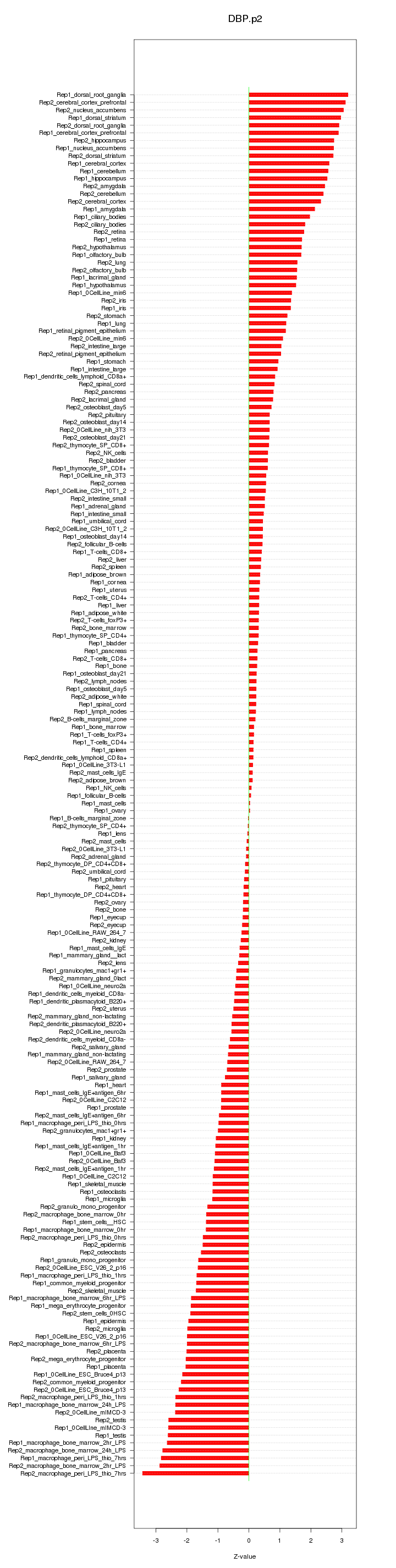
|
chr8_+_56040211
|
38.971
|
|
Gpm6a
|
glycoprotein m6a
|
|
chr8_+_56040553
|
38.369
|
|
Gpm6a
|
glycoprotein m6a
|
|
chr8_+_56040445
|
37.557
|
|
Gpm6a
|
glycoprotein m6a
|
|
chr1_-_166368439
|
31.532
|
|
Atp1b1
|
ATPase, Na+/K+ transporting, beta 1 polypeptide
|
|
chr16_+_7069971
|
29.678
|
|
Rbfox1
|
RNA binding protein, fox-1 homolog (C. elegans) 1
|
|
chr10_+_40657980
|
29.564
|
|
Wasf1
|
WASP family 1
|
|
chr14_+_84933363
|
27.001
|
|
Pcdh17
|
protocadherin 17
|
|
chrX_-_100381703
|
25.810
|
|
Nap1l2
|
nucleosome assembly protein 1-like 2
|
|
chrX_+_54043764
|
25.584
|
|
Fhl1
|
four and a half LIM domains 1
|
|
chr12_+_91570569
|
22.463
|
|
Nrxn3
|
neurexin III
|
|
chr16_+_7069927
|
21.307
|
NM_183188
|
Rbfox1
|
RNA binding protein, fox-1 homolog (C. elegans) 1
|
|
chr8_+_56040193
|
18.913
|
|
Gpm6a
|
glycoprotein m6a
|
|
chr9_+_14135884
|
18.441
|
|
Sesn3
|
sestrin 3
|
|
chr10_+_122701051
|
17.769
|
NM_182807
|
Fam19a2
|
family with sequence similarity 19, member A2
|
|
chr1_+_34364633
|
17.451
|
|
Dst
|
dystonin
|
|
chr5_+_72091188
|
17.090
|
NM_008069
|
Gabrb1
|
gamma-aminobutyric acid (GABA) A receptor, subunit beta 1
|
|
chr10_+_115614313
|
17.079
|
NM_001161839
NM_001161838
NM_001161840
|
Ptprr
|
protein tyrosine phosphatase, receptor type, R
|
|
chr8_+_56040080
|
16.727
|
NM_153581
|
Gpm6a
|
glycoprotein m6a
|
|
chr14_+_14286471
|
16.722
|
NM_001163032
|
Synpr
|
synaptoporin
|
|
chr5_-_122906470
|
16.536
|
|
Atp2a2
|
ATPase, Ca++ transporting, cardiac muscle, slow twitch 2
|
|
chr12_-_102988596
|
16.519
|
|
|
|
|
chr18_-_15562126
|
16.042
|
NM_009700
|
Aqp4
|
aquaporin 4
|
|
chr2_-_52414072
|
15.928
|
NM_146123
|
Cacnb4
|
calcium channel, voltage-dependent, beta 4 subunit
|
|
chr4_-_75779379
|
15.926
|
NM_001014288
|
Ptprd
|
protein tyrosine phosphatase, receptor type, D
|
|
chr16_+_7409879
|
15.744
|
|
Rbfox1
|
RNA binding protein, fox-1 homolog (C. elegans) 1
|
|
chr12_+_62624880
|
15.297
|
|
Lrfn5
|
leucine rich repeat and fibronectin type III domain containing 5
|
|
chr3_-_126646303
|
15.103
|
NM_001034168
|
Ank2
|
ankyrin 2, brain
|
|
chr15_-_37389073
|
14.935
|
NM_001170867
NM_001170868
|
Ncald
|
neurocalcin delta
|
|
chr18_+_37663917
|
14.898
|
NM_053145
|
Pcdhb20
|
protocadherin beta 20
|
|
chr14_-_55735114
