Project
GNF SymAtlas + NCI-60 cancer cell lines, human (Su, 2004; Ross, 2000)
Navigation
Downloads
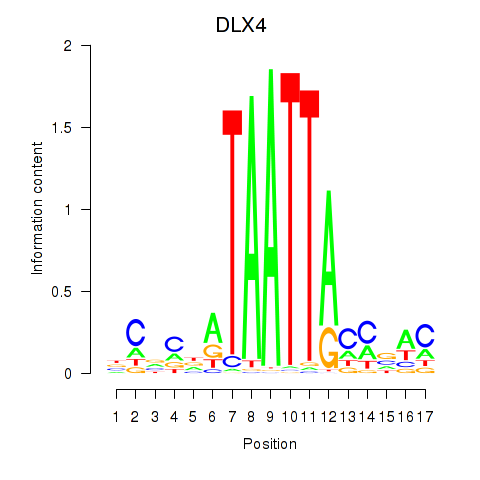
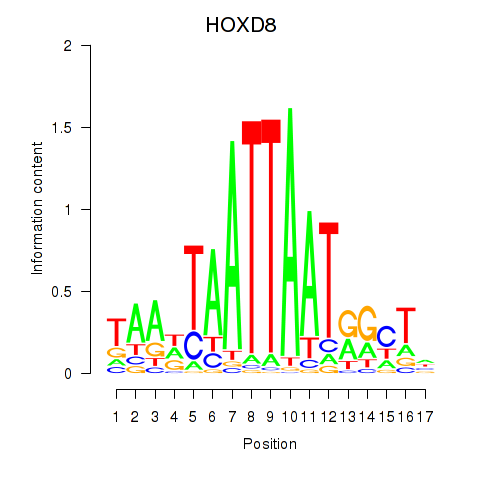
Results for DLX4_HOXD8
Z-value: 1.08
Transcription factors associated with DLX4_HOXD8
| Gene Symbol | Gene ID | Gene Info |
|---|---|---|
|
DLX4
|
ENSG00000108813.9 | distal-less homeobox 4 |
|
HOXD8
|
ENSG00000175879.7 | homeobox D8 |
Activity-expression correlation:
| Gene | Promoter | Pearson corr. coef. | P-value | Plot |
|---|---|---|---|---|
| DLX4 | hg19_v2_chr17_+_48046538_48046575 | 0.40 | 6.9e-10 | Click! |
Activity profile of DLX4_HOXD8 motif
Sorted Z-values of DLX4_HOXD8 motif
| Promoter | Log-likelihood | Transcript | Gene | Gene Info |
|---|---|---|---|---|
| chrX_+_13671225 | 17.76 |
ENST00000545566.1
ENST00000544987.1 ENST00000314720.4 |
TCEANC
|
transcription elongation factor A (SII) N-terminal and central domain containing |
| chr10_+_7745303 | 17.73 |
ENST00000429820.1
ENST00000379587.4 |
ITIH2
|
inter-alpha-trypsin inhibitor heavy chain 2 |
| chr3_-_138763734 | 16.49 |
ENST00000413199.1
ENST00000502927.2 |
PRR23C
|
proline rich 23C |
| chr6_-_52705641 | 14.91 |
ENST00000370989.2
|
GSTA5
|
glutathione S-transferase alpha 5 |
| chr1_-_158656488 | 13.10 |
ENST00000368147.4
|
SPTA1
|
spectrin, alpha, erythrocytic 1 (elliptocytosis 2) |
| chr1_+_233086326 | 12.99 |
ENST00000366628.5
ENST00000366627.4 |
NTPCR
|
nucleoside-triphosphatase, cancer-related |
| chr13_-_88323218 | 12.90 |
ENST00000436290.2
ENST00000453832.2 ENST00000606590.1 |
MIR4500HG
|
MIR4500 host gene (non-protein coding) |
| chr2_-_86333244 | 12.73 |
ENST00000263857.6
ENST00000409681.1 |
POLR1A
|
polymerase (RNA) I polypeptide A, 194kDa |
| chr11_-_5255861 | 12.52 |
ENST00000380299.3
|
HBD
|
hemoglobin, delta |
| chr2_-_89399845 | 11.80 |
ENST00000479981.1
|
IGKV1-16
|
immunoglobulin kappa variable 1-16 |
| chr18_+_71815743 | 10.92 |
ENST00000169551.6
ENST00000580087.1 |
TIMM21
|
translocase of inner mitochondrial membrane 21 homolog (yeast) |
| chr1_+_158801095 | 10.84 |
ENST00000368141.4
|
MNDA
|
myeloid cell nuclear differentiation antigen |
| chr16_+_58283814 | 10.48 |
ENST00000443128.2
ENST00000219299.4 |
CCDC113
|
coiled-coil domain containing 113 |
| chr11_-_59950486 | 10.11 |
ENST00000426738.2
ENST00000533023.1 ENST00000420732.2 |
MS4A6A
|
membrane-spanning 4-domains, subfamily A, member 6A |
| chr5_-_135290705 | 10.04 |
ENST00000274507.1
|
LECT2
|
leukocyte cell-derived chemotaxin 2 |
| chr2_-_89340242 | 9.97 |
ENST00000480492.1
|
IGKV1-12
|
immunoglobulin kappa variable 1-12 |
| chr10_+_7745232 | 9.97 |
ENST00000358415.4
|
ITIH2
|
inter-alpha-trypsin inhibitor heavy chain 2 |
| chr7_-_150020578 | 9.79 |
ENST00000478393.1
|
ACTR3C
|
ARP3 actin-related protein 3 homolog C (yeast) |
| chr17_+_56315936 | 9.72 |
ENST00000543544.1
|
LPO
|
lactoperoxidase |
| chr2_+_90211643 | 9.63 |
ENST00000390277.2
|
IGKV3D-11
|
immunoglobulin kappa variable 3D-11 |
| chr7_+_20686946 | 9.61 |
ENST00000443026.2
ENST00000406935.1 |
ABCB5
|
ATP-binding cassette, sub-family B (MDR/TAP), member 5 |
| chr9_-_21351377 | 9.26 |
ENST00000380210.1
|
IFNA6
|
interferon, alpha 6 |
| chr11_-_59950519 | 9.19 |
ENST00000528851.1
|
MS4A6A
|
membrane-spanning 4-domains, subfamily A, member 6A |
| chr4_+_74275057 | 9.17 |
ENST00000511370.1
|
ALB
|
albumin |
| chr11_-_59950622 | 9.12 |
ENST00000323961.3
ENST00000412309.2 |
MS4A6A
|
membrane-spanning 4-domains, subfamily A, member 6A |
| chr18_+_21693306 | 9.10 |
ENST00000540918.2
|
TTC39C
|
tetratricopeptide repeat domain 39C |
| chr11_-_85376121 | 8.96 |
ENST00000527447.1
|
CREBZF
|
CREB/ATF bZIP transcription factor |
| chr8_-_86253888 | 8.22 |
ENST00000522389.1
ENST00000432364.2 ENST00000517618.1 |
CA1
|
carbonic anhydrase I |
| chr19_+_42817527 | 8.17 |
ENST00000598766.1
|
TMEM145
|
transmembrane protein 145 |
| chr11_-_128894053 | 7.95 |
ENST00000392657.3
|
ARHGAP32
|
Rho GTPase activating protein 32 |
| chr4_-_74853897 | 7.87 |
ENST00000296028.3
|
PPBP
|
pro-platelet basic protein (chemokine (C-X-C motif) ligand 7) |
| chr12_-_91573132 | 7.87 |
ENST00000550563.1
ENST00000546370.1 |
DCN
|
decorin |
| chr10_-_71169031 | 7.81 |
ENST00000373307.1
|
TACR2
|
tachykinin receptor 2 |
| chr6_-_49712123 | 7.65 |
ENST00000263045.4
|
CRISP3
|
cysteine-rich secretory protein 3 |
| chr2_+_90248739 | 7.53 |
ENST00000468879.1
|
IGKV1D-43
|
immunoglobulin kappa variable 1D-43 |
| chr8_-_86290333 | 7.40 |
ENST00000521846.1
ENST00000523022.1 ENST00000524324.1 ENST00000519991.1 ENST00000520663.1 ENST00000517590.1 ENST00000522579.1 ENST00000522814.1 ENST00000522662.1 ENST00000523858.1 ENST00000519129.1 |
CA1
|
carbonic anhydrase I |
| chr3_+_10206545 | 7.39 |
ENST00000256458.4
|
IRAK2
|
interleukin-1 receptor-associated kinase 2 |
| chr17_+_56315787 | 7.34 |
ENST00000262290.4
ENST00000421678.2 |
LPO
|
lactoperoxidase |
| chr2_-_89292422 | 7.29 |
ENST00000495489.1
|
IGKV1-8
|
immunoglobulin kappa variable 1-8 |
| chr12_-_10282836 | 7.23 |
ENST00000304084.8
ENST00000353231.5 ENST00000525605.1 |
CLEC7A
|
C-type lectin domain family 7, member A |
| chr1_+_28199047 | 7.17 |
ENST00000373925.1
ENST00000328928.7 ENST00000373927.3 ENST00000427466.1 ENST00000442118.1 ENST00000373921.3 |
THEMIS2
|
thymocyte selection associated family member 2 |
| chr9_-_98079965 | 7.11 |
ENST00000289081.3
|
FANCC
|
Fanconi anemia, complementation group C |
| chr2_+_211342432 | 7.11 |
ENST00000430249.2
|
CPS1
|
carbamoyl-phosphate synthase 1, mitochondrial |
| chr6_-_32908765 | 7.10 |
ENST00000416244.2
|
HLA-DMB
|
major histocompatibility complex, class II, DM beta |
| chr2_+_90273679 | 7.05 |
ENST00000423080.2
|
IGKV3D-7
|
immunoglobulin kappa variable 3D-7 |
| chr4_+_69681710 | 7.00 |
ENST00000265403.7
ENST00000458688.2 |
UGT2B10
|
UDP glucuronosyltransferase 2 family, polypeptide B10 |
| chr3_+_35722487 | 6.91 |
ENST00000441454.1
|
ARPP21
|
cAMP-regulated phosphoprotein, 21kDa |
| chr1_-_183559693 | 6.81 |
ENST00000367535.3
ENST00000413720.1 ENST00000418089.1 |
NCF2
|
neutrophil cytosolic factor 2 |
| chr1_-_183560011 | 6.80 |
ENST00000367536.1
|
NCF2
|
neutrophil cytosolic factor 2 |
| chr17_-_64225508 | 6.79 |
ENST00000205948.6
|
APOH
|
apolipoprotein H (beta-2-glycoprotein I) |
| chr6_-_49712147 | 6.74 |
ENST00000433368.2
ENST00000354620.4 |
CRISP3
|
cysteine-rich secretory protein 3 |
| chr2_+_90139056 | 6.72 |
ENST00000492446.1
|
IGKV1D-16
|
immunoglobulin kappa variable 1D-16 |
| chr6_-_52710893 | 6.70 |
ENST00000284562.2
|
GSTA5
|
glutathione S-transferase alpha 5 |
| chr4_+_56815102 | 6.69 |
ENST00000257287.4
|
CEP135
|
centrosomal protein 135kDa |
| chr3_+_149192475 | 6.64 |
ENST00000465758.1
|
TM4SF4
|
transmembrane 4 L six family member 4 |
| chr6_+_26204825 | 6.55 |
ENST00000360441.4
|
HIST1H4E
|
histone cluster 1, H4e |
| chr21_-_15918618 | 6.52 |
ENST00000400564.1
ENST00000400566.1 |
SAMSN1
|
SAM domain, SH3 domain and nuclear localization signals 1 |
| chr7_+_156433573 | 6.46 |
ENST00000311822.8
|
RNF32
|
ring finger protein 32 |
| chr9_-_100684845 | 6.44 |
ENST00000375119.3
|
C9orf156
|
chromosome 9 open reading frame 156 |
| chr1_-_207226313 | 6.40 |
ENST00000367084.1
|
YOD1
|
YOD1 deubiquitinase |
| chr11_+_5710919 | 6.34 |
ENST00000379965.3
ENST00000425490.1 |
TRIM22
|
tripartite motif containing 22 |
| chr17_-_64216748 | 6.34 |
ENST00000585162.1
|
APOH
|
apolipoprotein H (beta-2-glycoprotein I) |
| chr19_+_42212501 | 6.33 |
ENST00000398599.4
|
CEACAM5
|
carcinoembryonic antigen-related cell adhesion molecule 5 |
| chr12_-_112123524 | 6.26 |
ENST00000327551.6
|
BRAP
|
BRCA1 associated protein |
| chr4_-_46911248 | 6.17 |
ENST00000355591.3
ENST00000505102.1 |
COX7B2
|
cytochrome c oxidase subunit VIIb2 |
| chr14_-_106926724 | 6.15 |
ENST00000434710.1
|
IGHV3-43
|
immunoglobulin heavy variable 3-43 |
| chr11_+_92085262 | 6.13 |
ENST00000298047.6
ENST00000409404.2 ENST00000541502.1 |
FAT3
|
FAT atypical cadherin 3 |
| chr2_+_25015968 | 6.10 |
ENST00000380834.2
ENST00000473706.1 |
CENPO
|
centromere protein O |
| chr4_+_155484155 | 6.05 |
ENST00000509493.1
|
FGB
|
fibrinogen beta chain |
| chr15_-_22448819 | 6.04 |
ENST00000604066.1
|
IGHV1OR15-1
|
immunoglobulin heavy variable 1/OR15-1 (non-functional) |
| chr4_+_69962212 | 6.02 |
ENST00000508661.1
|
UGT2B7
|
UDP glucuronosyltransferase 2 family, polypeptide B7 |
| chr4_-_46911223 | 6.00 |
ENST00000396533.1
|
COX7B2
|
cytochrome c oxidase subunit VIIb2 |
| chr12_-_11548496 | 5.99 |
ENST00000389362.4
ENST00000565533.1 ENST00000546254.1 |
PRB2
PRB1
|
proline-rich protein BstNI subfamily 2 proline-rich protein BstNI subfamily 1 |
| chr19_-_12721616 | 5.97 |
ENST00000311437.6
|
ZNF490
|
zinc finger protein 490 |
| chr3_-_39321512 | 5.94 |
ENST00000399220.2
|
CX3CR1
|
chemokine (C-X3-C motif) receptor 1 |
| chr1_+_196743912 | 5.94 |
ENST00000367425.4
|
CFHR3
|
complement factor H-related 3 |
| chr2_+_90077680 | 5.91 |
ENST00000390270.2
|
IGKV3D-20
|
immunoglobulin kappa variable 3D-20 |
| chr4_+_69962185 | 5.85 |
ENST00000305231.7
|
UGT2B7
|
UDP glucuronosyltransferase 2 family, polypeptide B7 |
| chr2_-_89327228 | 5.79 |
ENST00000483158.1
|
IGKV3-11
|
immunoglobulin kappa variable 3-11 |
| chr6_+_161123270 | 5.76 |
ENST00000366924.2
ENST00000308192.9 ENST00000418964.1 |
PLG
|
plasminogen |
| chr2_-_89513402 | 5.70 |
ENST00000498435.1
|
IGKV1-27
|
immunoglobulin kappa variable 1-27 |
| chr6_-_32908792 | 5.69 |
ENST00000418107.2
|
HLA-DMB
|
major histocompatibility complex, class II, DM beta |
| chr4_-_100356291 | 5.60 |
ENST00000476959.1
ENST00000482593.1 |
ADH7
|
alcohol dehydrogenase 7 (class IV), mu or sigma polypeptide |
| chr12_-_11422739 | 5.53 |
ENST00000279573.7
|
PRB3
|
proline-rich protein BstNI subfamily 3 |
| chr8_+_94241867 | 5.46 |
ENST00000598428.1
|
AC016885.1
|
Uncharacterized protein |
| chr9_-_97402413 | 5.46 |
ENST00000414122.1
|
FBP1
|
fructose-1,6-bisphosphatase 1 |
| chr12_+_8995832 | 5.44 |
ENST00000541459.1
|
A2ML1
|
alpha-2-macroglobulin-like 1 |
| chrX_+_1710484 | 5.41 |
ENST00000313871.3
ENST00000381261.3 |
AKAP17A
|
A kinase (PRKA) anchor protein 17A |
| chr15_+_99791567 | 5.36 |
ENST00000558879.1
ENST00000301981.3 ENST00000422500.2 ENST00000447360.2 ENST00000442993.2 |
LRRC28
|
leucine rich repeat containing 28 |
| chr2_-_158345462 | 5.36 |
ENST00000439355.1
ENST00000540637.1 |
CYTIP
|
cytohesin 1 interacting protein |
| chr19_+_51728316 | 5.32 |
ENST00000436584.2
ENST00000421133.2 ENST00000391796.3 ENST00000262262.4 |
CD33
|
CD33 molecule |
| chr16_+_72090053 | 5.30 |
ENST00000576168.2
ENST00000567185.3 ENST00000567612.2 |
HP
|
haptoglobin |
| chr4_-_39033963 | 5.30 |
ENST00000381938.3
|
TMEM156
|
transmembrane protein 156 |
| chr2_+_90198535 | 5.29 |
ENST00000390276.2
|
IGKV1D-12
|
immunoglobulin kappa variable 1D-12 |
| chr12_-_7656357 | 5.29 |
ENST00000396620.3
ENST00000432237.2 ENST00000359156.4 |
CD163
|
CD163 molecule |
| chr16_-_25122785 | 5.28 |
ENST00000563962.1
ENST00000569920.1 |
RP11-449H11.1
|
RP11-449H11.1 |
| chr7_-_76829125 | 5.27 |
ENST00000248598.5
|
FGL2
|
fibrinogen-like 2 |
| chr1_+_196912902 | 5.22 |
ENST00000476712.2
ENST00000367415.5 |
CFHR2
|
complement factor H-related 2 |
| chr3_+_111717600 | 5.22 |
ENST00000273368.4
|
TAGLN3
|
transgelin 3 |
| chr11_-_59633951 | 5.19 |
ENST00000257264.3
|
TCN1
|
transcobalamin I (vitamin B12 binding protein, R binder family) |
| chr9_-_97402531 | 5.14 |
ENST00000415431.1
|
FBP1
|
fructose-1,6-bisphosphatase 1 |
| chr5_-_131892501 | 5.13 |
ENST00000450655.1
|
IL5
|
interleukin 5 (colony-stimulating factor, eosinophil) |
| chr14_-_71001708 | 5.13 |
ENST00000256389.3
|
ADAM20
|
ADAM metallopeptidase domain 20 |
| chr17_+_3118915 | 5.12 |
ENST00000304094.1
|
OR1A1
|
olfactory receptor, family 1, subfamily A, member 1 |
| chr4_-_70080449 | 5.08 |
ENST00000446444.1
|
UGT2B11
|
UDP glucuronosyltransferase 2 family, polypeptide B11 |
| chr2_-_89310012 | 5.08 |
ENST00000493819.1
|
IGKV1-9
|
immunoglobulin kappa variable 1-9 |
| chr17_-_73663245 | 5.07 |
ENST00000584999.1
ENST00000317905.5 ENST00000420326.2 ENST00000340830.5 |
RECQL5
|
RecQ protein-like 5 |
| chr7_-_115670792 | 5.07 |
ENST00000265440.7
ENST00000393485.1 |
TFEC
|
transcription factor EC |
| chr16_-_28634874 | 5.06 |
ENST00000395609.1
ENST00000350842.4 |
SULT1A1
|
sulfotransferase family, cytosolic, 1A, phenol-preferring, member 1 |
| chr14_+_21423611 | 5.03 |
ENST00000304625.2
|
RNASE2
|
ribonuclease, RNase A family, 2 (liver, eosinophil-derived neurotoxin) |
| chr19_+_15852203 | 5.03 |
ENST00000305892.1
|
OR10H3
|
olfactory receptor, family 10, subfamily H, member 3 |
| chr6_-_52668605 | 5.01 |
ENST00000334575.5
|
GSTA1
|
glutathione S-transferase alpha 1 |
| chr17_+_67498538 | 4.96 |
ENST00000589647.1
|
MAP2K6
|
mitogen-activated protein kinase kinase 6 |
| chr1_+_117963209 | 4.95 |
ENST00000449370.2
|
MAN1A2
|
mannosidase, alpha, class 1A, member 2 |
| chr2_+_90121477 | 4.94 |
ENST00000483379.1
|
IGKV1D-17
|
immunoglobulin kappa variable 1D-17 |
| chr16_-_66584059 | 4.94 |
ENST00000417693.3
ENST00000544898.1 ENST00000569718.1 ENST00000527284.1 ENST00000299697.7 ENST00000451102.2 |
TK2
|
thymidine kinase 2, mitochondrial |
| chr10_-_99052382 | 4.90 |
ENST00000453547.2
ENST00000316676.8 ENST00000358308.3 ENST00000466484.1 ENST00000358531.4 |
ARHGAP19-SLIT1
ARHGAP19
|
ARHGAP19-SLIT1 readthrough (NMD candidate) Rho GTPase activating protein 19 |
| chr11_-_89541743 | 4.87 |
ENST00000329758.1
|
TRIM49
|
tripartite motif containing 49 |
| chr19_+_45417921 | 4.85 |
ENST00000252491.4
ENST00000592885.1 ENST00000589781.1 |
APOC1
|
apolipoprotein C-I |
| chr12_+_113354341 | 4.83 |
ENST00000553152.1
|
OAS1
|
2'-5'-oligoadenylate synthetase 1, 40/46kDa |
| chr2_-_89619904 | 4.76 |
ENST00000498574.1
|
IGKV1-39
|
immunoglobulin kappa variable 1-39 (gene/pseudogene) |
| chr9_-_116837249 | 4.75 |
ENST00000466610.2
|
AMBP
|
alpha-1-microglobulin/bikunin precursor |
| chr12_-_114841703 | 4.70 |
ENST00000526441.1
|
TBX5
|
T-box 5 |
| chr7_-_115670804 | 4.70 |
ENST00000320239.7
|
TFEC
|
transcription factor EC |
| chr7_+_6522922 | 4.58 |
ENST00000601673.1
|
FLJ20306
|
CDNA FLJ20306 fis, clone HEP06881; Putative uncharacterized protein FLJ20306; Uncharacterized protein |
| chr16_-_71610985 | 4.57 |
ENST00000355962.4
|
TAT
|
tyrosine aminotransferase |
| chr16_+_33020496 | 4.56 |
ENST00000565407.2
|
IGHV3OR16-8
|
immunoglobulin heavy variable 3/OR16-8 (non-functional) |
| chr1_-_33430286 | 4.55 |
ENST00000373456.7
ENST00000356990.5 ENST00000235150.4 |
RNF19B
|
ring finger protein 19B |
| chr4_-_8873531 | 4.42 |
ENST00000400677.3
|
HMX1
|
H6 family homeobox 1 |
| chr17_+_1674982 | 4.42 |
ENST00000572048.1
ENST00000573763.1 |
SERPINF1
|
serpin peptidase inhibitor, clade F (alpha-2 antiplasmin, pigment epithelium derived factor), member 1 |
| chr2_+_166095898 | 4.42 |
ENST00000424833.1
ENST00000375437.2 ENST00000357398.3 |
SCN2A
|
sodium channel, voltage-gated, type II, alpha subunit |
| chr1_-_13390765 | 4.38 |
ENST00000357367.2
|
PRAMEF8
|
PRAME family member 8 |
| chrX_-_6453159 | 4.36 |
ENST00000381089.3
ENST00000398729.1 |
VCX3A
|
variable charge, X-linked 3A |
| chr11_+_89764274 | 4.35 |
ENST00000448984.1
ENST00000432771.1 |
TRIM49C
|
tripartite motif containing 49C |
| chr11_+_3011093 | 4.35 |
ENST00000332881.2
|
AC131971.1
|
HCG1782999; PRO0943; Uncharacterized protein |
| chr4_-_120243545 | 4.35 |
ENST00000274024.3
|
FABP2
|
fatty acid binding protein 2, intestinal |
| chr12_+_9102632 | 4.35 |
ENST00000539240.1
|
KLRG1
|
killer cell lectin-like receptor subfamily G, member 1 |
| chr6_+_32407619 | 4.33 |
ENST00000395388.2
ENST00000374982.5 |
HLA-DRA
|
major histocompatibility complex, class II, DR alpha |
| chr2_-_89417335 | 4.30 |
ENST00000490686.1
|
IGKV1-17
|
immunoglobulin kappa variable 1-17 |
| chrY_-_6740649 | 4.26 |
ENST00000383036.1
ENST00000383037.4 |
AMELY
|
amelogenin, Y-linked |
| chr22_+_23054174 | 4.26 |
ENST00000390308.2
|
IGLV3-21
|
immunoglobulin lambda variable 3-21 |
| chr16_+_10479906 | 4.24 |
ENST00000562527.1
ENST00000396560.2 ENST00000396559.1 ENST00000562102.1 ENST00000543967.1 ENST00000569939.1 ENST00000569900.1 |
ATF7IP2
|
activating transcription factor 7 interacting protein 2 |
| chr3_+_98482175 | 4.24 |
ENST00000485391.1
ENST00000492254.1 |
ST3GAL6
|
ST3 beta-galactoside alpha-2,3-sialyltransferase 6 |
| chr22_+_23161491 | 4.23 |
ENST00000390316.2
|
IGLV3-9
|
immunoglobulin lambda variable 3-9 (gene/pseudogene) |
| chr2_-_100939195 | 4.23 |
ENST00000393437.3
|
LONRF2
|
LON peptidase N-terminal domain and ring finger 2 |
| chr1_+_197170592 | 4.20 |
ENST00000535699.1
|
CRB1
|
crumbs homolog 1 (Drosophila) |
| chr5_+_140213815 | 4.20 |
ENST00000525929.1
ENST00000378125.3 |
PCDHA7
|
protocadherin alpha 7 |
| chr14_-_25078864 | 4.19 |
ENST00000216338.4
ENST00000557220.2 ENST00000382548.4 |
GZMH
|
granzyme H (cathepsin G-like 2, protein h-CCPX) |
| chr19_-_22034809 | 4.18 |
ENST00000594012.1
ENST00000595461.1 ENST00000596899.1 |
ZNF43
|
