Project
GNF SymAtlas + NCI-60 cancer cell lines, human (Su, 2004; Ross, 2000)
Navigation
Downloads
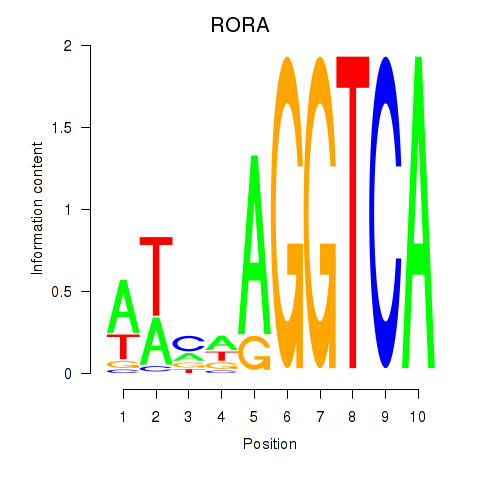
Results for RORA
Z-value: 0.98
Transcription factors associated with RORA
| Gene Symbol | Gene ID | Gene Info |
|---|---|---|
|
RORA
|
ENSG00000069667.11 | RAR related orphan receptor A |
Activity-expression correlation:
| Gene | Promoter | Pearson corr. coef. | P-value | Plot |
|---|---|---|---|---|
| RORA | hg19_v2_chr15_-_60884706_60884743, hg19_v2_chr15_-_61521495_61521518 | -0.05 | 4.6e-01 | Click! |
Activity profile of RORA motif
Sorted Z-values of RORA motif
| Promoter | Log-likelihood | Transcript | Gene | Gene Info |
|---|---|---|---|---|
| chr20_+_3776371 | 30.51 |
ENST00000245960.5
|
CDC25B
|
cell division cycle 25B |
| chr20_+_3776936 | 29.46 |
ENST00000439880.2
|
CDC25B
|
cell division cycle 25B |
| chr16_-_88717423 | 27.08 |
ENST00000568278.1
ENST00000569359.1 ENST00000567174.1 |
CYBA
|
cytochrome b-245, alpha polypeptide |
| chr1_+_43148625 | 25.26 |
ENST00000436427.1
|
YBX1
|
Y box binding protein 1 |
| chr14_-_58893832 | 22.30 |
ENST00000556007.2
|
TIMM9
|
translocase of inner mitochondrial membrane 9 homolog (yeast) |
| chr7_+_112063192 | 20.89 |
ENST00000005558.4
|
IFRD1
|
interferon-related developmental regulator 1 |
| chr1_-_150207017 | 20.17 |
ENST00000369119.3
|
ANP32E
|
acidic (leucine-rich) nuclear phosphoprotein 32 family, member E |
| chr6_-_41039567 | 19.49 |
ENST00000468811.1
|
OARD1
|
O-acyl-ADP-ribose deacylase 1 |
| chr12_-_53320245 | 19.47 |
ENST00000552150.1
|
KRT8
|
keratin 8 |
| chr9_-_35103105 | 19.40 |
ENST00000452248.2
ENST00000356493.5 |
STOML2
|
stomatin (EPB72)-like 2 |
| chr1_-_153522562 | 18.55 |
ENST00000368714.1
|
S100A4
|
S100 calcium binding protein A4 |
| chr17_+_49230897 | 17.57 |
ENST00000393196.3
ENST00000336097.3 ENST00000480143.1 ENST00000511355.1 ENST00000013034.3 ENST00000393198.3 ENST00000608447.1 ENST00000393193.2 ENST00000376392.6 ENST00000555572.1 |
NME1
NME1-NME2
NME2
|
NME/NM23 nucleoside diphosphate kinase 1 NME1-NME2 readthrough NME/NM23 nucleoside diphosphate kinase 2 |
| chr7_+_128399002 | 17.27 |
ENST00000493278.1
|
CALU
|
calumenin |
| chr10_+_94050913 | 15.99 |
ENST00000358935.2
|
MARCH5
|
membrane-associated ring finger (C3HC4) 5 |
| chr10_-_65028817 | 15.43 |
ENST00000542921.1
|
JMJD1C
|
jumonji domain containing 1C |
| chrX_-_129299638 | 15.31 |
ENST00000535724.1
ENST00000346424.2 |
AIFM1
|
apoptosis-inducing factor, mitochondrion-associated, 1 |
| chr10_+_81107271 | 14.43 |
ENST00000448165.1
|
PPIF
