Project
GNF SymAtlas + NCI-60 cancer cell lines, human (Su, 2004; Ross, 2000)
Navigation
Downloads
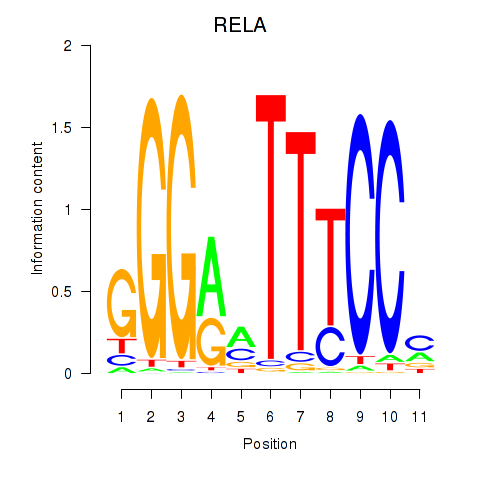
Results for RELA
Z-value: 1.01
Transcription factors associated with RELA
| Gene Symbol | Gene ID | Gene Info |
|---|---|---|
|
RELA
|
ENSG00000173039.14 | RELA proto-oncogene, NF-kB subunit |
Activity-expression correlation:
| Gene | Promoter | Pearson corr. coef. | P-value | Plot |
|---|---|---|---|---|
| RELA | hg19_v2_chr11_-_65430554_65430579 | 0.57 | 5.2e-20 | Click! |
Activity profile of RELA motif
Sorted Z-values of RELA motif
| Promoter | Log-likelihood | Transcript | Gene | Gene Info |
|---|---|---|---|---|
| chr14_-_35873856 | 32.56 |
ENST00000553342.1
ENST00000216797.5 ENST00000557140.1 |
NFKBIA
|
nuclear factor of kappa light polypeptide gene enhancer in B-cells inhibitor, alpha |
| chr17_-_4852332 | 26.02 |
ENST00000572383.1
|
PFN1
|
profilin 1 |
| chr6_+_138188551 | 24.43 |
ENST00000237289.4
ENST00000433680.1 |
TNFAIP3
|
tumor necrosis factor, alpha-induced protein 3 |
| chr4_-_174256276 | 23.85 |
ENST00000296503.5
|
HMGB2
|
high mobility group box 2 |
| chr5_-_150466692 | 20.21 |
ENST00000315050.7
ENST00000523338.1 ENST00000522100.1 |
TNIP1
|
TNFAIP3 interacting protein 1 |
| chr5_-_150460914 | 20.08 |
ENST00000389378.2
|
TNIP1
|
TNFAIP3 interacting protein 1 |
| chr11_+_102188224 | 17.83 |
ENST00000263464.3
|
BIRC3
|
baculoviral IAP repeat containing 3 |
| chr6_-_30712313 | 17.57 |
ENST00000376377.2
ENST00000259874.5 |
IER3
|
immediate early response 3 |
| chr5_-_150460539 | 16.14 |
ENST00000520931.1
ENST00000520695.1 ENST00000521591.1 ENST00000518977.1 |
TNIP1
|
TNFAIP3 interacting protein 1 |
| chr11_+_102188272 | 15.50 |
ENST00000532808.1
|
BIRC3
|
baculoviral IAP repeat containing 3 |
| chr4_+_74735102 | 15.24 |
ENST00000395761.3
|
CXCL1
|
chemokine (C-X-C motif) ligand 1 (melanoma growth stimulating activity, alpha) |
| chr22_-_37640277 | 14.81 |
ENST00000401529.3
ENST00000249071.6 |
RAC2
|
ras-related C3 botulinum toxin substrate 2 (rho family, small GTP binding protein Rac2) |
| chr7_-_93520259 | 14.45 |
ENST00000222543.5
|
TFPI2
|
tissue factor pathway inhibitor 2 |
| chr1_-_209825674 | 14.16 |
ENST00000367030.3
ENST00000356082.4 |
LAMB3
|
laminin, beta 3 |
| chr6_-_31550192 | 14.11 |
ENST00000429299.2
ENST00000446745.2 |
LTB
|
lymphotoxin beta (TNF superfamily, member 3) |
| chr4_+_103422471 | 14.02 |
ENST00000226574.4
ENST00000394820.4 |
NFKB1
|
nuclear factor of kappa light polypeptide gene enhancer in B-cells 1 |
| chr11_-_58345569 | 13.17 |
ENST00000528954.1
ENST00000528489.1 |
LPXN
|
leupaxin |
| chr4_-_103749179 | 13.16 |
ENST00000502690.1
|
UBE2D3
|
ubiquitin-conjugating enzyme E2D 3 |
| chr4_-_74964904 | 13.09 |
ENST00000508487.2
|
CXCL2
|
chemokine (C-X-C motif) ligand 2 |
| chr6_+_32605195 | 13.01 |
ENST00000374949.2
|
HLA-DQA1
|
major histocompatibility complex, class II, DQ alpha 1 |
| chr19_+_41725088 | 12.55 |
ENST00000301178.4
|
AXL
|
AXL receptor tyrosine kinase |
| chr10_+_104154229 | 12.24 |
ENST00000428099.1
ENST00000369966.3 |
NFKB2
|
nuclear factor of kappa light polypeptide gene enhancer in B-cells 2 (p49/p100) |
| chr4_-_103749205 | 12.20 |
ENST00000508249.1
|
UBE2D3
|
ubiquitin-conjugating enzyme E2D 3 |
| chr18_+_3252265 | 11.88 |
ENST00000580887.1
ENST00000536605.1 |
MYL12A
|
myosin, light chain 12A, regulatory, non-sarcomeric |
| chr9_+_36572851 | 11.63 |
ENST00000298048.2
ENST00000538311.1 ENST00000536987.1 ENST00000545008.1 ENST00000536860.1 ENST00000536329.1 ENST00000541717.1 ENST00000543751.1 |
MELK
|
maternal embryonic leucine zipper kinase |
| chr6_+_29691198 | 11.41 |
ENST00000440587.2
ENST00000434407.2 |
HLA-F
|
major histocompatibility complex, class I, F |
| chr6_+_29691056 | 11.05 |
ENST00000414333.1
ENST00000334668.4 ENST00000259951.7 |
HLA-F
|
major histocompatibility complex, class I, F |
| chr5_+_82767284 | 11.03 |
ENST00000265077.3
|
VCAN
|
versican |
| chr1_+_155657737 | 10.80 |
ENST00000471642.2
ENST00000471214.1 |
DAP3
|
death associated protein 3 |
| chr6_+_32605134 | 10.65 |
ENST00000343139.5
ENST00000395363.1 ENST00000496318.1 |
HLA-DQA1
|
major histocompatibility complex, class II, DQ alpha 1 |
| chr6_+_29910301 | 10.51 |
ENST00000376809.5
ENST00000376802.2 |
HLA-A
|
major histocompatibility complex, class I, A |
| chr14_-_24616426 | 10.49 |
ENST00000216802.5
|
PSME2
|
proteasome (prosome, macropain) activator subunit 2 (PA28 beta) |
| chr7_-_93520191 | 10.46 |
ENST00000545378.1
|
TFPI2
|
tissue factor pathway inhibitor 2 |
| chr9_-_125667494 | 10.42 |
ENST00000335387.5
ENST00000357244.2 ENST00000373665.2 |
RC3H2
|
ring finger and CCCH-type domains 2 |
| chr22_-_37640456 | 10.03 |
ENST00000405484.1
ENST00000441619.1 ENST00000406508.1 |
RAC2
|
ras-related C3 botulinum toxin substrate 2 (rho family, small GTP binding protein Rac2) |
| chr12_-_9913489 | 9.56 |
ENST00000228434.3
ENST00000536709.1 |
CD69
|
CD69 molecule |
| chr19_-_6591113 | 9.48 |
ENST00000423145.3
ENST00000245903.3 |
CD70
|
CD70 molecule |
| chr3_+_157154578 | 9.33 |
ENST00000295927.3
|
PTX3
|
pentraxin 3, long |
| chr21_+_34775772 | 9.22 |
ENST00000405436.1
|
IFNGR2
|
interferon gamma receptor 2 (interferon gamma transducer 1) |
| chr15_+_52311398 | 9.14 |
ENST00000261845.5
|
MAPK6
|
mitogen-activated protein kinase 6 |
| chr17_+_16318909 | 9.12 |
ENST00000577397.1
|
TRPV2
|
transient receptor potential cation channel, subfamily V, member 2 |
| chr1_-_32403903 | 9.03 |
ENST00000344035.6
ENST00000356536.3 |
PTP4A2
|
protein tyrosine phosphatase type IVA, member 2 |
| chr13_+_37581115 | 8.75 |
ENST00000481013.1
|
EXOSC8
|
exosome component 8 |
| chr1_-_65432171 | 8.66 |
ENST00000342505.4
|
JAK1
|
Janus kinase 1 |
| chr21_+_34775698 | 8.63 |
ENST00000381995.1
|
IFNGR2
|
interferon gamma receptor 2 (interferon gamma transducer 1) |
| chr17_+_16318850 | 8.44 |
ENST00000338560.7
|
TRPV2
|
transient receptor potential cation channel, subfamily V, member 2 |
| chr19_-_50143452 | 8.42 |
ENST00000246792.3
|
RRAS
|
related RAS viral (r-ras) oncogene homolog |
| chr17_+_34900737 | 8.38 |
ENST00000304718.4
ENST00000485685.2 |
GGNBP2
|
gametogenetin binding protein 2 |
| chr20_+_1115821 | 8.26 |
ENST00000435720.1
|
PSMF1
|
proteasome (prosome, macropain) inhibitor subunit 1 (PI31) |
| chr4_-_40631859 | 8.24 |
ENST00000295971.7
ENST00000319592.4 |
RBM47
|
RNA binding motif protein 47 |
| chr6_-_17706618 | 8.21 |
ENST00000262077.2
ENST00000537253.1 |
NUP153
|
nucleoporin 153kDa |
| chr4_-_185395672 | 8.14 |
ENST00000393593.3
|
IRF2
|
interferon regulatory factor 2 |
| chr17_-_7590745 | 7.99 |
ENST00000514944.1
ENST00000503591.1 ENST00000455263.2 ENST00000420246.2 ENST00000445888.2 ENST00000509690.1 ENST00000604348.1 ENST00000269305.4 |
TP53
|
tumor protein p53 |
| chr1_-_1822495 | 7.94 |
ENST00000378609.4
|
GNB1
|
guanine nucleotide binding protein (G protein), beta polypeptide 1 |
| chr21_+_34775181 | 7.85 |
ENST00000290219.6
|
IFNGR2
|
interferon gamma receptor 2 (interferon gamma transducer 1) |
| chr20_-_2451395 | 7.82 |
ENST00000339610.6
ENST00000381342.2 ENST00000438552.2 |
SNRPB
|
small nuclear ribonucleoprotein polypeptides B and B1 |
| chr12_-_54653313 | 7.63 |
ENST00000550411.1
ENST00000439541.2 |
CBX5
|
chromobox homolog 5 |
| chr4_+_41614720 | 7.58 |
ENST00000509277.1
|
LIMCH1
|
LIM and calponin homology domains 1 |
| chr16_+_50776021 | 7.48 |
ENST00000566679.2
ENST00000564634.1 ENST00000398568.2 |
CYLD
|
cylindromatosis (turban tumor syndrome) |
| chr1_+_44870866 | 7.36 |
ENST00000355387.2
ENST00000361799.2 |
RNF220
|
ring finger protein 220 |
| chr14_+_103589789 | 7.26 |
ENST00000558056.1
ENST00000560869.1 |
TNFAIP2
|
tumor necrosis factor, alpha-induced protein 2 |
| chr8_-_103668114 | 7.13 |
ENST00000285407.6
|
KLF10
|
Kruppel-like factor 10 |
| chr7_+_22766766 | 7.03 |