|
14.556
|
NM_177049
|
Jph4
|
junctophilin 4
|
|
chr7_-_82453216
|
13.765
|
NM_001109753
|
Sv2b
|
synaptic vesicle glycoprotein 2 b
|
|
chr17_-_90487479
|
13.392
|
|
Nrxn1
|
neurexin I
|
|
chr6_+_30758582
|
13.287
|
|
Copg2as2
|
coatomer protein complex, subunit gamma 2, antisense 2
|
|
chr19_+_23239333
|
13.248
|
|
Klf9
|
Kruppel-like factor 9
|
|
chr2_+_177910053
|
13.061
|
NM_001177791
|
Phactr3
|
phosphatase and actin regulator 3
|
|
chr14_+_28689289
|
12.757
|
|
Erc2
|
ELKS/RAB6-interacting/CAST family member 2
|
|
chr5_-_51945054
|
12.584
|
NM_008904
|
Ppargc1a
|
peroxisome proliferative activated receptor, gamma, coactivator 1 alpha
|
|
chr14_+_13073221
|
12.574
|
|
Ptprg
|
protein tyrosine phosphatase, receptor type, G
|
|
chr3_+_64913564
|
12.402
|
NM_010597
|
Kcnab1
|
potassium voltage-gated channel, shaker-related subfamily, beta member 1
|
|
chr3_+_52153283
|
12.286
|
|
Foxo1
|
forkhead box O1
|
|
chr3_+_17954324
|
12.213
|
NM_021560
|
Bhlhe22
|
basic helix-loop-helix family, member e22
|
|
chr9_+_75192367
|
12.187
|
|
Gnb5
|
guanine nucleotide binding protein (G protein), beta 5
|
|
chr3_+_136598485
|
12.079
|
|
Ppp3ca
|
protein phosphatase 3, catalytic subunit, alpha isoform
|
|
chr14_-_122138082
|
12.067
|
NM_001128307
NM_001128308
|
Dock9
|
dedicator of cytokinesis 9
|
|
chr11_-_36757743
|
11.956
|
NM_011856
|
Odz2
|
odd Oz/ten-m homolog 2 (Drosophila)
|
|
chr9_+_32031833
|
11.946
|
NM_177379
|
Arhgap32
|
Rho GTPase activating protein 32
|
|
chr9_-_112090150
|
11.836
|
NM_033264
NM_001177615
NM_001177620
NM_028755
|
Arpp21
|
cyclic AMP-regulated phosphoprotein, 21
|
|
chr5_-_122907002
|
11.762
|
|
Atp2a2
|
ATPase, Ca++ transporting, cardiac muscle, slow twitch 2
|
|
chr3_+_136598184
|
11.684
|
|
Ppp3ca
|
protein phosphatase 3, catalytic subunit, alpha isoform
|
|
chr11_+_31901708
|
11.593
|
|
Nsg2
|
neuron specific gene family member 2
|
|
chr8_-_67071497
|
11.554
|
|
|
|
|
chr9_+_27598853
|
11.545
|
NM_177906
|
Opcml
|
opioid binding protein/cell adhesion molecule-like
|
|
chr11_+_98629709
|
11.520
|
|
|
|
|
chr18_-_25912453
|
11.427
|
NM_001146292
NM_001146293
NM_001146294
NM_001146295
NM_001174074
NM_133195
|
Celf4
|
CUGBP, Elav-like family member 4
|
|
chr3_-_80606704
|
11.313
|
NM_001039195
NM_001083806
NM_013540
|
Gria2
|
glutamate receptor, ionotropic, AMPA2 (alpha 2)