|
peptidylprolyl isomerase F |
| chr14_-_102552659 | 13.95 |
ENST00000441629.2
|
HSP90AA1
|
heat shock protein 90kDa alpha (cytosolic), class A member 1 |
| chr2_+_198365122 | 13.49 |
ENST00000604458.1
|
HSPE1-MOB4
|
HSPE1-MOB4 readthrough |
| chr3_-_113465065 | 13.11 |
ENST00000497255.1
ENST00000478020.1 ENST00000240922.3 ENST00000493900.1 |
NAA50
|
N(alpha)-acetyltransferase 50, NatE catalytic subunit |
| chr7_-_140179276 | 13.06 |
ENST00000443720.2
ENST00000255977.2 |
MKRN1
|
makorin ring finger protein 1 |
| chr9_-_130966497 | 12.88 |
ENST00000393608.1
ENST00000372948.3 |
CIZ1
|
CDKN1A interacting zinc finger protein 1 |
| chr1_-_211752073 | 12.78 |
ENST00000367001.4
|
SLC30A1
|
solute carrier family 30 (zinc transporter), member 1 |
| chr18_-_19284724 | 11.82 |
ENST00000580981.1
ENST00000289119.2 |
ABHD3
|
abhydrolase domain containing 3 |
| chr19_-_39826639 | 11.73 |
ENST00000602185.1
ENST00000598034.1 ENST00000601387.1 ENST00000595636.1 ENST00000253054.8 ENST00000594700.1 ENST00000597595.1 |
GMFG
|
glia maturation factor, gamma |
| chr7_-_25164969 | 11.63 |
ENST00000305786.2
|
CYCS
|
cytochrome c, somatic |
| chr21_-_46330545 | 11.45 |
ENST00000320216.6
ENST00000397852.1 |
ITGB2
|
integrin, beta 2 (complement component 3 receptor 3 and 4 subunit) |
| chr16_+_21964662 | 11.41 |
ENST00000561553.1
ENST00000565331.1 |
UQCRC2
|
ubiquinol-cytochrome c reductase core protein II |
| chr2_-_86422095 | 10.99 |
ENST00000254636.5
|
IMMT
|
inner membrane protein, mitochondrial |
| chr11_+_118272328 | 10.91 |
ENST00000524422.1
|
ATP5L
|
ATP synthase, H+ transporting, mitochondrial Fo complex, subunit G |
| chr2_+_198365095 | 10.77 |
ENST00000409468.1
|
HSPE1
|
heat shock 10kDa protein 1 |
| chr3_-_141747950 | 10.14 |
ENST00000497579.1
|
TFDP2
|
transcription factor Dp-2 (E2F dimerization partner 2) |
| chr2_-_209118974 | 10.00 |
ENST00000415913.1
ENST00000415282.1 ENST00000446179.1 |
IDH1
|
isocitrate dehydrogenase 1 (NADP+), soluble |
| chr3_+_179322573 | 9.97 |
ENST00000493866.1
ENST00000472629.1 ENST00000482604.1 |
NDUFB5
|
NADH dehydrogenase (ubiquinone) 1 beta subcomplex, 5, 16kDa |
| chr9_+_134103496 | 9.54 |
ENST00000498010.1
ENST00000476004.1 ENST00000528406.1 |
NUP214
|
nucleoporin 214kDa |
| chr1_-_33502528 | 9.32 |
ENST00000354858.6
|
AK2
|
adenylate kinase 2 |
| chr1_-_32403903 | 9.21 |
ENST00000344035.6
ENST00000356536.3 |
PTP4A2
|
protein tyrosine phosphatase type IVA, member 2 |
| chr16_-_47177874 | 9.16 |
ENST00000562435.1
|
NETO2
|
neuropilin (NRP) and tolloid (TLL)-like 2 |
| chr10_-_103578182 | 8.99 |