ENST00000426291.1
ENST00000401651.1 ENST00000258743.5 ENST00000420258.2 ENST00000407492.1 ENST00000401630.3 ENST00000406575.1 |
IL6
|
interleukin 6 (interferon, beta 2) |
| chr2_+_228678550 | 6.78 |
ENST00000409189.3
ENST00000358813.4 |
CCL20
|
chemokine (C-C motif) ligand 20 |
| chr1_-_186649543 | 6.78 |
ENST00000367468.5
|
PTGS2
|
prostaglandin-endoperoxide synthase 2 (prostaglandin G/H synthase and cyclooxygenase) |
| chr7_+_143078652 | 6.70 |
ENST00000354434.4
ENST00000449423.2 |
ZYX
|
zyxin |
| chr10_-_43904235 | 6.63 |
ENST00000356053.3
|
HNRNPF
|
heterogeneous nuclear ribonucleoprotein F |
| chr2_+_161993412 | 6.63 |
ENST00000259075.2
ENST00000432002.1 |
TANK
|
TRAF family member-associated NFKB activator |
| chr7_-_16844611 | 6.51 |
ENST00000401412.1
ENST00000419304.2 |
AGR2
|
anterior gradient 2 |
| chr5_+_177019159 | 6.42 |
ENST00000332598.6
|
TMED9
|
transmembrane emp24 protein transport domain containing 9 |
| chr19_+_45504688 | 6.36 |
ENST00000221452.8
ENST00000540120.1 ENST00000505236.1 |
RELB
|
v-rel avian reticuloendotheliosis viral oncogene homolog B |
| chr10_+_22605374 | 6.31 |
ENST00000448361.1
|
COMMD3
|
COMM domain containing 3 |
| chr4_-_103749313 | 6.07 |
ENST00000394803.5
|
UBE2D3
|
ubiquitin-conjugating enzyme E2D 3 |
| chr1_-_113249678 | 6.02 |
ENST00000369633.2
ENST00000425265.2 ENST00000369632.2 ENST00000436685.2 |
RHOC
|
ras homolog family member C |
| chr9_-_127952032 | 6.00 |
ENST00000456642.1
ENST00000373546.3 ENST00000373547.4 |
PPP6C
|
protein phosphatase 6, catalytic subunit |
| chr1_-_209824643 | 5.93 |
ENST00000391911.1
ENST00000415782.1 |
LAMB3
|
laminin, beta 3 |
| chr1_-_113249734 | 5.81 |
ENST00000484054.3
ENST00000369636.2 ENST00000369637.1 ENST00000285735.2 ENST00000369638.2 |
RHOC
|
ras homolog family member C |
| chr19_+_41725140 | 5.79 |
ENST00000359092.3
|
AXL
|
AXL receptor tyrosine kinase |
| chr1_-_94079648 | 5.79 |
ENST00000370247.3
|
BCAR3
|
breast cancer anti-estrogen resistance 3 |
| chr14_-_69446034 | 5.64 |
ENST00000193403.6
|
ACTN1
|
actinin, alpha 1 |
| chr11_-_6633799 | 5.62 |
ENST00000299424.4
|
TAF10
|
TAF10 RNA polymerase II, TATA box binding protein (TBP)-associated factor, 30kDa |
| chr4_+_140222609 | 5.62 |
ENST00000296543.5
ENST00000398947.1 |
NAA15
|
N(alpha)-acetyltransferase 15, NatA auxiliary subunit |
| chr1_-_205744574 | 5.56 |
ENST00000367139.3
ENST00000235932.4 ENST00000437324.2 ENST00000414729.1 |
RAB7L1
|
RAB7, member RAS oncogene family-like 1 |
| chr6_-_32636145 | 5.38 |
ENST00000399084.1
|
HLA-DQB1
|
major histocompatibility complex, class II, DQ beta 1 |
| chr10_+_104155450 | 5.28 |
ENST00000471698.1
ENST00000189444.6 |
NFKB2
|
nuclear factor of kappa light polypeptide gene enhancer in B-cells 2 (p49/p100) |
| chr11_+_19798964 | 5.26 |
ENST00000527559.2
|
NAV2
|
neuron navigator 2 |
| chr1_+_26869597 | 5.26 |
ENST00000530003.1
|
RPS6KA1
|
ribosomal protein S6 kinase, 90kDa, polypeptide 1 |
| chr9_-_127952187 | 5.26 |
ENST00000451402.1
ENST00000415905.1 |
PPP6C
|
protein phosphatase 6, catalytic subunit |
| chr10_-_102089729 | 5.18 |
ENST00000465680.2
|
PKD2L1
|
polycystic kidney disease 2-like 1 |
| chr4_-_103749105 | 5.18 |
ENST00000394801.4
ENST00000394804.2 |
UBE2D3
|
ubiquitin-conjugating enzyme E2D 3 |
| chr2_-_89442621 | 5.17 |
ENST00000492167.1
|
IGKV3-20
|
immunoglobulin kappa variable 3-20 |
| chr4_-_103748880 | 5.09 |
ENST00000453744.2
ENST00000349311.8 |
UBE2D3
|
ubiquitin-conjugating enzyme E2D 3 |
| chr1_-_36615051 | 5.09 |
ENST00000373163.1
|
TRAPPC3
|
trafficking protein particle complex 3 |
| chr16_-_28223166 | 5.05 |
ENST00000304658.5
|
XPO6
|
exportin 6 |
| chr1_+_179050512 | 5.02 |
ENST00000367627.3
|
TOR3A
|
torsin family 3, member A |
| chr12_-_57023995 | 4.87 |
ENST00000549884.1
ENST00000546695.1 |
BAZ2A
|
bromodomain adjacent to zinc finger domain, 2A |
| chr22_-_28315115 | 4.84 |
ENST00000455418.3
ENST00000436663.1 ENST00000320996.10 ENST00000335272.5 |
PITPNB
|
phosphatidylinositol transfer protein, beta |
| chr18_-_12884259 | 4.82 |
ENST00000353319.4
ENST00000327283.3 |
PTPN2
|
protein tyrosine phosphatase, non-receptor type 2 |
| chr10_+_89419370 | 4.81 |
ENST00000361175.4
ENST00000456849.1 |
PAPSS2
|
3'-phosphoadenosine 5'-phosphosulfate synthase 2 |
| chr19_+_45251804 | 4.76 |
ENST00000164227.5
|
BCL3
|
B-cell CLL/lymphoma 3 |
| chr5_-_149792295 | 4.66 |
ENST00000518797.1
ENST00000524315.1 ENST00000009530.7 ENST00000377795.3 |
CD74
|
CD74 molecule, major histocompatibility complex, class II invariant chain |
| chr1_-_36615065 | 4.64 |
ENST00000373166.3
ENST00000373159.1 ENST00000373162.1 |
TRAPPC3
|
trafficking protein particle complex 3 |
| chr11_-_72853091 | 4.62 |
ENST00000311172.7
ENST00000409314.1 |
FCHSD2
|
FCH and double SH3 domains 2 |
| chr11_+_19799327 | 4.54 |
ENST00000540292.1
|
NAV2
|
neuron navigator 2 |
| chr17_+_21191341 | 4.52 |
ENST00000526076.2
ENST00000361818.5 ENST00000316920.6 |
MAP2K3
|
mitogen-activated protein kinase kinase 3 |
| chr11_+_6624955 | 4.50 |
ENST00000299421.4
ENST00000537806.1 |
ILK
|
integrin-linked kinase |
| chr1_-_44818599 | 4.44 |
ENST00000537474.1
|
ERI3
|
ERI1 exoribonuclease family member 3 |
| chr7_-_24797546 | 4.43 |
ENST00000414428.1
ENST00000419307.1 ENST00000342947.3 |
DFNA5
|
deafness, autosomal dominant 5 |
| chr10_+_22605304 | 4.43 |
ENST00000475460.2
ENST00000602390.1 ENST00000489125.2 ENST00000456711.1 ENST00000444869.1 |
COMMD3-BMI1
COMMD3
|
COMMD3-BMI1 readthrough COMM domain containing 3 |
| chr19_+_42364460 | 4.34 |
ENST00000593863.1
|
RPS19
|
ribosomal protein S19 |
| chr1_-_54304212 | 4.32 |
ENST00000540001.1
|
NDC1
|
NDC1 transmembrane nucleoporin |
| chr16_-_28222797 | 4.31 |
ENST00000569951.1
ENST00000565698.1 |
XPO6
|
exportin 6 |
| chr11_+_6625046 | 4.30 |
ENST00000396751.2
|
ILK
|
integrin-linked kinase |
| chr6_+_30297306 | 4.28 |
ENST00000420746.1
ENST00000513556.1 |
TRIM39
TRIM39-RPP21
|
tripartite motif containing 39 TRIM39-RPP21 readthrough |
| chr4_-_74864386 | 4.27 |
ENST00000296027.4
|
CXCL5
|
chemokine (C-X-C motif) ligand 5 |
| chr1_-_205744205 | 4.14 |
ENST00000446390.2
|
RAB7L1
|
RAB7, member RAS oncogene family-like 1 |
| chr3_+_184033135 | 4.10 |
ENST00000424196.1
|
EIF4G1
|
eukaryotic translation initiation factor 4 gamma, 1 |
| chr2_+_208394455 | 4.09 |
ENST00000430624.1
|
CREB1
|
cAMP responsive element binding protein 1 |
| chr7_-_24797032 | 4.08 |
ENST00000409970.1
ENST00000409775.3 |
DFNA5
|
deafness, autosomal dominant 5 |
| chr7_-_100881109 | 4.07 |
ENST00000308344.5
|
CLDN15
|
claudin 15 |
| chr7_-_47579188 | 4.07 |
ENST00000398879.1
ENST00000355730.3 ENST00000442536.2 ENST00000458317.2 |
TNS3
|
tensin 3 |
| chr22_-_37545972 | 3.90 |
ENST00000216223.5
|
IL2RB
|
interleukin 2 receptor, beta |
| chr12_-_57504069 | 3.88 |
ENST00000543873.2
ENST00000554663.1 ENST00000557635.1 |
STAT6
|
signal transducer and activator of transcription 6, interleukin-4 induced |
| chr20_-_43977055 | 3.86 |
ENST00000372733.3
ENST00000537976.1 |
SDC4
|
syndecan 4 |
| chr3_+_9439400 | 3.62 |
ENST00000450326.1
ENST00000402198.1 ENST00000402466.1 |
SETD5
|
SET domain containing 5 |
| chr14_+_103243813 | 3.62 |
ENST00000560371.1
ENST00000347662.4 ENST00000392745.2 ENST00000539721.1 ENST00000560463.1 |
TRAF3
|
TNF receptor-associated factor 3 |
| chr12_+_7055631 | 3.60 |
ENST00000543115.1
ENST00000399448.1 |
PTPN6
|
protein tyrosine phosphatase, non-receptor type 6 |
| chr22_+_41347363 | 3.51 |
ENST00000216225.8
|
RBX1
|
ring-box 1, E3 ubiquitin protein ligase |
| chr6_-_74231303 | 3.45 |
ENST00000309268.6
|
EEF1A1
|
eukaryotic translation elongation factor 1 alpha 1 |
| chrX_+_117861535 | 3.40 |
ENST00000371666.3
ENST00000371642.1 |
IL13RA1
|
interleukin 13 receptor, alpha 1 |
| chr1_+_235490659 | 3.37 |
ENST00000488594.1
|
GGPS1
|
geranylgeranyl diphosphate synthase 1 |
