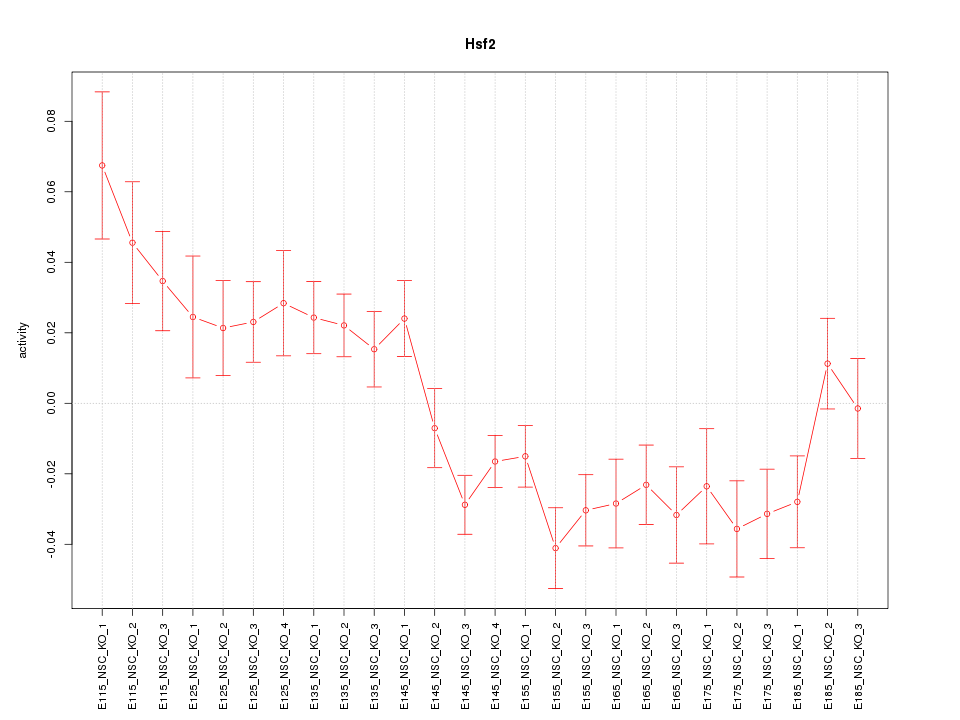
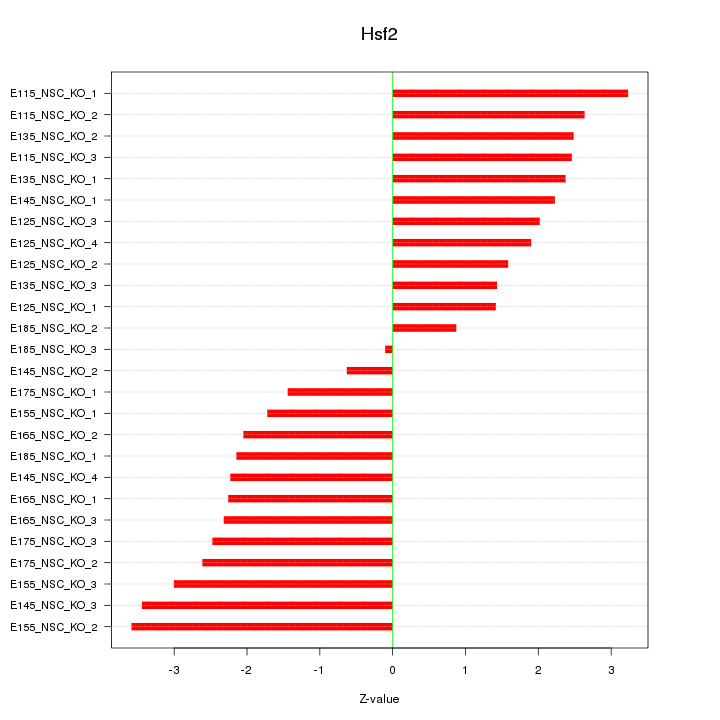
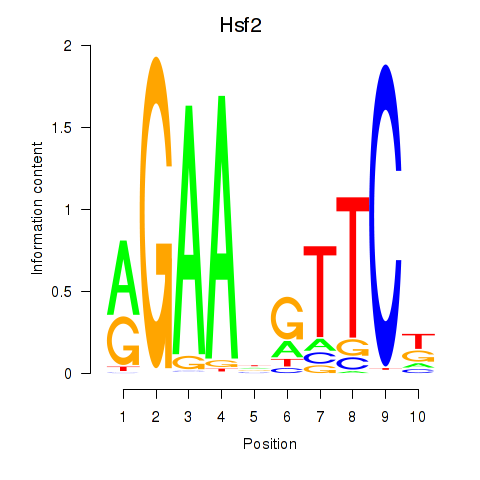
Motif ID: Hsf2
Z-value: 2.251
Transcription factors associated with Hsf2:
| Gene Symbol | Entrez ID | Gene Name |
|---|---|---|
| Hsf2 | ENSMUSG00000019878.7 | Hsf2 |
Activity-expression correlation:
| Gene Symbol | Promoter | Pearson corr. coef. | P-value | Plot |
|---|---|---|---|---|
| Hsf2 | mm10_v2_chr10_+_57486354_57486414 | 0.47 | 1.6e-02 | Click! |
Top targets:
Gene overrepresentation in biological_process category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 2.2 | 8.9 | GO:0021698 | cerebellar cortex structural organization(GO:0021698) |
| 2.2 | 8.7 | GO:0046552 | eye photoreceptor cell fate commitment(GO:0042706) photoreceptor cell fate commitment(GO:0046552) |
| 1.9 | 5.6 | GO:0002842 | T cell mediated immune response to tumor cell(GO:0002424) regulation of T cell mediated immune response to tumor cell(GO:0002840) positive regulation of T cell mediated immune response to tumor cell(GO:0002842) |
| 1.8 | 10.7 | GO:0045583 | regulation of cytotoxic T cell differentiation(GO:0045583) positive regulation of cytotoxic T cell differentiation(GO:0045585) |
| 1.5 | 4.6 | GO:0021759 | globus pallidus development(GO:0021759) |
| 1.5 | 6.0 | GO:2000974 | negative regulation of mechanoreceptor differentiation(GO:0045632) negative regulation of pro-B cell differentiation(GO:2000974) negative regulation of inner ear receptor cell differentiation(GO:2000981) |
| 1.4 | 4.2 | GO:0070315 | G1 to G0 transition involved in cell differentiation(GO:0070315) |
| 1.4 | 8.2 | GO:1903690 | negative regulation of wound healing, spreading of epidermal cells(GO:1903690) |
| 1.3 | 21.0 | GO:0048387 | negative regulation of retinoic acid receptor signaling pathway(GO:0048387) |
| 1.2 | 3.6 | GO:0030421 | defecation(GO:0030421) |
| 1.2 | 6.0 | GO:0021914 | negative regulation of smoothened signaling pathway involved in ventral spinal cord patterning(GO:0021914) |
| 1.1 | 3.4 | GO:1900275 | negative regulation of phospholipase C activity(GO:1900275) |
| 1.0 | 6.2 | GO:0035743 | CD4-positive, alpha-beta T cell cytokine production(GO:0035743) T-helper 2 cell cytokine production(GO:0035745) |
| 1.0 | 4.0 | GO:0002337 | B-1a B cell differentiation(GO:0002337) |
| 1.0 | 4.9 | GO:0035262 | gonad morphogenesis(GO:0035262) |
| 1.0 | 3.9 | GO:0048631 | regulation of skeletal muscle tissue growth(GO:0048631) |
| 0.8 | 4.2 | GO:0006348 | chromatin silencing at telomere(GO:0006348) |
| 0.8 | 3.4 | GO:0014846 | esophagus smooth muscle contraction(GO:0014846) |
| 0.8 | 2.5 | GO:0003360 | brainstem development(GO:0003360) |
| 0.8 | 1.6 | GO:0072076 | nephrogenic mesenchyme development(GO:0072076) |
| 0.8 | 4.0 | GO:0034421 | post-translational protein acetylation(GO:0034421) |
| 0.8 | 3.1 | GO:2000313 | fibroblast growth factor receptor signaling pathway involved in neural plate anterior/posterior pattern formation(GO:0060825) regulation of fibroblast growth factor receptor signaling pathway involved in neural plate anterior/posterior pattern formation(GO:2000313) |
| 0.8 | 3.0 | GO:0033326 | cerebrospinal fluid secretion(GO:0033326) |
| 0.7 | 2.2 | GO:0045819 | positive regulation of glycogen catabolic process(GO:0045819) |
| 0.7 | 2.9 | GO:0042276 | error-prone translesion synthesis(GO:0042276) |
| 0.7 | 2.0 | GO:0040009 | nucleolus to nucleoplasm transport(GO:0032066) regulation of growth rate(GO:0040009) |
| 0.7 | 2.0 | GO:0097461 | ferric iron import(GO:0033216) ferric iron import into cell(GO:0097461) |
| 0.6 | 3.9 | GO:0043137 | DNA replication, removal of RNA primer(GO:0043137) |
| 0.6 | 4.4 | GO:0043569 | negative regulation of insulin-like growth factor receptor signaling pathway(GO:0043569) |
| 0.6 | 5.5 | GO:0045719 | negative regulation of glycogen biosynthetic process(GO:0045719) negative regulation of glycogen metabolic process(GO:0070874) |
| 0.6 | 2.4 | GO:0060032 | notochord regression(GO:0060032) |
| 0.6 | 4.8 | GO:2000020 | positive regulation of male gonad development(GO:2000020) |
| 0.6 | 2.4 | GO:0045343 | MHC class I biosynthetic process(GO:0045341) regulation of MHC class I biosynthetic process(GO:0045343) positive regulation of MHC class I biosynthetic process(GO:0045345) |
| 0.6 | 1.8 | GO:1990046 | positive regulation of mitochondrial DNA replication(GO:0090297) regulation of cardiolipin metabolic process(GO:1900208) positive regulation of cardiolipin metabolic process(GO:1900210) stress-induced mitochondrial fusion(GO:1990046) |
| 0.6 | 4.1 | GO:0032534 | regulation of microvillus assembly(GO:0032534) |
| 0.6 | 2.8 | GO:0060040 | retinal bipolar neuron differentiation(GO:0060040) |
| 0.6 | 3.4 | GO:0071883 | activation of MAPK activity by adrenergic receptor signaling pathway(GO:0071883) |
| 0.5 | 4.3 | GO:0016081 | synaptic vesicle docking(GO:0016081) |
| 0.5 | 2.0 | GO:0007056 | spindle assembly involved in female meiosis(GO:0007056) positive regulation of oocyte development(GO:0060282) |
| 0.5 | 4.4 | GO:0006104 | succinyl-CoA metabolic process(GO:0006104) |
| 0.5 | 1.0 | GO:0006933 | negative regulation of cell adhesion involved in substrate-bound cell migration(GO:0006933) |
| 0.5 | 9.3 | GO:0060236 | regulation of mitotic spindle organization(GO:0060236) |
| 0.5 | 3.8 | GO:0014807 | regulation of somitogenesis(GO:0014807) |
| 0.5 | 1.9 | GO:0006203 | dGTP catabolic process(GO:0006203) dATP catabolic process(GO:0046061) |
| 0.5 | 1.4 | GO:0002408 | myeloid dendritic cell chemotaxis(GO:0002408) |
| 0.4 | 4.7 | GO:0002176 | male germ cell proliferation(GO:0002176) |
| 0.4 | 3.8 | GO:0014883 | transition between fast and slow fiber(GO:0014883) |
| 0.4 | 0.8 | GO:0061738 | late endosomal microautophagy(GO:0061738) |
| 0.4 | 7.8 | GO:0030213 | hyaluronan biosynthetic process(GO:0030213) |
| 0.4 | 1.6 | GO:0006529 | asparagine biosynthetic process(GO:0006529) |
| 0.4 | 1.2 | GO:0033566 | gamma-tubulin complex localization(GO:0033566) |
| 0.4 | 2.0 | GO:0002326 | B cell lineage commitment(GO:0002326) |
| 0.4 | 6.5 | GO:0000183 | chromatin silencing at rDNA(GO:0000183) |
| 0.4 | 3.6 | GO:0051574 | positive regulation of histone H3-K9 methylation(GO:0051574) |
| 0.4 | 2.0 | GO:0010668 | ectodermal cell differentiation(GO:0010668) |
| 0.4 | 1.2 | GO:0071603 | endothelial cell-cell adhesion(GO:0071603) |
| 0.4 | 10.6 | GO:0000413 | protein peptidyl-prolyl isomerization(GO:0000413) |
| 0.4 | 1.5 | GO:0007113 | endomitotic cell cycle(GO:0007113) |
| 0.4 | 5.3 | GO:0045793 | positive regulation of cell size(GO:0045793) |
| 0.4 | 1.1 | GO:0006434 | seryl-tRNA aminoacylation(GO:0006434) |
| 0.4 | 1.9 | GO:0060838 | lymphatic endothelial cell fate commitment(GO:0060838) regulation of transcription involved in lymphatic endothelial cell fate commitment(GO:0060849) |