zinc finger protein 43 |
| chr1_-_100231349 | 4.18 |
ENST00000287474.5
ENST00000414213.1 |
FRRS1
|
ferric-chelate reductase 1 |
| chr13_+_49551020 | 4.18 |
ENST00000541916.1
|
FNDC3A
|
fibronectin type III domain containing 3A |
| chr2_+_113816215 | 4.16 |
ENST00000346807.3
|
IL36RN
|
interleukin 36 receptor antagonist |
| chr3_-_194072019 | 4.15 |
ENST00000429275.1
ENST00000323830.3 |
CPN2
|
carboxypeptidase N, polypeptide 2 |
| chr2_+_234580499 | 4.14 |
ENST00000354728.4
|
UGT1A9
|
UDP glucuronosyltransferase 1 family, polypeptide A9 |
| chr12_-_11422630 | 4.13 |
ENST00000381842.3
ENST00000538488.1 |
PRB3
|
proline-rich protein BstNI subfamily 3 |
| chr11_-_117747434 | 4.11 |
ENST00000529335.2
ENST00000530956.1 ENST00000260282.4 |
FXYD6
|
FXYD domain containing ion transport regulator 6 |
| chr2_-_89459813 | 4.10 |
ENST00000390256.2
|
IGKV6-21
|
immunoglobulin kappa variable 6-21 (non-functional) |
| chr4_-_165898768 | 4.09 |
ENST00000329314.5
|
TRIM61
|
tripartite motif containing 61 |
| chr10_+_53806501 | 4.09 |
ENST00000373975.2
|
PRKG1
|
protein kinase, cGMP-dependent, type I |
| chr12_+_9822331 | 4.07 |
ENST00000545918.1
ENST00000543300.1 ENST00000261339.6 ENST00000466035.2 |
CLEC2D
|
C-type lectin domain family 2, member D |
| chr1_+_196788887 | 4.06 |
ENST00000320493.5
ENST00000367424.4 ENST00000367421.3 |
CFHR1
CFHR2
|
complement factor H-related 1 complement factor H-related 2 |
| chr4_-_76957214 | 4.04 |
ENST00000306621.3
|
CXCL11
|
chemokine (C-X-C motif) ligand 11 |
| chr2_+_234580525 | 4.04 |
ENST00000609637.1
|
UGT1A1
|
UDP glucuronosyltransferase 1 family, polypeptide A8 |
| chr12_-_10282742 | 4.04 |
ENST00000298523.5
ENST00000396484.2 ENST00000310002.4 |
CLEC7A
|
C-type lectin domain family 7, member A |
| chr22_+_23040274 | 4.03 |
ENST00000390306.2
|
IGLV2-23
|
immunoglobulin lambda variable 2-23 |
| chr1_+_153330322 | 3.99 |
ENST00000368738.3
|
S100A9
|
S100 calcium binding protein A9 |
| chr19_+_50016610 | 3.98 |
ENST00000596975.1
|
FCGRT
|
Fc fragment of IgG, receptor, transporter, alpha |
| chr9_-_116102530 | 3.98 |
ENST00000374195.3
ENST00000341761.4 |
WDR31
|
WD repeat domain 31 |
| chr3_+_46412345 | 3.94 |
ENST00000292303.4
|
CCR5
|
chemokine (C-C motif) receptor 5 (gene/pseudogene) |
| chr12_-_123201337 | 3.94 |
ENST00000528880.2
|
HCAR3
|
hydroxycarboxylic acid receptor 3 |
| chr5_-_140013275 | 3.93 |
ENST00000512545.1
ENST00000302014.6 ENST00000401743.2 |
CD14
|
CD14 molecule |
| chr15_+_75639296 | 3.93 |
ENST00000564500.1
ENST00000355059.4 ENST00000566752.1 |
NEIL1
|
nei endonuclease VIII-like 1 (E. coli) |
| chr5_+_156696362 | 3.92 |
ENST00000377576.3
|
CYFIP2
|
cytoplasmic FMR1 interacting protein 2 |
| chr11_-_117748138 | 3.92 |
ENST00000527717.1
|
FXYD6
|
FXYD domain containing ion transport regulator 6 |
| chr2_+_90108504 | 3.91 |
ENST00000390271.2
|
IGKV6D-41
|
immunoglobulin kappa variable 6D-41 (non-functional) |
| chr14_-_106552755 | 3.90 |
ENST00000390600.2
|
IGHV3-9
|
immunoglobulin heavy variable 3-9 |
| chr3_+_38537960 | 3.89 |
ENST00000453767.1
|
EXOG
|
endo/exonuclease (5'-3'), endonuclease G-like |
| chr5_-_20575959 | 3.88 |
ENST00000507958.1
|
CDH18
|
cadherin 18, type 2 |
| chr11_+_49050504 | 3.88 |
ENST00000332682.7
|
TRIM49B
|
tripartite motif containing 49B |
| chr6_+_72926145 | 3.88 |
ENST00000425662.2
ENST00000453976.2 |
RIMS1
|
regulating synaptic membrane exocytosis 1 |
| chr16_-_11375179 | 3.88 |
ENST00000312511.3
|
PRM1
|
protamine 1 |
| chr6_+_131894284 | 3.88 |
ENST00000368087.3
ENST00000356962.2 |
ARG1
|
arginase 1 |
| chr12_+_21284118 | 3.87 |
ENST00000256958.2
|
SLCO1B1
|
solute carrier organic anion transporter family, member 1B1 |
| chr14_-_107049312 | 3.86 |
ENST00000390627.2
|
IGHV3-53
|
immunoglobulin heavy variable 3-53 |
| chr14_-_106994333 | 3.85 |
ENST00000390624.2
|
IGHV3-48
|
immunoglobulin heavy variable 3-48 |
| chr12_-_102224457 | 3.85 |
ENST00000549165.1
ENST00000549940.1 ENST00000392919.4 |
GNPTAB
|
N-acetylglucosamine-1-phosphate transferase, alpha and beta subunits |
| chr10_+_91152303 | 3.84 |
ENST00000371804.3
|
IFIT1
|
interferon-induced protein with tetratricopeptide repeats 1 |
| chr8_+_27631903 | 3.84 |
ENST00000305188.8
|
ESCO2
|
establishment of sister chromatid cohesion N-acetyltransferase 2 |
| chrX_+_8432871 | 3.84 |
ENST00000381032.1
ENST00000453306.1 ENST00000444481.1 |
VCX3B
|
variable charge, X-linked 3B |
| chr14_+_22337014 | 3.82 |
ENST00000390436.2
|
TRAV13-1
|
T cell receptor alpha variable 13-1 |
| chr19_-_58326267 | 3.81 |
ENST00000391701.1
|
ZNF552
|
zinc finger protein 552 |
| chr1_-_45140074 | 3.79 |
ENST00000420706.1
ENST00000372235.3 ENST00000372242.3 ENST00000372243.3 ENST00000372244.3 |
TMEM53
|
transmembrane protein 53 |
| chr20_+_3801162 | 3.75 |
ENST00000379573.2
ENST00000379567.2 ENST00000455742.1 ENST00000246041.2 |
AP5S1
|
adaptor-related protein complex 5, sigma 1 subunit |
| chr7_+_80275953 | 3.74 |
ENST00000538969.1
ENST00000544133.1 ENST00000433696.2 |
CD36
|
CD36 molecule (thrombospondin receptor) |
| chr6_-_133055815 | 3.74 |
ENST00000509351.1
ENST00000417437.2 ENST00000414302.2 ENST00000423615.2 ENST00000427187.2 ENST00000275223.3 ENST00000519686.2 |
VNN3
|
vanin 3 |
| chr12_+_81110684 | 3.73 |
ENST00000228644.3
|
MYF5
|
myogenic factor 5 |
| chr1_+_160709029 | 3.73 |
ENST00000444090.2
ENST00000441662.2 |
SLAMF7
|
SLAM family member 7 |
| chr6_-_135271219 | 3.72 |
ENST00000367847.2
ENST00000367845.2 |
ALDH8A1
|
aldehyde dehydrogenase 8 family, member A1 |
| chr1_-_182360498 | 3.68 |
ENST00000417584.2
|
GLUL
|
glutamate-ammonia ligase |
| chr20_-_35580240 | 3.66 |
ENST00000262878.4
|
SAMHD1
|
SAM domain and HD domain 1 |
Network of associatons between targets according to the STRING database.
First level regulatory network of DLX4_HOXD8
Gene Ontology Analysis
Gene overrepresentation in biological_process category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 4.3 | 17.1 | GO:0002503 | peptide antigen assembly with MHC class II protein complex(GO:0002503) |
| 3.8 | 3.8 | GO:0071168 | protein localization to chromatin(GO:0071168) |
| 3.1 | 9.2 | GO:0019836 | hemolysis by symbiont of host erythrocytes(GO:0019836) hemolysis in other organism(GO:0044179) hemolysis in other organism involved in symbiotic interaction(GO:0052331) |
| 2.8 | 2.8 | GO:0038123 | toll-like receptor TLR1:TLR2 signaling pathway(GO:0038123) detection of bacterial lipoprotein(GO:0042494) detection of triacyl bacterial lipopeptide(GO:0042495) detection of bacterial lipopeptide(GO:0070340) response to triacyl bacterial lipopeptide(GO:0071725) cellular response to triacyl bacterial lipopeptide(GO:0071727) |
| 2.8 | 11.1 | GO:1904749 | regulation of protein localization to nucleolus(GO:1904749) |
| 2.6 | 10.6 | GO:0005986 | sucrose biosynthetic process(GO:0005986) |
| 2.6 | 7.9 | GO:0002881 | negative regulation of chronic inflammatory response to non-antigenic stimulus(GO:0002881) |
| 2.6 | 7.8 | GO:0014057 | positive regulation of acetylcholine secretion, neurotransmission(GO:0014057) |
| 2.5 | 7.4 | GO:1990414 | replication-born double-strand break repair via sister chromatid exchange(GO:1990414) |
| 2.4 | 9.6 | GO:0048749 | compound eye development(GO:0048749) |
| 2.4 | 25.9 | GO:0051918 | negative regulation of fibrinolysis(GO:0051918) |
| 2.3 | 7.0 | GO:2000296 | negative regulation of hydrogen peroxide catabolic process(GO:2000296) |
| 2.3 | 9.2 | GO:0019470 | 4-hydroxyproline catabolic process(GO:0019470) |
| 2.3 | 6.8 | GO:0006711 | estrogen catabolic process(GO:0006711) |
| 2.2 | 6.7 | GO:0045643 | regulation of eosinophil differentiation(GO:0045643) positive regulation of eosinophil differentiation(GO:0045645) |
| 2.2 | 6.7 | GO:0010430 | fatty acid omega-oxidation(GO:0010430) |
| 2.1 | 6.4 | GO:1990168 | protein K33-linked deubiquitination(GO:1990168) |
| 1.9 | 5.7 | GO:0002522 | leukocyte migration involved in immune response(GO:0002522) |
| 1.9 | 18.8 | GO:0043152 | induction of bacterial agglutination(GO:0043152) |
| 1.9 | 1.9 | GO:1903487 | regulation of lactation(GO:1903487) |
| 1.8 | 1.8 | GO:0072599 | establishment of protein localization to endoplasmic reticulum(GO:0072599) |
| 1.8 | 3.6 | GO:0002581 | negative regulation of antigen processing and presentation of peptide or polysaccharide antigen via MHC class II(GO:0002581) |
| 1.7 | 15.6 | GO:0038124 | toll-like receptor TLR6:TLR2 signaling pathway(GO:0038124) response to diacyl bacterial lipopeptide(GO:0071724) cellular response to diacyl bacterial lipopeptide(GO:0071726) |
| 1.7 | 5.1 | GO:1902995 | regulation of phospholipid efflux(GO:1902994) positive regulation of phospholipid efflux(GO:1902995) |
| 1.7 | 5.0 | GO:0072709 | cellular response to sorbitol(GO:0072709) |
| 1.6 | 4.7 | GO:0003218 | cardiac left ventricle formation(GO:0003218) |
| 1.6 | 32.7 | GO:0052695 | cellular glucuronidation(GO:0052695) |
| 1.5 | 6.1 | GO:0046104 | thymidine metabolic process(GO:0046104) pyrimidine deoxyribonucleoside metabolic process(GO:0046125) |
| 1.5 | 7.5 | GO:0071279 | cellular response to cobalt ion(GO:0071279) |
| 1.4 | 1.4 | GO:0006145 | purine nucleobase catabolic process(GO:0006145) |
| 1.4 | 4.1 | GO:0071301 | cellular response to vitamin B1(GO:0071301) response to formaldehyde(GO:1904404) |
| 1.3 | 4.0 | GO:0010902 | positive regulation of very-low-density lipoprotein particle remodeling(GO:0010902) |
| 1.3 | 4.0 | GO:1990637 | response to prolactin(GO:1990637) |
| 1.3 | 4.0 | GO:0035606 | peptidyl-cysteine S-trans-nitrosylation(GO:0035606) |
| 1.3 | 7.8 | GO:0035609 | C-terminal protein deglutamylation(GO:0035609) |
| 1.3 | 3.8 | GO:0051097 | negative regulation of helicase activity(GO:0051097) |
| 1.3 | 5.1 | GO:0071461 | cellular response to redox state(GO:0071461) |
| 1.2 | 4.9 | GO:0032049 | cardiolipin biosynthetic process(GO:0032049) |
| 1.2 | 4.9 | GO:0010900 | negative regulation of phosphatidylcholine catabolic process(GO:0010900) |
| 1.2 | 12.0 | GO:0015939 | pantothenate metabolic process(GO:0015939) |
| 1.2 | 7.1 | GO:0034201 | response to oleic acid(GO:0034201) |
| 1.1 | 3.3 | GO:1904479 | cellular response to bile acid(GO:1903413) negative regulation of intestinal absorption(GO:1904479) response to iron ion starvation(GO:1990641) |
| 1.1 | 3.3 | GO:0060474 | positive regulation of sperm motility involved in capacitation(GO:0060474) |
| 1.1 | 1.1 | GO:0002434 | immune complex clearance(GO:0002434) immune complex clearance by monocytes and macrophages(GO:0002436) regulation of immune complex clearance by monocytes and macrophages(GO:0090264) negative regulation of eosinophil activation(GO:1902567) |
| 1.0 | 2.1 | GO:0014055 | acetylcholine secretion, neurotransmission(GO:0014055) regulation of acetylcholine secretion, neurotransmission(GO:0014056) |
| 1.0 | 3.1 | GO:0030451 | regulation of complement activation, alternative pathway(GO:0030451) negative regulation of complement activation, alternative pathway(GO:0045957) |
| 1.0 | 14.4 | GO:0045919 | positive regulation of cytolysis(GO:0045919) |
| 1.0 | 3.1 | GO:0002752 | cell surface pattern recognition receptor signaling pathway(GO:0002752) |
| 1.0 | 146.0 | GO:0006958 | complement activation, classical pathway(GO:0006958) |
| 1.0 | 6.8 | GO:0044800 | fusion of virus membrane with host plasma membrane(GO:0019064) membrane fusion involved in viral entry into host cell(GO:0039663) multi-organism membrane fusion(GO:0044800) |
| 1.0 | 5.8 | GO:1903385 | regulation of homophilic cell adhesion(GO:1903385) |
| 1.0 | 4.8 | GO:1904844 | response to L-glutamine(GO:1904844) cellular response to L-glutamine(GO:1904845) |
| 1.0 | 2.9 | GO:0045897 | positive regulation of transcription during mitosis(GO:0045897) |
| 0.9 | 3.7 | GO:0006542 | glutamine biosynthetic process(GO:0006542) |
| 0.9 | 2.8 | GO:0000294 | nuclear-transcribed mRNA catabolic process, endonucleolytic cleavage-dependent decay(GO:0000294) |
| 0.9 | 3.7 | GO:0006203 | dGTP catabolic process(GO:0006203) |
| 0.9 | 7.3 | GO:0051388 | positive regulation of neurotrophin TRK receptor signaling pathway(GO:0051388) |
| 0.9 | 3.6 | GO:0032053 | ciliary basal body organization(GO:0032053) |
| 0.9 | 2.7 | GO:0035915 | pore formation in membrane of other organism(GO:0035915) |
| 0.9 | 3.5 | GO:0006072 | glycerol-3-phosphate metabolic process(GO:0006072) |
| 0.9 | 6.9 | GO:1900165 | negative regulation of interleukin-6 secretion(GO:1900165) |
| 0.9 | 6.9 | GO:1902525 | regulation of protein monoubiquitination(GO:1902525) |
| 0.8 | 2.5 | GO:0006001 | fructose catabolic process(GO:0006001) fructose catabolic process to hydroxyacetone phosphate and glyceraldehyde-3-phosphate(GO:0061624) |
| 0.8 | 12.5 | GO:0015671 | oxygen transport(GO:0015671) |
| 0.8 | 4.2 | GO:1902714 | negative regulation of interferon-gamma secretion(GO:1902714) |
| 0.8 | 2.5 | GO:0061534 | gamma-aminobutyric acid secretion, neurotransmission(GO:0061534) |
| 0.8 | 9.1 | GO:1900747 | negative regulation of vascular endothelial growth factor signaling pathway(GO:1900747) |
| 0.8 | 4.9 | GO:1904381 | Golgi apparatus mannose trimming(GO:1904381) |
| 0.8 | 6.5 | GO:0009635 | response to herbicide(GO:0009635) |
| 0.8 | 4.0 | GO:0071918 | urea transmembrane transport(GO:0071918) |
| 0.8 | 12.9 | GO:0009812 | flavonoid metabolic process(GO:0009812) |
| 0.8 | 4.8 | GO:0002933 | lipid hydroxylation(GO:0002933) |
| 0.8 | 2.4 | GO:0009107 | lipoate biosynthetic process(GO:0009107) |
| 0.8 | 5.5 | GO:1990253 | cellular response to leucine starvation(GO:1990253) |
| 0.8 | 2.3 | GO:1901877 | regulation of calcium ion binding(GO:1901876) negative regulation of calcium ion binding(GO:1901877) |
| 0.8 | 4.7 | GO:2000158 | positive regulation of ubiquitin-specific protease activity(GO:2000158) |
| 0.8 | 11.3 | GO:0006910 | phagocytosis, recognition(GO:0006910) |
| 0.7 | 2.9 | GO:0006842 | tricarboxylic acid transport(GO:0006842) citrate transport(GO:0015746) |
| 0.7 | 2.9 | GO:0032074 | negative regulation of nuclease activity(GO:0032074) |
| 0.7 | 2.2 | GO:0044782 | cilium organization(GO:0044782) |
| 0.7 | 2.2 | GO:0097359 | UDP-glucosylation(GO:0097359) |
| 0.7 | 2.2 | GO:0070837 | dehydroascorbic acid transport(GO:0070837) |
| 0.7 | 2.8 | GO:0060133 | somatotropin secreting cell development(GO:0060133) |
| 0.7 | 2.1 | GO:0071557 | histone H3-K27 demethylation(GO:0071557) |
| 0.7 | 7.0 | GO:0042905 | 9-cis-retinoic acid biosynthetic process(GO:0042904) 9-cis-retinoic acid metabolic process(GO:0042905) |
| 0.7 | 0.7 | GO:1903939 | regulation of TORC2 signaling(GO:1903939) |
| 0.7 | 3.5 | GO:0060729 | intestinal epithelial structure maintenance(GO:0060729) |
| 0.7 | 2.1 | GO:0010133 | proline catabolic process to glutamate(GO:0010133) |
| 0.7 | 2.7 | GO:0072658 | positive regulation of cell communication by electrical coupling(GO:0010650) maintenance of protein location in membrane(GO:0072658) maintenance of protein location in plasma membrane(GO:0072660) positive regulation of membrane depolarization during cardiac muscle cell action potential(GO:1900827) |
| 0.7 | 10.2 | GO:0002523 | leukocyte migration involved in inflammatory response(GO:0002523) |
| 0.7 | 1.4 | GO:2001183 | negative regulation of interleukin-12 secretion(GO:2001183) |
| 0.7 | 4.0 | GO:0006741 | NADP biosynthetic process(GO:0006741) |
| 0.7 | 2.0 | GO:0072200 | apoptotic process involved in endocardial cushion morphogenesis(GO:0003277) intermediate mesoderm development(GO:0048389) intermediate mesoderm morphogenesis(GO:0048390) intermediate mesoderm formation(GO:0048391) intermediate mesodermal cell differentiation(GO:0048392) regulation of cardiac muscle fiber development(GO:0055018) positive regulation of cardiac muscle fiber development(GO:0055020) bud dilation involved in lung branching(GO:0060503) regulation of transcription from RNA polymerase II promoter involved in kidney development(GO:0060994) BMP signaling pathway involved in ureter morphogenesis(GO:0061149) renal system segmentation(GO:0061150) BMP signaling pathway involved in renal system segmentation(GO:0061151) pulmonary artery endothelial tube morphogenesis(GO:0061155) regulation of transcription from RNA polymerase II promoter involved in mesonephros development(GO:0061216) pattern specification involved in mesonephros development(GO:0061227) BMP signaling pathway involved in nephric duct formation(GO:0071893) negative regulation of branch elongation involved in ureteric bud branching(GO:0072096) negative regulation of branch elongation involved in ureteric bud branching by BMP signaling pathway(GO:0072097) anterior/posterior pattern specification involved in kidney development(GO:0072098) anterior/posterior pattern specification involved in ureteric bud development(GO:0072099) specification of ureteric bud anterior/posterior symmetry(GO:0072100) specification of ureteric bud anterior/posterior symmetry by BMP signaling pathway(GO:0072101) ureter urothelium development(GO:0072190) ureter epithelial cell differentiation(GO:0072192) ureter morphogenesis(GO:0072197) negative regulation of mesenchymal cell proliferation involved in ureter development(GO:0072200) positive regulation of cell proliferation involved in outflow tract morphogenesis(GO:1901964) cardiac jelly development(GO:1905072) regulation of metanephric S-shaped body morphogenesis(GO:2000004) negative regulation of metanephric S-shaped body morphogenesis(GO:2000005) regulation of metanephric comma-shaped body morphogenesis(GO:2000006) negative regulation of metanephric comma-shaped body morphogenesis(GO:2000007) |