|
|
chr7_-_82453166
|
11.206
|
|
Sv2b
|
synaptic vesicle glycoprotein 2 b
|
|
chr7_-_91520991
|
11.202
|
|
Arnt2
|
aryl hydrocarbon receptor nuclear translocator 2
|
|
chr3_-_80606500
|
11.055
|
|
Gria2
|
glutamate receptor, ionotropic, AMPA2 (alpha 2)
|
|
chr15_+_4325490
|
11.047
|
NM_177355
|
Plcxd3
|
phosphatidylinositol-specific phospholipase C, X domain containing 3
|
|
chr17_+_44090139
|
11.008
|
NM_030598
|
Rcan2
|
regulator of calcineurin 2
|
|
chr17_+_44090222
|
11.005
|
|
Rcan2
|
regulator of calcineurin 2
|
|
chr2_+_163843744
|
10.988
|
|
|
|
|
chr14_-_94287950
|
10.971
|
NM_001081377
|
Pcdh9
|
protocadherin 9
|
|
chrX_-_100381973
|
10.860
|
NM_008671
|
Nap1l2
|
nucleosome assembly protein 1-like 2
|
|
chrX_-_149471740
|
10.825
|
NM_023788
|
Mageh1
|
melanoma antigen, family H, 1
|
|
chr12_-_102988893
|
10.797
|
|
Fbln5
|
fibulin 5
|
|
chr8_+_12949417
|
10.587
|
NM_001159485
NM_001159486
|
Mcf2l
|
mcf.2 transforming sequence-like
|
|
chrX_+_155897136
|
10.474
|
|
Mtap7d2
|
MAP7 domain containing 2
|
|
chr3_+_13946371
|
10.405
|
NM_001163329
NM_001163330
|
Ralyl
|
RALY RNA binding protein-like
|
|
chr11_+_56825258
|
10.373
|
|
Gria1
|
glutamate receptor, ionotropic, AMPA1 (alpha 1)
|
|
chr13_+_118391270
|
10.329
|
|
Hcn1
|
hyperpolarization-activated, cyclic nucleotide-gated K+ 1
|
|
chr6_-_141804691
|
10.318
|
NM_030687
|
Slco1a4
|
solute carrier organic anion transporter family, member 1a4
|
|
chr12_+_71008808
|
10.118
|
|
Atl1
|
atlastin GTPase 1
|
|
chr19_-_40588193
|
10.066
|
|
Sorbs1
|
sorbin and SH3 domain containing 1
|
|
chr11_-_22185840
|
10.043
|
NM_153078
|
Ehbp1
|
EH domain binding protein 1
|
|
chr1_+_87804349
|
9.973
|
|
Itm2c
|
integral membrane protein 2C
|
|
chr7_+_129776837
|
9.963
|
|
Prkcb
|
protein kinase C, beta
|
|
chr14_+_56130147
|
9.840
|
NM_009947
|
Cpne6
|
copine VI
|
|
chr18_+_37644390
|
9.657
|
|
Pcdhb17
|
protocadherin beta 17
|
|
chr5_-_104542751
|
9.642
|
|
Sparcl1
|
SPARC-like 1
|
|
chr9_-_49607164
|
9.516
|
NM_001081445
NM_001113204
NM_010875
|
Ncam1
|
neural cell adhesion molecule 1
|
|
chr5_-_104542641
|
9.508
|
|
Sparcl1
|
SPARC-like 1
|
|
chr17_+_44090341
|
9.463
|
|
Rcan2
|
regulator of calcineurin 2
|
|
chr2_-_77541328
|
9.448
|
NM_001113399
|
Zfp385b
|
zinc finger protein 385B
|
|
chr3_-_80606641