| 0.4 | 1.1 | GO:1903225 | negative regulation of endodermal cell differentiation(GO:1903225) |
| 0.4 | 4.3 | GO:0015825 | L-serine transport(GO:0015825) |
| 0.4 | 4.6 | GO:0051451 | myoblast migration(GO:0051451) |
| 0.3 | 2.4 | GO:0007296 | vitellogenesis(GO:0007296) |
| 0.3 | 4.5 | GO:0045603 | positive regulation of endothelial cell differentiation(GO:0045603) |
| 0.3 | 3.5 | GO:1902913 | positive regulation of melanocyte differentiation(GO:0045636) positive regulation of neuroepithelial cell differentiation(GO:1902913) |
| 0.3 | 1.0 | GO:2000564 | regulation of MyD88-dependent toll-like receptor signaling pathway(GO:0034124) CD8-positive, alpha-beta T cell proliferation(GO:0035740) negative regulation of regulatory T cell differentiation(GO:0045590) regulation of CD8-positive, alpha-beta T cell proliferation(GO:2000564) |
| 0.3 | 3.9 | GO:0070986 | left/right axis specification(GO:0070986) |
| 0.3 | 2.6 | GO:0007195 | adenylate cyclase-inhibiting dopamine receptor signaling pathway(GO:0007195) |
| 0.3 | 6.4 | GO:0035873 | lactate transport(GO:0015727) lactate transmembrane transport(GO:0035873) plasma membrane lactate transport(GO:0035879) |
| 0.3 | 1.3 | GO:0002024 | diet induced thermogenesis(GO:0002024) adaptive thermogenesis(GO:1990845) |
| 0.3 | 3.4 | GO:0030238 | male sex determination(GO:0030238) |
| 0.3 | 2.7 | GO:0009992 | cellular water homeostasis(GO:0009992) |
| 0.3 | 3.9 | GO:0046643 | regulation of gamma-delta T cell differentiation(GO:0045586) regulation of gamma-delta T cell activation(GO:0046643) |
| 0.3 | 2.7 | GO:0060907 | positive regulation of macrophage cytokine production(GO:0060907) |
| 0.3 | 7.5 | GO:0046596 | regulation of viral entry into host cell(GO:0046596) |
| 0.3 | 5.8 | GO:0051044 | positive regulation of membrane protein ectodomain proteolysis(GO:0051044) |
| 0.3 | 0.8 | GO:1904154 | protein localization to endoplasmic reticulum exit site(GO:0070973) positive regulation of retrograde protein transport, ER to cytosol(GO:1904154) |
| 0.3 | 2.2 | GO:0031053 | primary miRNA processing(GO:0031053) |
| 0.3 | 1.9 | GO:1900262 | regulation of DNA-directed DNA polymerase activity(GO:1900262) positive regulation of DNA-directed DNA polymerase activity(GO:1900264) |
| 0.3 | 0.8 | GO:0036091 | positive regulation of transcription from RNA polymerase II promoter in response to oxidative stress(GO:0036091) |
| 0.3 | 1.3 | GO:0060018 | astrocyte fate commitment(GO:0060018) |
| 0.3 | 0.8 | GO:0035021 | negative regulation of Rac protein signal transduction(GO:0035021) |
| 0.3 | 3.5 | GO:0021891 | olfactory bulb interneuron development(GO:0021891) |
| 0.3 | 1.3 | GO:0003420 | regulation of growth plate cartilage chondrocyte proliferation(GO:0003420) |
| 0.3 | 2.1 | GO:0006004 | fucose metabolic process(GO:0006004) |
| 0.3 | 4.4 | GO:0005980 | polysaccharide catabolic process(GO:0000272) glycogen catabolic process(GO:0005980) glucan catabolic process(GO:0009251) cellular polysaccharide catabolic process(GO:0044247) |
| 0.3 | 1.0 | GO:0051866 | general adaptation syndrome(GO:0051866) |
| 0.2 | 1.2 | GO:0045053 | protein retention in Golgi apparatus(GO:0045053) |
| 0.2 | 0.7 | GO:0010757 | negative regulation of plasminogen activation(GO:0010757) |
| 0.2 | 0.7 | GO:0032790 | ribosome disassembly(GO:0032790) |
| 0.2 | 0.9 | GO:0070213 | protein auto-ADP-ribosylation(GO:0070213) |
| 0.2 | 1.6 | GO:0014816 | skeletal muscle satellite cell differentiation(GO:0014816) |
| 0.2 | 2.6 | GO:0046349 | amino sugar biosynthetic process(GO:0046349) |
| 0.2 | 1.6 | GO:0097501 | stress response to metal ion(GO:0097501) |
| 0.2 | 2.1 | GO:0098535 | de novo centriole assembly(GO:0098535) |
| 0.2 | 0.2 | GO:0001923 | B-1 B cell differentiation(GO:0001923) |
| 0.2 | 1.1 | GO:0018171 | peptidyl-cysteine oxidation(GO:0018171) |
| 0.2 | 1.6 | GO:0042769 | DNA damage response, detection of DNA damage(GO:0042769) |
| 0.2 | 2.2 | GO:0036120 | cellular response to platelet-derived growth factor stimulus(GO:0036120) |
| 0.2 | 0.7 | GO:0006597 | spermine biosynthetic process(GO:0006597) |
| 0.2 | 0.9 | GO:1901165 | positive regulation of trophoblast cell migration(GO:1901165) |
| 0.2 | 0.4 | GO:0034238 | macrophage fusion(GO:0034238) regulation of macrophage fusion(GO:0034239) |
| 0.2 | 0.7 | GO:0000055 | ribosomal large subunit export from nucleus(GO:0000055) |
| 0.2 | 2.4 | GO:1990173 | protein localization to nuclear body(GO:1903405) protein localization to Cajal body(GO:1904867) regulation of protein localization to Cajal body(GO:1904869) positive regulation of protein localization to Cajal body(GO:1904871) protein localization to nucleoplasm(GO:1990173) |
| 0.2 | 4.1 | GO:0042753 | positive regulation of circadian rhythm(GO:0042753) |
| 0.2 | 0.8 | GO:0070125 | mitochondrial translational elongation(GO:0070125) |
| 0.2 | 1.0 | GO:2000298 | regulation of Rho-dependent protein serine/threonine kinase activity(GO:2000298) |
| 0.2 | 1.2 | GO:0009235 | cobalamin metabolic process(GO:0009235) |
| 0.2 | 0.6 | GO:0097089 | methyl-branched fatty acid metabolic process(GO:0097089) |
| 0.2 | 0.8 | GO:0033580 | protein glycosylation at cell surface(GO:0033575) protein galactosylation at cell surface(GO:0033580) protein galactosylation(GO:0042125) |
| 0.2 | 2.2 | GO:0006610 | ribosomal protein import into nucleus(GO:0006610) |
| 0.2 | 2.7 | GO:0010972 | negative regulation of G2/M transition of mitotic cell cycle(GO:0010972) |
| 0.2 | 2.9 | GO:0097062 | dendritic spine maintenance(GO:0097062) |
| 0.2 | 10.5 | GO:0045600 | positive regulation of fat cell differentiation(GO:0045600) |
| 0.2 | 1.5 | GO:0010587 | miRNA catabolic process(GO:0010587) |
| 0.2 | 3.6 | GO:0045662 | negative regulation of myoblast differentiation(GO:0045662) |
| 0.2 | 1.1 | GO:0003433 | chondrocyte development involved in endochondral bone morphogenesis(GO:0003433) |
| 0.2 | 0.7 | GO:0044861 | protein transport into plasma membrane raft(GO:0044861) |
| 0.2 | 0.5 | GO:0006421 | asparaginyl-tRNA aminoacylation(GO:0006421) |
| 0.2 | 1.8 | GO:0001522 | pseudouridine synthesis(GO:0001522) |
| 0.2 | 0.9 | GO:0044314 | protein K27-linked ubiquitination(GO:0044314) |
| 0.2 | 1.8 | GO:0033234 | negative regulation of protein sumoylation(GO:0033234) |
| 0.2 | 0.7 | GO:1902268 | negative regulation of polyamine transmembrane transport(GO:1902268) |
| 0.2 | 1.3 | GO:0033353 | S-adenosylmethionine cycle(GO:0033353) |
| 0.2 | 1.4 | GO:0045899 | positive regulation of RNA polymerase II transcriptional preinitiation complex assembly(GO:0045899) |
| 0.2 | 2.0 | GO:0032287 | peripheral nervous system myelin maintenance(GO:0032287) |
| 0.2 | 1.1 | GO:0031119 | tRNA pseudouridine synthesis(GO:0031119) |
| 0.2 | 2.0 | GO:0018126 | protein hydroxylation(GO:0018126) |
| 0.2 | 0.5 | GO:0015920 | lipopolysaccharide transport(GO:0015920) |