| 0.7 | 6.6 | GO:0006957 | complement activation, alternative pathway(GO:0006957) |
| 0.7 | 2.0 | GO:0019442 | tryptophan catabolic process to acetyl-CoA(GO:0019442) |
| 0.7 | 5.3 | GO:0048630 | skeletal muscle tissue growth(GO:0048630) |
| 0.7 | 4.6 | GO:0006572 | tyrosine catabolic process(GO:0006572) |
| 0.6 | 2.6 | GO:0035633 | maintenance of blood-brain barrier(GO:0035633) |
| 0.6 | 1.9 | GO:0070105 | positive regulation of interleukin-6-mediated signaling pathway(GO:0070105) |
| 0.6 | 2.6 | GO:0002765 | immune response-inhibiting signal transduction(GO:0002765) immune response-inhibiting cell surface receptor signaling pathway(GO:0002767) |
| 0.6 | 1.9 | GO:0071934 | thiamine transport(GO:0015888) thiamine transmembrane transport(GO:0071934) |
| 0.6 | 1.9 | GO:0035356 | cellular triglyceride homeostasis(GO:0035356) |
| 0.6 | 1.9 | GO:0019287 | isopentenyl diphosphate biosynthetic process, mevalonate pathway(GO:0019287) |
| 0.6 | 2.5 | GO:1904566 | response to 1-oleoyl-sn-glycerol 3-phosphate(GO:1904565) cellular response to 1-oleoyl-sn-glycerol 3-phosphate(GO:1904566) |
| 0.6 | 1.8 | GO:0090187 | positive regulation of pancreatic juice secretion(GO:0090187) |
| 0.6 | 6.0 | GO:0043950 | positive regulation of cAMP-mediated signaling(GO:0043950) |
| 0.6 | 1.2 | GO:0030323 | respiratory tube development(GO:0030323) |
| 0.6 | 1.8 | GO:0036371 | protein localization to M-band(GO:0036309) protein localization to T-tubule(GO:0036371) |
| 0.6 | 2.9 | GO:2000639 | regulation of SREBP signaling pathway(GO:2000638) negative regulation of SREBP signaling pathway(GO:2000639) |
| 0.6 | 4.0 | GO:0042246 | tissue regeneration(GO:0042246) |
| 0.6 | 1.7 | GO:0036060 | filtration diaphragm assembly(GO:0036058) slit diaphragm assembly(GO:0036060) |
| 0.6 | 1.1 | GO:0052314 | phytoalexin metabolic process(GO:0052314) |
| 0.6 | 4.0 | GO:0002414 | immunoglobulin transcytosis in epithelial cells(GO:0002414) |
| 0.6 | 2.3 | GO:1900245 | positive regulation of MDA-5 signaling pathway(GO:1900245) |
| 0.6 | 2.8 | GO:0033031 | positive regulation of neutrophil apoptotic process(GO:0033031) |
| 0.6 | 2.2 | GO:0032661 | regulation of interleukin-18 production(GO:0032661) |
| 0.6 | 2.8 | GO:0017121 | phospholipid scrambling(GO:0017121) |
| 0.6 | 2.2 | GO:0007525 | somatic muscle development(GO:0007525) |
| 0.6 | 14.4 | GO:0045730 | respiratory burst(GO:0045730) |
| 0.5 | 1.6 | GO:0010738 | regulation of protein kinase A signaling(GO:0010738) |
| 0.5 | 2.2 | GO:2001206 | positive regulation of osteoclast development(GO:2001206) |
| 0.5 | 3.2 | GO:0070383 | DNA cytosine deamination(GO:0070383) |
| 0.5 | 2.6 | GO:0030070 | insulin processing(GO:0030070) |
| 0.5 | 2.1 | GO:0097089 | methyl-branched fatty acid metabolic process(GO:0097089) |
| 0.5 | 6.3 | GO:0008627 | intrinsic apoptotic signaling pathway in response to osmotic stress(GO:0008627) |
| 0.5 | 8.9 | GO:0010457 | centriole-centriole cohesion(GO:0010457) |
| 0.5 | 1.6 | GO:0001812 | positive regulation of type I hypersensitivity(GO:0001812) |
| 0.5 | 7.3 | GO:0071688 | striated muscle myosin thick filament assembly(GO:0071688) |
| 0.5 | 5.2 | GO:0015889 | cobalamin transport(GO:0015889) |
| 0.5 | 2.6 | GO:2000301 | negative regulation of synaptic vesicle exocytosis(GO:2000301) |
| 0.5 | 19.0 | GO:0050869 | negative regulation of B cell activation(GO:0050869) |
| 0.5 | 2.6 | GO:2000348 | regulation of CD40 signaling pathway(GO:2000348) |
| 0.5 | 1.5 | GO:0015855 | pyrimidine nucleobase transport(GO:0015855) purine nucleobase transmembrane transport(GO:1904823) |
| 0.5 | 8.7 | GO:0034374 | low-density lipoprotein particle remodeling(GO:0034374) |
| 0.5 | 2.0 | GO:0045629 | negative regulation of T-helper 2 cell differentiation(GO:0045629) |
| 0.5 | 10.0 | GO:0035589 | G-protein coupled purinergic nucleotide receptor signaling pathway(GO:0035589) |
| 0.5 | 1.5 | GO:0051697 | protein delipidation(GO:0051697) |
| 0.5 | 3.9 | GO:0098532 | histone H3-K27 trimethylation(GO:0098532) |
| 0.5 | 4.9 | GO:0051611 | negative regulation of neurotransmitter uptake(GO:0051581) serotonin uptake(GO:0051610) regulation of serotonin uptake(GO:0051611) negative regulation of serotonin uptake(GO:0051612) |
| 0.5 | 1.9 | GO:0007198 | adenylate cyclase-inhibiting serotonin receptor signaling pathway(GO:0007198) |
| 0.5 | 2.4 | GO:0001550 | ovarian cumulus expansion(GO:0001550) fused antrum stage(GO:0048165) |
| 0.5 | 3.4 | GO:2000503 | positive regulation of natural killer cell chemotaxis(GO:2000503) |
| 0.5 | 1.9 | GO:0042360 | vitamin E metabolic process(GO:0042360) |
| 0.5 | 23.9 | GO:0050911 | detection of chemical stimulus involved in sensory perception of smell(GO:0050911) |
| 0.5 | 1.4 | GO:0051919 | positive regulation of fibrinolysis(GO:0051919) |
| 0.5 | 8.0 | GO:0006265 | DNA topological change(GO:0006265) |
| 0.5 | 1.4 | GO:0045900 | negative regulation of translational elongation(GO:0045900) |
| 0.5 | 2.3 | GO:0070944 | neutrophil mediated cytotoxicity(GO:0070942) neutrophil mediated killing of symbiont cell(GO:0070943) neutrophil mediated killing of bacterium(GO:0070944) |
| 0.5 | 1.4 | GO:0060023 | soft palate development(GO:0060023) |
| 0.4 | 0.9 | GO:0000394 | RNA splicing, via endonucleolytic cleavage and ligation(GO:0000394) tRNA splicing, via endonucleolytic cleavage and ligation(GO:0006388) |
| 0.4 | 4.0 | GO:0050713 | negative regulation of interleukin-1 beta secretion(GO:0050713) |
| 0.4 | 1.8 | GO:0010966 | regulation of phosphate transport(GO:0010966) |
| 0.4 | 1.3 | GO:0015722 | canalicular bile acid transport(GO:0015722) |
| 0.4 | 4.0 | GO:0010886 | positive regulation of cholesterol storage(GO:0010886) |
| 0.4 | 5.7 | GO:0034162 | toll-like receptor 9 signaling pathway(GO:0034162) |
| 0.4 | 1.3 | GO:1990502 | dense core granule maturation(GO:1990502) |
| 0.4 | 6.5 | GO:0007597 | blood coagulation, intrinsic pathway(GO:0007597) |
| 0.4 | 1.3 | GO:0045938 | positive regulation of circadian sleep/wake cycle, sleep(GO:0045938) |
| 0.4 | 0.8 | GO:0010760 | negative regulation of macrophage chemotaxis(GO:0010760) |
| 0.4 | 0.8 | GO:0031860 | regulation of mitotic recombination(GO:0000019) telomeric 3' overhang formation(GO:0031860) |
| 0.4 | 2.1 | GO:1903301 | positive regulation of glucokinase activity(GO:0033133) positive regulation of hexokinase activity(GO:1903301) |
| 0.4 | 1.7 | GO:0010936 | negative regulation of macrophage cytokine production(GO:0010936) |
| 0.4 | 1.7 | GO:0060054 | positive regulation of epithelial cell proliferation involved in wound healing(GO:0060054) |
| 0.4 | 6.2 | GO:0060087 | relaxation of vascular smooth muscle(GO:0060087) |
| 0.4 | 1.7 | GO:1990418 | response to insulin-like growth factor stimulus(GO:1990418) |
| 0.4 | 1.2 | GO:0002314 | germinal center B cell differentiation(GO:0002314) |
| 0.4 | 1.2 | GO:1902958 | neuron intrinsic apoptotic signaling pathway in response to hydrogen peroxide(GO:0036482) positive regulation of cellular respiration(GO:1901857) positive regulation of mitochondrial electron transport, NADH to ubiquinone(GO:1902958) regulation of hydrogen peroxide-induced neuron intrinsic apoptotic signaling pathway(GO:1903383) negative regulation of hydrogen peroxide-induced neuron intrinsic apoptotic signaling pathway(GO:1903384) |
| 0.4 | 9.9 | GO:0042744 | hydrogen peroxide catabolic process(GO:0042744) |
| 0.4 | 2.1 | GO:1903237 | negative regulation of leukocyte tethering or rolling(GO:1903237) |
| 0.4 | 3.3 | GO:0070995 | NADPH oxidation(GO:0070995) |
| 0.4 | 8.6 | GO:0034505 | tooth mineralization(GO:0034505) |
| 0.4 | 1.2 | GO:0002101 | tRNA wobble cytosine modification(GO:0002101) |
| 0.4 | 5.7 | GO:0018298 | protein-chromophore linkage(GO:0018298) |
| 0.4 | 1.6 | GO:0006663 | platelet activating factor biosynthetic process(GO:0006663) |
| 0.4 | 4.7 | GO:0070327 | thyroid hormone transport(GO:0070327) |
| 0.4 | 1.6 | GO:0046271 | phenylpropanoid catabolic process(GO:0046271) |
| 0.4 | 2.7 | GO:0007000 | transcription initiation from RNA polymerase I promoter for nuclear large rRNA transcript(GO:0001180) nucleolus organization(GO:0007000) |
| 0.4 | 1.5 | GO:0030822 | positive regulation of cyclic nucleotide catabolic process(GO:0030807) positive regulation of cAMP catabolic process(GO:0030822) positive regulation of purine nucleotide catabolic process(GO:0033123) |
| 0.4 | 8.8 | GO:0030502 | negative regulation of bone mineralization(GO:0030502) |
| 0.4 | 0.8 | GO:0002331 | pre-B cell differentiation(GO:0002329) pre-B cell allelic exclusion(GO:0002331) |
| 0.4 | 8.3 | GO:0006749 | glutathione metabolic process(GO:0006749) |
| 0.4 | 1.5 | GO:0035093 | spermatogenesis, exchange of chromosomal proteins(GO:0035093) |
| 0.4 | 56.4 | GO:0002377 | immunoglobulin production(GO:0002377) |
| 0.4 | 2.6 | GO:0042723 | thiamine metabolic process(GO:0006772) thiamine-containing compound metabolic process(GO:0042723) |
| 0.4 | 1.1 | GO:0001694 | histamine biosynthetic process(GO:0001694) |
| 0.4 | 2.5 | GO:0071896 | protein localization to adherens junction(GO:0071896) |
| 0.4 | 6.4 | GO:0097151 | positive regulation of inhibitory postsynaptic potential(GO:0097151) modulation of inhibitory postsynaptic potential(GO:0098828) |
| 0.4 | 2.1 | GO:1903244 | positive regulation of cardiac muscle adaptation(GO:0010615) positive regulation of cardiac muscle hypertrophy in response to stress(GO:1903244) |
| 0.4 | 4.6 | GO:0033617 | mitochondrial respiratory chain complex IV assembly(GO:0033617) mitochondrial respiratory chain complex IV biogenesis(GO:0097034) |
| 0.4 | 1.8 | GO:0035672 | oligopeptide transmembrane transport(GO:0035672) |
| 0.3 | 2.8 | GO:0001920 | negative regulation of receptor recycling(GO:0001920) |
| 0.3 | 1.7 | GO:0007197 | adenylate cyclase-inhibiting G-protein coupled acetylcholine receptor signaling pathway(GO:0007197) |
| 0.3 | 3.1 | GO:0042574 | retinal metabolic process(GO:0042574) |
| 0.3 | 1.7 | GO:0010519 | negative regulation of phospholipase activity(GO:0010519) minus-end-directed organelle transport along microtubule(GO:0072385) regulation of phospholipase C-activating G-protein coupled receptor signaling pathway(GO:1900736) |
| 0.3 | 3.7 | GO:0048743 | positive regulation of skeletal muscle fiber development(GO:0048743) |
| 0.3 | 2.4 | GO:0000160 | phosphorelay signal transduction system(GO:0000160) |
| 0.3 | 1.0 | GO:0061713 | neural crest cell migration involved in heart formation(GO:0003147) cell migration involved in heart formation(GO:0060974) anterior neural tube closure(GO:0061713) |
| 0.3 | 2.7 | GO:0006528 | asparagine metabolic process(GO:0006528) |
| 0.3 | 3.6 | GO:1903608 | protein localization to cytoplasmic stress granule(GO:1903608) |
| 0.3 | 1.6 | GO:0072233 | thick ascending limb development(GO:0072023) metanephric thick ascending limb development(GO:0072233) |
| 0.3 | 1.3 | GO:0070309 | lens fiber cell morphogenesis(GO:0070309) |
| 0.3 | 0.6 | GO:0015074 | DNA integration(GO:0015074) |
| 0.3 | 3.8 | GO:0036158 | outer dynein arm assembly(GO:0036158) |
| 0.3 | 3.1 | GO:0046951 | ketone body biosynthetic process(GO:0046951) |
| 0.3 | 0.9 | GO:0032627 | interleukin-23 production(GO:0032627) regulation of interleukin-23 production(GO:0032667) |
| 0.3 | 1.8 | GO:0060282 | positive regulation of oocyte development(GO:0060282) |
| 0.3 | 1.8 | GO:0055059 | asymmetric neuroblast division(GO:0055059) |
| 0.3 | 1.8 | GO:0015811 | L-cystine transport(GO:0015811) |
| 0.3 | 2.1 | GO:0034720 | histone H3-K4 demethylation(GO:0034720) |
| 0.3 | 1.5 | GO:0042539 | hypotonic salinity response(GO:0042539) cellular hypotonic salinity response(GO:0071477) |
| 0.3 | 2.7 | GO:0014894 | response to muscle inactivity involved in regulation of muscle adaptation(GO:0014877) response to denervation involved in regulation of muscle adaptation(GO:0014894) |
| 0.3 | 0.6 | GO:0010701 | positive regulation of norepinephrine secretion(GO:0010701) |
| 0.3 | 6.2 | GO:0035435 | phosphate ion transmembrane transport(GO:0035435) |
| 0.3 | 2.6 | GO:0006068 | ethanol catabolic process(GO:0006068) |
| 0.3 | 15.4 | GO:0015701 | bicarbonate transport(GO:0015701) |
| 0.3 | 1.2 | GO:0014807 | regulation of somitogenesis(GO:0014807) |
| 0.3 | 1.7 | GO:0035914 | skeletal muscle cell differentiation(GO:0035914) |
| 0.3 | 4.8 | GO:0043651 | linoleic acid metabolic process(GO:0043651) |
| 0.3 | 1.4 | GO:0050955 | thermoception(GO:0050955) |
| 0.3 | 1.9 | GO:0010735 | positive regulation of transcription via serum response element binding(GO:0010735) |
| 0.3 | 1.7 | GO:0035331 | negative regulation of hippo signaling(GO:0035331) |
| 0.3 | 3.3 | GO:0071447 | cellular response to hydroperoxide(GO:0071447) |
| 0.3 | 3.8 | GO:0007016 | cytoskeletal anchoring at plasma membrane(GO:0007016) |
| 0.3 | 1.4 | GO:0047484 | regulation of response to osmotic stress(GO:0047484) |
| 0.3 | 3.5 | GO:0097186 | amelogenesis(GO:0097186) |
| 0.3 | 1.1 | GO:0060748 | tertiary branching involved in mammary gland duct morphogenesis(GO:0060748) |
| 0.3 | 10.9 | GO:0016266 | O-glycan processing(GO:0016266) |
| 0.3 | 1.0 | GO:0007249 | I-kappaB kinase/NF-kappaB signaling(GO:0007249) |
| 0.3 | 1.0 | GO:0060455 | negative regulation of gastric acid secretion(GO:0060455) |
| 0.3 | 2.3 | GO:0032264 | IMP salvage(GO:0032264) |
| 0.3 | 0.5 | GO:0050917 | sensory perception of umami taste(GO:0050917) |
| 0.3 | 9.7 | GO:0006779 | porphyrin-containing compound biosynthetic process(GO:0006779) |
| 0.3 | 2.5 | GO:0043249 | erythrocyte maturation(GO:0043249) |
| 0.2 | 1.0 | GO:0006127 | glycerophosphate shuttle(GO:0006127) |
| 0.2 | 0.5 | GO:2000630 | positive regulation of miRNA metabolic process(GO:2000630) |
| 0.2 | 6.4 | GO:0097503 | sialylation(GO:0097503) |
| 0.2 | 1.5 | GO:0033504 | floor plate development(GO:0033504) |
| 0.2 | 1.0 | GO:0010585 | glutamine secretion(GO:0010585) L-glutamine import(GO:0036229) L-glutamine import into cell(GO:1903803) |
| 0.2 | 1.7 | GO:0008218 | bioluminescence(GO:0008218) |
| 0.2 | 2.7 | GO:1900112 | regulation of histone H3-K9 trimethylation(GO:1900112) |
| 0.2 | 2.4 | GO:0050942 | positive regulation of pigment cell differentiation(GO:0050942) |
| 0.2 | 3.9 | GO:0042448 | progesterone metabolic process(GO:0042448) |
| 0.2 | 1.2 | GO:0010898 | positive regulation of triglyceride catabolic process(GO:0010898) |
| 0.2 | 1.0 | GO:0045964 | positive regulation of catecholamine metabolic process(GO:0045915) positive regulation of dopamine metabolic process(GO:0045964) |
| 0.2 | 8.4 | GO:0042462 | eye photoreceptor cell development(GO:0042462) |
| 0.2 | 8.4 | GO:0050829 | defense response to Gram-negative bacterium(GO:0050829) |
| 0.2 | 2.2 | GO:1902172 | keratinocyte apoptotic process(GO:0097283) regulation of keratinocyte apoptotic process(GO:1902172) |
| 0.2 | 1.6 | GO:0015705 | iodide transport(GO:0015705) |
| 0.2 | 2.0 | GO:0009650 | UV protection(GO:0009650) |
| 0.2 | 0.9 | GO:0002634 | regulation of germinal center formation(GO:0002634) primary heart field specification(GO:0003138) sinoatrial valve development(GO:0003172) sinoatrial valve morphogenesis(GO:0003185) |
| 0.2 | 4.2 | GO:0060746 | maternal behavior(GO:0042711) parental behavior(GO:0060746) |
| 0.2 | 1.1 | GO:0070544 | histone H3-K36 demethylation(GO:0070544) |
| 0.2 | 2.3 | GO:0055001 | muscle cell development(GO:0055001) |
| 0.2 | 1.4 | GO:0044351 | macropinocytosis(GO:0044351) |
| 0.2 | 0.2 | GO:1902358 | sulfate transmembrane transport(GO:1902358) |
| 0.2 | 8.6 | GO:0036297 | interstrand cross-link repair(GO:0036297) |
| 0.2 | 1.6 | GO:0097194 | execution phase of apoptosis(GO:0097194) |
| 0.2 | 1.4 | GO:0009249 | protein lipoylation(GO:0009249) |
| 0.2 | 0.4 | GO:0044205 | 'de novo' UMP biosynthetic process(GO:0044205) |
| 0.2 | 1.2 | GO:0006776 | vitamin A metabolic process(GO:0006776) |
| 0.2 | 6.8 | GO:0034219 | carbohydrate transmembrane transport(GO:0034219) |
| 0.2 | 1.4 | GO:0048014 | Tie signaling pathway(GO:0048014) |
| 0.2 | 0.6 | GO:1904263 | positive regulation of TORC1 signaling(GO:1904263) |
| 0.2 | 0.4 | GO:0090086 | negative regulation of protein deubiquitination(GO:0090086) |
| 0.2 | 1.2 | GO:0042982 | amyloid precursor protein metabolic process(GO:0042982) |
| 0.2 | 0.8 | GO:0032218 | riboflavin transport(GO:0032218) |
| 0.2 | 4.5 | GO:0045332 | lipid translocation(GO:0034204) phospholipid translocation(GO:0045332) |
| 0.2 | 11.7 | GO:0030212 | hyaluronan metabolic process(GO:0030212) |