| chr2_-_191885686 | 3.35 |
ENST00000432058.1
|
STAT1
|
signal transducer and activator of transcription 1, 91kDa |
| chr3_+_9439579 | 3.29 |
ENST00000406341.1
|
SETD5
|
SET domain containing 5 |
| chr1_-_54303934 | 3.24 |
ENST00000537333.1
|
NDC1
|
NDC1 transmembrane nucleoporin |
| chr15_+_85923797 | 3.22 |
ENST00000559362.1
|
AKAP13
|
A kinase (PRKA) anchor protein 13 |
| chrX_-_131352152 | 3.16 |
ENST00000342983.2
|
RAP2C
|
RAP2C, member of RAS oncogene family |
| chr11_-_3862206 | 3.11 |
ENST00000351018.4
|
RHOG
|
ras homolog family member G |
| chr1_-_153935983 | 3.09 |
ENST00000537590.1
ENST00000356205.4 |
SLC39A1
|
solute carrier family 39 (zinc transporter), member 1 |
| chr4_-_74904398 | 3.04 |
ENST00000296026.4
|
CXCL3
|
chemokine (C-X-C motif) ligand 3 |
| chr6_-_29527702 | 2.99 |
ENST00000377050.4
|
UBD
|
ubiquitin D |
| chr1_-_153935938 | 2.96 |
ENST00000368621.1
ENST00000368623.3 |
SLC39A1
|
solute carrier family 39 (zinc transporter), member 1 |
| chrX_-_48755030 | 2.84 |
ENST00000490755.2
ENST00000465150.2 ENST00000495490.2 |
TIMM17B
|
translocase of inner mitochondrial membrane 17 homolog B (yeast) |
| chr1_+_101185290 | 2.75 |
ENST00000370119.4
ENST00000347652.2 ENST00000294728.2 ENST00000370115.1 |
VCAM1
|
vascular cell adhesion molecule 1 |
| chr6_+_31540056 | 2.74 |
ENST00000418386.2
|
LTA
|
lymphotoxin alpha |
| chr2_-_203776864 | 2.73 |
ENST00000261015.4
|
WDR12
|
WD repeat domain 12 |
| chr3_+_53195136 | 2.73 |
ENST00000394729.2
ENST00000330452.3 |
PRKCD
|
protein kinase C, delta |
| chr5_+_14143728 | 2.71 |
ENST00000344204.4
ENST00000537187.1 |
TRIO
|
trio Rho guanine nucleotide exchange factor |
| chr19_-_7766991 | 2.69 |
ENST00000597921.1
ENST00000346664.5 |
FCER2
|
Fc fragment of IgE, low affinity II, receptor for (CD23) |
| chr2_+_208394616 | 2.69 |
ENST00000432329.2
ENST00000353267.3 ENST00000445803.1 |
CREB1
|
cAMP responsive element binding protein 1 |
| chr3_-_158390282 | 2.66 |
ENST00000264265.3
|
LXN
|
latexin |
| chr4_-_187644930 | 2.65 |
ENST00000441802.2
|
FAT1
|
FAT atypical cadherin 1 |
| chr4_+_74702214 | 2.61 |
ENST00000226317.5
ENST00000515050.1 |
CXCL6
|
chemokine (C-X-C motif) ligand 6 |
| chr6_-_74230741 | 2.61 |
ENST00000316292.9
|
EEF1A1
|
eukaryotic translation elongation factor 1 alpha 1 |
| chr19_+_42381337 | 2.58 |
ENST00000597454.1
ENST00000444740.2 |
CD79A
|
CD79a molecule, immunoglobulin-associated alpha |
| chr19_+_42364313 | 2.45 |
ENST00000601492.1
ENST00000600467.1 ENST00000221975.2 |
RPS19
|
ribosomal protein S19 |
| chr17_-_61777090 | 2.45 |
ENST00000578061.1
|
LIMD2
|
LIM domain containing 2 |
| chr19_+_2476116 | 2.43 |
ENST00000215631.4
ENST00000587345.1 |
GADD45B
|
growth arrest and DNA-damage-inducible, beta |
| chr9_-_125667618 | 2.41 |
ENST00000423239.2
|
RC3H2
|
ring finger and CCCH-type domains 2 |
| chr19_+_42381173 | 2.36 |
ENST00000221972.3
|
CD79A
|
CD79a molecule, immunoglobulin-associated alpha |
| chr12_+_49761224 | 2.34 |
ENST00000553127.1
ENST00000321898.6 |
SPATS2
|
spermatogenesis associated, serine-rich 2 |
| chr5_+_139781393 | 2.27 |
ENST00000360839.2
ENST00000297183.6 ENST00000421134.1 ENST00000394723.3 ENST00000511151.1 |
ANKHD1
|
ankyrin repeat and KH domain containing 1 |
| chr2_-_136743169 | 2.26 |
ENST00000264161.4
|
DARS
|
aspartyl-tRNA synthetase |
| chr4_+_169418195 | 2.25 |
ENST00000261509.6
ENST00000335742.7 |
PALLD
|
palladin, cytoskeletal associated protein |
| chr12_-_76425368 | 2.25 |
ENST00000602540.1
|
PHLDA1
|
pleckstrin homology-like domain, family A, member 1 |
| chr1_+_63833261 | 2.23 |
ENST00000371108.4
|
ALG6
|
ALG6, alpha-1,3-glucosyltransferase |
| chr5_+_112312416 | 2.22 |
ENST00000389063.2
|
DCP2
|
decapping mRNA 2 |
| chrX_-_48827976 | 2.18 |
ENST00000218176.3
|
KCND1
|
potassium voltage-gated channel, Shal-related subfamily, member 1 |
| chr1_-_155658085 | 2.15 |
ENST00000311573.5
ENST00000438245.2 |
YY1AP1
|
YY1 associated protein 1 |
| chr1_-_27216729 | 2.12 |
ENST00000431781.2
ENST00000374135.4 |
GPN2
|
GPN-loop GTPase 2 |
| chr2_+_44396000 | 2.09 |
ENST00000409895.4
ENST00000409432.3 ENST00000282412.4 ENST00000378551.2 ENST00000345249.4 |
PPM1B
|
protein phosphatase, Mg2+/Mn2+ dependent, 1B |
| chr2_+_233925064 | 2.08 |
ENST00000359570.5
ENST00000538935.1 |
INPP5D
|
inositol polyphosphate-5-phosphatase, 145kDa |
| chr1_+_8021954 | 2.05 |
ENST00000377491.1
ENST00000377488.1 |
PARK7
|
parkinson protein 7 |
| chr10_-_102090243 | 2.01 |
ENST00000338519.3
ENST00000353274.3 ENST00000318222.3 |
PKD2L1
|
polycystic kidney disease 2-like 1 |
| chr4_-_76928641 | 1.99 |
ENST00000264888.5
|
CXCL9
|
chemokine (C-X-C motif) ligand 9 |
| chr12_+_49761273 | 1.99 |
ENST00000551540.1
ENST00000552918.1 ENST00000548777.1 ENST00000547865.1 ENST00000552171.1 |
SPATS2
|
spermatogenesis associated, serine-rich 2 |
| chr2_+_162016916 | 1.99 |
ENST00000405852.1
|
TANK
|
TRAF family member-associated NFKB activator |
| chr5_+_139781445 | 1.98 |
ENST00000532219.1
ENST00000394722.3 |
ANKHD1-EIF4EBP3
ANKHD1
|
ANKHD1-EIF4EBP3 readthrough ankyrin repeat and KH domain containing 1 |
| chr11_+_18287721 | 1.98 |
ENST00000356524.4
|
SAA1
|
serum amyloid A1 |
| chrX_-_119709637 | 1.95 |
ENST00000404115.3
|
CUL4B
|
cullin 4B |
| chr15_+_57884199 | 1.95 |
ENST00000587652.1
ENST00000380568.3 ENST00000380565.4 ENST00000380563.2 |
GCOM1
MYZAP
POLR2M
|
GRINL1A complex locus 1 myocardial zonula adherens protein polymerase (RNA) II (DNA directed) polypeptide M |
| chr15_+_57884086 | 1.95 |
ENST00000380569.2
ENST00000380561.2 ENST00000574161.1 ENST00000572390.1 ENST00000396180.1 ENST00000380560.2 |
GCOM1
|
GRINL1A complex locus 1 |
| chr12_+_53662073 | 1.93 |
ENST00000553219.1
ENST00000257934.4 |
ESPL1
|
extra spindle pole bodies homolog 1 (S. cerevisiae) |
| chr11_-_44331679 | 1.86 |
ENST00000329255.3
|
ALX4
|
ALX homeobox 4 |
| chr14_+_96968707 | 1.83 |
ENST00000216277.8
ENST00000557320.1 ENST00000557471.1 |
PAPOLA
|
poly(A) polymerase alpha |
| chr1_-_54303949 | 1.82 |
ENST00000234725.8
|
NDC1
|
NDC1 transmembrane nucleoporin |
| chr11_+_18287801 | 1.81 |
ENST00000532858.1
ENST00000405158.2 |
SAA1
|
serum amyloid A1 |
| chr2_+_208394794 | 1.79 |
ENST00000536726.1
ENST00000374397.4 ENST00000452474.1 |
CREB1
|
cAMP responsive element binding protein 1 |
| chr1_+_221054584 | 1.78 |
ENST00000549319.1
|
HLX
|
H2.0-like homeobox |
| chrX_+_48755183 | 1.77 |
ENST00000376563.1
ENST00000376566.4 |
PQBP1
|
polyglutamine binding protein 1 |
| chr1_-_151319710 | 1.76 |
ENST00000290524.4
ENST00000437327.1 ENST00000452513.2 ENST00000368870.2 ENST00000452671.2 |
RFX5
|
regulatory factor X, 5 (influences HLA class II expression) |
| chr7_+_134464376 | 1.75 |
ENST00000454108.1
ENST00000361675.2 |
CALD1
|
caldesmon 1 |
| chr14_+_24439148 | 1.72 |
ENST00000543805.1
ENST00000534993.1 |
DHRS4L2
|
dehydrogenase/reductase (SDR family) member 4 like 2 |
| chr19_+_42363917 | 1.71 |
ENST00000598742.1
|
RPS19
|
ribosomal protein S19 |
| chr4_+_41614909 | 1.71 |
ENST00000509454.1
ENST00000396595.3 ENST00000381753.4 |
LIMCH1
|
LIM and calponin homology domains 1 |
| chr20_-_36156293 | 1.71 |
ENST00000373537.2
ENST00000414542.2 |
BLCAP
|
bladder cancer associated protein |
| chr12_-_28124903 | 1.71 |
ENST00000395872.1
ENST00000354417.3 ENST00000201015.4 |
PTHLH
|
parathyroid hormone-like hormone |
| chr11_+_10476851 | 1.68 |
ENST00000396553.2
|
AMPD3
|
adenosine monophosphate deaminase 3 |
| chr12_-_6716534 | 1.68 |
ENST00000544484.1
ENST00000309577.6 ENST00000357008.2 |
CHD4
|
chromodomain helicase DNA binding protein 4 |
| chr12_-_6716569 | 1.64 |
ENST00000544040.1
ENST00000545942.1 |
CHD4
|
chromodomain helicase DNA binding protein 4 |
| chr15_-_51914810 | 1.58 |
ENST00000543779.2
ENST00000449909.3 |
DMXL2
|
Dmx-like 2 |
Network of associatons between targets according to the STRING database.