| 0.2 | 0.8 | GO:0032472 | Golgi calcium ion transport(GO:0032472) |
| 0.1 | 4.0 | GO:0000154 | rRNA modification(GO:0000154) |
| 0.1 | 0.7 | GO:0007089 | traversing start control point of mitotic cell cycle(GO:0007089) |
| 0.1 | 1.0 | GO:0034393 | positive regulation of smooth muscle cell apoptotic process(GO:0034393) |
| 0.1 | 1.1 | GO:0048311 | mitochondrion distribution(GO:0048311) |
| 0.1 | 0.7 | GO:0031282 | regulation of guanylate cyclase activity(GO:0031282) |
| 0.1 | 1.6 | GO:0043517 | positive regulation of DNA damage response, signal transduction by p53 class mediator(GO:0043517) |
| 0.1 | 1.2 | GO:0006611 | protein export from nucleus(GO:0006611) |
| 0.1 | 0.8 | GO:0000056 | ribosomal small subunit export from nucleus(GO:0000056) |
| 0.1 | 1.5 | GO:0000244 | spliceosomal tri-snRNP complex assembly(GO:0000244) |
| 0.1 | 1.6 | GO:0038166 | angiotensin-activated signaling pathway(GO:0038166) |
| 0.1 | 0.2 | GO:0019471 | 4-hydroxyproline metabolic process(GO:0019471) |
| 0.1 | 0.8 | GO:0042572 | retinol metabolic process(GO:0042572) |
| 0.1 | 0.2 | GO:0032829 | regulation of CD4-positive, CD25-positive, alpha-beta regulatory T cell differentiation(GO:0032829) |
| 0.1 | 1.4 | GO:1990403 | embryonic brain development(GO:1990403) |
| 0.1 | 0.3 | GO:2000819 | regulation of nucleotide-excision repair(GO:2000819) |
| 0.1 | 0.5 | GO:0060334 | regulation of interferon-gamma-mediated signaling pathway(GO:0060334) |
| 0.1 | 4.5 | GO:0031572 | G2 DNA damage checkpoint(GO:0031572) |
| 0.1 | 0.4 | GO:0090370 | negative regulation of cholesterol efflux(GO:0090370) |
| 0.1 | 1.8 | GO:0050860 | negative regulation of T cell receptor signaling pathway(GO:0050860) |
| 0.1 | 1.1 | GO:0046036 | GTP biosynthetic process(GO:0006183) CTP biosynthetic process(GO:0006241) CTP metabolic process(GO:0046036) |
| 0.1 | 1.1 | GO:0030813 | positive regulation of nucleotide catabolic process(GO:0030813) positive regulation of glycolytic process(GO:0045821) positive regulation of cofactor metabolic process(GO:0051194) positive regulation of coenzyme metabolic process(GO:0051197) |
| 0.1 | 0.2 | GO:2000418 | positive regulation of eosinophil migration(GO:2000418) |
| 0.1 | 0.4 | GO:0035616 | histone H2B conserved C-terminal lysine deubiquitination(GO:0035616) |
| 0.1 | 1.9 | GO:0045947 | negative regulation of translational initiation(GO:0045947) |
| 0.1 | 0.4 | GO:0006627 | protein processing involved in protein targeting to mitochondrion(GO:0006627) |
| 0.1 | 0.4 | GO:0035752 | lysosomal lumen pH elevation(GO:0035752) |
| 0.1 | 0.5 | GO:1904338 | regulation of dopaminergic neuron differentiation(GO:1904338) |
| 0.1 | 0.3 | GO:0046077 | dUDP biosynthetic process(GO:0006227) pyrimidine nucleoside diphosphate biosynthetic process(GO:0009139) pyrimidine ribonucleoside diphosphate metabolic process(GO:0009193) pyrimidine deoxyribonucleoside diphosphate metabolic process(GO:0009196) pyrimidine deoxyribonucleoside diphosphate biosynthetic process(GO:0009197) UDP metabolic process(GO:0046048) dUDP metabolic process(GO:0046077) |
| 0.1 | 0.8 | GO:0042989 | sequestering of actin monomers(GO:0042989) |
| 0.1 | 0.6 | GO:0031584 | activation of phospholipase D activity(GO:0031584) |
| 0.1 | 0.3 | GO:0090285 | negative regulation of protein glycosylation in Golgi(GO:0090285) |
| 0.1 | 2.1 | GO:0006359 | regulation of transcription from RNA polymerase III promoter(GO:0006359) |
| 0.1 | 1.0 | GO:0045906 | negative regulation of vasoconstriction(GO:0045906) |
| 0.1 | 2.8 | GO:0051642 | centrosome localization(GO:0051642) |
| 0.1 | 2.3 | GO:0010842 | retina layer formation(GO:0010842) |
| 0.1 | 1.9 | GO:0070536 | protein K63-linked deubiquitination(GO:0070536) |
| 0.1 | 0.4 | GO:0051490 | negative regulation of filopodium assembly(GO:0051490) |
| 0.1 | 7.3 | GO:0050772 | positive regulation of axonogenesis(GO:0050772) |
| 0.1 | 0.3 | GO:1903984 | regulation of TRAIL-activated apoptotic signaling pathway(GO:1903121) positive regulation of TRAIL-activated apoptotic signaling pathway(GO:1903984) negative regulation of metalloendopeptidase activity(GO:1904684) negative regulation of metallopeptidase activity(GO:1905049) |
| 0.1 | 1.5 | GO:0016226 | iron-sulfur cluster assembly(GO:0016226) metallo-sulfur cluster assembly(GO:0031163) |
| 0.1 | 0.6 | GO:0035405 | histone-threonine phosphorylation(GO:0035405) |
| 0.1 | 4.0 | GO:0002066 | columnar/cuboidal epithelial cell development(GO:0002066) |
| 0.1 | 1.7 | GO:0048246 | macrophage chemotaxis(GO:0048246) |
| 0.1 | 4.1 | GO:0061077 | chaperone-mediated protein folding(GO:0061077) |
| 0.1 | 0.9 | GO:0000479 | endonucleolytic cleavage of tricistronic rRNA transcript (SSU-rRNA, 5.8S rRNA, LSU-rRNA)(GO:0000479) |
| 0.1 | 0.8 | GO:0070544 | histone H3-K36 demethylation(GO:0070544) |
| 0.1 | 0.5 | GO:0061299 | retina vasculature morphogenesis in camera-type eye(GO:0061299) |
| 0.1 | 2.7 | GO:0006801 | superoxide metabolic process(GO:0006801) |
| 0.1 | 0.3 | GO:0060479 | lung epithelium development(GO:0060428) lung cell differentiation(GO:0060479) lung epithelial cell differentiation(GO:0060487) |
| 0.1 | 1.6 | GO:0016180 | snRNA processing(GO:0016180) |
| 0.1 | 0.3 | GO:0048304 | positive regulation of isotype switching to IgG isotypes(GO:0048304) |
| 0.1 | 1.0 | GO:0046855 | phosphorylated carbohydrate dephosphorylation(GO:0046838) inositol phosphate dephosphorylation(GO:0046855) |
| 0.1 | 0.4 | GO:0042670 | retinal cone cell differentiation(GO:0042670) retinal cone cell development(GO:0046549) |
| 0.1 | 2.8 | GO:0060612 | adipose tissue development(GO:0060612) |
| 0.1 | 0.3 | GO:0015015 | heparan sulfate proteoglycan biosynthetic process, enzymatic modification(GO:0015015) |
| 0.1 | 1.4 | GO:0060259 | regulation of feeding behavior(GO:0060259) |
| 0.1 | 0.9 | GO:0006607 | NLS-bearing protein import into nucleus(GO:0006607) |
| 0.1 | 1.1 | GO:0045292 | mRNA cis splicing, via spliceosome(GO:0045292) |
| 0.1 | 1.0 | GO:0031018 | endocrine pancreas development(GO:0031018) |
| 0.1 | 0.3 | GO:0006564 | L-serine biosynthetic process(GO:0006564) |
| 0.1 | 0.6 | GO:0090557 | establishment of endothelial intestinal barrier(GO:0090557) |
| 0.1 | 0.3 | GO:0070535 | histone H2A K63-linked ubiquitination(GO:0070535) |
| 0.1 | 0.4 | GO:0015879 | carnitine transport(GO:0015879) |
| 0.1 | 0.3 | GO:0019065 | receptor-mediated endocytosis of virus by host cell(GO:0019065) endocytosis involved in viral entry into host cell(GO:0075509) |
| 0.1 | 1.1 | GO:0000028 | ribosomal small subunit assembly(GO:0000028) |
| 0.1 | 1.0 | GO:0045116 | protein neddylation(GO:0045116) |
| 0.1 | 1.4 | GO:0045103 | intermediate filament-based process(GO:0045103) |
| 0.1 | 0.5 | GO:0090244 | Wnt signaling pathway involved in somitogenesis(GO:0090244) |
| 0.1 | 1.4 | GO:0006509 | membrane protein ectodomain proteolysis(GO:0006509) |