| 0.2 | 3.7 | GO:0042135 | neurotransmitter catabolic process(GO:0042135) |
| 0.2 | 2.1 | GO:0032196 | transposition(GO:0032196) |
| 0.2 | 1.2 | GO:0061737 | leukotriene signaling pathway(GO:0061737) |
| 0.2 | 4.0 | GO:0034080 | CENP-A containing nucleosome assembly(GO:0034080) CENP-A containing chromatin organization(GO:0061641) |
| 0.2 | 4.3 | GO:0008210 | estrogen metabolic process(GO:0008210) |
| 0.2 | 0.6 | GO:0002678 | positive regulation of chronic inflammatory response(GO:0002678) |
| 0.2 | 0.2 | GO:0072053 | renal inner medulla development(GO:0072053) renal outer medulla development(GO:0072054) |
| 0.2 | 1.9 | GO:0019934 | cGMP-mediated signaling(GO:0019934) |
| 0.2 | 8.6 | GO:0018149 | peptide cross-linking(GO:0018149) |
| 0.2 | 5.3 | GO:0019433 | triglyceride catabolic process(GO:0019433) |
| 0.2 | 2.0 | GO:0060285 | cilium-dependent cell motility(GO:0060285) |
| 0.2 | 1.6 | GO:0038028 | insulin receptor signaling pathway via phosphatidylinositol 3-kinase(GO:0038028) |
| 0.2 | 2.1 | GO:2000271 | positive regulation of fibroblast apoptotic process(GO:2000271) |
| 0.2 | 0.5 | GO:0006655 | phosphatidylglycerol biosynthetic process(GO:0006655) |
| 0.2 | 1.8 | GO:0060219 | camera-type eye photoreceptor cell differentiation(GO:0060219) |
| 0.2 | 4.5 | GO:0035728 | response to hepatocyte growth factor(GO:0035728) |
| 0.2 | 0.7 | GO:0042746 | regulation of circadian sleep/wake cycle, wakefulness(GO:0010840) circadian sleep/wake cycle, wakefulness(GO:0042746) |
| 0.2 | 2.2 | GO:0044065 | regulation of respiratory system process(GO:0044065) |
| 0.2 | 9.9 | GO:0006501 | C-terminal protein lipidation(GO:0006501) |
| 0.2 | 1.4 | GO:0001865 | NK T cell differentiation(GO:0001865) |
| 0.2 | 1.7 | GO:0035457 | cellular response to interferon-alpha(GO:0035457) |
| 0.2 | 2.2 | GO:0045793 | positive regulation of cell size(GO:0045793) |
| 0.2 | 0.8 | GO:0015886 | heme transport(GO:0015886) |
| 0.2 | 5.6 | GO:0016056 | rhodopsin mediated signaling pathway(GO:0016056) |
| 0.2 | 1.3 | GO:0000050 | urea cycle(GO:0000050) |
| 0.2 | 1.8 | GO:0048172 | regulation of short-term neuronal synaptic plasticity(GO:0048172) |
| 0.2 | 1.3 | GO:0032026 | response to magnesium ion(GO:0032026) |
| 0.2 | 3.5 | GO:0006646 | phosphatidylethanolamine biosynthetic process(GO:0006646) |
| 0.2 | 2.4 | GO:0042340 | keratan sulfate catabolic process(GO:0042340) |
| 0.2 | 0.9 | GO:0003344 | pericardium morphogenesis(GO:0003344) |
| 0.2 | 2.9 | GO:0016558 | protein import into peroxisome matrix(GO:0016558) |
| 0.2 | 1.5 | GO:2000049 | positive regulation of cell-cell adhesion mediated by cadherin(GO:2000049) |
| 0.2 | 2.1 | GO:0006004 | fucose metabolic process(GO:0006004) |
| 0.2 | 1.7 | GO:0008612 | peptidyl-lysine modification to peptidyl-hypusine(GO:0008612) |
| 0.2 | 0.6 | GO:0046464 | neutral lipid catabolic process(GO:0046461) acylglycerol catabolic process(GO:0046464) |
| 0.1 | 1.2 | GO:0032534 | regulation of microvillus assembly(GO:0032534) |
| 0.1 | 2.4 | GO:0036344 | platelet formation(GO:0030220) platelet morphogenesis(GO:0036344) |
| 0.1 | 0.6 | GO:0030035 | microspike assembly(GO:0030035) |
| 0.1 | 6.3 | GO:0070206 | protein trimerization(GO:0070206) |
| 0.1 | 2.0 | GO:0031297 | replication fork processing(GO:0031297) |
| 0.1 | 0.4 | GO:0034343 | type III interferon production(GO:0034343) regulation of type III interferon production(GO:0034344) |
| 0.1 | 2.2 | GO:0060046 | regulation of acrosome reaction(GO:0060046) |
| 0.1 | 0.4 | GO:0061365 | positive regulation of triglyceride lipase activity(GO:0061365) |
| 0.1 | 3.0 | GO:0019835 | cytolysis(GO:0019835) |
| 0.1 | 0.7 | GO:0072221 | distal convoluted tubule development(GO:0072025) metanephric distal convoluted tubule development(GO:0072221) |
| 0.1 | 0.8 | GO:0016191 | synaptic vesicle uncoating(GO:0016191) |
| 0.1 | 1.0 | GO:0044245 | polysaccharide digestion(GO:0044245) |
| 0.1 | 1.1 | GO:1900017 | positive regulation of cytokine production involved in inflammatory response(GO:1900017) |
| 0.1 | 1.1 | GO:0048251 | elastic fiber assembly(GO:0048251) |
| 0.1 | 0.9 | GO:0006552 | leucine catabolic process(GO:0006552) |
| 0.1 | 0.4 | GO:1904425 | negative regulation of GTP binding(GO:1904425) |
| 0.1 | 0.7 | GO:0060088 | auditory receptor cell stereocilium organization(GO:0060088) |
| 0.1 | 0.7 | GO:0033227 | dsRNA transport(GO:0033227) |
| 0.1 | 0.7 | GO:0038095 | Fc-epsilon receptor signaling pathway(GO:0038095) |
| 0.1 | 20.1 | GO:0007156 | homophilic cell adhesion via plasma membrane adhesion molecules(GO:0007156) |
| 0.1 | 2.8 | GO:0008211 | glucocorticoid metabolic process(GO:0008211) |
| 0.1 | 1.8 | GO:1904714 | regulation of chaperone-mediated autophagy(GO:1904714) |
| 0.1 | 1.2 | GO:0086024 | adrenergic receptor signaling pathway involved in positive regulation of heart rate(GO:0086024) |
| 0.1 | 0.9 | GO:0035095 | behavioral response to nicotine(GO:0035095) |
| 0.1 | 2.5 | GO:0043923 | positive regulation by host of viral transcription(GO:0043923) |
| 0.1 | 4.9 | GO:0032729 | positive regulation of interferon-gamma production(GO:0032729) |
| 0.1 | 3.9 | GO:0006298 | mismatch repair(GO:0006298) |
| 0.1 | 1.8 | GO:0050832 | defense response to fungus(GO:0050832) |
| 0.1 | 0.1 | GO:0030521 | androgen receptor signaling pathway(GO:0030521) |
| 0.1 | 1.1 | GO:0002517 | T cell tolerance induction(GO:0002517) |
| 0.1 | 6.2 | GO:0042267 | natural killer cell mediated cytotoxicity(GO:0042267) |
| 0.1 | 0.6 | GO:0050860 | negative regulation of T cell receptor signaling pathway(GO:0050860) |
| 0.1 | 3.1 | GO:0006911 | phagocytosis, engulfment(GO:0006911) |
| 0.1 | 1.5 | GO:0006995 | cellular response to nitrogen starvation(GO:0006995) cellular response to nitrogen levels(GO:0043562) |
| 0.1 | 2.0 | GO:0019373 | epoxygenase P450 pathway(GO:0019373) |
| 0.1 | 4.8 | GO:0019228 | neuronal action potential(GO:0019228) |
| 0.1 | 3.2 | GO:0042417 | dopamine metabolic process(GO:0042417) |
| 0.1 | 2.1 | GO:0090022 | regulation of neutrophil chemotaxis(GO:0090022) |
| 0.1 | 0.8 | GO:0043116 | negative regulation of vascular permeability(GO:0043116) |
| 0.1 | 0.6 | GO:0038065 | collagen-activated tyrosine kinase receptor signaling pathway(GO:0038063) collagen-activated signaling pathway(GO:0038065) |
| 0.1 | 2.8 | GO:0051084 | 'de novo' posttranslational protein folding(GO:0051084) |
| 0.1 | 4.4 | GO:0006953 | acute-phase response(GO:0006953) |
| 0.1 | 0.3 | GO:1904177 | regulation of adipose tissue development(GO:1904177) |
| 0.1 | 0.8 | GO:0031665 | negative regulation of lipopolysaccharide-mediated signaling pathway(GO:0031665) |
| 0.1 | 1.6 | GO:0002323 | natural killer cell activation involved in immune response(GO:0002323) |
| 0.1 | 0.7 | GO:2001214 | positive regulation of vasculogenesis(GO:2001214) |
| 0.1 | 1.3 | GO:0010991 | negative regulation of SMAD protein complex assembly(GO:0010991) |
| 0.1 | 1.0 | GO:0019932 | second-messenger-mediated signaling(GO:0019932) |
| 0.1 | 1.9 | GO:0045725 | positive regulation of glycogen biosynthetic process(GO:0045725) |
| 0.1 | 0.3 | GO:0038043 | interleukin-5-mediated signaling pathway(GO:0038043) |
| 0.1 | 4.7 | GO:0007214 | gamma-aminobutyric acid signaling pathway(GO:0007214) |
| 0.1 | 0.8 | GO:0042989 | sequestering of actin monomers(GO:0042989) |
| 0.1 | 0.5 | GO:1904879 | positive regulation of calcium ion transmembrane transport via high voltage-gated calcium channel(GO:1904879) |
| 0.1 | 0.9 | GO:0090344 | negative regulation of cell aging(GO:0090344) |
| 0.1 | 1.6 | GO:0070328 | acylglycerol homeostasis(GO:0055090) triglyceride homeostasis(GO:0070328) |
| 0.1 | 1.4 | GO:0048762 | mesenchymal cell differentiation(GO:0048762) |
| 0.1 | 3.0 | GO:0007194 | negative regulation of adenylate cyclase activity(GO:0007194) |
| 0.1 | 2.2 | GO:0097120 | receptor localization to synapse(GO:0097120) |
| 0.1 | 1.4 | GO:1901673 | regulation of mitotic spindle assembly(GO:1901673) |
| 0.1 | 0.9 | GO:0006685 | sphingomyelin catabolic process(GO:0006685) |
| 0.1 | 1.1 | GO:0022038 | corpus callosum development(GO:0022038) |
| 0.1 | 1.0 | GO:0060021 | palate development(GO:0060021) |
| 0.1 | 0.1 | GO:1902908 | regulation of melanosome transport(GO:1902908) |
| 0.1 | 0.7 | GO:0046777 | protein autophosphorylation(GO:0046777) |
| 0.1 | 1.3 | GO:0045824 | negative regulation of innate immune response(GO:0045824) |
| 0.1 | 1.8 | GO:0021895 | cerebral cortex neuron differentiation(GO:0021895) |
| 0.1 | 2.5 | GO:0016338 | calcium-independent cell-cell adhesion via plasma membrane cell-adhesion molecules(GO:0016338) |
| 0.1 | 0.9 | GO:1990118 | sodium ion import across plasma membrane(GO:0098719) sodium ion import into cell(GO:1990118) |
| 0.1 | 1.2 | GO:0051450 | myoblast proliferation(GO:0051450) regulation of myoblast proliferation(GO:2000291) |
| 0.1 | 0.7 | GO:1903432 | regulation of TORC1 signaling(GO:1903432) |
| 0.1 | 0.8 | GO:0016180 | snRNA processing(GO:0016180) |
| 0.1 | 0.9 | GO:0043578 | nuclear matrix organization(GO:0043578) nuclear matrix anchoring at nuclear membrane(GO:0090292) |
| 0.1 | 0.5 | GO:0060215 | primitive hemopoiesis(GO:0060215) regulation of cell differentiation involved in embryonic placenta development(GO:0060800) |
| 0.1 | 2.1 | GO:0042554 | superoxide anion generation(GO:0042554) |
| 0.1 | 1.0 | GO:0045662 | negative regulation of myoblast differentiation(GO:0045662) |
| 0.1 | 1.1 | GO:0030038 | contractile actin filament bundle assembly(GO:0030038) stress fiber assembly(GO:0043149) |
| 0.1 | 2.0 | GO:0006198 | cAMP catabolic process(GO:0006198) |
| 0.1 | 3.0 | GO:1902475 | L-alpha-amino acid transmembrane transport(GO:1902475) |
| 0.1 | 1.4 | GO:0048147 | negative regulation of fibroblast proliferation(GO:0048147) |
| 0.1 | 1.5 | GO:0031915 | positive regulation of synaptic plasticity(GO:0031915) |
| 0.1 | 1.3 | GO:0042789 | mRNA transcription from RNA polymerase II promoter(GO:0042789) cellular response to thyroid hormone stimulus(GO:0097067) |
| 0.1 | 2.9 | GO:0007205 | protein kinase C-activating G-protein coupled receptor signaling pathway(GO:0007205) |
| 0.1 | 0.9 | GO:0010992 | ubiquitin homeostasis(GO:0010992) |
| 0.1 | 1.0 | GO:0051639 | actin filament network formation(GO:0051639) |
| 0.1 | 0.8 | GO:0019375 | galactosylceramide biosynthetic process(GO:0006682) galactolipid biosynthetic process(GO:0019375) |
| 0.1 | 1.2 | GO:0060009 | Sertoli cell development(GO:0060009) |
| 0.1 | 0.6 | GO:0060745 | mammary gland branching involved in pregnancy(GO:0060745) |
| 0.1 | 0.8 | GO:0002755 | MyD88-dependent toll-like receptor signaling pathway(GO:0002755) |
| 0.1 | 0.2 | GO:0044240 | multicellular organism lipid catabolic process(GO:0044240) |
| 0.1 | 3.0 | GO:0007595 | lactation(GO:0007595) |
| 0.1 | 0.7 | GO:0042348 | regulation of NF-kappaB import into nucleus(GO:0042345) NF-kappaB import into nucleus(GO:0042348) |
| 0.1 | 3.5 | GO:0050776 | regulation of immune response(GO:0050776) |
| 0.1 | 0.5 | GO:0034142 | toll-like receptor 4 signaling pathway(GO:0034142) |
| 0.1 | 1.5 | GO:0007409 | axonogenesis(GO:0007409) |
| 0.1 | 1.6 | GO:0070979 | protein K11-linked ubiquitination(GO:0070979) |
| 0.1 | 1.8 | GO:0003333 | amino acid transmembrane transport(GO:0003333) |
| 0.1 | 1.2 | GO:1990001 | inhibition of cysteine-type endopeptidase activity involved in apoptotic process(GO:1990001) |
| 0.1 | 1.2 | GO:0018345 | protein palmitoylation(GO:0018345) |
| 0.1 | 1.0 | GO:0044243 | multicellular organism catabolic process(GO:0044243) |
| 0.1 | 0.6 | GO:0039536 | negative regulation of RIG-I signaling pathway(GO:0039536) |
| 0.1 | 0.4 | GO:0031960 | response to corticosteroid(GO:0031960) response to glucocorticoid(GO:0051384) |
| 0.1 | 1.0 | GO:0035082 | axoneme assembly(GO:0035082) |
| 0.1 | 0.1 | GO:0035754 | B cell chemotaxis(GO:0035754) |
| 0.1 | 0.6 | GO:0003351 | epithelial cilium movement(GO:0003351) |
| 0.1 | 1.2 | GO:0051290 | protein heterotetramerization(GO:0051290) |
| 0.1 | 3.9 | GO:0043124 | negative regulation of I-kappaB kinase/NF-kappaB signaling(GO:0043124) |
| 0.1 | 1.8 | GO:0015695 | organic cation transport(GO:0015695) |
| 0.1 | 1.9 | GO:0016925 | protein sumoylation(GO:0016925) |
| 0.1 | 2.0 | GO:0006968 | cellular defense response(GO:0006968) |
| 0.1 | 11.5 | GO:0002250 | adaptive immune response(GO:0002250) |
| 0.1 | 0.5 | GO:0045059 | positive thymic T cell selection(GO:0045059) |
| 0.1 | 0.5 | GO:1901475 | pyruvate transport(GO:0006848) pyruvate transmembrane transport(GO:1901475) |
| 0.1 | 3.2 | GO:0045814 | negative regulation of gene expression, epigenetic(GO:0045814) |
| 0.1 | 0.7 | GO:0071361 | cellular response to ethanol(GO:0071361) |
| 0.1 | 0.1 | GO:0019249 | lactate biosynthetic process(GO:0019249) |
| 0.1 | 1.3 | GO:0032007 | negative regulation of TOR signaling(GO:0032007) |
| 0.1 | 1.1 | GO:0070935 | 3'-UTR-mediated mRNA stabilization(GO:0070935) |
| 0.1 | 0.2 | GO:0001514 | selenocysteine incorporation(GO:0001514) translational readthrough(GO:0006451) |
| 0.1 | 0.1 | GO:0050717 | positive regulation of interleukin-1 alpha secretion(GO:0050717) |
| 0.1 | 0.7 | GO:0006020 | inositol metabolic process(GO:0006020) |
| 0.1 | 0.2 | GO:0071545 | inositol phosphate catabolic process(GO:0071545) |
| 0.1 | 1.6 | GO:0042026 | protein refolding(GO:0042026) |
| 0.1 | 0.8 | GO:0050930 | induction of positive chemotaxis(GO:0050930) |
| 0.1 | 2.8 | GO:0070098 | chemokine-mediated signaling pathway(GO:0070098) |
| 0.1 | 0.4 | GO:0006465 | signal peptide processing(GO:0006465) |
| 0.1 | 0.2 | GO:0044339 | canonical Wnt signaling pathway involved in osteoblast differentiation(GO:0044339) |
| 0.1 | 0.6 | GO:0098789 | pre-mRNA cleavage required for polyadenylation(GO:0098789) |
| 0.0 | 0.2 | GO:2001106 | regulation of Rho guanyl-nucleotide exchange factor activity(GO:2001106) |
| 0.0 | 0.4 | GO:0002097 | tRNA wobble base modification(GO:0002097) |
| 0.0 | 0.1 | GO:1903566 | positive regulation of protein localization to cilium(GO:1903566) |
| 0.0 | 1.1 | GO:0046839 | phospholipid dephosphorylation(GO:0046839) phosphatidylinositol dephosphorylation(GO:0046856) |
| 0.0 | 0.1 | GO:0006537 | glutamate biosynthetic process(GO:0006537) |
| 0.0 | 0.4 | GO:0006384 | transcription initiation from RNA polymerase III promoter(GO:0006384) |
| 0.0 | 1.5 | GO:0051170 | nuclear import(GO:0051170) |
| 0.0 | 1.8 | GO:0001558 | regulation of cell growth(GO:0001558) |
| 0.0 | 0.8 | GO:0046606 | negative regulation of centrosome cycle(GO:0046606) |
| 0.0 | 0.5 | GO:0042036 | negative regulation of cytokine biosynthetic process(GO:0042036) |
| 0.0 | 1.6 | GO:0031294 | lymphocyte costimulation(GO:0031294) T cell costimulation(GO:0031295) |
| 0.0 | 0.5 | GO:0035520 | monoubiquitinated protein deubiquitination(GO:0035520) |
| 0.0 | 0.4 | GO:0098915 | membrane repolarization during ventricular cardiac muscle cell action potential(GO:0098915) |
| 0.0 | 1.7 | GO:0001895 | retina homeostasis(GO:0001895) |
| 0.0 | 0.5 | GO:0045742 | positive regulation of epidermal growth factor receptor signaling pathway(GO:0045742) |
| 0.0 | 1.6 | GO:0032233 | positive regulation of actin filament bundle assembly(GO:0032233) |
| 0.0 | 0.9 | GO:0016254 | preassembly of GPI anchor in ER membrane(GO:0016254) |
| 0.0 | 0.3 | GO:0032020 | ISG15-protein conjugation(GO:0032020) |
| 0.0 | 1.5 | GO:0010107 | potassium ion import(GO:0010107) |
| 0.0 | 1.7 | GO:0035196 | production of miRNAs involved in gene silencing by miRNA(GO:0035196) |
| 0.0 | 0.9 | GO:0007212 | dopamine receptor signaling pathway(GO:0007212) |
| 0.0 | 0.2 | GO:0021759 | globus pallidus development(GO:0021759) |
| 0.0 | 0.2 | GO:0010212 | response to ionizing radiation(GO:0010212) |
| 0.0 | 0.2 | GO:0035590 | purinergic nucleotide receptor signaling pathway(GO:0035590) |
| 0.0 | 0.1 | GO:0008616 | queuosine biosynthetic process(GO:0008616) queuosine metabolic process(GO:0046116) |
| 0.0 | 0.6 | GO:0055026 | negative regulation of cardiac muscle tissue development(GO:0055026) |
| 0.0 | 0.9 | GO:0060512 | prostate gland morphogenesis(GO:0060512) |
| 0.0 | 1.1 | GO:0001938 | positive regulation of endothelial cell proliferation(GO:0001938) |
| 0.0 | 3.1 | GO:0060333 | interferon-gamma-mediated signaling pathway(GO:0060333) |
| 0.0 | 2.6 | GO:0031214 | biomineral tissue development(GO:0031214) |
| 0.0 | 1.9 | GO:0032922 | circadian regulation of gene expression(GO:0032922) |
| 0.0 | 0.3 | GO:0034498 | early endosome to Golgi transport(GO:0034498) |
| 0.0 | 1.6 | GO:2000649 | regulation of sodium ion transmembrane transporter activity(GO:2000649) |
| 0.0 | 0.5 | GO:0032515 | negative regulation of phosphoprotein phosphatase activity(GO:0032515) |
| 0.0 | 0.2 | GO:0016584 | nucleosome positioning(GO:0016584) |
| 0.0 | 0.8 | GO:0006641 | triglyceride metabolic process(GO:0006641) |
| 0.0 | 0.6 | GO:0032755 | positive regulation of interleukin-6 production(GO:0032755) |
| 0.0 | 0.9 | GO:0007548 | sex differentiation(GO:0007548) |
| 0.0 | 0.3 | GO:0048387 | negative regulation of retinoic acid receptor signaling pathway(GO:0048387) |