ENST00000439817.1
|
MGEA5
|
meningioma expressed antigen 5 (hyaluronidase) |
| chr1_-_33502441 | 8.81 |
ENST00000548033.1
ENST00000487289.1 ENST00000373449.2 ENST00000480134.1 ENST00000467905.1 |
AK2
|
adenylate kinase 2 |
| chr1_-_205719295 | 8.67 |
ENST00000367142.4
|
NUCKS1
|
nuclear casein kinase and cyclin-dependent kinase substrate 1 |
| chr2_-_207024134 | 8.63 |
ENST00000457011.1
ENST00000440274.1 ENST00000432169.1 |
NDUFS1
|
NADH dehydrogenase (ubiquinone) Fe-S protein 1, 75kDa (NADH-coenzyme Q reductase) |
| chr17_-_42144949 | 8.59 |
ENST00000591247.1
|
LSM12
|
LSM12 homolog (S. cerevisiae) |
| chr5_-_140027175 | 8.44 |
ENST00000512088.1
|
NDUFA2
|
NADH dehydrogenase (ubiquinone) 1 alpha subcomplex, 2, 8kDa |
| chr10_-_103578162 | 8.30 |
ENST00000361464.3
ENST00000357797.5 ENST00000370094.3 |
MGEA5
|
meningioma expressed antigen 5 (hyaluronidase) |
| chr1_+_206858328 | 8.05 |
ENST00000367103.3
|
MAPKAPK2
|
mitogen-activated protein kinase-activated protein kinase 2 |
| chr11_-_64511575 | 7.45 |
ENST00000431822.1
ENST00000377486.3 ENST00000394432.3 |
RASGRP2
|
RAS guanyl releasing protein 2 (calcium and DAG-regulated) |
| chr7_-_107642348 | 7.30 |
ENST00000393561.1
|
LAMB1
|
laminin, beta 1 |
| chr2_+_207024306 | 7.23 |
ENST00000236957.5
ENST00000392221.1 ENST00000392222.2 ENST00000445505.1 |
EEF1B2
|
eukaryotic translation elongation factor 1 beta 2 |
| chr19_-_39926268 | 7.19 |
ENST00000599705.1
|
RPS16
|
ribosomal protein S16 |
| chr12_-_6484376 | 7.17 |
ENST00000360168.3
ENST00000358945.3 |
SCNN1A
|
sodium channel, non-voltage-gated 1 alpha subunit |
| chr2_+_109237717 | 6.68 |
ENST00000409441.1
|
LIMS1
|
LIM and senescent cell antigen-like domains 1 |
| chr10_-_65028938 | 6.63 |
ENST00000402544.1
|
JMJD1C
|
jumonji domain containing 1C |
| chrX_+_153770421 | 6.61 |
ENST00000369609.5
ENST00000369607.1 |
IKBKG
|
inhibitor of kappa light polypeptide gene enhancer in B-cells, kinase gamma |
| chr17_-_48785216 | 6.59 |
ENST00000285243.6
|
ANKRD40
|
ankyrin repeat domain 40 |
| chr7_-_45151272 | 6.46 |
ENST00000461363.1
ENST00000495078.1 ENST00000494076.1 ENST00000478532.1 ENST00000258770.3 ENST00000361278.3 |
TBRG4
|
transforming growth factor beta regulator 4 |
| chr1_+_43148059 | 6.21 |
ENST00000321358.7
ENST00000332220.6 |
YBX1
|
Y box binding protein 1 |
| chr10_+_72575643 | 6.19 |
ENST00000373202.3
|
SGPL1
|
sphingosine-1-phosphate lyase 1 |
| chr5_+_140027355 | 6.17 |
ENST00000417647.2
ENST00000507593.1 ENST00000508301.1 |
IK
|
IK cytokine, down-regulator of HLA II |
| chr7_-_123197733 | 6.16 |