| 0.1 | 2.6 | GO:0050830 | defense response to Gram-positive bacterium(GO:0050830) |
| 0.1 | 0.3 | GO:0019249 | lactate biosynthetic process from pyruvate(GO:0019244) lactate biosynthetic process(GO:0019249) |
| 0.1 | 0.7 | GO:0050872 | white fat cell differentiation(GO:0050872) |
| 0.1 | 0.6 | GO:0070914 | UV-damage excision repair(GO:0070914) |
| 0.1 | 0.3 | GO:0030826 | regulation of cGMP biosynthetic process(GO:0030826) |
| 0.0 | 0.6 | GO:0042744 | hydrogen peroxide catabolic process(GO:0042744) |
| 0.0 | 0.2 | GO:1903847 | regulation of aorta morphogenesis(GO:1903847) positive regulation of aorta morphogenesis(GO:1903849) |
| 0.0 | 1.0 | GO:0000462 | maturation of SSU-rRNA from tricistronic rRNA transcript (SSU-rRNA, 5.8S rRNA, LSU-rRNA)(GO:0000462) |
| 0.0 | 2.1 | GO:0010501 | RNA secondary structure unwinding(GO:0010501) |
| 0.0 | 0.9 | GO:0042474 | middle ear morphogenesis(GO:0042474) |
| 0.0 | 0.4 | GO:1900364 | negative regulation of mRNA polyadenylation(GO:1900364) |
| 0.0 | 0.5 | GO:0034498 | early endosome to Golgi transport(GO:0034498) |
| 0.0 | 0.3 | GO:0061179 | negative regulation of insulin secretion involved in cellular response to glucose stimulus(GO:0061179) |
| 0.0 | 0.4 | GO:0031339 | negative regulation of vesicle fusion(GO:0031339) |
| 0.0 | 1.8 | GO:0048636 | positive regulation of striated muscle tissue development(GO:0045844) positive regulation of muscle organ development(GO:0048636) positive regulation of muscle tissue development(GO:1901863) |
| 0.0 | 0.1 | GO:0001830 | trophectodermal cell fate commitment(GO:0001830) |
| 0.0 | 2.3 | GO:0006513 | protein monoubiquitination(GO:0006513) |
| 0.0 | 0.3 | GO:0060766 | negative regulation of androgen receptor signaling pathway(GO:0060766) |
| 0.0 | 1.6 | GO:2001238 | positive regulation of extrinsic apoptotic signaling pathway(GO:2001238) |
| 0.0 | 0.1 | GO:0019043 | establishment of viral latency(GO:0019043) |
| 0.0 | 0.6 | GO:0043968 | histone H2A acetylation(GO:0043968) |
| 0.0 | 0.1 | GO:0072488 | ammonium transmembrane transport(GO:0072488) |
| 0.0 | 1.0 | GO:0015909 | long-chain fatty acid transport(GO:0015909) |
| 0.0 | 0.3 | GO:0021683 | cerebellar granular layer morphogenesis(GO:0021683) |
| 0.0 | 0.8 | GO:0043029 | T cell homeostasis(GO:0043029) |
| 0.0 | 0.1 | GO:0032048 | cardiolipin metabolic process(GO:0032048) |
| 0.0 | 0.5 | GO:0003301 | physiological muscle hypertrophy(GO:0003298) physiological cardiac muscle hypertrophy(GO:0003301) cell growth involved in cardiac muscle cell development(GO:0061049) |
| 0.0 | 0.9 | GO:0046470 | phosphatidylcholine metabolic process(GO:0046470) |
| 0.0 | 2.9 | GO:0051028 | mRNA transport(GO:0051028) |
| 0.0 | 2.3 | GO:0030148 | sphingolipid biosynthetic process(GO:0030148) |
| 0.0 | 0.5 | GO:1902230 | negative regulation of intrinsic apoptotic signaling pathway in response to DNA damage(GO:1902230) |
| 0.0 | 3.1 | GO:0060348 | bone development(GO:0060348) |
| 0.0 | 1.4 | GO:0006635 | fatty acid beta-oxidation(GO:0006635) |
| 0.0 | 0.9 | GO:0009225 | nucleotide-sugar metabolic process(GO:0009225) |
| 0.0 | 3.5 | GO:0007519 | skeletal muscle tissue development(GO:0007519) |
| 0.0 | 1.6 | GO:0010923 | negative regulation of phosphatase activity(GO:0010923) |
| 0.0 | 0.8 | GO:0048286 | lung alveolus development(GO:0048286) |
| 0.0 | 0.2 | GO:0006532 | aspartate biosynthetic process(GO:0006532) |
| 0.0 | 0.5 | GO:0030488 | tRNA methylation(GO:0030488) |
| 0.0 | 0.6 | GO:0007157 | heterophilic cell-cell adhesion via plasma membrane cell adhesion molecules(GO:0007157) |
| 0.0 | 0.7 | GO:0017145 | stem cell division(GO:0017145) |
| 0.0 | 1.8 | GO:0007517 | muscle organ development(GO:0007517) |
| 0.0 | 0.4 | GO:0006220 | pyrimidine nucleotide metabolic process(GO:0006220) |
| 0.0 | 1.0 | GO:0007127 | meiosis I(GO:0007127) |
| 0.0 | 0.9 | GO:0045638 | negative regulation of myeloid cell differentiation(GO:0045638) |
| 0.0 | 0.1 | GO:0051482 | positive regulation of cytosolic calcium ion concentration involved in phospholipase C-activating G-protein coupled signaling pathway(GO:0051482) |
| 0.0 | 0.7 | GO:0006418 | tRNA aminoacylation for protein translation(GO:0006418) |
| 0.0 | 0.4 | GO:0051057 | positive regulation of small GTPase mediated signal transduction(GO:0051057) |
| 0.0 | 0.2 | GO:2000279 | negative regulation of DNA biosynthetic process(GO:2000279) |
| 0.0 | 0.4 | GO:0006284 | base-excision repair(GO:0006284) |
| 0.0 | 0.7 | GO:0050728 | negative regulation of inflammatory response(GO:0050728) |
| 0.0 | 1.0 | GO:0050796 | regulation of insulin secretion(GO:0050796) |
| 0.0 | 0.2 | GO:0071786 | endoplasmic reticulum tubular network organization(GO:0071786) |
| 0.0 | 0.4 | GO:1902653 | cholesterol biosynthetic process(GO:0006695) secondary alcohol biosynthetic process(GO:1902653) |
| 0.0 | 0.0 | GO:1903336 | negative regulation of vacuolar transport(GO:1903336) |
| 0.0 | 0.1 | GO:0060148 | positive regulation of posttranscriptional gene silencing(GO:0060148) |
Gene overrepresentation in cellular_component category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 2.1 | 10.7 | GO:0097226 | sperm mitochondrial sheath(GO:0097226) |
| 1.1 | 4.4 | GO:0036454 | insulin-like growth factor binding protein complex(GO:0016942) growth factor complex(GO:0036454) insulin-like growth factor ternary complex(GO:0042567) |
| 1.0 | 2.0 | GO:0042585 | germinal vesicle(GO:0042585) |
| 0.9 | 5.6 | GO:0046696 | lipopolysaccharide receptor complex(GO:0046696) |
| 0.8 | 2.5 | GO:0097059 | CNTFR-CLCF1 complex(GO:0097059) |
| 0.8 | 6.1 | GO:0005818 | aster(GO:0005818) |
| 0.7 | 4.5 | GO:0031618 | nuclear pericentric heterochromatin(GO:0031618) |
| 0.7 | 4.4 | GO:0030062 | mitochondrial tricarboxylic acid cycle enzyme complex(GO:0030062) |
| 0.7 | 3.7 | GO:0030314 | junctional membrane complex(GO:0030314) |
| 0.7 | 3.5 | GO:0008622 | epsilon DNA polymerase complex(GO:0008622) |
| 0.5 | 4.3 | GO:0044300 | cerebellar mossy fiber(GO:0044300) |
| 0.5 | 2.7 | GO:0033553 | rDNA heterochromatin(GO:0033553) |
| 0.5 | 7.9 | GO:0045180 | basal cortex(GO:0045180) |
| 0.5 | 2.6 | GO:0031523 | Myb complex(GO:0031523) |
| 0.5 | 6.4 | GO:0030061 | mitochondrial crista(GO:0030061) |
| 0.5 | 2.4 | GO:0002193 | MAML1-RBP-Jkappa- ICN1 complex(GO:0002193) |
| 0.5 | 4.5 | GO:0070531 | BRCA1-A complex(GO:0070531) |
| 0.4 | 1.8 | GO:0002111 | BRCA2-BRAF35 complex(GO:0002111) |
| 0.4 | 1.3 | GO:0030905 | retromer, tubulation complex(GO:0030905) |
| 0.4 | 2.0 | GO:0032389 | MutLalpha complex(GO:0032389) |
| 0.4 | 3.2 | GO:0031428 | box C/D snoRNP complex(GO:0031428) |
| 0.4 | 1.1 | GO:0005588 | collagen type V trimer(GO:0005588) |
| 0.4 | 2.1 | GO:0005677 | chromatin silencing complex(GO:0005677) |
| 0.3 | 3.7 | GO:0031080 | nuclear pore outer ring(GO:0031080) |