|
9.441
|
|
Gria2
|
glutamate receptor, ionotropic, AMPA2 (alpha 2)
|
|
chr3_+_13471681
|
9.207
|
|
Ralyl
|
RALY RNA binding protein-like
|
|
chr18_-_31476527
|
9.174
|
|
Rit2
|
Ras-like without CAAX 2
|
|
chr10_-_80477630
|
9.150
|
NM_010319
|
Gng7
|
guanine nucleotide binding protein (G protein), gamma 7
|
|
chr17_+_86577946
|
9.090
|
|
|
|
|
chr5_-_104542729
|
9.086
|
|
Sparcl1
|
SPARC-like 1
|
|
chr3_-_103541868
|
9.070
|
|
Olfml3
|
olfactomedin-like 3
|
|
chr11_-_20729367
|
9.050
|
|
1110067D22Rik
|
RIKEN cDNA 1110067D22 gene
|
|
chr10_-_77734341
|
9.014
|
|
|
|
|
chr7_+_97535516
|
8.984
|
|
Sytl2
|
synaptotagmin-like 2
|
|
chr11_-_41813792
|
8.804
|
NM_177408
|
Gabrg2
|
gamma-aminobutyric acid (GABA) A receptor, subunit gamma 2
|
|
chr6_-_53770687
|
8.735
|
NM_025817
|
Tril
|
TLR4 interactor with leucine-rich repeats
|
|
chr6_+_54251485
|
8.679
|
|
|
|
|
chr4_-_60594496
|
8.601
|
|
Mup1
|
major urinary protein 1
|
|
chr5_-_45190857
|
8.524
|
NM_001077398
NM_010698
|
Ldb2
|
LIM domain binding 2
|
|
chr13_-_111070225
|
8.442
|
|
Rab3c
|
RAB3C, member RAS oncogene family
|
|
chr6_+_120723785
|
8.438
|
NM_001081048
|
Slc25a18
|
solute carrier family 25 (mitochondrial carrier), member 18
|
|
chr10_+_111708176
|
8.433
|
NM_001025581
|
Kcnc2
|
potassium voltage gated channel, Shaw-related subfamily, member 2
|
|
chr3_-_83967952
|
8.358
|
|
Trim2
|
tripartite motif-containing 2
|
|
chr5_+_151062363
|
8.203
|
NM_172887
|
Fry
|
furry homolog (Drosophila)
|
|
chr9_-_29770699
|
8.196
|
NM_172290
|
Ntm
|
neurotrimin
|
|
chr12_+_30219769
|
8.146
|
NM_001093776
|
Myt1l
|
myelin transcription factor 1-like
|
|
chr7_+_97535490
|
8.103
|
NM_001040087
NM_001040088
|
Sytl2
|
synaptotagmin-like 2
|
|
chr13_+_59234737
|
8.078
|
|
Ntrk2
|
neurotrophic tyrosine kinase, receptor, type 2
|
|
chr5_+_31167731
|
8.066
|
|
Mapre3
|
microtubule-associated protein, RP/EB family, member 3
|
|
chr7_-_134164487
|
7.997
|
|
Prrt2
|
proline-rich transmembrane protein 2
|
|
chr3_-_146444377
|
7.973
|
NM_001164200
|
Prkacb
|
protein kinase, cAMP dependent, catalytic, beta
|
|
chr5_-_104542598
|
7.948
|
|
Sparcl1
|
SPARC-like 1
|
|
chr17_-_85463965
|
7.940
|
|
|
|
|
chr7_-_108025317
|
7.925
|
|
Arhgef17
|
Rho guanine nucleotide exchange factor (GEF) 17
|
|
chr4_-_117881591
|
7.923
|
|
Ptprf
|