| 0.0 | 0.4 | GO:0000188 | inactivation of MAPK activity(GO:0000188) |
| 0.0 | 0.4 | GO:0032228 | regulation of synaptic transmission, GABAergic(GO:0032228) |
| 0.0 | 4.7 | GO:0006906 | vesicle fusion(GO:0006906) |
| 0.0 | 0.6 | GO:0001881 | receptor recycling(GO:0001881) |
| 0.0 | 0.6 | GO:0010762 | regulation of fibroblast migration(GO:0010762) |
| 0.0 | 0.4 | GO:0010447 | response to acidic pH(GO:0010447) |
| 0.0 | 0.5 | GO:0050729 | positive regulation of inflammatory response(GO:0050729) |
| 0.0 | 0.3 | GO:0033189 | response to vitamin A(GO:0033189) |
| 0.0 | 0.6 | GO:0007264 | small GTPase mediated signal transduction(GO:0007264) |
| 0.0 | 0.3 | GO:0051480 | regulation of cytosolic calcium ion concentration(GO:0051480) |
| 0.0 | 0.4 | GO:0030728 | ovulation(GO:0030728) |
| 0.0 | 1.0 | GO:0090503 | RNA phosphodiester bond hydrolysis, exonucleolytic(GO:0090503) |
| 0.0 | 0.3 | GO:0007519 | skeletal muscle tissue development(GO:0007519) skeletal muscle organ development(GO:0060538) |
| 0.0 | 0.7 | GO:0045087 | innate immune response(GO:0045087) |
| 0.0 | 0.5 | GO:0001755 | neural crest cell migration(GO:0001755) |
| 0.0 | 0.3 | GO:1901750 | leukotriene D4 metabolic process(GO:1901748) leukotriene D4 biosynthetic process(GO:1901750) |
| 0.0 | 0.7 | GO:0042632 | cholesterol homeostasis(GO:0042632) sterol homeostasis(GO:0055092) |
| 0.0 | 0.2 | GO:0050951 | detection of temperature stimulus(GO:0016048) sensory perception of temperature stimulus(GO:0050951) |
| 0.0 | 0.5 | GO:0002244 | hematopoietic progenitor cell differentiation(GO:0002244) |
| 0.0 | 0.6 | GO:0030517 | negative regulation of axon extension(GO:0030517) |
| 0.0 | 0.1 | GO:0071985 | multivesicular body sorting pathway(GO:0071985) |
| 0.0 | 0.4 | GO:0010172 | embryonic body morphogenesis(GO:0010172) |
| 0.0 | 0.8 | GO:1904837 | beta-catenin-TCF complex assembly(GO:1904837) |
| 0.0 | 0.5 | GO:0032411 | positive regulation of transporter activity(GO:0032411) |
| 0.0 | 0.3 | GO:0032780 | negative regulation of ATPase activity(GO:0032780) |
| 0.0 | 0.2 | GO:0060338 | regulation of type I interferon-mediated signaling pathway(GO:0060338) |
| 0.0 | 0.4 | GO:0042059 | negative regulation of epidermal growth factor receptor signaling pathway(GO:0042059) |
| 0.0 | 0.1 | GO:0030007 | cellular potassium ion homeostasis(GO:0030007) response to carbon monoxide(GO:0034465) smooth muscle contraction involved in micturition(GO:0060083) |
| 0.0 | 0.9 | GO:0048704 | embryonic skeletal system morphogenesis(GO:0048704) |
| 0.0 | 0.3 | GO:0030154 | cell differentiation(GO:0030154) |
| 0.0 | 0.4 | GO:0001958 | endochondral ossification(GO:0001958) replacement ossification(GO:0036075) |
| 0.0 | 0.4 | GO:0007585 | respiratory gaseous exchange(GO:0007585) |
| 0.0 | 0.1 | GO:0006336 | DNA replication-independent nucleosome assembly(GO:0006336) DNA replication-independent nucleosome organization(GO:0034724) |
Gene overrepresentation in cellular_component category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 1.9 | 5.8 | GO:0033150 | perinuclear theca(GO:0033011) cytoskeletal calyx(GO:0033150) |
| 1.9 | 13.1 | GO:0032437 | cuticular plate(GO:0032437) |
| 1.8 | 7.0 | GO:0031838 | haptoglobin-hemoglobin complex(GO:0031838) |
| 1.7 | 13.6 | GO:0032010 | phagolysosome(GO:0032010) |
| 1.5 | 19.7 | GO:0005577 | fibrinogen complex(GO:0005577) |
| 1.5 | 8.8 | GO:0031232 | extrinsic component of external side of plasma membrane(GO:0031232) |
| 1.3 | 3.9 | GO:0097679 | other organism cytoplasm(GO:0097679) |
| 1.2 | 3.6 | GO:0071751 | IgA immunoglobulin complex(GO:0071745) IgA immunoglobulin complex, circulating(GO:0071746) monomeric IgA immunoglobulin complex(GO:0071748) polymeric IgA immunoglobulin complex(GO:0071749) secretory IgA immunoglobulin complex(GO:0071751) |
| 1.2 | 5.9 | GO:0046696 | lipopolysaccharide receptor complex(GO:0046696) |
| 1.1 | 2.3 | GO:0005927 | muscle tendon junction(GO:0005927) |
| 1.1 | 23.8 | GO:0042613 | MHC class II protein complex(GO:0042613) |
| 1.1 | 7.4 | GO:0043240 | Fanconi anaemia nuclear complex(GO:0043240) |
| 1.0 | 12.5 | GO:0005833 | hemoglobin complex(GO:0005833) |
| 1.0 | 25.0 | GO:0042627 | chylomicron(GO:0042627) |
| 0.9 | 5.5 | GO:0034448 | EGO complex(GO:0034448) Gtr1-Gtr2 GTPase complex(GO:1990131) |
| 0.9 | 2.8 | GO:0035354 | Toll-like receptor 1-Toll-like receptor 2 protein complex(GO:0035354) |
| 0.9 | 0.9 | GO:1904949 | cation-transporting ATPase complex(GO:0090533) ATPase dependent transmembrane transport complex(GO:0098533) ATPase complex(GO:1904949) |
| 0.9 | 4.3 | GO:0005579 | membrane attack complex(GO:0005579) |
| 0.8 | 4.1 | GO:0005602 | complement component C1 complex(GO:0005602) |
| 0.8 | 37.1 | GO:0042571 | immunoglobulin complex, circulating(GO:0042571) |
| 0.8 | 3.1 | GO:0097224 | sperm connecting piece(GO:0097224) |
| 0.7 | 3.5 | GO:0000801 | central element(GO:0000801) |
| 0.7 | 2.8 | GO:0044530 | supraspliceosomal complex(GO:0044530) |
| 0.7 | 107.0 | GO:0072562 | blood microparticle(GO:0072562) |
| 0.6 | 4.5 | GO:0044194 | cytolytic granule(GO:0044194) |
| 0.6 | 0.6 | GO:0036019 | endolysosome(GO:0036019) endolysosome membrane(GO:0036020) |
| 0.6 | 4.4 | GO:0043203 | axon hillock(GO:0043203) |
| 0.5 | 6.9 | GO:0043020 | NADPH oxidase complex(GO:0043020) |
| 0.5 | 3.6 | GO:0097165 | nuclear stress granule(GO:0097165) |
| 0.5 | 1.0 | GO:0031314 | extrinsic component of mitochondrial inner membrane(GO:0031314) |
| 0.5 | 2.0 | GO:0030430 | host cell cytoplasm(GO:0030430) host cell cytoplasm part(GO:0033655) |
| 0.5 | 5.8 | GO:0005744 | mitochondrial inner membrane presequence translocase complex(GO:0005744) |
| 0.5 | 8.1 | GO:0033270 | paranode region of axon(GO:0033270) |
| 0.5 | 13.2 | GO:0034451 | centriolar satellite(GO:0034451) |
| 0.5 | 2.3 | GO:0034688 | integrin alphaM-beta2 complex(GO:0034688) |
| 0.5 | 2.3 | GO:0048787 | presynaptic active zone membrane(GO:0048787) |
| 0.5 | 1.4 | GO:0005583 | fibrillar collagen trimer(GO:0005583) banded collagen fibril(GO:0098643) |
| 0.4 | 1.3 | GO:0044423 | virion(GO:0019012) virion part(GO:0044423) |
| 0.4 | 7.6 | GO:0005736 | DNA-directed RNA polymerase I complex(GO:0005736) |
| 0.4 | 2.9 | GO:1990452 | Parkin-FBXW7-Cul1 ubiquitin ligase complex(GO:1990452) |
| 0.4 | 3.3 | GO:0045179 | apical cortex(GO:0045179) |
| 0.4 | 9.0 | GO:0031233 | intrinsic component of external side of plasma membrane(GO:0031233) |
| 0.4 | 5.6 | GO:0000788 | nuclear nucleosome(GO:0000788) |
| 0.4 | 11.4 | GO:0031092 | platelet alpha granule membrane(GO:0031092) |
| 0.4 | 1.1 | GO:0005879 | axonemal microtubule(GO:0005879) |
| 0.4 | 1.9 | GO:0005899 | insulin receptor complex(GO:0005899) |
| 0.4 | 2.6 | GO:0033269 | internode region of axon(GO:0033269) |
| 0.3 | 2.1 | GO:0097442 | CA3 pyramidal cell dendrite(GO:0097442) |
| 0.3 | 1.7 | GO:0005947 | mitochondrial alpha-ketoglutarate dehydrogenase complex(GO:0005947) |
| 0.3 | 5.7 | GO:0045277 | respiratory chain complex IV(GO:0045277) |
| 0.3 | 2.7 | GO:0014802 | terminal cisterna(GO:0014802) |
| 0.3 | 4.3 | GO:0001741 | XY body(GO:0001741) |
| 0.3 | 4.6 | GO:0005589 | collagen type VI trimer(GO:0005589) collagen beaded filament(GO:0098647) |
| 0.3 | 7.8 | GO:0005892 | acetylcholine-gated channel complex(GO:0005892) |
| 0.3 | 10.7 | GO:1990777 | plasma lipoprotein particle(GO:0034358) lipoprotein particle(GO:1990777) |
| 0.3 | 0.9 | GO:0008282 | ATP-sensitive potassium channel complex(GO:0008282) |
| 0.3 | 7.6 | GO:0032982 | myosin filament(GO:0032982) |
| 0.3 | 2.3 | GO:0016012 | sarcoglycan complex(GO:0016012) |
| 0.3 | 0.6 | GO:0030485 | smooth muscle contractile fiber(GO:0030485) |
| 0.3 | 4.4 | GO:0042622 | photoreceptor outer segment membrane(GO:0042622) |
| 0.3 | 0.3 | GO:0043541 | UDP-N-acetylglucosamine transferase complex(GO:0043541) |
| 0.3 | 1.6 | GO:0098560 | cytoplasmic side of late endosome membrane(GO:0098560) |
| 0.3 | 3.1 | GO:0032045 | guanyl-nucleotide exchange factor complex(GO:0032045) |
| 0.3 | 3.6 | GO:0005858 | axonemal dynein complex(GO:0005858) |
| 0.3 | 6.9 | GO:0044298 | neuronal cell body membrane(GO:0032809) cell body membrane(GO:0044298) |
| 0.3 | 3.0 | GO:0043190 | ATP-binding cassette (ABC) transporter complex(GO:0043190) |
| 0.2 | 2.5 | GO:0008024 | cyclin/CDK positive transcription elongation factor complex(GO:0008024) |
| 0.2 | 1.2 | GO:0033263 | CORVET complex(GO:0033263) |
| 0.2 | 1.4 | GO:0001673 | male germ cell nucleus(GO:0001673) |
| 0.2 | 3.7 | GO:0097386 | glial cell projection(GO:0097386) |
| 0.2 | 2.3 | GO:1990712 | HFE-transferrin receptor complex(GO:1990712) |
| 0.2 | 3.4 | GO:0005662 | DNA replication factor A complex(GO:0005662) |
| 0.2 | 1.1 | GO:0097179 | protease inhibitor complex(GO:0097179) |
| 0.2 | 1.5 | GO:0070554 | synaptobrevin 2-SNAP-25-syntaxin-3-complexin complex(GO:0070554) |
| 0.2 | 0.8 | GO:1990031 | pinceau fiber(GO:1990031) |
| 0.2 | 2.6 | GO:0097524 | sperm plasma membrane(GO:0097524) |
| 0.2 | 16.3 | GO:1904724 | tertiary granule lumen(GO:1904724) |
| 0.2 | 1.3 | GO:0046581 | intercellular canaliculus(GO:0046581) |
| 0.2 | 2.7 | GO:0043194 | axon initial segment(GO:0043194) |
| 0.2 | 1.5 | GO:0097504 | Gemini of coiled bodies(GO:0097504) |
| 0.2 | 17.8 | GO:0035578 | azurophil granule lumen(GO:0035578) |
| 0.2 | 9.2 | GO:0001533 | cornified envelope(GO:0001533) |
| 0.2 | 0.7 | GO:0005726 | perichromatin fibrils(GO:0005726) |
| 0.2 | 1.5 | GO:0061574 | ASAP complex(GO:0061574) |
| 0.2 | 1.0 | GO:0009331 | glycerol-3-phosphate dehydrogenase complex(GO:0009331) |
| 0.2 | 2.6 | GO:0032300 | mismatch repair complex(GO:0032300) |
| 0.2 | 4.3 | GO:0090665 | dystrophin-associated glycoprotein complex(GO:0016010) glycoprotein complex(GO:0090665) |
| 0.2 | 4.1 | GO:0005779 | integral component of peroxisomal membrane(GO:0005779) intrinsic component of peroxisomal membrane(GO:0031231) |
| 0.1 | 6.4 | GO:0048786 | presynaptic active zone(GO:0048786) |
| 0.1 | 3.0 | GO:0031430 | M band(GO:0031430) |
| 0.1 | 2.5 | GO:0031235 | intrinsic component of the cytoplasmic side of the plasma membrane(GO:0031235) |
| 0.1 | 0.9 | GO:0044754 | autolysosome(GO:0044754) |
| 0.1 | 1.7 | GO:0030285 | integral component of synaptic vesicle membrane(GO:0030285) intrinsic component of synaptic vesicle membrane(GO:0098563) |
| 0.1 | 2.4 | GO:0000242 | pericentriolar material(GO:0000242) |
| 0.1 | 2.2 | GO:0044224 | juxtaparanode region of axon(GO:0044224) |
| 0.1 | 7.8 | GO:0005902 | microvillus(GO:0005902) |
| 0.1 | 1.8 | GO:0032585 | multivesicular body membrane(GO:0032585) |
| 0.1 | 15.4 | GO:0031225 | anchored component of membrane(GO:0031225) |
| 0.1 | 2.5 | GO:0034663 | endoplasmic reticulum chaperone complex(GO:0034663) |
| 0.1 | 181.7 | GO:0005615 | extracellular space(GO:0005615) |
| 0.1 | 1.6 | GO:0097025 | MPP7-DLG1-LIN7 complex(GO:0097025) |
| 0.1 | 15.7 | GO:0043204 | perikaryon(GO:0043204) |
| 0.1 | 1.6 | GO:0031083 | BLOC-1 complex(GO:0031083) |
| 0.1 | 9.2 | GO:0016323 | basolateral plasma membrane(GO:0016323) |
| 0.1 | 18.6 | GO:0009897 | external side of plasma membrane(GO:0009897) |
| 0.1 | 4.4 | GO:0031093 | platelet alpha granule lumen(GO:0031093) |
| 0.1 | 6.2 | GO:0016235 | aggresome(GO:0016235) |
| 0.1 | 3.3 | GO:0042101 | T cell receptor complex(GO:0042101) |
| 0.1 | 1.4 | GO:0097440 | apical dendrite(GO:0097440) |
| 0.1 | 2.7 | GO:0097381 | photoreceptor disc membrane(GO:0097381) |
| 0.1 | 2.0 | GO:0005922 | connexon complex(GO:0005922) |
| 0.1 | 1.2 | GO:0001518 | voltage-gated sodium channel complex(GO:0001518) |
| 0.1 | 1.1 | GO:0001527 | microfibril(GO:0001527) fibril(GO:0043205) |
| 0.1 | 1.1 | GO:0000815 | ESCRT III complex(GO:0000815) |
| 0.1 | 3.2 | GO:0005680 | anaphase-promoting complex(GO:0005680) |
| 0.1 | 2.1 | GO:0005849 | mRNA cleavage factor complex(GO:0005849) |
| 0.1 | 3.2 | GO:0005581 | collagen trimer(GO:0005581) |
| 0.1 | 1.4 | GO:0005885 | Arp2/3 protein complex(GO:0005885) |
| 0.1 | 0.6 | GO:0034098 | VCP-NPL4-UFD1 AAA ATPase complex(GO:0034098) |
| 0.1 | 2.1 | GO:0008074 | guanylate cyclase complex, soluble(GO:0008074) |
| 0.1 | 0.7 | GO:0005869 | dynactin complex(GO:0005869) |
| 0.1 | 0.8 | GO:0032039 | integrator complex(GO:0032039) |
| 0.1 | 3.0 | GO:0005891 | voltage-gated calcium channel complex(GO:0005891) |
| 0.1 | 1.9 | GO:0031672 | A band(GO:0031672) |
| 0.1 | 0.7 | GO:0030868 | smooth endoplasmic reticulum membrane(GO:0030868) smooth endoplasmic reticulum part(GO:0097425) |
| 0.1 | 0.7 | GO:0031931 | TORC1 complex(GO:0031931) |
| 0.1 | 0.8 | GO:0044666 | MLL3/4 complex(GO:0044666) |
| 0.1 | 2.8 | GO:0005719 | nuclear euchromatin(GO:0005719) |
| 0.1 | 0.9 | GO:0034992 | microtubule organizing center attachment site(GO:0034992) LINC complex(GO:0034993) |
| 0.1 | 0.5 | GO:0042788 | polysomal ribosome(GO:0042788) |
| 0.1 | 7.6 | GO:0005814 | centriole(GO:0005814) |
| 0.1 | 0.5 | GO:0002177 | manchette(GO:0002177) |
| 0.1 | 0.2 | GO:0070876 | SOSS complex(GO:0070876) |
| 0.1 | 1.3 | GO:0032420 | stereocilium(GO:0032420) |
| 0.1 | 5.4 | GO:0008076 | voltage-gated potassium channel complex(GO:0008076) potassium channel complex(GO:0034705) |
| 0.1 | 0.2 | GO:0031213 | RSF complex(GO:0031213) |
| 0.1 | 0.7 | GO:0017146 | NMDA selective glutamate receptor complex(GO:0017146) |
| 0.1 | 0.5 | GO:0070419 | nonhomologous end joining complex(GO:0070419) |
| 0.1 | 0.4 | GO:0000127 | transcription factor TFIIIC complex(GO:0000127) |
| 0.1 | 5.6 | GO:0035579 | specific granule membrane(GO:0035579) |
| 0.1 | 0.4 | GO:0005787 | signal peptidase complex(GO:0005787) |
| 0.1 | 1.6 | GO:0042734 | presynaptic membrane(GO:0042734) |
| 0.1 | 7.6 | GO:0005578 | proteinaceous extracellular matrix(GO:0005578) |
| 0.1 | 3.0 | GO:0009295 | nucleoid(GO:0009295) mitochondrial nucleoid(GO:0042645) |
| 0.1 | 1.5 | GO:0005801 | cis-Golgi network(GO:0005801) |
| 0.1 | 0.3 | GO:0071437 | invadopodium(GO:0071437) |
| 0.0 | 1.2 | GO:1902711 | GABA-A receptor complex(GO:1902711) |
| 0.0 | 1.7 | GO:0043198 | dendritic shaft(GO:0043198) |
| 0.0 | 1.9 | GO:0034707 | chloride channel complex(GO:0034707) |
| 0.0 | 5.0 | GO:0045211 | postsynaptic membrane(GO:0045211) |
| 0.0 | 0.5 | GO:1990454 | L-type voltage-gated calcium channel complex(GO:1990454) |
| 0.0 | 0.3 | GO:0042406 | extrinsic component of endoplasmic reticulum membrane(GO:0042406) |
| 0.0 | 1.8 | GO:0031526 | brush border membrane(GO:0031526) |
| 0.0 | 0.4 | GO:0000243 | commitment complex(GO:0000243) |
| 0.0 | 1.5 | GO:0000421 | autophagosome membrane(GO:0000421) |
| 0.0 | 6.0 | GO:0031234 | extrinsic component of cytoplasmic side of plasma membrane(GO:0031234) |
| 0.0 | 1.9 | GO:0015030 | Cajal body(GO:0015030) |
| 0.0 | 0.3 | GO:0000120 | RNA polymerase I transcription factor complex(GO:0000120) |
| 0.0 | 1.0 | GO:0036064 | ciliary basal body(GO:0036064) |
| 0.0 | 0.6 | GO:0031904 | endosome lumen(GO:0031904) |
| 0.0 | 2.2 | GO:0031901 | early endosome membrane(GO:0031901) |
| 0.0 | 3.1 | GO:0005769 | early endosome(GO:0005769) |
| 0.0 | 40.4 | GO:0031226 | intrinsic component of plasma membrane(GO:0031226) |
| 0.0 | 0.2 | GO:0008385 | IkappaB kinase complex(GO:0008385) |
| 0.0 | 2.1 | GO:0099572 | postsynaptic density(GO:0014069) excitatory synapse(GO:0060076) postsynaptic specialization(GO:0099572) |
| 0.0 | 0.4 | GO:0098576 | lumenal side of membrane(GO:0098576) |
| 0.0 | 0.1 | GO:0031417 | NatC complex(GO:0031417) |
| 0.0 | 0.7 | GO:0043195 | terminal bouton(GO:0043195) |
| 0.0 | 59.7 | GO:0016021 | integral component of membrane(GO:0016021) |
| 0.0 | 0.6 | GO:0000786 | nucleosome(GO:0000786) |
| 0.0 | 0.3 | GO:0033202 | Ino80 complex(GO:0031011) DNA helicase complex(GO:0033202) |
Gene overrepresentation in molecular_function category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 3.7 | 11.0 | GO:0098519 | nucleotide phosphatase activity, acting on free nucleotides(GO:0098519) |
| 3.4 | 17.2 | GO:0060230 | lipoprotein lipase activator activity(GO:0060230) |
| 2.6 | 10.6 | GO:0042132 | fructose 1,6-bisphosphate 1-phosphatase activity(GO:0042132) |
| 2.6 | 7.9 | GO:0016495 | C-X3-C chemokine receptor activity(GO:0016495) |
| 2.4 | 12.0 | GO:0017159 | pantetheine hydrolase activity(GO:0017159) |
| 2.4 | 7.1 | GO:0004088 | carbamoyl-phosphate synthase (ammonia) activity(GO:0004087) carbamoyl-phosphate synthase (glutamine-hydrolyzing) activity(GO:0004088) |
| 2.2 | 6.7 | GO:0005137 | interleukin-5 receptor binding(GO:0005137) |
| 2.1 | 6.4 | GO:1904455 | ubiquitin-specific protease activity involved in negative regulation of ERAD pathway(GO:1904455) |
| 2.1 | 14.6 | GO:0047894 | flavonol 3-sulfotransferase activity(GO:0047894) |
| 2.0 | 6.1 | GO:0004797 | deoxycytidine kinase activity(GO:0004137) thymidine kinase activity(GO:0004797) |
| 2.0 | 8.0 | GO:0004031 | aldehyde oxidase activity(GO:0004031) |
| 2.0 | 7.8 | GO:1904492 | Ac-Asp-Glu binding(GO:1904492) tetrahydrofolyl-poly(glutamate) polymer binding(GO:1904493) |
| 2.0 | 15.6 | GO:0004064 | arylesterase activity(GO:0004064) |