| 0.3 | 0.6 | GO:0036056 | filtration diaphragm(GO:0036056) slit diaphragm(GO:0036057) |
| 0.3 | 4.1 | GO:0097539 | ciliary transition fiber(GO:0097539) |
| 0.3 | 2.1 | GO:0098536 | deuterosome(GO:0098536) |
| 0.2 | 1.9 | GO:0031390 | Ctf18 RFC-like complex(GO:0031390) |
| 0.2 | 2.8 | GO:0071598 | neuronal ribonucleoprotein granule(GO:0071598) |
| 0.2 | 1.5 | GO:0030891 | VCB complex(GO:0030891) |
| 0.2 | 2.4 | GO:0005832 | chaperonin-containing T-complex(GO:0005832) |
| 0.2 | 0.8 | GO:0043202 | lysosomal lumen(GO:0043202) |
| 0.2 | 0.6 | GO:0071008 | U2-type post-mRNA release spliceosomal complex(GO:0071008) |
| 0.2 | 3.3 | GO:0005665 | DNA-directed RNA polymerase II, core complex(GO:0005665) |
| 0.2 | 2.3 | GO:0043240 | Fanconi anaemia nuclear complex(GO:0043240) |
| 0.2 | 0.8 | GO:0005784 | Sec61 translocon complex(GO:0005784) translocon complex(GO:0071256) |
| 0.2 | 1.1 | GO:0097422 | tubular endosome(GO:0097422) |
| 0.2 | 7.4 | GO:0000307 | cyclin-dependent protein kinase holoenzyme complex(GO:0000307) |
| 0.2 | 10.2 | GO:0016328 | lateral plasma membrane(GO:0016328) |
| 0.2 | 9.4 | GO:0016459 | myosin complex(GO:0016459) |
| 0.2 | 0.3 | GO:0016222 | procollagen-proline 4-dioxygenase complex(GO:0016222) |
| 0.2 | 4.6 | GO:0051233 | spindle midzone(GO:0051233) |
| 0.2 | 3.6 | GO:0035098 | ESC/E(Z) complex(GO:0035098) |
| 0.1 | 1.4 | GO:0031595 | nuclear proteasome complex(GO:0031595) |
| 0.1 | 0.9 | GO:0030688 | preribosome, small subunit precursor(GO:0030688) |
| 0.1 | 0.4 | GO:0042720 | mitochondrial inner membrane peptidase complex(GO:0042720) |
| 0.1 | 1.8 | GO:0042101 | T cell receptor complex(GO:0042101) |
| 0.1 | 1.0 | GO:0045261 | mitochondrial proton-transporting ATP synthase complex, catalytic core F(1)(GO:0000275) proton-transporting ATP synthase complex, catalytic core F(1)(GO:0045261) |
| 0.1 | 1.6 | GO:0030126 | COPI vesicle coat(GO:0030126) |
| 0.1 | 1.6 | GO:0032039 | integrator complex(GO:0032039) |
| 0.1 | 1.4 | GO:0032593 | insulin-responsive compartment(GO:0032593) |
| 0.1 | 1.3 | GO:0008385 | IkappaB kinase complex(GO:0008385) |
| 0.1 | 0.8 | GO:0060091 | kinocilium(GO:0060091) |
| 0.1 | 1.7 | GO:0005640 | nuclear outer membrane(GO:0005640) |
| 0.1 | 1.2 | GO:0034663 | endoplasmic reticulum chaperone complex(GO:0034663) |
| 0.1 | 1.0 | GO:0005671 | Ada2/Gcn5/Ada3 transcription activator complex(GO:0005671) |
| 0.1 | 0.7 | GO:0031091 | platelet alpha granule(GO:0031091) |
| 0.1 | 0.9 | GO:0034518 | mRNA cap binding complex(GO:0005845) RNA cap binding complex(GO:0034518) |
| 0.1 | 1.3 | GO:0005666 | DNA-directed RNA polymerase III complex(GO:0005666) |
| 0.1 | 2.7 | GO:0000788 | nuclear nucleosome(GO:0000788) |
| 0.1 | 2.3 | GO:0005689 | U12-type spliceosomal complex(GO:0005689) |
| 0.1 | 4.5 | GO:0031985 | Golgi cisterna(GO:0031985) |
| 0.1 | 0.7 | GO:0044352 | pinosome(GO:0044352) macropinosome(GO:0044354) |
| 0.1 | 4.1 | GO:0005882 | intermediate filament(GO:0005882) |
| 0.1 | 0.5 | GO:0042589 | zymogen granule membrane(GO:0042589) |
| 0.1 | 1.5 | GO:0046540 | U4/U6 x U5 tri-snRNP complex(GO:0046540) |
| 0.1 | 1.6 | GO:0032839 | dendrite cytoplasm(GO:0032839) |
| 0.1 | 1.6 | GO:0005921 | gap junction(GO:0005921) |
| 0.1 | 6.2 | GO:0030136 | clathrin-coated vesicle(GO:0030136) |
| 0.1 | 8.1 | GO:0031234 | extrinsic component of cytoplasmic side of plasma membrane(GO:0031234) |
| 0.1 | 2.9 | GO:0022627 | cytosolic small ribosomal subunit(GO:0022627) |
| 0.1 | 0.5 | GO:0042382 | paraspeckles(GO:0042382) |
| 0.1 | 4.1 | GO:0030315 | T-tubule(GO:0030315) |
| 0.1 | 0.4 | GO:0044232 | organelle membrane contact site(GO:0044232) |
| 0.1 | 6.8 | GO:0000784 | nuclear chromosome, telomeric region(GO:0000784) |
| 0.1 | 10.8 | GO:0031965 | nuclear membrane(GO:0031965) |
| 0.1 | 2.9 | GO:0005643 | nuclear pore(GO:0005643) |
| 0.1 | 1.2 | GO:0000930 | gamma-tubulin complex(GO:0000930) |
| 0.1 | 1.2 | GO:0032588 | trans-Golgi network membrane(GO:0032588) |
| 0.1 | 0.1 | GO:0031074 | nucleocytoplasmic shuttling complex(GO:0031074) |
| 0.0 | 0.4 | GO:0071012 | catalytic step 1 spliceosome(GO:0071012) |
| 0.0 | 1.1 | GO:0071339 | MLL1/2 complex(GO:0044665) MLL1 complex(GO:0071339) |
| 0.0 | 2.0 | GO:0042645 | nucleoid(GO:0009295) mitochondrial nucleoid(GO:0042645) |
| 0.0 | 1.4 | GO:0005719 | nuclear euchromatin(GO:0005719) |
| 0.0 | 4.4 | GO:0090575 | RNA polymerase II transcription factor complex(GO:0090575) |
| 0.0 | 0.4 | GO:0042622 | photoreceptor outer segment membrane(GO:0042622) |
| 0.0 | 2.0 | GO:0005793 | endoplasmic reticulum-Golgi intermediate compartment(GO:0005793) |
| 0.0 | 0.3 | GO:0031464 | Cul4A-RING E3 ubiquitin ligase complex(GO:0031464) |
| 0.0 | 0.8 | GO:0030175 | filopodium(GO:0030175) |
| 0.0 | 1.2 | GO:0030667 | secretory granule membrane(GO:0030667) |
| 0.0 | 1.3 | GO:0031463 | Cul3-RING ubiquitin ligase complex(GO:0031463) |
| 0.0 | 1.2 | GO:0016529 | sarcoplasmic reticulum(GO:0016529) |
| 0.0 | 5.4 | GO:0005802 | trans-Golgi network(GO:0005802) |
| 0.0 | 2.0 | GO:0031526 | brush border membrane(GO:0031526) |
| 0.0 | 0.4 | GO:0031616 | spindle pole centrosome(GO:0031616) |
| 0.0 | 2.2 | GO:0005681 | spliceosomal complex(GO:0005681) |
| 0.0 | 1.3 | GO:0034707 | chloride channel complex(GO:0034707) |
| 0.0 | 0.6 | GO:0030134 | ER to Golgi transport vesicle(GO:0030134) |
| 0.0 | 0.6 | GO:0043189 | NuA4 histone acetyltransferase complex(GO:0035267) H4/H2A histone acetyltransferase complex(GO:0043189) H4 histone acetyltransferase complex(GO:1902562) |
| 0.0 | 0.2 | GO:0098827 | endoplasmic reticulum tubular network(GO:0071782) endoplasmic reticulum subcompartment(GO:0098827) |
| 0.0 | 0.4 | GO:0071203 | WASH complex(GO:0071203) |
| 0.0 | 1.1 | GO:0045171 | intercellular bridge(GO:0045171) |
| 0.0 | 0.3 | GO:0008074 | guanylate cyclase complex, soluble(GO:0008074) |
| 0.0 | 0.5 | GO:0001741 | XY body(GO:0001741) |
| 0.0 | 0.2 | GO:0031011 | Ino80 complex(GO:0031011) |
| 0.0 | 7.5 | GO:0005667 | transcription factor complex(GO:0005667) |
| 0.0 | 0.3 | GO:0035631 | CD40 receptor complex(GO:0035631) |
| 0.0 | 0.4 | GO:0031588 | nucleotide-activated protein kinase complex(GO:0031588) |
| 0.0 | 2.6 | GO:0031225 | anchored component of membrane(GO:0031225) |
| 0.0 | 1.5 | GO:0071013 | catalytic step 2 spliceosome(GO:0071013) |
| 0.0 | 1.7 | GO:0016607 | nuclear speck(GO:0016607) |
| 0.0 | 1.5 | GO:0005770 | late endosome(GO:0005770) |
| 0.0 | 10.7 | GO:0005730 | nucleolus(GO:0005730) |
| 0.0 | 0.3 | GO:0035861 | site of double-strand break(GO:0035861) |
| 0.0 | 0.4 | GO:0005876 | spindle microtubule(GO:0005876) |
| 0.0 | 0.6 | GO:0044295 | axonal growth cone(GO:0044295) |
| 0.0 | 1.1 | GO:0031968 | mitochondrial outer membrane(GO:0005741) organelle outer membrane(GO:0031968) |
Gene overrepresentation in molecular_function category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 1.8 | 10.6 | GO:0002135 | CTP binding(GO:0002135) |