protein tyrosine phosphatase, receptor type, F
|
|
chr11_-_41814008
|
7.849
|
NM_008073
|
Gabrg2
|
gamma-aminobutyric acid (GABA) A receptor, subunit gamma 2
|
|
chr15_+_78744328
|
7.722
|
NM_020271
|
Pdxp
|
pyridoxal (pyridoxine, vitamin B6) phosphatase
|
|
chr1_+_87804443
|
7.678
|
|
Itm2c
|
integral membrane protein 2C
|
|
chr19_+_26822892
|
7.671
|
NM_026003
|
Smarca2
|
SWI/SNF related, matrix associated, actin dependent regulator of chromatin, subfamily a, member 2
|
|
chr4_+_28740314
|
7.640
|
|
Epha7
|
Eph receptor A7
|
|
chr4_+_55093796
|
7.615
|
|
Zfp462
|
zinc finger protein 462
|
|
chr7_+_64846528
|
7.576
|
NM_008071
|
Gabrb3
|
gamma-aminobutyric acid (GABA) A receptor, subunit beta 3
|
|
chr12_+_50483869
|
7.564
|
NM_001160112
NM_008241
|
Foxg1
|
forkhead box G1
|
|
chr1_+_155756310
|
7.553
|
|
Glul
|
glutamate-ammonia ligase (glutamine synthetase)
|
|
chr9_-_49607017
|
7.461
|
|
Ncam1
|
neural cell adhesion molecule 1
|
|
chr15_-_8660729
|
7.455
|
NM_148938
|
Slc1a3
|
solute carrier family 1 (glial high affinity glutamate transporter), member 3
|
|
chr17_+_44090192
|
7.438
|
|
Rcan2
|
regulator of calcineurin 2
|
|
chr2_+_67955769
|
7.410
|
NM_020283
|
B3galt1
|
UDP-Gal:betaGlcNAc beta 1,3-galactosyltransferase, polypeptide 1
|
|
chr18_-_25912171
|
7.378
|
|
Celf4
|
CUGBP, Elav-like family member 4
|
|
chr7_-_149838254
|
7.367
|
|
Igf2
|
insulin-like growth factor 2
|
|
chr11_+_97702591
|
7.334
|
|
B230217C12Rik
|
RIKEN cDNA B230217C12 gene
|
|
chr2_-_127347102
|
7.329
|
NM_019789
|
Kcnip3
|
Kv channel interacting protein 3, calsenilin
|
|
chr10_+_5277477
|
7.318
|
NM_022027
|
Syne1
|
synaptic nuclear envelope 1
|
|
chr10_-_88720534
|
7.269
|
NM_178773
|
Ano4
|
anoctamin 4
|
|
chr15_-_12479631
|
7.245
|
|
Pdzd2
|
PDZ domain containing 2
|
|
chr2_-_127346987
|
7.216
|
|
Kcnip3
|
Kv channel interacting protein 3, calsenilin
|
|
chr7_+_5031809
|
7.078
|
NM_010147
|
Epn1
|
epsin 1
|
|
chr5_-_104543106
|
7.074
|
NM_010097
|
Sparcl1
|
SPARC-like 1
|
|
chrX_-_42143327
|
7.067
|
NM_001190718
NM_178739
|
Dcaf12l1
|
DDB1 and CUL4 associated factor 12-like 1
|
|
chr8_-_113679781
|
7.044
|
|
Glg1
|
golgi apparatus protein 1
|
|
chr7_-_151621960
|
7.013
|
|
Cttn
|
cortactin
|
|
chr5_-_103640272
|
7.011
|
NM_001081567
NM_009158
|
Mapk10
|
mitogen-activated protein kinase 10
|
|