ENST00000470123.1
ENST00000471770.1 |
NDUFA5
|
NADH dehydrogenase (ubiquinone) 1 alpha subcomplex, 5 |
| chr7_-_127983877 | 5.96 |
ENST00000415472.2
ENST00000478061.1 ENST00000223073.2 ENST00000459726.1 |
RBM28
|
RNA binding motif protein 28 |
| chr11_+_111957497 | 5.65 |
ENST00000375549.3
ENST00000528182.1 ENST00000528048.1 ENST00000528021.1 ENST00000526592.1 ENST00000525291.1 |
SDHD
|
succinate dehydrogenase complex, subunit D, integral membrane protein |
| chr7_-_150777949 | 5.59 |
ENST00000482571.1
|
FASTK
|
Fas-activated serine/threonine kinase |
| chr6_-_111927062 | 5.57 |
ENST00000359831.4
|
TRAF3IP2
|
TRAF3 interacting protein 2 |
| chr11_-_111957451 | 5.37 |
ENST00000504148.2
ENST00000541231.1 |
TIMM8B
|
translocase of inner mitochondrial membrane 8 homolog B (yeast) |
| chr7_-_150777874 | 5.29 |
ENST00000540185.1
|
FASTK
|
Fas-activated serine/threonine kinase |
| chr16_-_30457048 | 5.20 |
ENST00000500504.2
ENST00000542752.1 |
SEPHS2
|
selenophosphate synthetase 2 |
| chr11_-_111741994 | 5.16 |
ENST00000398006.2
|
ALG9
|
ALG9, alpha-1,2-mannosyltransferase |
| chr7_-_150777920 | 5.02 |
ENST00000353841.2
ENST00000297532.6 |
FASTK
|
Fas-activated serine/threonine kinase |
| chr19_+_46850251 | 4.79 |
ENST00000012443.4
|
PPP5C
|
protein phosphatase 5, catalytic subunit |
| chr10_+_99185917 | 4.72 |
ENST00000334828.5
|
PGAM1
|
phosphoglycerate mutase 1 (brain) |
| chr20_-_3748416 | 4.34 |
ENST00000399672.1
|
C20orf27
|
chromosome 20 open reading frame 27 |
| chr3_+_113465866 | 4.26 |
ENST00000273398.3
ENST00000538620.1 ENST00000496747.1 ENST00000475322.1 |
ATP6V1A
|
ATPase, H+ transporting, lysosomal 70kDa, V1 subunit A |
| chr11_+_66624527 | 4.21 |
ENST00000393952.3
|
LRFN4
|
leucine rich repeat and fibronectin type III domain containing 4 |
| chr3_+_179322481 | 4.21 |
ENST00000259037.3
|
NDUFB5
|
NADH dehydrogenase (ubiquinone) 1 beta subcomplex, 5, 16kDa |
| chr17_-_41132010 | 4.13 |
ENST00000409103.1
ENST00000360221.4 |
PTGES3L-AARSD1
|
PTGES3L-AARSD1 readthrough |
| chr7_+_73868439 | 3.97 |
ENST00000424337.2
|
GTF2IRD1
|
GTF2I repeat domain containing 1 |
| chr19_+_46850320 | 3.89 |
ENST00000391919.1
|
PPP5C
|
protein phosphatase 5, catalytic subunit |
| chr18_-_12377283 | 3.83 |
ENST00000269143.3
|
AFG3L2
|
AFG3-like AAA ATPase 2 |
| chr16_-_1821496 | 3.75 |
ENST00000564628.1
ENST00000563498.1 |
NME3
|
NME/NM23 nucleoside diphosphate kinase 3 |
| chr12_+_120875910 | 3.71 |
ENST00000551806.1
|
AL021546.6
|
Glutamyl-tRNA(Gln) amidotransferase subunit C, mitochondrial |
| chr8_-_134072593 | 3.68 |