| 1.2 | 6.1 | GO:0061676 | importin-alpha family protein binding(GO:0061676) |
| 1.2 | 4.9 | GO:0016018 | cyclosporin A binding(GO:0016018) |
| 1.1 | 2.3 | GO:0042813 | Wnt-activated receptor activity(GO:0042813) |
| 1.1 | 4.4 | GO:2001069 | glycogen binding(GO:2001069) |
| 1.0 | 4.0 | GO:0004468 | lysine N-acetyltransferase activity, acting on acetyl phosphate as donor(GO:0004468) |
| 1.0 | 21.0 | GO:0070410 | co-SMAD binding(GO:0070410) |
| 0.9 | 2.7 | GO:0047256 | beta-galactosyl-N-acetylglucosaminylgalactosylglucosyl-ceramide beta-1,3-acetylglucosaminyltransferase activity(GO:0008457) lactosylceramide 1,3-N-acetyl-beta-D-glucosaminyltransferase activity(GO:0047256) |
| 0.8 | 3.4 | GO:0045131 | pre-mRNA branch point binding(GO:0045131) |
| 0.8 | 2.4 | GO:0032052 | bile acid binding(GO:0032052) |
| 0.8 | 3.9 | GO:0017108 | 5'-flap endonuclease activity(GO:0017108) |
| 0.7 | 2.2 | GO:0000402 | open form four-way junction DNA binding(GO:0000401) crossed form four-way junction DNA binding(GO:0000402) |
| 0.7 | 2.1 | GO:0005118 | sevenless binding(GO:0005118) |
| 0.7 | 5.6 | GO:0043559 | insulin binding(GO:0043559) |
| 0.7 | 2.7 | GO:0004784 | superoxide dismutase activity(GO:0004784) oxidoreductase activity, acting on superoxide radicals as acceptor(GO:0016721) |
| 0.7 | 3.4 | GO:0008449 | N-acetylglucosamine-6-sulfatase activity(GO:0008449) |
| 0.7 | 2.6 | GO:0034988 | Fc-gamma receptor I complex binding(GO:0034988) |
| 0.7 | 2.0 | GO:0008823 | cupric reductase activity(GO:0008823) ferric-chelate reductase (NADPH) activity(GO:0052851) |
| 0.6 | 2.5 | GO:0031727 | CCR2 chemokine receptor binding(GO:0031727) |
| 0.6 | 11.4 | GO:0008307 | structural constituent of muscle(GO:0008307) |
| 0.6 | 4.4 | GO:0031995 | insulin-like growth factor II binding(GO:0031995) |
| 0.6 | 1.8 | GO:1901611 | phosphatidylglycerol binding(GO:1901611) cardiolipin binding(GO:1901612) |
| 0.6 | 2.8 | GO:0001069 | regulatory region RNA binding(GO:0001069) |
| 0.5 | 2.7 | GO:0032767 | copper-dependent protein binding(GO:0032767) |
| 0.5 | 2.1 | GO:0042806 | fucose binding(GO:0042806) |
| 0.5 | 4.1 | GO:0070290 | N-acylphosphatidylethanolamine-specific phospholipase D activity(GO:0070290) |
| 0.5 | 1.8 | GO:0001105 | RNA polymerase II transcription coactivator activity(GO:0001105) transcriptional activator activity, RNA polymerase II transcription factor binding(GO:0001190) transcriptional repressor activity, RNA polymerase II activating transcription factor binding(GO:0098811) |
| 0.4 | 2.1 | GO:0005134 | interleukin-2 receptor binding(GO:0005134) |
| 0.4 | 2.9 | GO:0015016 | [heparan sulfate]-glucosamine N-sulfotransferase activity(GO:0015016) |
| 0.4 | 1.7 | GO:0008475 | procollagen-lysine 5-dioxygenase activity(GO:0008475) |
| 0.4 | 1.6 | GO:0004066 | asparagine synthase (glutamine-hydrolyzing) activity(GO:0004066) |
| 0.4 | 2.5 | GO:0004897 | ciliary neurotrophic factor receptor activity(GO:0004897) |
| 0.4 | 4.4 | GO:0034943 | acyl-CoA ligase activity(GO:0003996) 3-oxo-2-(2'-pentenyl)cyclopentane-1-octanoic acid CoA ligase activity(GO:0010435) 3-isopropenyl-6-oxoheptanoyl-CoA synthetase activity(GO:0018854) 2-oxo-delta3-4,5,5-trimethylcyclopentenylacetyl-CoA synthetase activity(GO:0018855) benzoyl acetate-CoA ligase activity(GO:0018856) 2,4-dichlorobenzoate-CoA ligase activity(GO:0018857) pivalate-CoA ligase activity(GO:0034783) cyclopropanecarboxylate-CoA ligase activity(GO:0034793) adipate-CoA ligase activity(GO:0034796) citronellyl-CoA ligase activity(GO:0034823) mentha-1,3-dione-CoA ligase activity(GO:0034841) thiophene-2-carboxylate-CoA ligase activity(GO:0034842) 2,4,4-trimethylpentanoate-CoA ligase activity(GO:0034865) cis-2-methyl-5-isopropylhexa-2,5-dienoate-CoA ligase activity(GO:0034942) trans-2-methyl-5-isopropylhexa-2,5-dienoate-CoA ligase activity(GO:0034943) branched-chain acyl-CoA synthetase (ADP-forming) activity(GO:0043759) aryl-CoA synthetase (ADP-forming) activity(GO:0043762) 3-hydroxypropionyl-CoA synthetase activity(GO:0043955) perillic acid:CoA ligase (ADP-forming) activity(GO:0052685) perillic acid:CoA ligase (AMP-forming) activity(GO:0052686) (3R)-3-isopropenyl-6-oxoheptanoate:CoA ligase (ADP-forming) activity(GO:0052687) (3R)-3-isopropenyl-6-oxoheptanoate:CoA ligase (AMP-forming) activity(GO:0052688) pristanate-CoA ligase activity(GO:0070251) malonyl-CoA synthetase activity(GO:0090409) |
| 0.4 | 2.0 | GO:0051880 | Y-form DNA binding(GO:0000403) bubble DNA binding(GO:0000405) G-quadruplex DNA binding(GO:0051880) |
| 0.4 | 6.4 | GO:0015129 | lactate transmembrane transporter activity(GO:0015129) |
| 0.4 | 4.4 | GO:0030957 | Tat protein binding(GO:0030957) |
| 0.4 | 1.2 | GO:0005534 | galactose binding(GO:0005534) |
| 0.4 | 4.6 | GO:0001161 | intronic transcription regulatory region sequence-specific DNA binding(GO:0001161) |
| 0.4 | 5.8 | GO:0019855 | calcium channel inhibitor activity(GO:0019855) |
| 0.4 | 1.9 | GO:0004095 | carnitine O-palmitoyltransferase activity(GO:0004095) |
| 0.4 | 1.1 | GO:0004828 | serine-tRNA ligase activity(GO:0004828) |
| 0.4 | 5.2 | GO:0022889 | L-serine transmembrane transporter activity(GO:0015194) serine transmembrane transporter activity(GO:0022889) |
| 0.3 | 1.4 | GO:0031735 | CCR10 chemokine receptor binding(GO:0031735) |
| 0.3 | 1.4 | GO:0003941 | L-serine ammonia-lyase activity(GO:0003941) |
| 0.3 | 2.7 | GO:0015168 | glycerol transmembrane transporter activity(GO:0015168) glycerol channel activity(GO:0015254) |
| 0.3 | 2.0 | GO:0070644 | vitamin D response element binding(GO:0070644) |
| 0.3 | 1.3 | GO:1990460 | leptin receptor binding(GO:1990460) |
| 0.3 | 3.3 | GO:0008474 | palmitoyl-(protein) hydrolase activity(GO:0008474) palmitoyl hydrolase activity(GO:0098599) |
| 0.3 | 3.7 | GO:1990226 | histone methyltransferase binding(GO:1990226) |
| 0.3 | 1.5 | GO:0050265 | RNA uridylyltransferase activity(GO:0050265) |
| 0.3 | 6.8 | GO:0016676 | cytochrome-c oxidase activity(GO:0004129) heme-copper terminal oxidase activity(GO:0015002) oxidoreductase activity, acting on a heme group of donors, oxygen as acceptor(GO:0016676) |
| 0.3 | 1.2 | GO:0008802 | L-aminoadipate-semialdehyde dehydrogenase activity(GO:0004043) betaine-aldehyde dehydrogenase activity(GO:0008802) |
| 0.3 | 0.9 | GO:0052740 | phosphatidylserine 1-acylhydrolase activity(GO:0052739) 1-acyl-2-lysophosphatidylserine acylhydrolase activity(GO:0052740) |
| 0.3 | 3.4 | GO:0070097 | delta-catenin binding(GO:0070097) |
| 0.3 | 2.0 | GO:0035174 | histone serine kinase activity(GO:0035174) |
| 0.3 | 0.8 | GO:0035575 | histone demethylase activity (H4-K20 specific)(GO:0035575) |
| 0.3 | 0.8 | GO:0031686 | A1 adenosine receptor binding(GO:0031686) |
| 0.3 | 1.9 | GO:0032554 | purine deoxyribonucleotide binding(GO:0032554) |