| 1.9 | 11.7 | GO:0070892 | lipoteichoic acid receptor activity(GO:0070892) |
| 1.9 | 1.9 | GO:0071723 | lipopeptide binding(GO:0071723) |
| 1.7 | 5.2 | GO:0050528 | acyloxyacyl hydrolase activity(GO:0050528) |
| 1.7 | 12.0 | GO:0015562 | efflux transmembrane transporter activity(GO:0015562) |
| 1.6 | 7.8 | GO:0004995 | tachykinin receptor activity(GO:0004995) |
| 1.5 | 4.6 | GO:0070546 | L-phenylalanine aminotransferase activity(GO:0070546) L-phenylalanine:2-oxoglutarate aminotransferase activity(GO:0080130) |
| 1.4 | 2.8 | GO:0035663 | Toll-like receptor 2 binding(GO:0035663) |
| 1.4 | 13.6 | GO:0016176 | superoxide-generating NADPH oxidase activator activity(GO:0016176) |
| 1.4 | 4.1 | GO:0061663 | NEDD8 ligase activity(GO:0061663) |
| 1.3 | 4.0 | GO:0035662 | Toll-like receptor 4 binding(GO:0035662) |
| 1.3 | 8.0 | GO:0003917 | DNA topoisomerase type I activity(GO:0003917) |
| 1.3 | 3.9 | GO:0016019 | peptidoglycan receptor activity(GO:0016019) |
| 1.3 | 5.2 | GO:0004657 | proline dehydrogenase activity(GO:0004657) |
| 1.2 | 7.4 | GO:0009378 | four-way junction helicase activity(GO:0009378) |
| 1.2 | 8.6 | GO:0030345 | extracellular matrix structural constituent conferring compression resistance(GO:0030021) structural constituent of tooth enamel(GO:0030345) |
| 1.2 | 6.0 | GO:0048248 | CXCR3 chemokine receptor binding(GO:0048248) |
| 1.2 | 6.0 | GO:0032554 | purine deoxyribonucleotide binding(GO:0032554) |
| 1.2 | 4.8 | GO:0019862 | IgA binding(GO:0019862) |
| 1.2 | 7.0 | GO:0030492 | hemoglobin binding(GO:0030492) |
| 1.1 | 12.5 | GO:0005344 | oxygen transporter activity(GO:0005344) |
| 1.1 | 2.2 | GO:0047718 | androsterone dehydrogenase activity(GO:0047023) indanol dehydrogenase activity(GO:0047718) |
| 1.1 | 4.3 | GO:0001855 | complement component C4b binding(GO:0001855) |
| 1.1 | 4.2 | GO:0052798 | beta-galactoside alpha-2,3-sialyltransferase activity(GO:0052798) |
| 1.1 | 19.0 | GO:0008329 | signaling pattern recognition receptor activity(GO:0008329) pattern recognition receptor activity(GO:0038187) |
| 1.0 | 39.6 | GO:0015020 | glucuronosyltransferase activity(GO:0015020) |
| 1.0 | 3.1 | GO:0032767 | copper-dependent protein binding(GO:0032767) |
| 1.0 | 3.0 | GO:0016838 | carbon-oxygen lyase activity, acting on phosphates(GO:0016838) |
| 1.0 | 8.0 | GO:0034235 | GPI anchor binding(GO:0034235) |
| 1.0 | 3.0 | GO:0070326 | very-low-density lipoprotein particle receptor binding(GO:0070326) |
| 1.0 | 4.0 | GO:0019770 | IgG receptor activity(GO:0019770) |
| 1.0 | 1.0 | GO:0030627 | pre-mRNA 5'-splice site binding(GO:0030627) |
| 0.9 | 3.7 | GO:0016211 | glutamate-ammonia ligase activity(GO:0004356) ammonia ligase activity(GO:0016211) acid-ammonia (or amide) ligase activity(GO:0016880) |
| 0.9 | 2.7 | GO:0001181 | transcription factor activity, core RNA polymerase I binding(GO:0001181) |
| 0.9 | 2.7 | GO:0004802 | transketolase activity(GO:0004802) |
| 0.9 | 207.1 | GO:0003823 | antigen binding(GO:0003823) |
| 0.9 | 3.5 | GO:0051916 | granulocyte colony-stimulating factor binding(GO:0051916) |
| 0.9 | 11.3 | GO:0019957 | C-C chemokine binding(GO:0019957) |
| 0.9 | 3.5 | GO:0004060 | arylamine N-acetyltransferase activity(GO:0004060) |
| 0.8 | 2.5 | GO:0004833 | tryptophan 2,3-dioxygenase activity(GO:0004833) |
| 0.8 | 2.5 | GO:0008798 | beta-aspartyl-peptidase activity(GO:0008798) |
| 0.8 | 5.0 | GO:0004522 | ribonuclease A activity(GO:0004522) |
| 0.8 | 4.2 | GO:0005152 | interleukin-1 receptor antagonist activity(GO:0005152) |
| 0.8 | 4.1 | GO:0004692 | cGMP-dependent protein kinase activity(GO:0004692) |
| 0.8 | 4.0 | GO:0015204 | urea transmembrane transporter activity(GO:0015204) |
| 0.8 | 2.4 | GO:0016992 | lipoate synthase activity(GO:0016992) radical SAM enzyme activity(GO:0070283) |
| 0.8 | 3.1 | GO:0071987 | WD40-repeat domain binding(GO:0071987) |
| 0.8 | 4.7 | GO:0035800 | deubiquitinase activator activity(GO:0035800) |
| 0.8 | 3.8 | GO:0004468 | lysine N-acetyltransferase activity, acting on acetyl phosphate as donor(GO:0004468) |
| 0.7 | 2.2 | GO:0032093 | SAM domain binding(GO:0032093) |
| 0.7 | 2.9 | GO:0015137 | citrate transmembrane transporter activity(GO:0015137) tricarboxylic acid transmembrane transporter activity(GO:0015142) |
| 0.7 | 2.2 | GO:0001665 | alpha-N-acetylgalactosaminide alpha-2,6-sialyltransferase activity(GO:0001665) |
| 0.7 | 2.2 | GO:0003980 | UDP-glucose:glycoprotein glucosyltransferase activity(GO:0003980) |
| 0.7 | 2.2 | GO:0033300 | dehydroascorbic acid transporter activity(GO:0033300) |
| 0.7 | 3.6 | GO:0097199 | cysteine-type endopeptidase activity involved in apoptotic signaling pathway(GO:0097199) |
| 0.7 | 2.1 | GO:0071558 | histone demethylase activity (H3-K27 specific)(GO:0071558) |
| 0.7 | 4.2 | GO:0000293 | ferric-chelate reductase activity(GO:0000293) |
| 0.7 | 2.8 | GO:0017128 | phospholipid scramblase activity(GO:0017128) |
| 0.7 | 4.9 | GO:0060228 | phosphatidylcholine-sterol O-acyltransferase activator activity(GO:0060228) |
| 0.7 | 6.9 | GO:0001730 | 2'-5'-oligoadenylate synthetase activity(GO:0001730) |
| 0.7 | 2.1 | GO:0090556 | apolipoprotein A-I receptor activity(GO:0034188) phosphatidylserine-translocating ATPase activity(GO:0090556) |
| 0.6 | 3.2 | GO:0047844 | deoxycytidine deaminase activity(GO:0047844) |
| 0.6 | 1.9 | GO:0015403 | thiamine transmembrane transporter activity(GO:0015234) thiamine uptake transmembrane transporter activity(GO:0015403) uptake transmembrane transporter activity(GO:0015563) |
| 0.6 | 3.6 | GO:0045029 | UDP-activated nucleotide receptor activity(GO:0045029) |
| 0.6 | 1.8 | GO:0032089 | NACHT domain binding(GO:0032089) |
| 0.6 | 1.8 | GO:0008332 | low voltage-gated calcium channel activity(GO:0008332) |
| 0.6 | 4.7 | GO:0015349 | thyroid hormone transmembrane transporter activity(GO:0015349) |
| 0.6 | 2.3 | GO:0004165 | dodecenoyl-CoA delta-isomerase activity(GO:0004165) |
| 0.6 | 6.4 | GO:0045028 | G-protein coupled nucleotide receptor activity(GO:0001608) G-protein coupled purinergic nucleotide receptor activity(GO:0045028) |
| 0.6 | 1.8 | GO:0015440 | peptide-transporting ATPase activity(GO:0015440) |
| 0.6 | 1.7 | GO:0005549 | odorant binding(GO:0005549) |
| 0.6 | 9.2 | GO:0015643 | toxic substance binding(GO:0015643) |
| 0.6 | 1.7 | GO:0003826 | alpha-ketoacid dehydrogenase activity(GO:0003826) 3-methyl-2-oxobutanoate dehydrogenase (2-methylpropanoyl-transferring) activity(GO:0003863) |
| 0.6 | 1.7 | GO:0003829 | beta-1,3-galactosyl-O-glycosyl-glycoprotein beta-1,6-N-acetylglucosaminyltransferase activity(GO:0003829) |
| 0.6 | 1.7 | GO:0019961 | interferon binding(GO:0019961) |
| 0.5 | 4.9 | GO:0047499 | calcium-independent phospholipase A2 activity(GO:0047499) |
| 0.5 | 2.7 | GO:0046912 | transferase activity, transferring acyl groups, acyl groups converted into alkyl on transfer(GO:0046912) |
| 0.5 | 2.1 | GO:0004803 | transposase activity(GO:0004803) |
| 0.5 | 2.1 | GO:0022850 | serotonin-gated cation channel activity(GO:0022850) |
| 0.5 | 8.3 | GO:0030169 | low-density lipoprotein particle binding(GO:0030169) |
| 0.5 | 1.5 | GO:0005350 | pyrimidine nucleobase transmembrane transporter activity(GO:0005350) pyrimidine- and adenine-specific:sodium symporter activity(GO:0015389) |
| 0.5 | 5.1 | GO:0005094 | Rho GDP-dissociation inhibitor activity(GO:0005094) |
| 0.5 | 6.1 | GO:0001758 | retinal dehydrogenase activity(GO:0001758) |
| 0.5 | 1.5 | GO:0070052 | collagen V binding(GO:0070052) |
| 0.5 | 3.4 | GO:0031726 | CCR1 chemokine receptor binding(GO:0031726) |
| 0.5 | 2.9 | GO:0008467 | [heparan sulfate]-glucosamine 3-sulfotransferase 1 activity(GO:0008467) |
| 0.5 | 1.9 | GO:0005502 | 11-cis retinal binding(GO:0005502) |
| 0.5 | 2.4 | GO:0060961 | phospholipase D inhibitor activity(GO:0060961) |
| 0.5 | 2.3 | GO:0001851 | complement component C3b binding(GO:0001851) |
| 0.5 | 5.0 | GO:0045236 | CXCR chemokine receptor binding(GO:0045236) |
| 0.5 | 22.2 | GO:0004984 | olfactory receptor activity(GO:0004984) |
| 0.5 | 1.4 | GO:0031731 | CCR6 chemokine receptor binding(GO:0031731) |
| 0.4 | 1.3 | GO:0004947 | bradykinin receptor activity(GO:0004947) |
| 0.4 | 2.2 | GO:0001010 | transcription factor activity, sequence-specific DNA binding transcription factor recruiting(GO:0001010) |
| 0.4 | 3.9 | GO:0004321 | fatty-acyl-CoA synthase activity(GO:0004321) |
| 0.4 | 1.7 | GO:0004980 | melanocyte-stimulating hormone receptor activity(GO:0004980) |
| 0.4 | 1.7 | GO:0031871 | proteinase activated receptor binding(GO:0031871) |
| 0.4 | 7.6 | GO:0001054 | RNA polymerase I activity(GO:0001054) |
| 0.4 | 9.9 | GO:0043395 | heparan sulfate proteoglycan binding(GO:0043395) |
| 0.4 | 5.3 | GO:0043855 | intracellular cyclic nucleotide activated cation channel activity(GO:0005221) cyclic nucleotide-gated ion channel activity(GO:0043855) |
| 0.4 | 6.9 | GO:0016175 | superoxide-generating NADPH oxidase activity(GO:0016175) |
| 0.4 | 6.8 | GO:0070330 | aromatase activity(GO:0070330) |
| 0.4 | 1.2 | GO:0052871 | tocopherol omega-hydroxylase activity(GO:0052870) alpha-tocopherol omega-hydroxylase activity(GO:0052871) 20-hydroxy-leukotriene B4 omega oxidase activity(GO:0097258) 20-aldehyde-leukotriene B4 20-monooxygenase activity(GO:0097259) |
| 0.4 | 7.5 | GO:0005248 | voltage-gated sodium channel activity(GO:0005248) voltage-gated ion channel activity involved in regulation of postsynaptic membrane potential(GO:1905030) |
| 0.4 | 2.4 | GO:0000155 | phosphorelay sensor kinase activity(GO:0000155) |
| 0.4 | 1.6 | GO:0019763 | immunoglobulin receptor activity(GO:0019763) |
| 0.4 | 2.0 | GO:0001594 | trace-amine receptor activity(GO:0001594) |
| 0.4 | 1.6 | GO:0016712 | oxidoreductase activity, acting on paired donors, with incorporation or reduction of molecular oxygen, reduced flavin or flavoprotein as one donor, and incorporation of one atom of oxygen(GO:0016712) |
| 0.4 | 40.1 | GO:0004867 | serine-type endopeptidase inhibitor activity(GO:0004867) |
| 0.4 | 5.2 | GO:0016813 | hydrolase activity, acting on carbon-nitrogen (but not peptide) bonds, in linear amidines(GO:0016813) |
| 0.4 | 1.1 | GO:0004382 | guanosine-diphosphatase activity(GO:0004382) |
| 0.4 | 9.5 | GO:0004181 | metallocarboxypeptidase activity(GO:0004181) |
| 0.4 | 2.9 | GO:0050816 | phosphothreonine binding(GO:0050816) |
| 0.4 | 1.1 | GO:0017129 | triglyceride binding(GO:0017129) |
| 0.4 | 1.1 | GO:0031727 | CCR2 chemokine receptor binding(GO:0031727) |
| 0.3 | 2.1 | GO:0003873 | 6-phosphofructo-2-kinase activity(GO:0003873) fructose-2,6-bisphosphate 2-phosphatase activity(GO:0004331) |
| 0.3 | 3.8 | GO:0005436 | sodium:phosphate symporter activity(GO:0005436) |
| 0.3 | 1.7 | GO:0016907 | G-protein coupled acetylcholine receptor activity(GO:0016907) |
| 0.3 | 2.0 | GO:0005087 | Ran guanyl-nucleotide exchange factor activity(GO:0005087) |
| 0.3 | 1.7 | GO:0001848 | complement binding(GO:0001848) |
| 0.3 | 3.3 | GO:0003696 | satellite DNA binding(GO:0003696) |
| 0.3 | 3.6 | GO:0004571 | mannosyl-oligosaccharide 1,2-alpha-mannosidase activity(GO:0004571) |
| 0.3 | 3.3 | GO:0003720 | telomerase activity(GO:0003720) RNA-directed DNA polymerase activity(GO:0003964) |
| 0.3 | 1.3 | GO:0008336 | gamma-butyrobetaine dioxygenase activity(GO:0008336) |
| 0.3 | 2.9 | GO:0004499 | N,N-dimethylaniline monooxygenase activity(GO:0004499) |
| 0.3 | 3.5 | GO:0031419 | cobalamin binding(GO:0031419) |
| 0.3 | 0.3 | GO:0009008 | DNA-methyltransferase activity(GO:0009008) |
| 0.3 | 5.6 | GO:0008409 | 5'-3' exonuclease activity(GO:0008409) |
| 0.3 | 7.1 | GO:0097493 | structural molecule activity conferring elasticity(GO:0097493) |
| 0.3 | 2.5 | GO:0097322 | 7SK snRNA binding(GO:0097322) |
| 0.3 | 1.2 | GO:0055131 | C3HC4-type RING finger domain binding(GO:0055131) |
| 0.3 | 3.1 | GO:0032027 | myosin light chain binding(GO:0032027) |
| 0.3 | 5.1 | GO:0003906 | DNA-(apurinic or apyrimidinic site) lyase activity(GO:0003906) |
| 0.3 | 0.9 | GO:0018636 | phenanthrene 9,10-monooxygenase activity(GO:0018636) ketosteroid monooxygenase activity(GO:0047086) trans-1,2-dihydrobenzene-1,2-diol dehydrogenase activity(GO:0047115) |
| 0.3 | 2.1 | GO:0032453 | histone demethylase activity (H3-K4 specific)(GO:0032453) |
| 0.3 | 16.9 | GO:0004601 | peroxidase activity(GO:0004601) |
| 0.3 | 0.9 | GO:0001596 | angiotensin type I receptor activity(GO:0001596) |
| 0.3 | 5.7 | GO:0022848 | acetylcholine-gated cation channel activity(GO:0022848) |
| 0.3 | 1.7 | GO:0005499 | vitamin D binding(GO:0005499) |
| 0.3 | 2.5 | GO:0070915 | lysophosphatidic acid receptor activity(GO:0070915) |
| 0.3 | 2.5 | GO:1901702 | urate transmembrane transporter activity(GO:0015143) salt transmembrane transporter activity(GO:1901702) |
| 0.3 | 0.8 | GO:0034041 | sterol-transporting ATPase activity(GO:0034041) |
| 0.3 | 2.3 | GO:0005025 | transforming growth factor beta receptor activity, type I(GO:0005025) |
| 0.3 | 2.3 | GO:0003876 | AMP deaminase activity(GO:0003876) adenosine-phosphate deaminase activity(GO:0047623) |
| 0.3 | 0.8 | GO:0047275 | glucosaminylgalactosylglucosylceramide beta-galactosyltransferase activity(GO:0047275) |
| 0.3 | 0.8 | GO:0003976 | UDP-N-acetylglucosamine-lysosomal-enzyme N-acetylglucosaminephosphotransferase activity(GO:0003976) |
| 0.3 | 1.0 | GO:0061714 | folic acid receptor activity(GO:0061714) |
| 0.3 | 2.5 | GO:0016713 | oxidoreductase activity, acting on paired donors, with incorporation or reduction of molecular oxygen, reduced iron-sulfur protein as one donor, and incorporation of one atom of oxygen(GO:0016713) |
| 0.2 | 1.0 | GO:0052591 | sn-glycerol-3-phosphate:ubiquinone oxidoreductase activity(GO:0052590) sn-glycerol-3-phosphate:ubiquinone-8 oxidoreductase activity(GO:0052591) |
| 0.2 | 1.2 | GO:0016300 | tRNA (uracil) methyltransferase activity(GO:0016300) |
| 0.2 | 1.0 | GO:0050262 | ribosylnicotinamide kinase activity(GO:0050262) ribosylnicotinate kinase activity(GO:0061769) |
| 0.2 | 0.7 | GO:0001016 | transcription factor activity, RNA polymerase III transcription factor binding(GO:0001007) RNA polymerase III regulatory region DNA binding(GO:0001016) |
| 0.2 | 0.7 | GO:0004769 | 3-beta-hydroxy-delta5-steroid dehydrogenase activity(GO:0003854) steroid delta-isomerase activity(GO:0004769) |
| 0.2 | 0.9 | GO:0008417 | fucosyltransferase activity(GO:0008417) |
| 0.2 | 6.0 | GO:0016646 | oxidoreductase activity, acting on the CH-NH group of donors, NAD or NADP as acceptor(GO:0016646) |
| 0.2 | 0.9 | GO:0042030 | ATPase inhibitor activity(GO:0042030) |
| 0.2 | 0.9 | GO:0004465 | lipoprotein lipase activity(GO:0004465) |
| 0.2 | 0.7 | GO:0031877 | somatostatin receptor binding(GO:0031877) |
| 0.2 | 2.0 | GO:0039706 | co-receptor binding(GO:0039706) |
| 0.2 | 0.7 | GO:0019863 | IgE binding(GO:0019863) |
| 0.2 | 4.2 | GO:0016805 | dipeptidase activity(GO:0016805) |
| 0.2 | 0.9 | GO:0004376 | glycolipid mannosyltransferase activity(GO:0004376) |
| 0.2 | 2.6 | GO:0019911 | structural constituent of myelin sheath(GO:0019911) |
| 0.2 | 3.2 | GO:0004653 | polypeptide N-acetylgalactosaminyltransferase activity(GO:0004653) |
| 0.2 | 0.9 | GO:0016160 | amylase activity(GO:0016160) |
| 0.2 | 1.7 | GO:0034711 | inhibin binding(GO:0034711) |
| 0.2 | 1.1 | GO:1990430 | extracellular matrix protein binding(GO:1990430) |
| 0.2 | 3.1 | GO:0005243 | gap junction channel activity(GO:0005243) |
| 0.2 | 0.6 | GO:0005148 | prolactin receptor binding(GO:0005148) |
| 0.2 | 4.5 | GO:0004012 | phospholipid-translocating ATPase activity(GO:0004012) |
| 0.2 | 2.2 | GO:0008429 | phosphatidylethanolamine binding(GO:0008429) |
| 0.2 | 4.8 | GO:0019198 | transmembrane receptor protein tyrosine phosphatase activity(GO:0005001) transmembrane receptor protein phosphatase activity(GO:0019198) |
| 0.2 | 1.8 | GO:0032051 | clathrin light chain binding(GO:0032051) |
| 0.2 | 2.6 | GO:0030280 | structural constituent of epidermis(GO:0030280) |
| 0.2 | 5.7 | GO:0047555 | 3',5'-cyclic-GMP phosphodiesterase activity(GO:0047555) |
| 0.2 | 0.8 | GO:0032217 | riboflavin transporter activity(GO:0032217) |
| 0.2 | 1.6 | GO:0046920 | alpha-(1->3)-fucosyltransferase activity(GO:0046920) |
| 0.2 | 1.4 | GO:0030020 | extracellular matrix structural constituent conferring tensile strength(GO:0030020) |
| 0.2 | 3.6 | GO:0005068 | transmembrane receptor protein tyrosine kinase adaptor activity(GO:0005068) |
| 0.2 | 6.7 | GO:0031624 | ubiquitin conjugating enzyme binding(GO:0031624) |
| 0.2 | 5.5 | GO:0003951 | NAD+ kinase activity(GO:0003951) |
| 0.2 | 0.8 | GO:0016499 | orexin receptor activity(GO:0016499) |
| 0.2 | 2.3 | GO:0032393 | MHC class I receptor activity(GO:0032393) |
| 0.2 | 0.6 | GO:0016296 | oleoyl-[acyl-carrier-protein] hydrolase activity(GO:0004320) myristoyl-[acyl-carrier-protein] hydrolase activity(GO:0016295) palmitoyl-[acyl-carrier-protein] hydrolase activity(GO:0016296) acyl-[acyl-carrier-protein] hydrolase activity(GO:0016297) |
| 0.2 | 10.0 | GO:0001784 | phosphotyrosine binding(GO:0001784) |