ENST00000427060.2
|
SLA
|
Src-like-adaptor |
| chr10_+_75757863 | 3.62 |
ENST00000372755.3
ENST00000211998.4 ENST00000417648.2 |
VCL
|
vinculin |
| chr16_-_30537839 | 3.30 |
ENST00000380412.5
|
ZNF768
|
zinc finger protein 768 |
| chr12_+_56414851 | 3.26 |
ENST00000547167.1
|
IKZF4
|
IKAROS family zinc finger 4 (Eos) |
| chr1_+_212738676 | 3.22 |
ENST00000366981.4
ENST00000366987.2 |
ATF3
|
activating transcription factor 3 |
| chr5_+_169780485 | 3.01 |
ENST00000377360.4
|
KCNIP1
|
Kv channel interacting protein 1 |
| chr3_-_42845951 | 2.93 |
ENST00000418900.2
ENST00000430190.1 |
HIGD1A
|
HIG1 hypoxia inducible domain family, member 1A |
| chr3_-_27498235 | 2.93 |
ENST00000295736.5
ENST00000428386.1 ENST00000428179.1 |
SLC4A7
|
solute carrier family 4, sodium bicarbonate cotransporter, member 7 |
| chr17_+_42148225 | 2.73 |
ENST00000591696.1
|
G6PC3
|
glucose 6 phosphatase, catalytic, 3 |
| chr8_-_100905925 | 2.72 |
ENST00000518171.1
|
COX6C
|
cytochrome c oxidase subunit VIc |
| chr5_-_140027357 | 2.58 |
ENST00000252102.4
|
NDUFA2
|
NADH dehydrogenase (ubiquinone) 1 alpha subcomplex, 2, 8kDa |
| chr6_-_97345689 | 2.57 |
ENST00000316149.7
|
NDUFAF4
|
NADH dehydrogenase (ubiquinone) complex I, assembly factor 4 |
| chr1_+_104159999 | 2.56 |
ENST00000414303.2
ENST00000423678.1 |
AMY2A
|
amylase, alpha 2A (pancreatic) |
| chr10_+_112257596 | 2.51 |
ENST00000369583.3
|
DUSP5
|
dual specificity phosphatase 5 |
| chr3_-_24536253 | 2.40 |
ENST00000428492.1
ENST00000396671.2 ENST00000431815.1 ENST00000418247.1 ENST00000416420.1 ENST00000356447.4 |
THRB
|
thyroid hormone receptor, beta |
| chr2_-_207024233 | 2.34 |
ENST00000423725.1
ENST00000233190.6 |
NDUFS1
|
NADH dehydrogenase (ubiquinone) Fe-S protein 1, 75kDa (NADH-coenzyme Q reductase) |
| chr9_-_35685452 | 2.23 |
ENST00000607559.1
|
TPM2
|
tropomyosin 2 (beta) |
| chr17_+_40996590 | 2.18 |
ENST00000253799.3
ENST00000452774.2 |
AOC2
|
amine oxidase, copper containing 2 (retina-specific) |
| chr6_-_41703296 | 2.10 |
ENST00000373033.1
|
TFEB
|
transcription factor EB |
| chr19_-_11689752 | 2.07 |
ENST00000592659.1
ENST00000592828.1 ENST00000218758.5 ENST00000412435.2 |
ACP5
|
acid phosphatase 5, tartrate resistant |
| chr12_-_57039739 | 2.06 |
ENST00000552959.1
ENST00000551020.1 ENST00000553007.2 ENST00000552919.1 ENST00000552104.1 ENST00000262030.3 |
ATP5B
|
ATP synthase, H+ transporting, mitochondrial F1 complex, beta polypeptide |
| chr7_-_43965937 | 1.90 |
ENST00000455877.1
ENST00000223341.7 ENST00000447717.3 ENST00000426198.1 |
URGCP
|
upregulator of cell proliferation |