chr9_+_62225584
|
6.960
|
|
Anp32a
|
acidic (leucine-rich) nuclear phosphoprotein 32 family, member A
|
|
chr16_+_92086695
|
6.897
|
|
Slc5a3
|
solute carrier family 5 (inositol transporters), member 3
|
|
chr18_-_35374531
|
6.893
|
|
Lrrtm2
|
leucine rich repeat transmembrane neuronal 2
|
|
chr1_-_42749940
|
6.823
|
|
2610017I09Rik
|
RIKEN cDNA 2610017I09 gene
|
|
chr5_+_89012596
|
6.798
|
NM_027530
|
Rufy3
|
RUN and FYVE domain containing 3
|
|
chr13_+_118391126
|
6.763
|
NM_010408
|
Hcn1
|
hyperpolarization-activated, cyclic nucleotide-gated K+ 1
|
|
chr2_+_136539202
|
6.740
|
|
Snap25
|
synaptosomal-associated protein 25
|
|
chr3_-_30799013
|
6.678
|
|
Phc3
|
polyhomeotic-like 3 (Drosophila)
|
|
chrX_+_140100287
|
6.654
|
NM_001195049
|
Pak3
|
p21 protein (Cdc42/Rac)-activated kinase 3
|
|
chr1_+_51346086
|
6.654
|
|
Sdpr
|
serum deprivation response
|
|
chr7_+_64846558
|
6.647
|
|
Gabrb3
|
gamma-aminobutyric acid (GABA) A receptor, subunit beta 3
|
|
chr4_-_108311676
|
6.647
|
|
|
|
|
chr12_-_102988510
|
6.612
|
|
Fbln5
|
fibulin 5
|
|
chr1_+_51346108
|
6.577
|
|
Sdpr
|
serum deprivation response
|
|
chr18_-_31476721
|
6.496
|
NM_009065
|
Rit2
|
Ras-like without CAAX 2
|
|
chr5_-_24107718
|
6.476
|
NM_025891
|
Smarcd3
|
SWI/SNF related, matrix associated, actin dependent regulator of chromatin, subfamily d, member 3
|
|
chr9_-_83003967
|
6.439
|
|
|
|
|
chr16_+_13986691
|
6.409
|
NM_144518
|
2900011O08Rik
|
RIKEN cDNA 2900011O08 gene
|
|
chr3_-_103541882
|
6.392
|
NM_133859
|
Olfml3
|
olfactomedin-like 3
|
|
chr2_-_6466116
|
6.392
|
|
Celf2
|
CUGBP, Elav-like family member 2
|
|
chr5_+_89012547
|
6.382
|
|
Rufy3
|
RUN and FYVE domain containing 3
|
|
chrX_+_155897056
|
6.253
|
|
Mtap7d2
|
MAP7 domain containing 2
|
|
chr19_+_29999624
|
6.240
|
|
Il33
|
interleukin 33
|
|
chr5_+_44053634
|
6.147
|
NM_172274
|
Cc2d2a
|
coiled-coil and C2 domain containing 2A
|
|
chr19_-_44471675
|
6.129
|
|
Scd1
|
stearoyl-Coenzyme A desaturase 1
|
|
chr9_+_36651243
|
6.066
|
NM_183171
|
Fez1
|
fasciculation and elongation protein zeta 1 (zygin I)
|
|
chr1_+_87804881
|
6.045
|
|
Itm2c
|
integral membrane protein 2C
|
|
chr8_+_86520110
|
6.044
|
|
Prkaca
|
protein kinase, cAMP dependent, catalytic, alpha
|
|
chr18_+_23656024
|
6.025
|
|
Dtna
|
dystrobrevin alpha
|
|
chr18_+_23962468