| 0.2 | 1.8 | GO:0047498 | calcium-dependent phospholipase A2 activity(GO:0047498) |
| 0.2 | 3.6 | GO:0043325 | phosphatidylinositol-3,4-bisphosphate binding(GO:0043325) |
| 0.2 | 7.0 | GO:0005044 | scavenger receptor activity(GO:0005044) |
| 0.2 | 0.7 | GO:0004126 | cytidine deaminase activity(GO:0004126) |
| 0.2 | 0.5 | GO:0004605 | phosphatidate cytidylyltransferase activity(GO:0004605) |
| 0.2 | 0.7 | GO:0005220 | inositol 1,4,5-trisphosphate-sensitive calcium-release channel activity(GO:0005220) |
| 0.2 | 2.4 | GO:0015245 | fatty acid transporter activity(GO:0015245) very long-chain fatty acid-CoA ligase activity(GO:0031957) |
| 0.2 | 0.8 | GO:0051880 | G-quadruplex DNA binding(GO:0051880) |
| 0.2 | 2.1 | GO:0015279 | store-operated calcium channel activity(GO:0015279) |
| 0.2 | 4.5 | GO:0008200 | ion channel inhibitor activity(GO:0008200) channel inhibitor activity(GO:0016248) |
| 0.2 | 1.1 | GO:0051864 | histone demethylase activity (H3-K36 specific)(GO:0051864) |
| 0.2 | 4.5 | GO:0017075 | syntaxin-1 binding(GO:0017075) |
| 0.1 | 2.7 | GO:0015467 | G-protein activated inward rectifier potassium channel activity(GO:0015467) |
| 0.1 | 0.4 | GO:0008195 | phosphatidate phosphatase activity(GO:0008195) |
| 0.1 | 1.0 | GO:0050682 | AF-2 domain binding(GO:0050682) |
| 0.1 | 0.9 | GO:0010997 | anaphase-promoting complex binding(GO:0010997) |
| 0.1 | 2.1 | GO:0010857 | calcium-dependent protein serine/threonine kinase activity(GO:0009931) calcium-dependent protein kinase activity(GO:0010857) |
| 0.1 | 0.7 | GO:0033857 | diphosphoinositol-pentakisphosphate kinase activity(GO:0033857) |
| 0.1 | 2.5 | GO:0035591 | signaling adaptor activity(GO:0035591) |
| 0.1 | 1.0 | GO:0015186 | L-glutamine transmembrane transporter activity(GO:0015186) |
| 0.1 | 2.6 | GO:0030983 | mismatched DNA binding(GO:0030983) |
| 0.1 | 0.4 | GO:0005540 | hyaluronic acid binding(GO:0005540) |
| 0.1 | 3.0 | GO:0004115 | 3',5'-cyclic-AMP phosphodiesterase activity(GO:0004115) |
| 0.1 | 3.4 | GO:0004708 | MAP kinase kinase activity(GO:0004708) |
| 0.1 | 0.3 | GO:0008192 | RNA guanylyltransferase activity(GO:0008192) |
| 0.1 | 2.5 | GO:0004535 | poly(A)-specific ribonuclease activity(GO:0004535) |
| 0.1 | 1.6 | GO:0019864 | IgG binding(GO:0019864) |
| 0.1 | 1.5 | GO:0015379 | potassium:chloride symporter activity(GO:0015379) potassium ion symporter activity(GO:0022820) |
| 0.1 | 1.2 | GO:0004969 | histamine receptor activity(GO:0004969) |
| 0.1 | 2.0 | GO:0005092 | GDP-dissociation inhibitor activity(GO:0005092) |
| 0.1 | 3.0 | GO:0008191 | metalloendopeptidase inhibitor activity(GO:0008191) |
| 0.1 | 1.7 | GO:0022851 | GABA-gated chloride ion channel activity(GO:0022851) |
| 0.1 | 1.0 | GO:0004957 | prostaglandin E receptor activity(GO:0004957) |
| 0.1 | 0.5 | GO:0005477 | pyruvate secondary active transmembrane transporter activity(GO:0005477) |
| 0.1 | 0.4 | GO:0003858 | 3-hydroxybutyrate dehydrogenase activity(GO:0003858) |
| 0.1 | 1.5 | GO:0017034 | Rap guanyl-nucleotide exchange factor activity(GO:0017034) |
| 0.1 | 0.8 | GO:0015232 | heme transporter activity(GO:0015232) |
| 0.1 | 1.5 | GO:0034185 | apolipoprotein binding(GO:0034185) |
| 0.1 | 0.8 | GO:0004977 | melanocortin receptor activity(GO:0004977) |
| 0.1 | 1.8 | GO:0015101 | organic cation transmembrane transporter activity(GO:0015101) |
| 0.1 | 0.9 | GO:0015385 | sodium:proton antiporter activity(GO:0015385) potassium:proton antiporter activity(GO:0015386) |
| 0.1 | 0.8 | GO:0001517 | N-acetylglucosamine 6-O-sulfotransferase activity(GO:0001517) |
| 0.1 | 1.9 | GO:0043522 | leucine zipper domain binding(GO:0043522) |
| 0.1 | 1.6 | GO:0005132 | type I interferon receptor binding(GO:0005132) |
| 0.1 | 1.3 | GO:0034056 | estrogen response element binding(GO:0034056) |
| 0.1 | 5.3 | GO:0004869 | cysteine-type endopeptidase inhibitor activity(GO:0004869) |
| 0.1 | 2.5 | GO:0045503 | dynein light chain binding(GO:0045503) |
| 0.1 | 0.9 | GO:0004198 | calcium-dependent cysteine-type endopeptidase activity(GO:0004198) |
| 0.1 | 4.1 | GO:0043539 | protein serine/threonine kinase activator activity(GO:0043539) |
| 0.1 | 5.9 | GO:0017112 | Rab guanyl-nucleotide exchange factor activity(GO:0017112) |
| 0.1 | 0.7 | GO:0051033 | nucleic acid transmembrane transporter activity(GO:0051032) RNA transmembrane transporter activity(GO:0051033) |
| 0.1 | 3.0 | GO:0004385 | guanylate kinase activity(GO:0004385) |
| 0.1 | 26.6 | GO:0030674 | protein binding, bridging(GO:0030674) |
| 0.1 | 0.3 | GO:0004914 | interleukin-5 receptor activity(GO:0004914) |
| 0.1 | 2.3 | GO:0005095 | GTPase inhibitor activity(GO:0005095) |
| 0.1 | 2.6 | GO:0070182 | DNA polymerase binding(GO:0070182) |
| 0.1 | 1.2 | GO:0099589 | serotonin receptor activity(GO:0099589) |
| 0.1 | 0.4 | GO:0004346 | glucose-6-phosphatase activity(GO:0004346) sugar-terminal-phosphatase activity(GO:0050309) |
| 0.1 | 0.4 | GO:0016635 | oxidoreductase activity, acting on the CH-CH group of donors, quinone or related compound as acceptor(GO:0016635) |
| 0.1 | 0.6 | GO:0051525 | NFAT protein binding(GO:0051525) |
| 0.1 | 0.5 | GO:0034046 | poly(G) binding(GO:0034046) |
| 0.1 | 1.4 | GO:0050786 | RAGE receptor binding(GO:0050786) |
| 0.1 | 3.4 | GO:0042379 | chemokine receptor binding(GO:0042379) |
| 0.1 | 1.6 | GO:0004062 | aryl sulfotransferase activity(GO:0004062) |
| 0.1 | 1.4 | GO:0003785 | actin monomer binding(GO:0003785) |
| 0.1 | 1.5 | GO:0004806 | triglyceride lipase activity(GO:0004806) |
| 0.1 | 0.9 | GO:0004767 | sphingomyelin phosphodiesterase activity(GO:0004767) |
| 0.1 | 10.6 | GO:0004222 | metalloendopeptidase activity(GO:0004222) |
| 0.1 | 1.0 | GO:0089720 | caspase binding(GO:0089720) |
| 0.1 | 1.1 | GO:0052658 | inositol-1,4,5-trisphosphate 5-phosphatase activity(GO:0052658) |
| 0.1 | 1.0 | GO:0055056 | D-glucose transmembrane transporter activity(GO:0055056) |
| 0.1 | 2.1 | GO:0004383 | guanylate cyclase activity(GO:0004383) |
| 0.1 | 1.7 | GO:0031434 | mitogen-activated protein kinase kinase binding(GO:0031434) |
| 0.1 | 4.8 | GO:0050840 | extracellular matrix binding(GO:0050840) |
| 0.1 | 0.3 | GO:0044323 | retinoic acid-responsive element binding(GO:0044323) |
| 0.1 | 0.9 | GO:0008020 | G-protein coupled photoreceptor activity(GO:0008020) |
| 0.1 | 0.6 | GO:0008271 | secondary active sulfate transmembrane transporter activity(GO:0008271) |
| 0.1 | 2.7 | GO:0008139 | nuclear localization sequence binding(GO:0008139) |
| 0.1 | 4.4 | GO:0001105 | RNA polymerase II transcription coactivator activity(GO:0001105) |
| 0.1 | 1.1 | GO:0046934 | phosphatidylinositol-4,5-bisphosphate 3-kinase activity(GO:0046934) |
| 0.1 | 1.9 | GO:0034237 | protein kinase A regulatory subunit binding(GO:0034237) |
| 0.1 | 1.5 | GO:0051400 | BH domain binding(GO:0051400) |
| 0.1 | 25.1 | GO:0030695 | GTPase regulator activity(GO:0030695) |
| 0.1 | 2.2 | GO:0005212 | structural constituent of eye lens(GO:0005212) |
| 0.1 | 0.9 | GO:0043225 | anion transmembrane-transporting ATPase activity(GO:0043225) |
| 0.1 | 0.5 | GO:0086056 | voltage-gated calcium channel activity involved in AV node cell action potential(GO:0086056) |
| 0.1 | 1.9 | GO:0004033 | aldo-keto reductase (NADP) activity(GO:0004033) |
| 0.1 | 1.4 | GO:0000900 | translation repressor activity, nucleic acid binding(GO:0000900) |
| 0.1 | 0.7 | GO:0005432 | calcium:sodium antiporter activity(GO:0005432) |
| 0.1 | 0.1 | GO:0016639 | oxidoreductase activity, acting on the CH-NH2 group of donors, NAD or NADP as acceptor(GO:0016639) |
| 0.1 | 0.9 | GO:0097371 | MDM2/MDM4 family protein binding(GO:0097371) |
| 0.1 | 1.1 | GO:0046933 | proton-transporting ATP synthase activity, rotational mechanism(GO:0046933) |
| 0.1 | 0.4 | GO:0032052 | bile acid binding(GO:0032052) |
| 0.1 | 0.3 | GO:0004974 | leukotriene receptor activity(GO:0004974) |
| 0.1 | 0.3 | GO:0052740 | 1-acyl-2-lysophosphatidylserine acylhydrolase activity(GO:0052740) |
| 0.1 | 0.9 | GO:0060229 | lipase activator activity(GO:0060229) |
| 0.1 | 0.7 | GO:0042056 | chemoattractant activity(GO:0042056) |
| 0.1 | 0.4 | GO:1902282 | voltage-gated potassium channel activity involved in ventricular cardiac muscle cell action potential repolarization(GO:1902282) |
| 0.1 | 0.5 | GO:0035259 | glucocorticoid receptor binding(GO:0035259) |
| 0.1 | 0.8 | GO:0004065 | arylsulfatase activity(GO:0004065) |
| 0.1 | 10.1 | GO:0004252 | serine-type endopeptidase activity(GO:0004252) |
| 0.1 | 0.3 | GO:0042296 | ISG15 transferase activity(GO:0042296) |
| 0.1 | 1.5 | GO:0043014 | alpha-tubulin binding(GO:0043014) |
| 0.1 | 11.6 | GO:0005088 | Ras guanyl-nucleotide exchange factor activity(GO:0005088) |
| 0.1 | 1.2 | GO:0004890 | GABA-A receptor activity(GO:0004890) |
| 0.1 | 1.2 | GO:0005158 | insulin receptor binding(GO:0005158) |
| 0.1 | 1.3 | GO:0001190 | transcriptional activator activity, RNA polymerase II transcription factor binding(GO:0001190) transcriptional repressor activity, RNA polymerase II activating transcription factor binding(GO:0098811) |
| 0.0 | 0.9 | GO:0035198 | miRNA binding(GO:0035198) |
| 0.0 | 1.9 | GO:0051879 | Hsp90 protein binding(GO:0051879) |
| 0.0 | 2.1 | GO:0017080 | sodium channel regulator activity(GO:0017080) |
| 0.0 | 3.2 | GO:0004879 | RNA polymerase II transcription factor activity, ligand-activated sequence-specific DNA binding(GO:0004879) transcription factor activity, direct ligand regulated sequence-specific DNA binding(GO:0098531) |
| 0.0 | 0.6 | GO:0016018 | cyclosporin A binding(GO:0016018) |
| 0.0 | 0.8 | GO:0042800 | histone methyltransferase activity (H3-K4 specific)(GO:0042800) |
| 0.0 | 0.2 | GO:0052847 | endopolyphosphatase activity(GO:0000298) diphosphoinositol-polyphosphate diphosphatase activity(GO:0008486) bis(5'-adenosyl)-hexaphosphatase activity(GO:0034431) bis(5'-adenosyl)-pentaphosphatase activity(GO:0034432) inositol diphosphate tetrakisphosphate diphosphatase activity(GO:0052840) inositol bisdiphosphate tetrakisphosphate diphosphatase activity(GO:0052841) inositol diphosphate pentakisphosphate diphosphatase activity(GO:0052842) inositol-1-diphosphate-2,3,4,5,6-pentakisphosphate diphosphatase activity(GO:0052843) inositol-3-diphosphate-1,2,4,5,6-pentakisphosphate diphosphatase activity(GO:0052844) inositol-5-diphosphate-1,2,3,4,6-pentakisphosphate diphosphatase activity(GO:0052845) inositol-1,5-bisdiphosphate-2,3,4,6-tetrakisphosphate 1-diphosphatase activity(GO:0052846) inositol-1,5-bisdiphosphate-2,3,4,6-tetrakisphosphate 5-diphosphatase activity(GO:0052847) inositol-3,5-bisdiphosphate-2,3,4,6-tetrakisphosphate 5-diphosphatase activity(GO:0052848) |
| 0.0 | 1.2 | GO:0015002 | cytochrome-c oxidase activity(GO:0004129) heme-copper terminal oxidase activity(GO:0015002) oxidoreductase activity, acting on a heme group of donors, oxygen as acceptor(GO:0016676) |
| 0.0 | 1.5 | GO:0030276 | clathrin binding(GO:0030276) |
| 0.0 | 1.1 | GO:0016409 | palmitoyltransferase activity(GO:0016409) |
| 0.0 | 2.2 | GO:0030170 | pyridoxal phosphate binding(GO:0030170) |
| 0.0 | 1.6 | GO:0050699 | WW domain binding(GO:0050699) |
| 0.0 | 0.5 | GO:0051787 | misfolded protein binding(GO:0051787) |
| 0.0 | 0.7 | GO:0003995 | acyl-CoA dehydrogenase activity(GO:0003995) |
| 0.0 | 0.1 | GO:0004558 | alpha-1,4-glucosidase activity(GO:0004558) oligo-1,6-glucosidase activity(GO:0004574) |
| 0.0 | 1.9 | GO:0005254 | chloride channel activity(GO:0005254) |
| 0.0 | 0.4 | GO:0001206 | transcriptional repressor activity, RNA polymerase II distal enhancer sequence-specific binding(GO:0001206) |
| 0.0 | 1.5 | GO:0004402 | histone acetyltransferase activity(GO:0004402) peptide-lysine-N-acetyltransferase activity(GO:0061733) |
| 0.0 | 0.2 | GO:0003796 | lysozyme activity(GO:0003796) |
| 0.0 | 1.4 | GO:0070412 | R-SMAD binding(GO:0070412) |
| 0.0 | 1.0 | GO:0016410 | N-acyltransferase activity(GO:0016410) |
| 0.0 | 0.6 | GO:0004550 | nucleoside diphosphate kinase activity(GO:0004550) |
| 0.0 | 0.2 | GO:0016493 | C-C chemokine receptor activity(GO:0016493) |
| 0.0 | 0.5 | GO:0005246 | calcium channel regulator activity(GO:0005246) |
| 0.0 | 0.6 | GO:0003707 | steroid hormone receptor activity(GO:0003707) |
| 0.0 | 0.2 | GO:0031491 | nucleosome binding(GO:0031491) |
| 0.0 | 0.3 | GO:0005537 | mannose binding(GO:0005537) |
| 0.0 | 1.0 | GO:0004715 | non-membrane spanning protein tyrosine kinase activity(GO:0004715) |
| 0.0 | 0.1 | GO:0008479 | queuine tRNA-ribosyltransferase activity(GO:0008479) |
| 0.0 | 0.3 | GO:0036374 | glutathione hydrolase activity(GO:0036374) |
| 0.0 | 0.1 | GO:0035473 | lipase binding(GO:0035473) |
| 0.0 | 0.7 | GO:0004364 | glutathione transferase activity(GO:0004364) |
| 0.0 | 0.2 | GO:0048018 | receptor agonist activity(GO:0048018) |
| 0.0 | 0.2 | GO:0016502 | purinergic nucleotide receptor activity(GO:0001614) nucleotide receptor activity(GO:0016502) |
| 0.0 | 1.9 | GO:0004713 | protein tyrosine kinase activity(GO:0004713) |
| 0.0 | 24.7 | GO:0003700 | nucleic acid binding transcription factor activity(GO:0001071) transcription factor activity, sequence-specific DNA binding(GO:0003700) |
| 0.0 | 0.3 | GO:0051959 | dynein light intermediate chain binding(GO:0051959) |
| 0.0 | 0.6 | GO:0019209 | kinase activator activity(GO:0019209) |
| 0.0 | 0.3 | GO:0000146 | microfilament motor activity(GO:0000146) |
| 0.0 | 1.7 | GO:0005178 | integrin binding(GO:0005178) |
| 0.0 | 0.1 | GO:0060072 | large conductance calcium-activated potassium channel activity(GO:0060072) |
| 0.0 | 0.4 | GO:0015144 | carbohydrate transmembrane transporter activity(GO:0015144) carbohydrate transporter activity(GO:1901476) |
| 0.0 | 0.1 | GO:0008454 | alpha-1,3-mannosylglycoprotein 4-beta-N-acetylglucosaminyltransferase activity(GO:0008454) |
Gene overrepresentation in C2:CP category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.6 | 20.1 | PID TOLL ENDOGENOUS PATHWAY | Endogenous TLR signaling |
| 0.4 | 20.0 | PID INTEGRIN2 PATHWAY | Beta2 integrin cell surface interactions |
| 0.4 | 10.0 | SA G1 AND S PHASES | Cdk2, 4, and 6 bind cyclin D in G1, while cdk2/cyclin E promotes the G1/S transition. |
| 0.3 | 2.6 | PID KIT PATHWAY | Signaling events mediated by Stem cell factor receptor (c-Kit) |
| 0.3 | 82.6 | NABA ECM REGULATORS | Genes encoding enzymes and their regulators involved in the remodeling of the extracellular matrix |
| 0.3 | 27.1 | PID RAC1 PATHWAY | RAC1 signaling pathway |
| 0.3 | 7.0 | PID IL5 PATHWAY | IL5-mediated signaling events |
| 0.3 | 21.1 | PID HNF3B PATHWAY | FOXA2 and FOXA3 transcription factor networks |
| 0.3 | 33.9 | PID CMYB PATHWAY | C-MYB transcription factor network |
| 0.3 | 1.9 | PID IL6 7 PATHWAY | IL6-mediated signaling events |
| 0.2 | 12.9 | NABA PROTEOGLYCANS | Genes encoding proteoglycans |
| 0.2 | 9.8 | PID P38 ALPHA BETA PATHWAY | Regulation of p38-alpha and p38-beta |
| 0.2 | 19.1 | PID IL12 2PATHWAY | IL12-mediated signaling events |
| 0.2 | 0.9 | PID AR NONGENOMIC PATHWAY | Nongenotropic Androgen signaling |
| 0.2 | 1.7 | ST TYPE I INTERFERON PATHWAY | Type I Interferon (alpha/beta IFN) Pathway |
| 0.2 | 32.3 | NABA ECM AFFILIATED | Genes encoding proteins affiliated structurally or functionally to extracellular matrix proteins |
| 0.2 | 4.1 | ST G ALPHA S PATHWAY | G alpha s Pathway |
| 0.2 | 5.7 | PID RHODOPSIN PATHWAY | Visual signal transduction: Rods |
| 0.2 | 0.6 | PID S1P S1P4 PATHWAY | S1P4 pathway |
| 0.2 | 0.6 | SA G2 AND M PHASES | Cdc25 activates the cdc2/cyclin B complex to induce the G2/M transition. |
| 0.1 | 1.6 | PID TCR JNK PATHWAY | JNK signaling in the CD4+ TCR pathway |
| 0.1 | 6.8 | PID ALPHA SYNUCLEIN PATHWAY | Alpha-synuclein signaling |
| 0.1 | 1.4 | PID CONE PATHWAY | Visual signal transduction: Cones |
| 0.1 | 2.2 | PID ANGIOPOIETIN RECEPTOR PATHWAY | Angiopoietin receptor Tie2-mediated signaling |
| 0.1 | 3.7 | PID INTEGRIN A9B1 PATHWAY | Alpha9 beta1 integrin signaling events |
| 0.1 | 1.1 | PID ANTHRAX PATHWAY | Cellular roles of Anthrax toxin |
| 0.1 | 0.9 | PID IL27 PATHWAY | IL27-mediated signaling events |
| 0.1 | 2.3 | PID EPO PATHWAY | EPO signaling pathway |
| 0.1 | 2.9 | PID IFNG PATHWAY | IFN-gamma pathway |
| 0.1 | 2.9 | PID EPHRINB REV PATHWAY | Ephrin B reverse signaling |
| 0.1 | 1.9 | PID INSULIN PATHWAY | Insulin Pathway |
| 0.1 | 4.7 | PID ERBB2 ERBB3 PATHWAY | ErbB2/ErbB3 signaling events |
| 0.1 | 3.3 | PID IL8 CXCR2 PATHWAY | IL8- and CXCR2-mediated signaling events |
| 0.1 | 6.1 | PID PTP1B PATHWAY | Signaling events mediated by PTP1B |
| 0.1 | 2.5 | PID LYSOPHOSPHOLIPID PATHWAY | LPA receptor mediated events |
| 0.1 | 3.8 | PID PI3KCI PATHWAY | Class I PI3K signaling events |
| 0.1 | 2.7 | PID CXCR3 PATHWAY | CXCR3-mediated signaling events |
| 0.1 | 2.6 | PID HEDGEHOG 2PATHWAY | Signaling events mediated by the Hedgehog family |
| 0.1 | 6.1 | PID IL4 2PATHWAY | IL4-mediated signaling events |
| 0.1 | 2.2 | SA PTEN PATHWAY | PTEN is a tumor suppressor that dephosphorylates the lipid messenger phosphatidylinositol triphosphate. |
