
| 0.1 | 2.5 | GO:0003887 | DNA-directed DNA polymerase activity(GO:0003887) |
| 0.1 | 1.1 | GO:0031996 | thioesterase binding(GO:0031996) |
| 0.1 | 0.8 | GO:0004571 | mannosyl-oligosaccharide 1,2-alpha-mannosidase activity(GO:0004571) |
| 0.1 | 1.2 | GO:0070180 | large ribosomal subunit rRNA binding(GO:0070180) |
| 0.1 | 2.0 | GO:0016887 | ATPase activity(GO:0016887) |
| 0.1 | 1.0 | GO:0008301 | DNA binding, bending(GO:0008301) |
| 0.1 | 2.8 | GO:0001103 | RNA polymerase II repressing transcription factor binding(GO:0001103) |
| 0.1 | 2.6 | GO:0043425 | bHLH transcription factor binding(GO:0043425) |
| 0.1 | 1.6 | GO:0003841 | 1-acylglycerol-3-phosphate O-acyltransferase activity(GO:0003841) |
| 0.1 | 3.9 | GO:0031491 | nucleosome binding(GO:0031491) |
| 0.1 | 1.1 | GO:0001054 | RNA polymerase I activity(GO:0001054) |
| 0.1 | 1.9 | GO:0004407 | histone deacetylase activity(GO:0004407) |
| 0.1 | 1.9 | GO:0003785 | actin monomer binding(GO:0003785) |
| 0.1 | 4.7 | GO:0009019 | tRNA (guanine-N1-)-methyltransferase activity(GO:0009019) |
| 0.1 | 1.4 | GO:0001106 | RNA polymerase II transcription corepressor activity(GO:0001106) |
| 0.1 | 0.7 | GO:0019215 | intermediate filament binding(GO:0019215) |
| 0.1 | 1.3 | GO:0004003 | ATP-dependent DNA helicase activity(GO:0004003) |
| 0.1 | 0.3 | GO:0046975 | histone methyltransferase activity (H3-K36 specific)(GO:0046975) |
| 0.1 | 0.9 | GO:0070411 | I-SMAD binding(GO:0070411) |
| 0.0 | 4.5 | GO:0004843 | thiol-dependent ubiquitin-specific protease activity(GO:0004843) |
| 0.0 | 0.3 | GO:0004013 | adenosylhomocysteinase activity(GO:0004013) trialkylsulfonium hydrolase activity(GO:0016802) |
| 0.0 | 0.7 | GO:0070491 | repressing transcription factor binding(GO:0070491) |
| 0.0 | 1.2 | GO:0008536 | Ran GTPase binding(GO:0008536) |
| 0.0 | 0.5 | GO:0005149 | interleukin-1 receptor binding(GO:0005149) |
| 0.0 | 7.7 | GO:0001077 | transcriptional activator activity, RNA polymerase II core promoter proximal region sequence-specific binding(GO:0001077) |
| 0.0 | 5.5 | GO:0001047 | core promoter binding(GO:0001047) |
| 0.0 | 3.3 | GO:0003705 | transcription factor activity, RNA polymerase II distal enhancer sequence-specific binding(GO:0003705) |
| 0.0 | 2.2 | GO:0004386 | helicase activity(GO:0004386) |
| 0.0 | 0.7 | GO:0030506 | ankyrin binding(GO:0030506) |
| 0.0 | 0.9 | GO:0034945 | dihydrolipoamide branched chain acyltransferase activity(GO:0004147) palmitoleoyl [acyl-carrier-protein]-dependent acyltransferase activity(GO:0008951) serine O-acyltransferase activity(GO:0016412) O-succinyltransferase activity(GO:0016750) sinapoyltransferase activity(GO:0016752) O-sinapoyltransferase activity(GO:0016753) peptidyl-lysine N6-myristoyltransferase activity(GO:0018030) peptidyl-lysine N6-palmitoyltransferase activity(GO:0018031) benzoyl acetate-CoA thiolase activity(GO:0018711) 3-hydroxybutyryl-CoA thiolase