| 0.3 | 2.6 | GO:0042301 | phosphate ion binding(GO:0042301) |
| 0.3 | 2.5 | GO:0001055 | RNA polymerase II activity(GO:0001055) |
| 0.3 | 5.5 | GO:0001784 | phosphotyrosine binding(GO:0001784) |
| 0.3 | 1.0 | GO:0042731 | PH domain binding(GO:0042731) |
| 0.2 | 4.2 | GO:0010485 | H4 histone acetyltransferase activity(GO:0010485) |
| 0.2 | 1.5 | GO:0005173 | stem cell factor receptor binding(GO:0005173) |
| 0.2 | 1.7 | GO:0005049 | nuclear export signal receptor activity(GO:0005049) |
| 0.2 | 4.2 | GO:0005540 | hyaluronic acid binding(GO:0005540) |
| 0.2 | 1.9 | GO:0001972 | retinoic acid binding(GO:0001972) |
| 0.2 | 1.1 | GO:0004382 | guanosine-diphosphatase activity(GO:0004382) |
| 0.2 | 0.7 | GO:0030249 | guanylate cyclase regulator activity(GO:0030249) |
| 0.2 | 2.7 | GO:0001163 | RNA polymerase I regulatory region sequence-specific DNA binding(GO:0001163) RNA polymerase I CORE element sequence-specific DNA binding(GO:0001164) |
| 0.2 | 1.3 | GO:1990829 | C-rich single-stranded DNA binding(GO:1990829) |
| 0.2 | 1.4 | GO:0036402 | proteasome-activating ATPase activity(GO:0036402) |
| 0.2 | 6.1 | GO:0017147 | Wnt-protein binding(GO:0017147) |
| 0.2 | 3.4 | GO:0005078 | MAP-kinase scaffold activity(GO:0005078) |
| 0.2 | 1.3 | GO:0004013 | adenosylhomocysteinase activity(GO:0004013) trialkylsulfonium hydrolase activity(GO:0016802) |
| 0.2 | 4.9 | GO:0071837 | HMG box domain binding(GO:0071837) |
| 0.2 | 2.6 | GO:0019215 | intermediate filament binding(GO:0019215) |
| 0.2 | 2.7 | GO:0008574 | ATP-dependent microtubule motor activity, plus-end-directed(GO:0008574) |
| 0.2 | 4.1 | GO:0005528 | macrolide binding(GO:0005527) FK506 binding(GO:0005528) |
| 0.2 | 0.5 | GO:0004816 | asparagine-tRNA ligase activity(GO:0004816) |
| 0.2 | 1.9 | GO:0008190 | eukaryotic initiation factor 4E binding(GO:0008190) |
| 0.2 | 1.2 | GO:0031419 | cobalamin binding(GO:0031419) |
| 0.2 | 1.9 | GO:0003689 | DNA clamp loader activity(GO:0003689) protein-DNA loading ATPase activity(GO:0033170) |
| 0.2 | 0.7 | GO:0008073 | ornithine decarboxylase inhibitor activity(GO:0008073) |
| 0.2 | 0.9 | GO:0034511 | U3 snoRNA binding(GO:0034511) |
| 0.2 | 2.9 | GO:0009982 | pseudouridine synthase activity(GO:0009982) |
| 0.2 | 1.2 | GO:0043237 | laminin-1 binding(GO:0043237) |
| 0.2 | 1.0 | GO:0019788 | NEDD8 transferase activity(GO:0019788) |
| 0.2 | 1.3 | GO:0035242 | protein-arginine omega-N asymmetric methyltransferase activity(GO:0035242) |
| 0.2 | 7.4 | GO:0097472 | cyclin-dependent protein serine/threonine kinase activity(GO:0004693) cyclin-dependent protein kinase activity(GO:0097472) |
| 0.2 | 2.1 | GO:0000983 | transcription factor activity, RNA polymerase II core promoter sequence-specific(GO:0000983) |
| 0.2 | 6.2 | GO:0043027 | cysteine-type endopeptidase inhibitor activity involved in apoptotic process(GO:0043027) |
| 0.2 | 1.1 | GO:0034452 | dynactin binding(GO:0034452) |
| 0.2 | 1.4 | GO:0004128 | cytochrome-b5 reductase activity, acting on NAD(P)H(GO:0004128) |
| 0.1 | 1.2 | GO:0046790 | virion binding(GO:0046790) |
| 0.1 | 4.7 | GO:0005109 | frizzled binding(GO:0005109) |
| 0.1 | 1.3 | GO:0004634 | phosphopyruvate hydratase activity(GO:0004634) |
| 0.1 | 2.1 | GO:0030898 | actin-dependent ATPase activity(GO:0030898) |
| 0.1 | 0.8 | GO:0047276 | N-acetyllactosaminide 3-alpha-galactosyltransferase activity(GO:0047276) |
| 0.1 | 4.2 | GO:0032452 | histone demethylase activity(GO:0032452) |
| 0.1 | 0.8 | GO:0031957 | very long-chain fatty acid-CoA ligase activity(GO:0031957) |
| 0.1 | 0.4 | GO:0005124 | scavenger receptor binding(GO:0005124) |
| 0.1 | 4.5 | GO:0070888 | E-box binding(GO:0070888) |
| 0.1 | 4.8 | GO:0017112 | Rab guanyl-nucleotide exchange factor activity(GO:0017112) |
| 0.1 | 1.8 | GO:0000400 | four-way junction DNA binding(GO:0000400) |
| 0.1 | 1.4 | GO:0034946 | 2-oxoglutaryl-CoA thioesterase activity(GO:0034843) 2,4,4-trimethyl-3-oxopentanoyl-CoA thioesterase activity(GO:0034869) 3-isopropylbut-3-enoyl-CoA thioesterase activity(GO:0034946) glutaryl-CoA hydrolase activity(GO:0044466) |
| 0.1 | 1.4 | GO:0004467 | long-chain fatty acid-CoA ligase activity(GO:0004467) decanoate--CoA ligase activity(GO:0102391) |
| 0.1 | 0.5 | GO:0070891 | lipoteichoic acid binding(GO:0070891) |
| 0.1 | 3.1 | GO:0008139 | nuclear localization sequence binding(GO:0008139) |
| 0.1 | 1.5 | GO:0008641 | small protein activating enzyme activity(GO:0008641) |
| 0.1 | 0.4 | GO:0071987 | WD40-repeat domain binding(GO:0071987) |
| 0.1 | 4.6 | GO:0016538 | cyclin-dependent protein serine/threonine kinase regulator activity(GO:0016538) |
| 0.1 | 2.9 | GO:0003887 | DNA-directed DNA polymerase activity(GO:0003887) |
| 0.1 | 0.4 | GO:0070182 | DNA polymerase binding(GO:0070182) |
| 0.1 | 2.2 | GO:0005154 | epidermal growth factor receptor binding(GO:0005154) |
| 0.1 | 26.0 | GO:0001077 | transcriptional activator activity, RNA polymerase II core promoter proximal region sequence-specific binding(GO:0001077) |
| 0.1 | 0.3 | GO:0004127 | cytidylate kinase activity(GO:0004127) uridylate kinase activity(GO:0009041) |
| 0.1 | 0.3 | GO:0071633 | dihydroceramidase activity(GO:0071633) |
| 0.1 | 0.8 | GO:0000182 | rDNA binding(GO:0000182) |
| 0.1 | 5.4 | GO:0003743 | translation initiation factor activity(GO:0003743) |
| 0.1 | 1.3 | GO:0045125 | bioactive lipid receptor activity(GO:0045125) |
| 0.1 | 0.3 | GO:0004647 | phosphoserine phosphatase activity(GO:0004647) |
| 0.1 | 0.8 | GO:0004571 | mannosyl-oligosaccharide 1,2-alpha-mannosidase activity(GO:0004571) |
| 0.1 | 2.3 | GO:0030544 | Hsp70 protein binding(GO:0030544) |
| 0.1 | 1.3 | GO:0005355 | glucose transmembrane transporter activity(GO:0005355) |
| 0.1 | 1.9 | GO:0032183 | SUMO binding(GO:0032183) |
| 0.1 | 1.0 | GO:0046933 | proton-transporting ATP synthase activity, rotational mechanism(GO:0046933) |
| 0.1 | 2.4 | GO:0005112 | Notch binding(GO:0005112) |
| 0.1 | 0.6 | GO:0035184 | histone threonine kinase activity(GO:0035184) |
| 0.1 | 1.5 | GO:0030295 | protein kinase activator activity(GO:0030295) |
| 0.1 | 2.2 | GO:0070717 | poly-purine tract binding(GO:0070717) |
| 0.1 | 3.5 | GO:0051018 | protein kinase A binding(GO:0051018) |
| 0.1 | 1.3 | GO:0001056 | RNA polymerase III activity(GO:0001056) |
| 0.1 | 1.5 | GO:0003755 | peptidyl-prolyl cis-trans isomerase activity(GO:0003755) |
| 0.1 | 2.4 | GO:0043014 | alpha-tubulin binding(GO:0043014) |
| 0.1 | 0.9 | GO:0005160 | transforming growth factor beta receptor binding(GO:0005160) |
| 0.1 | 0.4 | GO:0005222 | intracellular cAMP activated cation channel activity(GO:0005222) |
| 0.1 | 0.3 | GO:0004459 | L-lactate dehydrogenase activity(GO:0004459) |
| 0.1 | 0.4 | GO:1990932 | 5.8S rRNA binding(GO:1990932) |
| 0.1 | 0.9 | GO:0031418 | L-ascorbic acid binding(GO:0031418) |
| 0.1 | 0.9 | GO:0016805 | dipeptidase activity(GO:0016805) |