| 0.1 | 0.8 | PID ARF6 DOWNSTREAM PATHWAY | Arf6 downstream pathway |
| 0.1 | 5.2 | PID FANCONI PATHWAY | Fanconi anemia pathway |
| 0.1 | 2.4 | SA CASPASE CASCADE | Apoptosis is mediated by caspases, cysteine proteases arranged in a proteolytic cascade. |
| 0.1 | 2.1 | PID SHP2 PATHWAY | SHP2 signaling |
| 0.1 | 2.3 | PID HDAC CLASSIII PATHWAY | Signaling events mediated by HDAC Class III |
| 0.1 | 3.0 | PID RXR VDR PATHWAY | RXR and RAR heterodimerization with other nuclear receptor |
| 0.1 | 1.3 | PID IL2 1PATHWAY | IL2-mediated signaling events |
| 0.1 | 16.3 | NABA ECM GLYCOPROTEINS | Genes encoding structural ECM glycoproteins |
| 0.1 | 1.3 | PID CIRCADIAN PATHWAY | Circadian rhythm pathway |
| 0.1 | 3.2 | PID AR TF PATHWAY | Regulation of Androgen receptor activity |
| 0.1 | 2.1 | PID LKB1 PATHWAY | LKB1 signaling events |
| 0.1 | 0.7 | ST P38 MAPK PATHWAY | p38 MAPK Pathway |
| 0.0 | 0.9 | PID AP1 PATHWAY | AP-1 transcription factor network |
| 0.0 | 1.9 | PID RHOA PATHWAY | RhoA signaling pathway |
| 0.0 | 1.6 | PID CD8 TCR DOWNSTREAM PATHWAY | Downstream signaling in naïve CD8+ T cells |
| 0.0 | 2.5 | PID BMP PATHWAY | BMP receptor signaling |
| 0.0 | 1.0 | PID GLYPICAN 1PATHWAY | Glypican 1 network |
| 0.0 | 0.7 | ST WNT CA2 CYCLIC GMP PATHWAY | Wnt/Ca2+/cyclic GMP signaling. |
| 0.0 | 2.6 | PID NOTCH PATHWAY | Notch signaling pathway |
| 0.0 | 0.6 | PID ER NONGENOMIC PATHWAY | Plasma membrane estrogen receptor signaling |
| 0.0 | 0.9 | PID MAPK TRK PATHWAY | Trk receptor signaling mediated by the MAPK pathway |
| 0.0 | 1.6 | PID RAC1 REG PATHWAY | Regulation of RAC1 activity |
| 0.0 | 0.6 | PID ENDOTHELIN PATHWAY | Endothelins |
| 0.0 | 0.7 | PID BCR 5PATHWAY | BCR signaling pathway |
| 0.0 | 1.8 | PID SMAD2 3NUCLEAR PATHWAY | Regulation of nuclear SMAD2/3 signaling |
| 0.0 | 7.5 | NABA SECRETED FACTORS | Genes encoding secreted soluble factors |
| 0.0 | 0.1 | SA PROGRAMMED CELL DEATH | Programmed cell death, or apoptosis, eliminates damaged or unneeded cells. |
Gene overrepresentation in C2:CP:REACTOME category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 1.8 | 47.7 | REACTOME GLUCURONIDATION | Genes involved in Glucuronidation |
| 1.4 | 6.8 | REACTOME BINDING AND ENTRY OF HIV VIRION | Genes involved in Binding and entry of HIV virion |
| 1.0 | 15.7 | REACTOME REVERSIBLE HYDRATION OF CARBON DIOXIDE | Genes involved in Reversible Hydration of Carbon Dioxide |
| 0.8 | 5.9 | REACTOME TRAF6 MEDIATED INDUCTION OF TAK1 COMPLEX | Genes involved in TRAF6 mediated induction of TAK1 complex |
| 0.8 | 6.7 | REACTOME ETHANOL OXIDATION | Genes involved in Ethanol oxidation |
| 0.8 | 8.2 | REACTOME IRAK2 MEDIATED ACTIVATION OF TAK1 COMPLEX UPON TLR7 8 OR 9 STIMULATION | Genes involved in IRAK2 mediated activation of TAK1 complex upon TLR7/8 or 9 stimulation |
| 0.7 | 18.1 | REACTOME HDL MEDIATED LIPID TRANSPORT | Genes involved in HDL-mediated lipid transport |
| 0.7 | 25.0 | REACTOME GLUTATHIONE CONJUGATION | Genes involved in Glutathione conjugation |
| 0.7 | 6.8 | REACTOME BETA DEFENSINS | Genes involved in Beta defensins |
| 0.7 | 10.6 | REACTOME COMMON PATHWAY | Genes involved in Common Pathway |
| 0.7 | 22.2 | REACTOME INTERACTION BETWEEN L1 AND ANKYRINS | Genes involved in Interaction between L1 and Ankyrins |
| 0.6 | 16.1 | REACTOME CYTOSOLIC SULFONATION OF SMALL MOLECULES | Genes involved in Cytosolic sulfonation of small molecules |
| 0.6 | 10.7 | REACTOME P2Y RECEPTORS | Genes involved in P2Y receptors |
| 0.5 | 25.1 | REACTOME COMPLEMENT CASCADE | Genes involved in Complement cascade |
| 0.5 | 29.0 | REACTOME CHEMOKINE RECEPTORS BIND CHEMOKINES | Genes involved in Chemokine receptors bind chemokines |
| 0.5 | 7.9 | REACTOME INTRINSIC PATHWAY | Genes involved in Intrinsic Pathway |
| 0.5 | 7.8 | REACTOME PRESYNAPTIC NICOTINIC ACETYLCHOLINE RECEPTORS | Genes involved in Presynaptic nicotinic acetylcholine receptors |
| 0.5 | 8.2 | REACTOME DOPAMINE NEUROTRANSMITTER RELEASE CYCLE | Genes involved in Dopamine Neurotransmitter Release Cycle |
| 0.4 | 8.4 | REACTOME TERMINATION OF O GLYCAN BIOSYNTHESIS | Genes involved in Termination of O-glycan biosynthesis |
| 0.4 | 22.1 | REACTOME OLFACTORY SIGNALING PATHWAY | Genes involved in Olfactory Signaling Pathway |
| 0.4 | 9.9 | REACTOME FANCONI ANEMIA PATHWAY | Genes involved in Fanconi Anemia pathway |
| 0.4 | 1.2 | REACTOME SYNTHESIS OF PA | Genes involved in Synthesis of PA |
| 0.4 | 9.7 | REACTOME REGULATION OF INSULIN LIKE GROWTH FACTOR IGF ACTIVITY BY INSULIN LIKE GROWTH FACTOR BINDING PROTEINS IGFBPS | Genes involved in Regulation of Insulin-like Growth Factor (IGF) Activity by Insulin-like Growth Factor Binding Proteins (IGFBPs) |
| 0.4 | 2.7 | REACTOME DEFENSINS | Genes involved in Defensins |
| 0.4 | 5.9 | REACTOME LIPOPROTEIN METABOLISM | Genes involved in Lipoprotein metabolism |
| 0.3 | 11.8 | REACTOME ABC FAMILY PROTEINS MEDIATED TRANSPORT | Genes involved in ABC-family proteins mediated transport |
| 0.3 | 8.9 | REACTOME CS DS DEGRADATION | Genes involved in CS/DS degradation |
| 0.3 | 6.2 | REACTOME AKT PHOSPHORYLATES TARGETS IN THE CYTOSOL | Genes involved in AKT phosphorylates targets in the cytosol |
| 0.3 | 5.8 | REACTOME XENOBIOTICS | Genes involved in Xenobiotics |
| 0.3 | 6.4 | REACTOME ACTIVATED TAK1 MEDIATES P38 MAPK ACTIVATION | Genes involved in activated TAK1 mediates p38 MAPK activation |
| 0.3 | 7.1 | REACTOME ACYL CHAIN REMODELLING OF PE | Genes involved in Acyl chain remodelling of PE |
| 0.3 | 4.1 | REACTOME NFKB ACTIVATION THROUGH FADD RIP1 PATHWAY MEDIATED BY CASPASE 8 AND10 | Genes involved in NF-kB activation through FADD/RIP-1 pathway mediated by caspase-8 and -10 |
| 0.3 | 11.3 | REACTOME CGMP EFFECTS | Genes involved in cGMP effects |
| 0.3 | 1.7 | REACTOME RECYCLING OF BILE ACIDS AND SALTS | Genes involved in Recycling of bile acids and salts |
| 0.3 | 3.3 | REACTOME ROLE OF DCC IN REGULATING APOPTOSIS | Genes involved in Role of DCC in regulating apoptosis |
| 0.2 | 10.6 | REACTOME RNA POL I TRANSCRIPTION INITIATION | Genes involved in RNA Polymerase I Transcription Initiation |
| 0.2 | 3.7 | REACTOME BILE SALT AND ORGANIC ANION SLC TRANSPORTERS | Genes involved in Bile salt and organic anion SLC transporters |
| 0.2 | 2.8 | REACTOME ELEVATION OF CYTOSOLIC CA2 LEVELS | Genes involved in Elevation of cytosolic Ca2+ levels |
| 0.2 | 7.5 | REACTOME IL RECEPTOR SHC SIGNALING | Genes involved in Interleukin receptor SHC signaling |
| 0.2 | 3.0 | REACTOME SEROTONIN RECEPTORS | Genes involved in Serotonin receptors |
| 0.2 | 12.1 | REACTOME RNA POL I PROMOTER OPENING | Genes involved in RNA Polymerase I Promoter Opening |
| 0.2 | 1.5 | REACTOME FORMATION OF RNA POL II ELONGATION COMPLEX | Genes involved in Formation of RNA Pol II elongation complex |
| 0.2 | 2.1 | REACTOME SYNTHESIS OF BILE ACIDS AND BILE SALTS VIA 24 HYDROXYCHOLESTEROL | Genes involved in Synthesis of bile acids and bile salts via 24-hydroxycholesterol |
| 0.2 | 2.8 | REACTOME DOWNSTREAM TCR SIGNALING | Genes involved in Downstream TCR signaling |
| 0.2 | 2.5 | REACTOME PEPTIDE LIGAND BINDING RECEPTORS | Genes involved in Peptide ligand-binding receptors |
| 0.2 | 2.1 | REACTOME NITRIC OXIDE STIMULATES GUANYLATE CYCLASE | Genes involved in Nitric oxide stimulates guanylate cyclase |
| 0.2 | 6.3 | REACTOME TRAF6 MEDIATED IRF7 ACTIVATION | Genes involved in TRAF6 mediated IRF7 activation |
| 0.2 | 4.0 | REACTOME FACILITATIVE NA INDEPENDENT GLUCOSE TRANSPORTERS | Genes involved in Facilitative Na+-independent glucose transporters |
| 0.2 | 5.7 | REACTOME KERATAN SULFATE BIOSYNTHESIS | Genes involved in Keratan sulfate biosynthesis |
| 0.2 | 5.4 | REACTOME MHC CLASS II ANTIGEN PRESENTATION | Genes involved in MHC class II antigen presentation |
| 0.2 | 1.4 | REACTOME TRAF6 MEDIATED IRF7 ACTIVATION IN TLR7 8 OR 9 SIGNALING | Genes involved in TRAF6 mediated IRF7 activation in TLR7/8 or 9 signaling |
| 0.2 | 2.4 | REACTOME PASSIVE TRANSPORT BY AQUAPORINS | Genes involved in Passive Transport by Aquaporins |
| 0.2 | 2.5 | REACTOME PLATELET CALCIUM HOMEOSTASIS | Genes involved in Platelet calcium homeostasis |
| 0.2 | 3.6 | REACTOME TRANSLOCATION OF ZAP 70 TO IMMUNOLOGICAL SYNAPSE | Genes involved in Translocation of ZAP-70 to Immunological synapse |
| 0.2 | 7.1 | REACTOME LATENT INFECTION OF HOMO SAPIENS WITH MYCOBACTERIUM TUBERCULOSIS | Genes involved in Latent infection of Homo sapiens with Mycobacterium tuberculosis |
| 0.2 | 5.4 | REACTOME REGULATION OF SIGNALING BY CBL | Genes involved in Regulation of signaling by CBL |
| 0.1 | 2.2 | REACTOME SPRY REGULATION OF FGF SIGNALING | Genes involved in Spry regulation of FGF signaling |
| 0.1 | 6.6 | REACTOME VOLTAGE GATED POTASSIUM CHANNELS | Genes involved in Voltage gated Potassium channels |
| 0.1 | 1.2 | REACTOME SIGNAL ATTENUATION | Genes involved in Signal attenuation |
| 0.1 | 2.7 | REACTOME INHIBITION OF VOLTAGE GATED CA2 CHANNELS VIA GBETA GAMMA SUBUNITS | Genes involved in Inhibition of voltage gated Ca2+ channels via Gbeta/gamma subunits |
| 0.1 | 1.8 | REACTOME APOPTOTIC CLEAVAGE OF CELL ADHESION PROTEINS | Genes involved in Apoptotic cleavage of cell adhesion proteins |
| 0.1 | 3.9 | REACTOME ADHERENS JUNCTIONS INTERACTIONS | Genes involved in Adherens junctions interactions |
| 0.1 | 1.9 | REACTOME REGULATION OF IFNA SIGNALING | Genes involved in Regulation of IFNA signaling |
| 0.1 | 5.7 | REACTOME INTERFERON ALPHA BETA SIGNALING | Genes involved in Interferon alpha/beta signaling |
| 0.1 | 2.0 | REACTOME GAP JUNCTION ASSEMBLY | Genes involved in Gap junction assembly |
| 0.1 | 6.2 | REACTOME GLUCONEOGENESIS | Genes involved in Gluconeogenesis |
| 0.1 | 19.0 | REACTOME SIGNALING BY RHO GTPASES | Genes involved in Signaling by Rho GTPases |
| 0.1 | 3.8 | REACTOME CELL CELL JUNCTION ORGANIZATION | Genes involved in Cell-cell junction organization |
| 0.1 | 0.7 | REACTOME ANDROGEN BIOSYNTHESIS | Genes involved in Androgen biosynthesis |
| 0.1 | 1.6 | REACTOME SYNTHESIS SECRETION AND INACTIVATION OF GLP1 | Genes involved in Synthesis, Secretion, and Inactivation of Glucagon-like Peptide-1 (GLP-1) |
| 0.1 | 4.0 | REACTOME AMINO ACID TRANSPORT ACROSS THE PLASMA MEMBRANE | Genes involved in Amino acid transport across the plasma membrane |
| 0.1 | 1.7 | REACTOME LIPID DIGESTION MOBILIZATION AND TRANSPORT | Genes involved in Lipid digestion, mobilization, and transport |
| 0.1 | 1.0 | REACTOME PROSTANOID LIGAND RECEPTORS | Genes involved in Prostanoid ligand receptors |
| 0.1 | 2.3 | REACTOME PURINE SALVAGE | Genes involved in Purine salvage |
| 0.1 | 3.8 | REACTOME CYTOCHROME P450 ARRANGED BY SUBSTRATE TYPE | Genes involved in Cytochrome P450 - arranged by substrate type |
| 0.1 | 16.0 | REACTOME G ALPHA Q SIGNALLING EVENTS | Genes involved in G alpha (q) signalling events |
| 0.1 | 1.0 | REACTOME DIGESTION OF DIETARY CARBOHYDRATE | Genes involved in Digestion of dietary carbohydrate |
| 0.1 | 1.0 | REACTOME PHASE II CONJUGATION | Genes involved in Phase II conjugation |
| 0.1 | 3.7 | REACTOME NUCLEAR SIGNALING BY ERBB4 | Genes involved in Nuclear signaling by ERBB4 |
| 0.1 | 11.3 | REACTOME RESPONSE TO ELEVATED PLATELET CYTOSOLIC CA2 | Genes involved in Response to elevated platelet cytosolic Ca2+ |
| 0.1 | 2.5 | REACTOME AMYLOIDS | Genes involved in Amyloids |
| 0.1 | 1.7 | REACTOME O LINKED GLYCOSYLATION OF MUCINS | Genes involved in O-linked glycosylation of mucins |
| 0.1 | 1.5 | REACTOME REGULATION OF GENE EXPRESSION IN BETA CELLS | Genes involved in Regulation of gene expression in beta cells |
| 0.1 | 0.9 | REACTOME OPSINS | Genes involved in Opsins |
| 0.1 | 2.5 | REACTOME PTM GAMMA CARBOXYLATION HYPUSINE FORMATION AND ARYLSULFATASE ACTIVATION | Genes involved in PTM: gamma carboxylation, hypusine formation and arylsulfatase activation |
| 0.1 | 2.3 | REACTOME CONVERSION FROM APC C CDC20 TO APC C CDH1 IN LATE ANAPHASE | Genes involved in Conversion from APC/C:Cdc20 to APC/C:Cdh1 in late anaphase |
| 0.1 | 1.2 | REACTOME TRYPTOPHAN CATABOLISM | Genes involved in Tryptophan catabolism |
| 0.1 | 1.1 | REACTOME COSTIMULATION BY THE CD28 FAMILY | Genes involved in Costimulation by the CD28 family |
| 0.1 | 1.9 | REACTOME AMINE LIGAND BINDING RECEPTORS | Genes involved in Amine ligand-binding receptors |
| 0.1 | 0.6 | REACTOME THROMBIN SIGNALLING THROUGH PROTEINASE ACTIVATED RECEPTORS PARS | Genes involved in Thrombin signalling through proteinase activated receptors (PARs) |
| 0.1 | 1.7 | REACTOME SIGNALING BY BMP | Genes involved in Signaling by BMP |
| 0.1 | 2.7 | REACTOME PYRIMIDINE METABOLISM | Genes involved in Pyrimidine metabolism |
| 0.1 | 2.9 | REACTOME NOTCH1 INTRACELLULAR DOMAIN REGULATES TRANSCRIPTION | Genes involved in NOTCH1 Intracellular Domain Regulates Transcription |
| 0.1 | 1.9 | REACTOME ACTIVATED NOTCH1 TRANSMITS SIGNAL TO THE NUCLEUS | Genes involved in Activated NOTCH1 Transmits Signal to the Nucleus |
| 0.1 | 1.7 | REACTOME AMINE COMPOUND SLC TRANSPORTERS | Genes involved in Amine compound SLC transporters |
| 0.1 | 1.1 | REACTOME THE NLRP3 INFLAMMASOME | Genes involved in The NLRP3 inflammasome |
| 0.1 | 6.6 | REACTOME NUCLEAR RECEPTOR TRANSCRIPTION PATHWAY | Genes involved in Nuclear Receptor transcription pathway |
| 0.1 | 2.9 | REACTOME LIGAND GATED ION CHANNEL TRANSPORT | Genes involved in Ligand-gated ion channel transport |
| 0.1 | 1.2 | REACTOME PHASE1 FUNCTIONALIZATION OF COMPOUNDS | Genes involved in Phase 1 - Functionalization of compounds |
| 0.1 | 0.1 | REACTOME SYNTHESIS OF BILE ACIDS AND BILE SALTS | Genes involved in Synthesis of bile acids and bile salts |
| 0.1 | 1.9 | REACTOME CHOLESTEROL BIOSYNTHESIS | Genes involved in Cholesterol biosynthesis |
| 0.1 | 1.5 | REACTOME SIGNALING BY NODAL | Genes involved in Signaling by NODAL |
| 0.1 | 0.9 | REACTOME CALNEXIN CALRETICULIN CYCLE | Genes involved in Calnexin/calreticulin cycle |
| 0.0 | 2.1 | REACTOME ION TRANSPORT BY P TYPE ATPASES | Genes involved in Ion transport by P-type ATPases |
| 0.0 | 1.5 | REACTOME MEIOTIC RECOMBINATION | Genes involved in Meiotic Recombination |
| 0.0 | 0.9 | REACTOME CIRCADIAN REPRESSION OF EXPRESSION BY REV ERBA | Genes involved in Circadian Repression of Expression by REV-ERBA |
| 0.0 | 1.1 | REACTOME IL1 SIGNALING | Genes involved in Interleukin-1 signaling |
| 0.0 | 0.9 | REACTOME CASPASE MEDIATED CLEAVAGE OF CYTOSKELETAL PROTEINS | Genes involved in Caspase-mediated cleavage of cytoskeletal proteins |
| 0.0 | 0.5 | REACTOME INHIBITION OF INSULIN SECRETION BY ADRENALINE NORADRENALINE | Genes involved in Inhibition of Insulin Secretion by Adrenaline/Noradrenaline |
| 0.0 | 3.7 | REACTOME G ALPHA I SIGNALLING EVENTS | Genes involved in G alpha (i) signalling events |
| 0.0 | 0.9 | REACTOME SYNTHESIS OF GLYCOSYLPHOSPHATIDYLINOSITOL GPI | Genes involved in Synthesis of glycosylphosphatidylinositol (GPI) |
| 0.0 | 0.4 | REACTOME GLUCOSE TRANSPORT | Genes involved in Glucose transport |
| 0.0 | 2.6 | REACTOME STRIATED MUSCLE CONTRACTION | Genes involved in Striated Muscle Contraction |
| 0.0 | 0.9 | REACTOME AMINO ACID SYNTHESIS AND INTERCONVERSION TRANSAMINATION | Genes involved in Amino acid synthesis and interconversion (transamination) |
| 0.0 | 0.9 | REACTOME NEGATIVE REGULATORS OF RIG I MDA5 SIGNALING | Genes involved in Negative regulators of RIG-I/MDA5 signaling |
| 0.0 | 1.9 | REACTOME TOLL RECEPTOR CASCADES | Genes involved in Toll Receptor Cascades |
| 0.0 | 9.6 | REACTOME GENERIC TRANSCRIPTION PATHWAY | Genes involved in Generic Transcription Pathway |
| 0.0 | 0.7 | REACTOME SIGNALING BY FGFR1 FUSION MUTANTS | Genes involved in Signaling by FGFR1 fusion mutants |
| 0.0 | 0.5 | REACTOME FGFR1 LIGAND BINDING AND ACTIVATION | Genes involved in FGFR1 ligand binding and activation |
| 0.0 | 2.1 | REACTOME TRANSPORT OF INORGANIC CATIONS ANIONS AND AMINO ACIDS OLIGOPEPTIDES | Genes involved in Transport of inorganic cations/anions and amino acids/oligopeptides |
| 0.0 | 0.5 | REACTOME SIGNALING BY HIPPO | Genes involved in Signaling by Hippo |
| 0.0 | 0.7 | REACTOME RNA POL III TRANSCRIPTION INITIATION FROM TYPE 3 PROMOTER | Genes involved in RNA Polymerase III Transcription Initiation From Type 3 Promoter |
| 0.0 | 0.9 | REACTOME CIRCADIAN CLOCK | Genes involved in Circadian Clock |
| 0.0 | 0.4 | REACTOME SYNTHESIS OF PE | Genes involved in Synthesis of PE |
| 0.0 | 1.6 | REACTOME TRANSPORT OF MATURE TRANSCRIPT TO CYTOPLASM | Genes involved in Transport of Mature Transcript to Cytoplasm |
| 0.0 | 0.2 | REACTOME CREB PHOSPHORYLATION THROUGH THE ACTIVATION OF CAMKII | Genes involved in CREB phosphorylation through the activation of CaMKII |
| 0.0 | 1.4 | REACTOME G ALPHA S SIGNALLING EVENTS | Genes involved in G alpha (s) signalling events |
| 0.0 | 0.2 | REACTOME RETROGRADE NEUROTROPHIN SIGNALLING | Genes involved in Retrograde neurotrophin signalling |
| 0.0 | 0.3 | REACTOME APOPTOTIC CLEAVAGE OF CELLULAR PROTEINS | Genes involved in Apoptotic cleavage of cellular proteins |