|
5.978
|
NM_153058
|
Mapre2
|
microtubule-associated protein, RP/EB family, member 2
|
|
chr12_+_118997623
|
5.965
|
|
Rapgef5
|
Rap guanine nucleotide exchange factor (GEF) 5
|
|
chr10_-_63552994
|
5.957
|
NM_178678
|
Lrrtm3
|
leucine rich repeat transmembrane neuronal 3
|
|
chr18_+_63868368
|
5.956
|
|
Wdr7
|
WD repeat domain 7
|
|
chr7_-_134164627
|
5.942
|
NM_001102563
|
Prrt2
|
proline-rich transmembrane protein 2
|
|
chr5_-_77543903
|
5.906
|
|
Hopx
|
HOP homeobox
|
|
chr14_-_70923865
|
5.893
|
NM_011359
|
Sftpc
|
surfactant associated protein C
|
|
chr3_+_67696141
|
5.863
|
NM_001113419
NM_177585
|
Iqcj-schip1
Iqcj
|
IQ motif containing J-schwannomin interacting protein 1 read-through transcript
IQ motif containing J
|
|
chr2_+_28048511
|
5.855
|
|
Olfm1
|
olfactomedin 1
|
|
chr9_-_4796141
|
5.845
|
NM_001113180
NM_001113181
NM_019691
|
Gria4
|
glutamate receptor, ionotropic, AMPA4 (alpha 4)
|
|
chr12_+_50484907
|
5.832
|
|
Foxg1
|
forkhead box G1
|
|
chr2_-_57905140
|
5.831
|
NM_029972
|
Ermn
|
ermin, ERM-like protein
|
|
chr16_-_42340613
|
5.824
|
NM_008083
|
Gap43
|
growth associated protein 43
|
|
chr2_-_110790259
|
5.796
|
NM_001128103
NM_001081556
|
Ano3
|
anoctamin 3
|
|
chr12_+_30687597
|
5.767
|
|
Pxdn
|
peroxidasin homolog (Drosophila)
|
|
chr11_-_11798101
|
5.764
|
|
Ddc
|
dopa decarboxylase
|
|
chr14_-_109313447
|
5.746
|
NM_199065
|
Slitrk1
|
SLIT and NTRK-like family, member 1
|
|
chr15_-_90764873
|
5.745
|
|
Kif21a
|
kinesin family member 21A
|
|
chr5_-_77544147
|
5.731
|
NM_175606
|
Hopx
|
HOP homeobox
|
|
chr3_+_17954500
|
5.726
|
|
Bhlhe22
|
basic helix-loop-helix family, member e22
|
|
chr16_-_85030543
|
5.671
|
|
App
|
amyloid beta (A4) precursor protein
|
|
chr11_-_11798138
|
5.637
|
NM_001190448
|
Ddc
|
dopa decarboxylase
|
|
chr10_-_30375572
|
5.588
|
NM_001111267
|
Ncoa7
|
nuclear receptor coactivator 7
|
|
chr6_-_124414668
|
5.565
|
NM_153508
|
Clstn3
|
calsyntenin 3
|
|
chr12_+_37968225
|
5.505
|
NM_178767
|
Tmem195
|
transmembrane protein 195
|
|
chr10_+_96942181
|
5.477
|
|
Dcn
|
decorin
|
|
chr18_+_37644674
|
5.454
|
NM_053142
|
Pcdhb17
|
protocadherin beta 17
|
|
chr2_+_179777363
|
5.412
|
|
Ss18l1
|
synovial sarcoma translocation gene on chromosome 18-like 1
|
|
chr19_-_44472087
|
5.396
|
|
Scd1
|
stearoyl-Coenzyme A desaturase 1
|