First level regulatory network of RELA
Gene Ontology Analysis
Gene overrepresentation in biological_process category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 18.8 | 56.4 | GO:0085032 | modulation of signal transduction in other organism(GO:0044501) modulation by symbiont of host signal transduction pathway(GO:0052027) modulation of signal transduction in other organism involved in symbiotic interaction(GO:0052250) modulation by symbiont of host I-kappaB kinase/NF-kappaB cascade(GO:0085032) |
| 9.5 | 57.0 | GO:0070427 | nucleotide-binding oligomerization domain containing 1 signaling pathway(GO:0070427) |
| 5.6 | 22.3 | GO:0002268 | follicular dendritic cell differentiation(GO:0002268) |
| 4.3 | 12.8 | GO:0061470 | T follicular helper cell differentiation(GO:0061470) |
| 3.7 | 33.3 | GO:0070424 | regulation of nucleotide-binding oligomerization domain containing signaling pathway(GO:0070424) |
| 3.3 | 33.0 | GO:0002480 | antigen processing and presentation of exogenous peptide antigen via MHC class I, TAP-independent(GO:0002480) |
| 3.1 | 9.4 | GO:0071790 | spindle pole body duplication(GO:0030474) spindle pole body organization(GO:0051300) spindle pole body localization(GO:0070631) establishment of spindle pole body localization(GO:0070632) spindle pole body localization to nuclear envelope(GO:0071789) establishment of spindle pole body localization to nuclear envelope(GO:0071790) |
| 3.1 | 28.1 | GO:0045075 | interleukin-12 biosynthetic process(GO:0042090) regulation of interleukin-12 biosynthetic process(GO:0045075) |
| 3.1 | 18.3 | GO:0048549 | positive regulation of pinocytosis(GO:0048549) |
| 2.8 | 8.4 | GO:0033140 | negative regulation of peptidyl-serine phosphorylation of STAT protein(GO:0033140) |
| 2.7 | 8.0 | GO:0051097 | negative regulation of helicase activity(GO:0051097) oligodendrocyte apoptotic process(GO:0097252) |
| 2.6 | 26.0 | GO:1900029 | positive regulation of ruffle assembly(GO:1900029) |
| 2.5 | 9.8 | GO:0021564 | vagus nerve development(GO:0021564) |
| 2.3 | 7.0 | GO:0002384 | hepatic immune response(GO:0002384) response to prolactin(GO:1990637) regulation of STAT protein import into nucleus(GO:2000364) positive regulation of STAT protein import into nucleus(GO:2000366) |
| 2.3 | 6.8 | GO:0045362 | regulation of interleukin-1 biosynthetic process(GO:0045360) positive regulation of interleukin-1 biosynthetic process(GO:0045362) |
| 2.3 | 24.8 | GO:0060753 | regulation of mast cell chemotaxis(GO:0060753) |
| 2.2 | 8.8 | GO:0034476 | U1 snRNA 3'-end processing(GO:0034473) U5 snRNA 3'-end processing(GO:0034476) |
| 2.1 | 8.5 | GO:0060265 | positive regulation of respiratory burst involved in inflammatory response(GO:0060265) |
| 2.1 | 21.2 | GO:0045654 | positive regulation of megakaryocyte differentiation(GO:0045654) |
| 2.1 | 16.8 | GO:0050859 | negative regulation of B cell receptor signaling pathway(GO:0050859) |
| 2.0 | 13.9 | GO:0071352 | interleukin-2-mediated signaling pathway(GO:0038110) cellular response to interleukin-2(GO:0071352) |
| 2.0 | 11.8 | GO:1902766 | skeletal muscle satellite cell migration(GO:1902766) |
| 1.9 | 9.3 | GO:0052199 | negative regulation of catalytic activity in other organism involved in symbiotic interaction(GO:0052199) |
| 1.7 | 6.8 | GO:2001280 | positive regulation of prostaglandin biosynthetic process(GO:0031394) positive regulation of platelet-derived growth factor production(GO:0090362) positive regulation of unsaturated fatty acid biosynthetic process(GO:2001280) |
| 1.6 | 6.5 | GO:1903899 | positive regulation of PERK-mediated unfolded protein response(GO:1903899) |
| 1.6 | 6.4 | GO:0035964 | COPI-coated vesicle budding(GO:0035964) Golgi transport vesicle coating(GO:0048200) COPI coating of Golgi vesicle(GO:0048205) |
| 1.6 | 4.7 | GO:1903093 | regulation of protein K48-linked deubiquitination(GO:1903093) negative regulation of protein K48-linked deubiquitination(GO:1903094) negative regulation of ubiquitin-specific protease activity(GO:2000157) |
| 1.4 | 1.4 | GO:1902523 | positive regulation of protein K63-linked ubiquitination(GO:1902523) |
| 1.4 | 8.6 | GO:2000158 | positive regulation of ubiquitin-specific protease activity(GO:2000158) |
| 1.4 | 8.2 | GO:0016554 | cytidine to uridine editing(GO:0016554) |
| 1.3 | 34.5 | GO:0090023 | positive regulation of neutrophil chemotaxis(GO:0090023) |
| 1.3 | 1.3 | GO:0060585 | regulation of prostaglandin-endoperoxide synthase activity(GO:0060584) positive regulation of prostaglandin-endoperoxide synthase activity(GO:0060585) |
| 1.2 | 3.7 | GO:1990108 | protein linear deubiquitination(GO:1990108) |
| 1.2 | 4.8 | GO:0000103 | sulfate assimilation(GO:0000103) |
| 1.1 | 3.4 | GO:0033386 | geranylgeranyl diphosphate metabolic process(GO:0033385) geranylgeranyl diphosphate biosynthetic process(GO:0033386) |
| 1.1 | 3.3 | GO:0006422 | aspartyl-tRNA aminoacylation(GO:0006422) |
| 1.1 | 5.4 | GO:0002925 | positive regulation of humoral immune response mediated by circulating immunoglobulin(GO:0002925) |
| 1.0 | 7.2 | GO:0050915 | sensory perception of sour taste(GO:0050915) |
| 1.0 | 3.9 | GO:0002296 | T-helper 1 cell lineage commitment(GO:0002296) |
| 0.9 | 5.6 | GO:0017196 | N-terminal peptidyl-methionine acetylation(GO:0017196) |
| 0.9 | 29.0 | GO:0060334 | regulation of interferon-gamma-mediated signaling pathway(GO:0060334) |
| 0.9 | 41.7 | GO:0070979 | protein K11-linked ubiquitination(GO:0070979) |
| 0.9 | 3.6 | GO:0008063 | Toll signaling pathway(GO:0008063) |
| 0.9 | 5.1 | GO:1904550 | chemotaxis to arachidonic acid(GO:0034670) response to arachidonic acid(GO:1904550) |
| 0.8 | 3.3 | GO:0072553 | terminal button organization(GO:0072553) |
| 0.8 | 2.4 | GO:1900169 | regulation of glucocorticoid mediated signaling pathway(GO:1900169) |
| 0.8 | 3.9 | GO:1903553 | positive regulation of extracellular exosome assembly(GO:1903553) |
| 0.8 | 4.6 | GO:2000491 | positive regulation of hepatic stellate cell activation(GO:2000491) |
| 0.7 | 24.9 | GO:0071498 | cellular response to fluid shear stress(GO:0071498) |
| 0.7 | 2.1 | GO:1990022 | RNA polymerase II complex import to nucleus(GO:0044376) RNA polymerase III complex localization to nucleus(GO:1990022) |
| 0.7 | 2.1 | GO:0045659 | regulation of neutrophil differentiation(GO:0045658) negative regulation of neutrophil differentiation(GO:0045659) |
| 0.7 | 20.1 | GO:0031581 | hemidesmosome assembly(GO:0031581) |
| 0.7 | 2.7 | GO:0060697 | positive regulation of phospholipid catabolic process(GO:0060697) |
| 0.7 | 7.3 | GO:0051601 | exocyst localization(GO:0051601) |
| 0.7 | 2.0 | GO:0051685 | maintenance of ER location(GO:0051685) |
| 0.6 | 8.2 | GO:0046831 | regulation of RNA export from nucleus(GO:0046831) |
| 0.6 | 3.0 | GO:0070842 | aggresome assembly(GO:0070842) |
| 0.6 | 1.8 | GO:0045629 | negative regulation of T-helper 2 cell differentiation(GO:0045629) |
| 0.6 | 1.1 | GO:2000110 | negative regulation of macrophage apoptotic process(GO:2000110) |
| 0.6 | 4.5 | GO:0035897 | proteolysis in other organism(GO:0035897) |
| 0.5 | 10.8 | GO:0045663 | positive regulation of myoblast differentiation(GO:0045663) |
| 0.5 | 4.5 | GO:2000507 | positive regulation of energy homeostasis(GO:2000507) |
| 0.5 | 2.4 | GO:0032929 | negative regulation of superoxide anion generation(GO:0032929) |
| 0.5 | 5.6 | GO:0070365 | hepatocyte differentiation(GO:0070365) |
| 0.5 | 7.9 | GO:0007213 | G-protein coupled acetylcholine receptor signaling pathway(GO:0007213) |
| 0.5 | 2.7 | GO:0007185 | transmembrane receptor protein tyrosine phosphatase signaling pathway(GO:0007185) |
| 0.4 | 11.0 | GO:0030208 | dermatan sulfate biosynthetic process(GO:0030208) |
| 0.4 | 39.1 | GO:0060333 | interferon-gamma-mediated signaling pathway(GO:0060333) |
| 0.4 | 1.2 | GO:1902299 | pre-replicative complex assembly involved in nuclear cell cycle DNA replication(GO:0006267) pre-replicative complex assembly(GO:0036388) pre-replicative complex assembly involved in cell cycle DNA replication(GO:1902299) |
| 0.4 | 5.6 | GO:0051639 | actin filament network formation(GO:0051639) |