activity(GO:0018712) 3-ketopimelyl-CoA thiolase activity(GO:0018713) N-palmitoyltransferase activity(GO:0019105) acyl-CoA N-acyltransferase activity(GO:0019186) protein-cysteine S-myristoyltransferase activity(GO:0019705) glucosaminyl-phosphotidylinositol O-acyltransferase activity(GO:0032216) ergosterol O-acyltransferase activity(GO:0034737) lanosterol O-acyltransferase activity(GO:0034738) naphthyl-2-oxomethyl-succinyl-CoA succinyl transferase activity(GO:0034848) 2,4,4-trimethyl-3-oxopentanoyl-CoA 2-C-propanoyl transferase activity(GO:0034851) 2-methylhexanoyl-CoA C-acetyltransferase activity(GO:0034915) butyryl-CoA 2-C-propionyltransferase activity(GO:0034919) 2,6-dimethyl-5-methylene-3-oxo-heptanoyl-CoA C-acetyltransferase activity(GO:0034945) L-2-aminoadipate N-acetyltransferase activity(GO:0043741) keto acid formate lyase activity(GO:0043806) azetidine-2-carboxylic acid acetyltransferase activity(GO:0046941) peptidyl-lysine N-acetyltransferase activity, acting on acetyl phosphate as donor(GO:0052858) acetyl-CoA:L-lysine N6-acetyltransferase(GO:0090595) |
| 0.0 | 1.2 | GO:0004197 | cysteine-type endopeptidase activity(GO:0004197) |
| 0.0 | 6.2 | GO:0003735 | structural constituent of ribosome(GO:0003735) |
| 0.0 | 0.7 | GO:0035326 | enhancer binding(GO:0035326) |
| 0.0 | 1.2 | GO:0035064 | methylated histone binding(GO:0035064) |
| 0.0 | 0.2 | GO:0030515 | snoRNA binding(GO:0030515) |
| 0.0 | 2.5 | GO:0042393 | histone binding(GO:0042393) |
| 0.0 | 0.2 | GO:0101005 | thiol-dependent ubiquitinyl hydrolase activity(GO:0036459) ubiquitinyl hydrolase activity(GO:0101005) |
| 0.0 | 21.4 | GO:0044822 | poly(A) RNA binding(GO:0044822) |
| 0.0 | 0.4 | GO:0004298 | threonine-type endopeptidase activity(GO:0004298) threonine-type peptidase activity(GO:0070003) |
| 0.0 | 0.4 | GO:0003743 | translation initiation factor activity(GO:0003743) |
| 0.0 | 0.6 | GO:0001104 | RNA polymerase II transcription cofactor activity(GO:0001104) |
| 0.0 | 3.6 | GO:0003723 | RNA binding(GO:0003723) |
| 0.0 | 0.1 | GO:0005290 | L-histidine transmembrane transporter activity(GO:0005290) |
| 0.0 | 5.5 | GO:0046982 | protein heterodimerization activity(GO:0046982) |
| 0.0 | 2.9 | GO:0043774 | UDP-N-acetylmuramoylalanyl-D-glutamyl-2,6-diaminopimelate-D-alanyl-D-alanine ligase activity(GO:0008766) ribosomal S6-glutamic acid ligase activity(GO:0018169) coenzyme F420-0 gamma-glutamyl ligase activity(GO:0043773) coenzyme F420-2 alpha-glutamyl ligase activity(GO:0043774) protein-glycine ligase activity(GO:0070735) protein-glycine ligase activity, initiating(GO:0070736) protein-glycine ligase activity, elongating(GO:0070737) tubulin-glycine ligase activity(GO:0070738) |
| 0.0 | 3.2 | GO:0003682 | chromatin binding(GO:0003682) |
| 0.0 | 1.5 | GO:0001948 | glycoprotein binding(GO:0001948) |
| 0.0 | 0.2 | GO:0008308 | voltage-gated anion channel activity(GO:0008308) |