| 0.1 | 0.3 | GO:0051022 | Rho GDP-dissociation inhibitor binding(GO:0051022) |
| 0.1 | 0.2 | GO:0004069 | L-aspartate:2-oxoglutarate aminotransferase activity(GO:0004069) |
| 0.1 | 1.6 | GO:0005385 | zinc ion transmembrane transporter activity(GO:0005385) |
| 0.1 | 1.2 | GO:0017134 | fibroblast growth factor binding(GO:0017134) |
| 0.1 | 1.5 | GO:0005201 | extracellular matrix structural constituent(GO:0005201) |
| 0.1 | 1.6 | GO:0003746 | translation elongation factor activity(GO:0003746) |
| 0.1 | 1.1 | GO:0070064 | proline-rich region binding(GO:0070064) |
| 0.1 | 0.7 | GO:0051010 | microtubule plus-end binding(GO:0051010) |
| 0.1 | 0.2 | GO:0047066 | phospholipid-hydroperoxide glutathione peroxidase activity(GO:0047066) |
| 0.0 | 1.0 | GO:0017025 | TBP-class protein binding(GO:0017025) |
| 0.0 | 2.4 | GO:0031491 | nucleosome binding(GO:0031491) |
| 0.0 | 4.9 | GO:0001228 | transcriptional activator activity, RNA polymerase II transcription regulatory region sequence-specific binding(GO:0001228) |
| 0.0 | 0.9 | GO:0003950 | NAD+ ADP-ribosyltransferase activity(GO:0003950) |
| 0.0 | 0.1 | GO:0005087 | Ran guanyl-nucleotide exchange factor activity(GO:0005087) |
| 0.0 | 0.7 | GO:0050253 | prenylcysteine methylesterase activity(GO:0010296) 1-oxa-2-oxocycloheptane lactonase activity(GO:0018731) sulfolactone hydrolase activity(GO:0018732) butyrolactone hydrolase activity(GO:0018734) endosulfan lactone lactonase activity(GO:0034892) L-ascorbate 6-phosphate lactonase activity(GO:0035460) Ser-tRNA(Thr) hydrolase activity(GO:0043905) Ala-tRNA(Pro) hydrolase activity(GO:0043906) Cys-tRNA(Pro) hydrolase activity(GO:0043907) Ser(Gly)-tRNA(Ala) hydrolase activity(GO:0043908) all-trans-retinyl-palmitate hydrolase, all-trans-retinol forming activity(GO:0047376) retinyl-palmitate esterase activity(GO:0050253) mannosyl-oligosaccharide 1,6-alpha-mannosidase activity(GO:0052767) mannosyl-oligosaccharide 1,3-alpha-mannosidase activity(GO:0052768) methyl indole-3-acetate esterase activity(GO:0080030) methyl salicylate esterase activity(GO:0080031) methyl jasmonate esterase activity(GO:0080032) |
| 0.0 | 2.7 | GO:0008186 | ATP-dependent RNA helicase activity(GO:0004004) RNA-dependent ATPase activity(GO:0008186) |
| 0.0 | 0.4 | GO:0042288 | MHC class I protein binding(GO:0042288) |
| 0.0 | 0.4 | GO:0005149 | interleukin-1 receptor binding(GO:0005149) |
| 0.0 | 0.8 | GO:0070016 | armadillo repeat domain binding(GO:0070016) |
| 0.0 | 0.5 | GO:0001221 | transcription cofactor binding(GO:0001221) |
| 0.0 | 0.7 | GO:0004190 | aspartic-type endopeptidase activity(GO:0004190) aspartic-type peptidase activity(GO:0070001) |
| 0.0 | 0.2 | GO:0003854 | 3-beta-hydroxy-delta5-steroid dehydrogenase activity(GO:0003854) |
| 0.0 | 0.8 | GO:0043325 | phosphatidylinositol-3,4-bisphosphate binding(GO:0043325) |
| 0.0 | 0.2 | GO:0016176 | superoxide-generating NADPH oxidase activator activity(GO:0016176) |
| 0.0 | 1.2 | GO:0001227 | transcriptional repressor activity, RNA polymerase II transcription regulatory region sequence-specific binding(GO:0001227) |
| 0.0 | 0.4 | GO:0070636 | DNA-(apurinic or apyrimidinic site) lyase activity(GO:0003906) single-strand selective uracil DNA N-glycosylase activity(GO:0017065) nicotinamide riboside hydrolase activity(GO:0070635) nicotinic acid riboside hydrolase activity(GO:0070636) deoxyribonucleoside 5'-monophosphate N-glycosidase activity(GO:0070694) |
| 0.0 | 0.9 | GO:0019789 | SUMO transferase activity(GO:0019789) |
| 0.0 | 2.2 | GO:0005518 | collagen binding(GO:0005518) |
| 0.0 | 3.6 | GO:0001078 | transcriptional repressor activity, RNA polymerase II core promoter proximal region sequence-specific binding(GO:0001078) |
| 0.0 | 4.5 | GO:0017137 | Rab GTPase binding(GO:0017137) |
| 0.0 | 3.2 | GO:0008083 | growth factor activity(GO:0008083) |
| 0.0 | 0.6 | GO:0000980 | RNA polymerase II distal enhancer sequence-specific DNA binding(GO:0000980) |
| 0.0 | 0.2 | GO:0070191 | methionine-R-sulfoxide reductase activity(GO:0070191) |
| 0.0 | 0.3 | GO:0016308 | 1-phosphatidylinositol-4-phosphate 5-kinase activity(GO:0016308) |
| 0.0 | 1.9 | GO:0019955 | cytokine binding(GO:0019955) |
| 0.0 | 0.8 | GO:0003785 | actin monomer binding(GO:0003785) |
| 0.0 | 3.0 | GO:0003774 | motor activity(GO:0003774) |
| 0.0 | 0.4 | GO:0016208 | AMP binding(GO:0016208) |
| 0.0 | 4.3 | GO:0008237 | metallopeptidase activity(GO:0008237) |
| 0.0 | 0.1 | GO:0001639 | PLC activating G-protein coupled glutamate receptor activity(GO:0001639) |
| 0.0 | 0.3 | GO:0004383 | guanylate cyclase activity(GO:0004383) |
| 0.0 | 1.9 | GO:0061659 | ubiquitin protein ligase activity(GO:0061630) ubiquitin-like protein ligase activity(GO:0061659) |
| 0.0 | 0.3 | GO:0008191 | metalloendopeptidase inhibitor activity(GO:0008191) |
| 0.0 | 0.1 | GO:0070883 | pre-miRNA binding(GO:0070883) |
| 0.0 | 0.7 | GO:0005504 | fatty acid binding(GO:0005504) |
| 0.0 | 0.7 | GO:0051879 | Hsp90 protein binding(GO:0051879) |
| 0.0 | 0.6 | GO:0016765 | transferase activity, transferring alkyl or aryl (other than methyl) groups(GO:0016765) |
| 0.0 | 1.0 | GO:0031593 | polyubiquitin binding(GO:0031593) |
| 0.0 | 1.0 | GO:0005179 | hormone activity(GO:0005179) |
| 0.0 | 1.7 | GO:0016423 | tRNA (guanine) methyltransferase activity(GO:0016423) |
| 0.0 | 0.9 | GO:0016836 | hydro-lyase activity(GO:0016836) |
| 0.0 | 0.8 | GO:0005125 | cytokine activity(GO:0005125) |
| 0.0 | 0.7 | GO:0004812 | aminoacyl-tRNA ligase activity(GO:0004812) |
| 0.0 | 1.0 | GO:0035254 | glutamate receptor binding(GO:0035254) |
| 0.0 | 0.1 | GO:0004614 | phosphoglucomutase activity(GO:0004614) |
| 0.0 | 0.3 | GO:0005234 | extracellular-glutamate-gated ion channel activity(GO:0005234) |
| 0.0 | 0.3 | GO:0017069 | snRNA binding(GO:0017069) |
| 0.0 | 0.2 | GO:0004550 | nucleoside diphosphate kinase activity(GO:0004550) |
| 0.0 | 0.3 | GO:0048487 | beta-tubulin binding(GO:0048487) |
| 0.0 | 0.3 | GO:0034930 | N-acetylglucosamine 6-O-sulfotransferase activity(GO:0001517) heparan sulfate 2-O-sulfotransferase activity(GO:0004394) HNK-1 sulfotransferase activity(GO:0016232) heparan sulfate 6-O-sulfotransferase activity(GO:0017095) trans-9R,10R-dihydrodiolphenanthrene sulfotransferase activity(GO:0018721) 1-phenanthrol sulfotransferase activity(GO:0018722) 3-phenanthrol sulfotransferase activity(GO:0018723) 4-phenanthrol sulfotransferase activity(GO:0018724) trans-3,4-dihydrodiolphenanthrene sulfotransferase activity(GO:0018725) 9-phenanthrol sulfotransferase activity(GO:0018726) 2-phenanthrol sulfotransferase activity(GO:0018727) phenanthrol sulfotransferase activity(GO:0019111) 1-hydroxypyrene sulfotransferase activity(GO:0034930) proteoglycan sulfotransferase activity(GO:0050698) cholesterol sulfotransferase activity(GO:0051922) hydroxyjasmonate sulfotransferase activity(GO:0080131) |