| 0.4 | 3.5 | GO:0010265 | SCF complex assembly(GO:0010265) |
| 0.3 | 2.1 | GO:0006499 | N-terminal protein myristoylation(GO:0006499) protein myristoylation(GO:0018377) |
| 0.3 | 10.0 | GO:0008334 | histone mRNA metabolic process(GO:0008334) |
| 0.3 | 6.1 | GO:1904714 | regulation of chaperone-mediated autophagy(GO:1904714) |
| 0.3 | 7.5 | GO:0043011 | myeloid dendritic cell differentiation(GO:0043011) |
| 0.3 | 4.3 | GO:2000059 | negative regulation of protein ubiquitination involved in ubiquitin-dependent protein catabolic process(GO:2000059) |
| 0.3 | 1.7 | GO:0006196 | AMP catabolic process(GO:0006196) |
| 0.3 | 1.4 | GO:0045359 | interferon-beta biosynthetic process(GO:0045350) regulation of interferon-beta biosynthetic process(GO:0045357) positive regulation of interferon-beta biosynthetic process(GO:0045359) |
| 0.3 | 6.1 | GO:0071577 | zinc II ion transmembrane transport(GO:0071577) |
| 0.3 | 3.0 | GO:0045842 | positive regulation of mitotic metaphase/anaphase transition(GO:0045842) positive regulation of mitotic sister chromatid separation(GO:1901970) positive regulation of metaphase/anaphase transition of cell cycle(GO:1902101) |
| 0.3 | 17.6 | GO:0045773 | positive regulation of axon extension(GO:0045773) |
| 0.3 | 1.3 | GO:0045078 | positive regulation of interferon-gamma biosynthetic process(GO:0045078) |
| 0.3 | 0.5 | GO:0002232 | leukocyte chemotaxis involved in inflammatory response(GO:0002232) |
| 0.2 | 2.7 | GO:0000463 | maturation of LSU-rRNA from tricistronic rRNA transcript (SSU-rRNA, 5.8S rRNA, LSU-rRNA)(GO:0000463) |
| 0.2 | 2.7 | GO:0070070 | proton-transporting V-type ATPase complex assembly(GO:0070070) vacuolar proton-transporting V-type ATPase complex assembly(GO:0070072) |
| 0.2 | 5.8 | GO:0002089 | lens morphogenesis in camera-type eye(GO:0002089) |
| 0.2 | 19.8 | GO:0048208 | vesicle coating(GO:0006901) vesicle targeting, rough ER to cis-Golgi(GO:0048207) COPII vesicle coating(GO:0048208) |
| 0.2 | 3.2 | GO:0090557 | establishment of endothelial intestinal barrier(GO:0090557) |
| 0.2 | 0.2 | GO:2000620 | positive regulation of histone H4-K16 acetylation(GO:2000620) |
| 0.2 | 2.8 | GO:0030150 | protein import into mitochondrial matrix(GO:0030150) |
| 0.2 | 11.6 | GO:0008631 | intrinsic apoptotic signaling pathway in response to oxidative stress(GO:0008631) |
| 0.2 | 18.8 | GO:0006521 | regulation of cellular amino acid metabolic process(GO:0006521) |
| 0.2 | 1.4 | GO:0006689 | ganglioside catabolic process(GO:0006689) |
| 0.2 | 5.2 | GO:0003094 | glomerular filtration(GO:0003094) |
| 0.2 | 9.1 | GO:0060999 | positive regulation of dendritic spine development(GO:0060999) |
| 0.2 | 2.0 | GO:0035518 | histone H2A monoubiquitination(GO:0035518) |
| 0.1 | 8.5 | GO:0060113 | inner ear receptor cell differentiation(GO:0060113) |
| 0.1 | 3.1 | GO:1900027 | regulation of ruffle assembly(GO:1900027) |
| 0.1 | 12.0 | GO:0070527 | platelet aggregation(GO:0070527) |
| 0.1 | 10.8 | GO:0070125 | mitochondrial translational elongation(GO:0070125) mitochondrial translational termination(GO:0070126) |
| 0.1 | 0.6 | GO:0071802 | negative regulation of podosome assembly(GO:0071802) |
| 0.1 | 0.4 | GO:1901842 | negative regulation of high voltage-gated calcium channel activity(GO:1901842) |
| 0.1 | 4.3 | GO:0050716 | positive regulation of interleukin-1 secretion(GO:0050716) |
| 0.1 | 2.2 | GO:0006488 | dolichol-linked oligosaccharide biosynthetic process(GO:0006488) |
| 0.1 | 0.6 | GO:0050861 | positive regulation of B cell receptor signaling pathway(GO:0050861) |
| 0.1 | 2.7 | GO:0050965 | detection of temperature stimulus involved in sensory perception(GO:0050961) detection of temperature stimulus involved in sensory perception of pain(GO:0050965) |
| 0.1 | 1.6 | GO:0006564 | L-serine biosynthetic process(GO:0006564) |
| 0.1 | 4.1 | GO:0048286 | lung alveolus development(GO:0048286) |
| 0.1 | 2.4 | GO:1900745 | positive regulation of p38MAPK cascade(GO:1900745) |
| 0.1 | 2.5 | GO:0003334 | keratinocyte development(GO:0003334) |
| 0.1 | 0.6 | GO:0055059 | asymmetric neuroblast division(GO:0055059) |
| 0.1 | 6.5 | GO:0035065 | regulation of histone acetylation(GO:0035065) |
| 0.1 | 4.1 | GO:0051865 | protein autoubiquitination(GO:0051865) |
| 0.1 | 1.2 | GO:0042276 | error-prone translesion synthesis(GO:0042276) |
| 0.1 | 4.9 | GO:0006305 | DNA alkylation(GO:0006305) DNA methylation(GO:0006306) |
| 0.1 | 1.8 | GO:0032331 | negative regulation of chondrocyte differentiation(GO:0032331) |
| 0.1 | 1.1 | GO:0019852 | L-ascorbic acid metabolic process(GO:0019852) |
| 0.1 | 1.5 | GO:0050858 | negative regulation of antigen receptor-mediated signaling pathway(GO:0050858) |
| 0.1 | 4.3 | GO:0006611 | protein export from nucleus(GO:0006611) |
| 0.1 | 4.8 | GO:0015914 | phospholipid transport(GO:0015914) |
| 0.1 | 13.7 | GO:2001020 | regulation of response to DNA damage stimulus(GO:2001020) |
| 0.1 | 2.6 | GO:0030262 | apoptotic nuclear changes(GO:0030262) |
| 0.1 | 7.1 | GO:0042752 | regulation of circadian rhythm(GO:0042752) |
| 0.1 | 8.4 | GO:0051896 | regulation of protein kinase B signaling(GO:0051896) |
| 0.1 | 15.9 | GO:0031032 | actomyosin structure organization(GO:0031032) |
| 0.1 | 1.2 | GO:0099500 | synaptic vesicle fusion to presynaptic active zone membrane(GO:0031629) vesicle fusion to plasma membrane(GO:0099500) |
| 0.1 | 2.4 | GO:0007274 | neuromuscular synaptic transmission(GO:0007274) |
| 0.1 | 0.2 | GO:0071847 | TNFSF11-mediated signaling pathway(GO:0071847) |
| 0.1 | 4.9 | GO:0050853 | B cell receptor signaling pathway(GO:0050853) |
| 0.0 | 1.8 | GO:0031440 | regulation of mRNA 3'-end processing(GO:0031440) |
| 0.0 | 6.1 | GO:0008543 | fibroblast growth factor receptor signaling pathway(GO:0008543) |
| 0.0 | 0.3 | GO:0014051 | gamma-aminobutyric acid secretion(GO:0014051) |
| 0.0 | 2.2 | GO:0044839 | cell cycle G2/M phase transition(GO:0044839) |
| 0.0 | 1.8 | GO:0048814 | regulation of dendrite morphogenesis(GO:0048814) |
| 0.0 | 0.4 | GO:0048535 | lymph node development(GO:0048535) |
| 0.0 | 5.0 | GO:0097191 | extrinsic apoptotic signaling pathway(GO:0097191) |
| 0.0 | 1.1 | GO:0032418 | lysosome localization(GO:0032418) |
| 0.0 | 0.6 | GO:0022400 | regulation of rhodopsin mediated signaling pathway(GO:0022400) |
| 0.0 | 0.9 | GO:0048704 | embryonic skeletal system morphogenesis(GO:0048704) |
| 0.0 | 0.8 | GO:0045454 | cell redox homeostasis(GO:0045454) |
| 0.0 | 1.0 | GO:2000134 | negative regulation of G1/S transition of mitotic cell cycle(GO:2000134) |
Gene overrepresentation in cellular_component category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 6.0 | 72.1 | GO:0033256 | I-kappaB/NF-kappaB complex(GO:0033256) |
| 3.1 | 9.4 | GO:0070762 | nuclear pore transmembrane ring(GO:0070762) |
| 3.0 | 33.0 | GO:0042612 | MHC class I protein complex(GO:0042612) |
| 2.5 | 20.1 | GO:0005610 | laminin-5 complex(GO:0005610) |
| 2.0 | 13.7 | GO:0032584 | growth cone membrane(GO:0032584) |
| 1.7 | 7.0 | GO:0005896 | interleukin-6 receptor complex(GO:0005896) |
| 1.7 | 10.5 | GO:0008537 | proteasome activator complex(GO:0008537) |
| 1.7 | 5.2 | GO:0071749 | IgA immunoglobulin complex(GO:0071745) IgA immunoglobulin complex, circulating(GO:0071746) monomeric IgA immunoglobulin complex(GO:0071748) polymeric IgA immunoglobulin complex(GO:0071749) secretory IgA immunoglobulin complex(GO:0071751) |
| 1.6 | 33.7 | GO:0042611 | MHC protein complex(GO:0042611) MHC class II protein complex(GO:0042613) |
| 1.2 | 18.3 | GO:0033643 | host cell part(GO:0033643) |
| 1.2 | 3.5 | GO:0043224 | nuclear SCF ubiquitin ligase complex(GO:0043224) |
| 1.0 | 7.8 | GO:0071204 | U7 snRNP(GO:0005683) histone pre-mRNA 3'end processing complex(GO:0071204) |
| 1.0 | 4.9 | GO:0033553 | rDNA heterochromatin(GO:0033553) |
| 0.9 | 2.7 | GO:0071065 | alpha9-beta1 integrin-vascular cell adhesion molecule-1 complex(GO:0071065) |
| 0.9 | 2.7 | GO:0043291 | RAVE complex(GO:0043291) |
| 0.9 | 9.8 | GO:0005614 | interstitial matrix(GO:0005614) |
| 0.9 | 1.8 | GO:0071598 | neuronal ribonucleoprotein granule(GO:0071598) |
| 0.8 | 9.7 | GO:0030008 | TRAPP complex(GO:0030008) |
| 0.8 | 5.6 | GO:0031415 | NatA complex(GO:0031415) |
| 0.8 | 8.8 | GO:0000176 | nuclear exosome (RNase complex)(GO:0000176) |
| 0.7 | 5.1 | GO:1990589 | ATF4-CREB1 transcription factor complex(GO:1990589) |
| 0.7 | 4.9 | GO:0019815 | B cell receptor complex(GO:0019815) |
| 0.7 | 6.9 | GO:0016593 | Cdc73/Paf1 complex(GO:0016593) |
| 0.7 | 8.2 | GO:0044615 | nuclear pore nuclear basket(GO:0044615) |
| 0.6 | 5.6 | GO:0000125 | PCAF complex(GO:0000125) |
| 0.6 | 6.1 | GO:0005853 | eukaryotic translation elongation factor 1 complex(GO:0005853) |
| 0.5 | 7.6 | GO:0031618 | nuclear pericentric heterochromatin(GO:0031618) |
| 0.5 | 7.9 | GO:0042622 | photoreceptor outer segment membrane(GO:0042622) |
| 0.5 | 7.3 | GO:0000145 | exocyst(GO:0000145) |
| 0.5 | 2.7 | GO:0070545 | PeBoW complex(GO:0070545) |
| 0.4 | 3.0 | GO:0030478 | actin cap(GO:0030478) |
| 0.4 | 10.8 | GO:0000314 | organellar small ribosomal subunit(GO:0000314) mitochondrial small ribosomal subunit(GO:0005763) |
| 0.4 | 9.1 | GO:0032156 | septin cytoskeleton(GO:0032156) |
| 0.4 | 1.2 | GO:0005656 | nuclear pre-replicative complex(GO:0005656) pre-replicative complex(GO:0036387) |
| 0.4 | 5.6 | GO:0005916 | fascia adherens(GO:0005916) |
| 0.4 | 1.4 | GO:0005889 | hydrogen:potassium-exchanging ATPase complex(GO:0005889) |
| 0.3 | 11.8 | GO:0032420 | stereocilium(GO:0032420) |
| 0.3 | 3.6 | GO:0042105 | alpha-beta T cell receptor complex(GO:0042105) |
| 0.3 | 24.8 | GO:0030670 | phagocytic vesicle membrane(GO:0030670) |
| 0.3 | 14.7 | GO:0002102 | podosome(GO:0002102) |
| 0.3 | 2.0 | GO:0031465 | Cul4B-RING E3 ubiquitin ligase complex(GO:0031465) |
| 0.3 | 21.4 | GO:1904724 | tertiary granule lumen(GO:1904724) |
| 0.3 | 8.3 | GO:0005839 | proteasome core complex(GO:0005839) |
| 0.2 | 2.8 | GO:0005744 | mitochondrial inner membrane presequence translocase complex(GO:0005744) |
| 0.2 | 4.5 | GO:0016281 | eukaryotic translation initiation factor 4F complex(GO:0016281) |
| 0.2 | 3.8 | GO:0071682 | endocytic vesicle lumen(GO:0071682) |
| 0.2 | 6.4 | GO:0030140 | trans-Golgi network transport vesicle(GO:0030140) |
| 0.2 | 8.0 | GO:0005669 | transcription factor TFIID complex(GO:0005669) |
| 0.2 | 2.2 | GO:0031332 | RISC complex(GO:0016442) RNAi effector complex(GO:0031332) |
| 0.2 | 14.4 | GO:0097517 | stress fiber(GO:0001725) contractile actin filament bundle(GO:0097517) |
| 0.2 | 3.3 | GO:0017101 | aminoacyl-tRNA synthetase multienzyme complex(GO:0017101) |
| 0.2 | 11.9 | GO:0016459 | myosin complex(GO:0016459) |
| 0.1 | 8.5 | GO:0022627 | cytosolic small ribosomal subunit(GO:0022627) |
| 0.1 | 14.9 | GO:0005796 | Golgi lumen(GO:0005796) |
| 0.1 | 2.2 | GO:0031011 | Ino80 complex(GO:0031011) DNA helicase complex(GO:0033202) |
| 0.1 | 19.7 | GO:0072562 | blood microparticle(GO:0072562) |
| 0.1 | 12.4 | GO:1904813 | ficolin-1-rich granule(GO:0101002) ficolin-1-rich granule lumen(GO:1904813) |
| 0.1 | 1.3 | GO:0034098 | VCP-NPL4-UFD1 AAA ATPase complex(GO:0034098) |
| 0.1 | 6.7 | GO:0060170 | ciliary membrane(GO:0060170) |
| 0.1 | 5.1 | GO:0016328 | lateral plasma membrane(GO:0016328) |
| 0.1 | 20.2 | GO:0000793 | condensed chromosome(GO:0000793) |
| 0.1 | 3.7 | GO:0097542 | ciliary tip(GO:0097542) |
| 0.1 | 1.2 | GO:0042575 | DNA polymerase complex(GO:0042575) |
| 0.1 | 4.6 | GO:0031594 | neuromuscular junction(GO:0031594) |
| 0.1 | 36.4 | GO:0010008 | endosome membrane(GO:0010008) |
| 0.1 | 29.8 | GO:0045121 | membrane raft(GO:0045121) membrane microdomain(GO:0098857) |
| 0.1 | 3.0 | GO:0016235 | aggresome(GO:0016235) |
| 0.1 | 5.0 | GO:0005788 | endoplasmic reticulum lumen(GO:0005788) |
| 0.1 | 9.0 | GO:0005769 | early endosome(GO:0005769) |
| 0.1 | 6.6 | GO:0071013 | catalytic step 2 spliceosome(GO:0071013) |
| 0.1 | 27.3 | GO:0005925 | focal adhesion(GO:0005925) |
| 0.1 | 4.2 | GO:0035578 | azurophil granule lumen(GO:0035578) |
| 0.1 | 20.5 | GO:0005764 | lytic vacuole(GO:0000323) lysosome(GO:0005764) |
| 0.1 | 0.5 | GO:0072559 | NLRP3 inflammasome complex(GO:0072559) |
| 0.1 | 1.4 | GO:0008180 | COP9 signalosome(GO:0008180) |
| 0.0 | 1.2 | GO:0031201 | SNARE complex(GO:0031201) |
| 0.0 | 4.1 | GO:0005793 | endoplasmic reticulum-Golgi intermediate compartment(GO:0005793) |
| 0.0 | 2.4 | GO:0005884 | actin filament(GO:0005884) |
| 0.0 | 16.5 | GO:0030659 | cytoplasmic vesicle membrane(GO:0030659) |
| 0.0 | 3.9 | GO:0031674 | I band(GO:0031674) |
| 0.0 | 10.8 | GO:0005578 | proteinaceous extracellular matrix(GO:0005578) |
| 0.0 | 6.9 | GO:0031410 | cytoplasmic vesicle(GO:0031410) |
| 0.0 | 4.7 | GO:0005739 | mitochondrion(GO:0005739) |
| 0.0 | 1.1 | GO:0031519 | PcG protein complex(GO:0031519) |
| 0.0 | 3.0 | GO:0032993 | protein-DNA complex(GO:0032993) |
| 0.0 | 1.9 | GO:0000932 | cytoplasmic mRNA processing body(GO:0000932) |
| 0.0 | 3.1 | GO:0030425 | dendrite(GO:0030425) |
Gene overrepresentation in molecular_function category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 6.0 | 23.8 | GO:0044378 | non-sequence-specific DNA binding, bending(GO:0044378) |
| 5.1 | 25.7 | GO:0004906 | interferon-gamma receptor activity(GO:0004906) |
| 3.7 | 22.5 | GO:0046979 | TAP2 binding(GO:0046979) |
| 2.8 | 28.1 | GO:0061578 | Lys63-specific deubiquitinase activity(GO:0061578) |
| 2.3 | 6.8 | GO:0031731 | CCR6 chemokine receptor binding(GO:0031731) |
| 2.3 | 9.0 | GO:0004727 | prenylated protein tyrosine phosphatase activity(GO:0004727) |
| 2.2 | 26.0 | GO:0000774 | adenyl-nucleotide exchange factor activity(GO:0000774) |
| 2.1 | 29.0 | GO:0032395 | MHC class II receptor activity(GO:0032395) |
| 2.0 | 7.9 | GO:0047391 | alkylglycerophosphoethanolamine phosphodiesterase activity(GO:0047391) |
| 1.7 | 6.8 | GO:0004666 | prostaglandin-endoperoxide synthase activity(GO:0004666) arachidonate 15-lipoxygenase activity(GO:0050473) |
| 1.6 | 26.1 | GO:0045236 | CXCR chemokine receptor binding(GO:0045236) |
| 1.6 | 4.8 | GO:0097677 | STAT family protein binding(GO:0097677) |
| 1.6 | 4.8 | GO:0004020 | adenylylsulfate kinase activity(GO:0004020) sulfate adenylyltransferase activity(GO:0004779) sulfate adenylyltransferase (ATP) activity(GO:0004781) |
| 1.6 | 7.8 | GO:0071208 | histone pre-mRNA DCP binding(GO:0071208) |
| 1.5 | 10.5 | GO:0061133 | endopeptidase activator activity(GO:0061133) |
| 1.4 | 8.6 | GO:0035800 | deubiquitinase activator activity(GO:0035800) |
| 1.2 | 9.3 | GO:0001849 | complement component C1q binding(GO:0001849) |
| 1.2 | 4.7 | GO:0042289 | MHC class II protein binding(GO:0042289) |
| 1.1 | 18.3 | GO:0032036 | myosin heavy chain binding(GO:0032036) |
| 1.1 | 56.4 | GO:0051019 | mitogen-activated protein kinase binding(GO:0051019) |
| 1.1 | 40.8 | GO:0008139 | nuclear localization sequence binding(GO:0008139) |
| 1.0 | 9.8 | GO:0031730 | CCR5 chemokine receptor binding(GO:0031730) |
| 0.9 | 41.7 | GO:0061631 | ubiquitin conjugating enzyme activity(GO:0061631) |
| 0.9 | 8.0 | GO:0000990 | transcription factor activity, core RNA polymerase binding(GO:0000990) |
| 0.9 | 5.1 | GO:0001225 | RNA polymerase II transcription coactivator binding(GO:0001225) |
| 0.8 | 3.4 | GO:0016532 | superoxide dismutase copper chaperone activity(GO:0016532) |
| 0.8 | 38.2 | GO:0043027 | cysteine-type endopeptidase inhibitor activity involved in apoptotic process(GO:0043027) |
| 0.8 | 7.0 | GO:0005138 | interleukin-6 receptor binding(GO:0005138) |
| 0.7 | 2.1 | GO:0016314 | phosphatidylinositol-3,4,5-trisphosphate 3-phosphatase activity(GO:0016314) |
| 0.7 | 3.4 | GO:0004699 | calcium-independent protein kinase C activity(GO:0004699) |
| 0.7 | 3.4 | GO:0004161 | dimethylallyltranstransferase activity(GO:0004161) geranyltranstransferase activity(GO:0004337) |
| 0.7 | 3.3 | GO:0004815 | aspartate-tRNA ligase activity(GO:0004815) |
| 0.6 | 30.0 | GO:0005164 | tumor necrosis factor receptor binding(GO:0005164) |
| 0.6 | 10.5 | GO:0030881 | beta-2-microglobulin binding(GO:0030881) |
| 0.6 | 11.3 | GO:0070628 | proteasome binding(GO:0070628) |
| 0.5 | 9.1 | GO:0004707 | MAP kinase activity(GO:0004707) |
| 0.5 | 7.2 | GO:0008527 | taste receptor activity(GO:0008527) |
| 0.5 | 9.4 | GO:0017056 | structural constituent of nuclear pore(GO:0017056) |
| 0.4 | 3.5 | GO:0019788 | NEDD8 transferase activity(GO:0019788) |
| 0.4 | 5.6 | GO:0004596 | peptide alpha-N-acetyltransferase activity(GO:0004596) |
| 0.4 | 1.3 | GO:0036435 | K48-linked polyubiquitin binding(GO:0036435) |
| 0.4 | 3.9 | GO:0070053 | thrombospondin receptor activity(GO:0070053) |
| 0.4 | 11.0 | GO:0005540 | hyaluronic acid binding(GO:0005540) |
| 0.3 | 2.7 | GO:0008131 | primary amine oxidase activity(GO:0008131) |
| 0.3 | 2.7 | GO:0019863 | IgE binding(GO:0019863) |
| 0.3 | 5.6 | GO:0017166 | vinculin binding(GO:0017166) |
| 0.3 | 4.5 | GO:0004708 | MAP kinase kinase activity(GO:0004708) |
| 0.3 | 6.1 | GO:0005385 | zinc ion transmembrane transporter activity(GO:0005385) |
| 0.3 | 1.6 | GO:0004647 | phosphoserine phosphatase activity(GO:0004647) |
| 0.3 | 4.5 | GO:0008190 | eukaryotic initiation factor 4E binding(GO:0008190) |
| 0.3 | 10.1 | GO:0008536 | Ran GTPase binding(GO:0008536) |
| 0.3 | 6.5 | GO:0002162 | dystroglycan binding(GO:0002162) |
| 0.3 | 2.8 | GO:0015450 | P-P-bond-hydrolysis-driven protein transmembrane transporter activity(GO:0015450) |
| 0.3 | 16.2 | GO:0042805 | actinin binding(GO:0042805) |
| 0.3 | 2.2 | GO:0005250 | A-type (transient outward) potassium channel activity(GO:0005250) |
| 0.3 | 4.9 | GO:0001164 | RNA polymerase I regulatory region DNA binding(GO:0001013) RNA polymerase I regulatory region sequence-specific DNA binding(GO:0001163) RNA polymerase I CORE element sequence-specific DNA binding(GO:0001164) |
| 0.3 | 23.4 | GO:0005201 | extracellular matrix structural constituent(GO:0005201) |
| 0.3 | 1.3 | GO:0031708 | endothelin B receptor binding(GO:0031708) |
| 0.3 | 8.5 | GO:0017134 | fibroblast growth factor binding(GO:0017134) |
| 0.2 | 2.2 | GO:0050072 | m7G(5')pppN diphosphatase activity(GO:0050072) |
| 0.2 | 1.8 | GO:0004652 | polynucleotide adenylyltransferase activity(GO:0004652) |
| 0.2 | 6.1 | GO:0003746 | translation elongation factor activity(GO:0003746) |
| 0.2 | 1.7 | GO:0047623 | AMP deaminase activity(GO:0003876) adenosine-phosphate deaminase activity(GO:0047623) |
| 0.2 | 1.0 | GO:0008420 | CTD phosphatase activity(GO:0008420) |
| 0.2 | 2.4 | GO:0008199 | ferric iron binding(GO:0008199) |
| 0.2 | 6.3 | GO:0017025 | TBP-class protein binding(GO:0017025) |
| 0.2 | 1.4 | GO:0004563 | beta-N-acetylhexosaminidase activity(GO:0004563) |
| 0.2 | 9.0 | GO:0017112 | Rab guanyl-nucleotide exchange factor activity(GO:0017112) |
| 0.2 | 12.7 | GO:0019003 | GDP binding(GO:0019003) |
| 0.2 | 2.0 | GO:0042834 | peptidoglycan binding(GO:0042834) |
| 0.2 | 6.2 | GO:0004532 | exoribonuclease activity(GO:0004532) |
| 0.1 | 3.6 | GO:0019198 | transmembrane receptor protein tyrosine phosphatase activity(GO:0005001) transmembrane receptor protein phosphatase activity(GO:0019198) |
| 0.1 | 5.6 | GO:0004527 | exonuclease activity(GO:0004527) |
| 0.1 | 2.2 | GO:0046527 | glucosyltransferase activity(GO:0046527) |
| 0.1 | 14.1 | GO:0005262 | calcium channel activity(GO:0005262) |
| 0.1 | 14.8 | GO:0000149 | SNARE binding(GO:0000149) |
| 0.1 | 2.7 | GO:0008191 | metalloendopeptidase inhibitor activity(GO:0008191) |
| 0.1 | 3.0 | GO:0005523 | tropomyosin binding(GO:0005523) |
| 0.1 | 28.6 | GO:0003924 | GTPase activity(GO:0003924) |
| 0.1 | 3.8 | GO:0042056 | chemoattractant activity(GO:0042056) |
| 0.1 | 7.0 | GO:0035064 | methylated histone binding(GO:0035064) |
| 0.1 | 0.4 | GO:0019976 | interleukin-2 receptor activity(GO:0004911) interleukin-2 binding(GO:0019976) |
| 0.1 | 1.7 | GO:0051428 | peptide hormone receptor binding(GO:0051428) |
| 0.1 | 3.3 | GO:0001103 | RNA polymerase II repressing transcription factor binding(GO:0001103) |
| 0.1 | 5.1 | GO:0004402 | histone acetyltransferase activity(GO:0004402) |
| 0.1 | 1.9 | GO:0071837 | HMG box domain binding(GO:0071837) |
| 0.1 | 11.0 | GO:0001046 | core promoter sequence-specific DNA binding(GO:0001046) |
| 0.1 | 6.4 | GO:0001047 | core promoter binding(GO:0001047) |
| 0.1 | 1.2 | GO:0003688 | DNA replication origin binding(GO:0003688) |
| 0.1 | 5.8 | GO:0005070 | SH3/SH2 adaptor activity(GO:0005070) |
| 0.1 | 9.8 | GO:0008201 | heparin binding(GO:0008201) |
| 0.1 | 5.7 | GO:0005089 | Rho guanyl-nucleotide exchange factor activity(GO:0005089) |
| 0.1 | 8.8 | GO:0017124 | SH3 domain binding(GO:0017124) |
| 0.1 | 10.8 | GO:0003735 | structural constituent of ribosome(GO:0003735) |
| 0.0 | 1.1 | GO:0015002 | cytochrome-c oxidase activity(GO:0004129) heme-copper terminal oxidase activity(GO:0015002) oxidoreductase activity, acting on a heme group of donors, oxygen as acceptor(GO:0016676) |
| 0.0 | 1.5 | GO:0004198 | calcium-dependent cysteine-type endopeptidase activity(GO:0004198) |
| 0.0 | 2.0 | GO:0003899 | DNA-directed RNA polymerase activity(GO:0003899) |
| 0.0 | 0.4 | GO:0016494 | C-X-C chemokine receptor activity(GO:0016494) |
| 0.0 | 0.1 | GO:0004692 | cGMP-dependent protein kinase activity(GO:0004692) |
| 0.0 | 1.7 | GO:0004715 | non-membrane spanning protein tyrosine kinase activity(GO:0004715) |
| 0.0 | 2.0 | GO:0003684 | damaged DNA binding(GO:0003684) |
| 0.0 | 0.1 | GO:0043184 | vascular endothelial growth factor receptor binding(GO:0005172) vascular endothelial growth factor receptor 2 binding(GO:0043184) |
| 0.0 | 1.4 | GO:0043021 | ribonucleoprotein complex binding(GO:0043021) |
| 0.0 | 0.2 | GO:0055056 | D-glucose transmembrane transporter activity(GO:0055056) |
| 0.0 | 4.0 | GO:0030246 | carbohydrate binding(GO:0030246) |
| 0.0 | 0.2 | GO:0003682 | chromatin binding(GO:0003682) |
| 0.0 | 2.6 | GO:0044822 | mRNA binding(GO:0003729) poly(A) RNA binding(GO:0044822) |
| 0.0 | 0.7 | GO:0003727 | single-stranded RNA binding(GO:0003727) |
Gene overrepresentation in C2:CP category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 2.7 | 111.8 | PID NFKAPPAB CANONICAL PATHWAY | Canonical NF-kappaB pathway |
| 1.2 | 12.1 | ST IL 13 PATHWAY | Interleukin 13 (IL-13) Pathway |
| 1.1 | 46.3 | ST TUMOR NECROSIS FACTOR PATHWAY | Tumor Necrosis Factor Pathway. |
| 0.9 | 20.1 | PID INTEGRIN4 PATHWAY | Alpha6 beta4 integrin-ligand interactions |
| 0.7 | 10.8 | PID TRAIL PATHWAY | TRAIL signaling pathway |
| 0.6 | 16.2 | PID TOLL ENDOGENOUS PATHWAY | Endogenous TLR signaling |
| 0.6 | 32.4 | PID IL8 CXCR2 PATHWAY | IL8- and CXCR2-mediated signaling events |
| 0.6 | 20.9 | PID PRL SIGNALING EVENTS PATHWAY | Signaling events mediated by PRL |
| 0.5 | 24.4 | PID IL23 PATHWAY | IL23-mediated signaling events |
| 0.5 | 18.2 | ST GRANULE CELL SURVIVAL PATHWAY | Granule Cell Survival Pathway is a specific case of more general PAC1 Receptor Pathway. |
| 0.5 | 7.3 | PID ANTHRAX PATHWAY | Cellular roles of Anthrax toxin |
| 0.4 | 26.7 | ST FAS SIGNALING PATHWAY | Fas Signaling Pathway |
| 0.4 | 8.8 | SA PTEN PATHWAY | PTEN is a tumor suppressor that dephosphorylates the lipid messenger phosphatidylinositol triphosphate. |
| 0.4 | 6.8 | PID S1P S1P1 PATHWAY | S1P1 pathway |
| 0.4 | 22.6 | PID IL12 2PATHWAY | IL12-mediated signaling events |
| 0.4 | 5.3 | PID LYMPH ANGIOGENESIS PATHWAY | VEGFR3 signaling in lymphatic endothelium |
| 0.3 | 9.3 | ST ADRENERGIC | Adrenergic Pathway |
| 0.3 | 11.8 | PID THROMBIN PAR1 PATHWAY | PAR1-mediated thrombin signaling events |
| 0.3 | 9.2 | PID SMAD2 3PATHWAY | Regulation of cytoplasmic and nuclear SMAD2/3 signaling |
| 0.3 | 10.6 | PID SYNDECAN 4 PATHWAY | Syndecan-4-mediated signaling events |
| 0.2 | 18.7 | PID DELTA NP63 PATHWAY | Validated transcriptional targets of deltaNp63 isoforms |
| 0.2 | 1.6 | PID NFKAPPAB ATYPICAL PATHWAY | Atypical NF-kappaB pathway |
| 0.2 | 10.4 | PID BCR 5PATHWAY | BCR signaling pathway |
| 0.2 | 4.8 | PID IFNG PATHWAY | IFN-gamma pathway |
| 0.2 | 1.2 | PID IL27 PATHWAY | IL27-mediated signaling events |
| 0.2 | 6.2 | PID RAS PATHWAY | Regulation of Ras family activation |
| 0.2 | 5.6 | PID MYC PATHWAY | C-MYC pathway |
| 0.1 | 7.6 | PID AURORA B PATHWAY | Aurora B signaling |
| 0.1 | 5.8 | PID CDC42 REG PATHWAY | Regulation of CDC42 activity |
| 0.1 | 3.5 | PID HIF1A PATHWAY | Hypoxic and oxygen homeostasis regulation of HIF-1-alpha |
| 0.1 | 2.7 | ST ERK1 ERK2 MAPK PATHWAY | ERK1/ERK2 MAPK Pathway |
| 0.1 | 3.3 | PID RAC1 REG PATHWAY | Regulation of RAC1 activity |
| 0.1 | 1.7 | PID HEDGEHOG 2PATHWAY | Signaling events mediated by the Hedgehog family |
| 0.1 | 3.1 | PID ILK PATHWAY | Integrin-linked kinase signaling |
| 0.1 | 1.9 | PID ENDOTHELIN PATHWAY | Endothelins |
| 0.0 | 4.5 | PID MYC ACTIV PATHWAY | Validated targets of C-MYC transcriptional activation |
| 0.0 | 1.3 | PID ALPHA SYNUCLEIN PATHWAY | Alpha-synuclein signaling |
Gene overrepresentation in C2:CP:REACTOME category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 3.3 | 33.0 | REACTOME ENDOSOMAL VACUOLAR PATHWAY | Genes involved in Endosomal/Vacuolar pathway |
| 1.9 | 46.1 | REACTOME REGULATION OF IFNG SIGNALING | Genes involved in Regulation of IFNG signaling |
| 1.8 | 26.5 | REACTOME NFKB IS ACTIVATED AND SIGNALS SURVIVAL | Genes involved in NF-kB is activated and signals survival |
| 1.4 | 72.1 | REACTOME NEGATIVE REGULATORS OF RIG I MDA5 SIGNALING | Genes involved in Negative regulators of RIG-I/MDA5 signaling |
| 1.2 | 23.8 | REACTOME APOPTOSIS INDUCED DNA FRAGMENTATION | Genes involved in Apoptosis induced DNA fragmentation |
| 1.1 | 16.7 | REACTOME TAK1 ACTIVATES NFKB BY PHOSPHORYLATION AND ACTIVATION OF IKKS COMPLEX | Genes involved in TAK1 activates NFkB by phosphorylation and activation of IKKs complex |
| 1.0 | 23.7 | REACTOME TRANSLOCATION OF ZAP 70 TO IMMUNOLOGICAL SYNAPSE | Genes involved in Translocation of ZAP-70 to Immunological synapse |
| 1.0 | 8.8 | REACTOME MRNA DECAY BY 3 TO 5 EXORIBONUCLEASE | Genes involved in mRNA Decay by 3' to 5' Exoribonuclease |
| 0.9 | 44.2 | REACTOME SEMA4D IN SEMAPHORIN SIGNALING | Genes involved in Sema4D in semaphorin signaling |
| 0.8 | 34.8 | REACTOME NOD1 2 SIGNALING PATHWAY | Genes involved in NOD1/2 Signaling Pathway |
| 0.8 | 4.5 | REACTOME ACTIVATED TAK1 MEDIATES P38 MAPK ACTIVATION | Genes involved in activated TAK1 mediates p38 MAPK activation |
| 0.7 | 7.0 | REACTOME IL 6 SIGNALING | Genes involved in Interleukin-6 signaling |
| 0.5 | 26.0 | REACTOME SIGNALING BY ROBO RECEPTOR | Genes involved in Signaling by Robo receptor |
| 0.5 | 34.3 | REACTOME CHEMOKINE RECEPTORS BIND CHEMOKINES | Genes involved in Chemokine receptors bind chemokines |
| 0.5 | 13.4 | REACTOME CELL EXTRACELLULAR MATRIX INTERACTIONS | Genes involved in Cell-extracellular matrix interactions |
| 0.5 | 7.8 | REACTOME SLBP DEPENDENT PROCESSING OF REPLICATION DEPENDENT HISTONE PRE MRNAS | Genes involved in SLBP Dependent Processing of Replication-Dependent Histone Pre-mRNAs |
| 0.4 | 14.9 | REACTOME A TETRASACCHARIDE LINKER SEQUENCE IS REQUIRED FOR GAG SYNTHESIS | Genes involved in A tetrasaccharide linker sequence is required for GAG synthesis |
| 0.4 | 3.5 | REACTOME VIF MEDIATED DEGRADATION OF APOBEC3G | Genes involved in Vif-mediated degradation of APOBEC3G |
| 0.4 | 6.1 | REACTOME ZINC TRANSPORTERS | Genes involved in Zinc transporters |
| 0.4 | 26.7 | REACTOME AUTODEGRADATION OF THE E3 UBIQUITIN LIGASE COP1 | Genes involved in Autodegradation of the E3 ubiquitin ligase COP1 |
| 0.4 | 9.8 | REACTOME TRAF6 MEDIATED IRF7 ACTIVATION | Genes involved in TRAF6 mediated IRF7 activation |
| 0.3 | 1.4 | REACTOME RIP MEDIATED NFKB ACTIVATION VIA DAI | Genes involved in RIP-mediated NFkB activation via DAI |
| 0.3 | 7.9 | REACTOME G BETA GAMMA SIGNALLING THROUGH PLC BETA | Genes involved in G beta:gamma signalling through PLC beta |
| 0.3 | 2.2 | REACTOME DESTABILIZATION OF MRNA BY BRF1 | Genes involved in Destabilization of mRNA by Butyrate Response Factor 1 (BRF1) |
| 0.3 | 10.4 | REACTOME NUCLEAR EVENTS KINASE AND TRANSCRIPTION FACTOR ACTIVATION | Genes involved in Nuclear Events (kinase and transcription factor activation) |
| 0.2 | 8.2 | REACTOME TRANSPORT OF RIBONUCLEOPROTEINS INTO THE HOST NUCLEUS | Genes involved in Transport of Ribonucleoproteins into the Host Nucleus |
| 0.2 | 4.5 | REACTOME MTORC1 MEDIATED SIGNALLING | Genes involved in mTORC1-mediated signalling |
| 0.2 | 4.8 | REACTOME CYTOSOLIC SULFONATION OF SMALL MOLECULES | Genes involved in Cytosolic sulfonation of small molecules |
| 0.1 | 1.8 | REACTOME PROCESSING OF CAPPED INTRONLESS PRE MRNA | Genes involved in Processing of Capped Intronless Pre-mRNA |
| 0.1 | 5.7 | REACTOME NRAGE SIGNALS DEATH THROUGH JNK | Genes involved in NRAGE signals death through JNK |
| 0.1 | 8.5 | REACTOME FORMATION OF THE TERNARY COMPLEX AND SUBSEQUENTLY THE 43S COMPLEX | Genes involved in Formation of the ternary complex, and subsequently, the 43S complex |
| 0.1 | 4.9 | REACTOME ANTIGEN ACTIVATES B CELL RECEPTOR LEADING TO GENERATION OF SECOND MESSENGERS | Genes involved in Antigen Activates B Cell Receptor Leading to Generation of Second Messengers |
| 0.1 | 1.2 | REACTOME CDC6 ASSOCIATION WITH THE ORC ORIGIN COMPLEX | Genes involved in CDC6 association with the ORC:origin complex |
| 0.1 | 5.6 | REACTOME RNA POL II TRANSCRIPTION PRE INITIATION AND PROMOTER OPENING | Genes involved in RNA Polymerase II Transcription Pre-Initiation And Promoter Opening |
| 0.1 | 3.3 | REACTOME CYTOSOLIC TRNA AMINOACYLATION | Genes involved in Cytosolic tRNA aminoacylation |
| 0.1 | 2.2 | REACTOME BIOSYNTHESIS OF THE N GLYCAN PRECURSOR DOLICHOL LIPID LINKED OLIGOSACCHARIDE LLO AND TRANSFER TO A NASCENT PROTEIN | Genes involved in Biosynthesis of the N-glycan precursor (dolichol lipid-linked oligosaccharide, LLO) and transfer to a nascent protein |
| 0.1 | 4.2 | REACTOME INTERFERON GAMMA SIGNALING | Genes involved in Interferon gamma signaling |
| 0.1 | 6.8 | REACTOME PHASE1 FUNCTIONALIZATION OF COMPOUNDS | Genes involved in Phase 1 - Functionalization of compounds |
| 0.1 | 4.7 | REACTOME PI METABOLISM | Genes involved in PI Metabolism |
| 0.1 | 2.1 | REACTOME ANTIVIRAL MECHANISM BY IFN STIMULATED GENES | Genes involved in Antiviral mechanism by IFN-stimulated genes |
| 0.1 | 1.7 | REACTOME PURINE SALVAGE | Genes involved in Purine salvage |
| 0.1 | 3.0 | REACTOME SMOOTH MUSCLE CONTRACTION | Genes involved in Smooth Muscle Contraction |
| 0.1 | 6.1 | REACTOME PEPTIDE CHAIN ELONGATION | Genes involved in Peptide chain elongation |
| 0.1 | 1.6 | REACTOME AMINO ACID SYNTHESIS AND INTERCONVERSION TRANSAMINATION | Genes involved in Amino acid synthesis and interconversion (transamination) |
| 0.0 | 3.5 | REACTOME METABOLISM OF VITAMINS AND COFACTORS | Genes involved in Metabolism of vitamins and cofactors |
| 0.0 | 4.7 | REACTOME MHC CLASS II ANTIGEN PRESENTATION | Genes involved in MHC class II antigen presentation |
| 0.0 | 2.8 | REACTOME MITOCHONDRIAL PROTEIN IMPORT | Genes involved in Mitochondrial Protein Import |
| 0.0 | 1.2 | REACTOME PROTEOLYTIC CLEAVAGE OF SNARE COMPLEX PROTEINS | Genes involved in Proteolytic cleavage of SNARE complex proteins |
| 0.0 | 0.5 | REACTOME THE NLRP3 INFLAMMASOME | Genes involved in The NLRP3 inflammasome |
| 0.0 | 1.4 | REACTOME GLYCOSPHINGOLIPID METABOLISM | Genes involved in Glycosphingolipid metabolism |
| 0.0 | 2.9 | REACTOME MRNA SPLICING | Genes involved in mRNA Splicing |
| 0.0 | 0.3 | REACTOME ROLE OF SECOND MESSENGERS IN NETRIN1 SIGNALING | Genes involved in Role of second messengers in netrin-1 signaling |
| 0.0 | 0.7 | REACTOME CHOLESTEROL BIOSYNTHESIS | Genes involved in Cholesterol biosynthesis |
| 0.0 | 4.1 | REACTOME ANTIGEN PROCESSING UBIQUITINATION PROTEASOME DEGRADATION | Genes involved in Antigen processing: Ubiquitination & Proteasome degradation |
| 0.0 | 3.0 | REACTOME FACTORS INVOLVED IN MEGAKARYOCYTE DEVELOPMENT AND PLATELET PRODUCTION | Genes involved in Factors involved in megakaryocyte development and platelet production |
| 0.0 | 0.6 | REACTOME EFFECTS OF PIP2 HYDROLYSIS | Genes involved in Effects of PIP2 hydrolysis |
| 0.0 | 1.8 | REACTOME G ALPHA Q SIGNALLING EVENTS | Genes involved in G alpha (q) signalling events |
| 0.0 | 1.2 | REACTOME DNA REPAIR | Genes involved in DNA Repair |
| 0.0 | 0.2 | REACTOME FACILITATIVE NA INDEPENDENT GLUCOSE TRANSPORTERS | Genes involved in Facilitative Na+-independent glucose transporters |