| chr16_-_30538079 | 1.89 |
ENST00000562803.1
|
ZNF768
|
zinc finger protein 768 |
| chr2_+_162016804 | 1.76 |
ENST00000392749.2
ENST00000440506.1 |
TANK
|
TRAF family member-associated NFKB activator |
| chr19_+_45251804 | 1.74 |
ENST00000164227.5
|
BCL3
|
B-cell CLL/lymphoma 3 |
| chr16_+_31366536 | 1.70 |
ENST00000562522.1
|
ITGAX
|
integrin, alpha X (complement component 3 receptor 4 subunit) |
| chr20_+_49126881 | 1.68 |
ENST00000371621.3
ENST00000541713.1 |
PTPN1
|
protein tyrosine phosphatase, non-receptor type 1 |
| chr11_+_128563652 | 1.54 |
ENST00000527786.2
|
FLI1
|
Fli-1 proto-oncogene, ETS transcription factor |
| chr9_+_139717847 | 1.46 |
ENST00000436380.1
|
RABL6
|
RAB, member RAS oncogene family-like 6 |
| chr10_-_14050522 | 1.45 |
ENST00000342409.2
|
FRMD4A
|
FERM domain containing 4A |
| chr16_+_29991673 | 1.42 |
ENST00000416441.2
|
TAOK2
|
TAO kinase 2 |
| chr19_+_16059818 | 1.35 |
ENST00000322107.1
|
OR10H4
|
olfactory receptor, family 10, subfamily H, member 4 |
| chr10_+_104178946 | 1.33 |
ENST00000432590.1
|
FBXL15
|
F-box and leucine-rich repeat protein 15 |
| chr17_+_48423453 | 1.30 |
ENST00000017003.2
ENST00000509778.1 ENST00000507602.1 |
XYLT2
|
xylosyltransferase II |
| chr1_-_185286461 | 1.24 |
ENST00000367498.3
|
IVNS1ABP
|
influenza virus NS1A binding protein |
| chr8_-_100905850 | 1.18 |
ENST00000520271.1
ENST00000522940.1 ENST00000523016.1 ENST00000517682.2 ENST00000297564.2 |
COX6C
|
cytochrome c oxidase subunit VIc |
| chr19_+_48898132 | 1.18 |
ENST00000263269.3
|
GRIN2D
|
glutamate receptor, ionotropic, N-methyl D-aspartate 2D |
| chr19_-_49552006 | 1.13 |
ENST00000391869.3
|
CGB1
|
chorionic gonadotropin, beta polypeptide 1 |
| chr15_-_75017711 | 1.06 |
ENST00000567032.1
ENST00000564596.1 ENST00000566503.1 ENST00000395049.4 ENST00000395048.2 ENST00000379727.3 |
CYP1A1
|
cytochrome P450, family 1, subfamily A, polypeptide 1 |
| chr12_+_69139886 | 1.03 |
ENST00000398004.2
|
SLC35E3
|
solute carrier family 35, member E3 |
| chr12_+_93963590 | 0.95 |
ENST00000340600.2
|
SOCS2
|
suppressor of cytokine signaling 2 |
| chrX_-_108725301 | 0.93 |
ENST00000218006.2
|
GUCY2F
|
guanylate cyclase 2F, retinal |
| chr16_+_31366455 | 0.83 |
ENST00000268296.4
|
ITGAX
|
integrin, alpha X (complement component 3 receptor 4 subunit) |
| chr2_+_162016916 | 0.82 |
ENST00000405852.1
|
TANK
|
TRAF family member-associated NFKB activator |
| chr6_-_43021437 | 0.56 |
ENST00000265348.3
|
CUL7
|
cullin 7 |
| chr3_-_49377446 | 0.54 |
ENST00000351842.4
ENST00000416417.1 ENST00000415188.1 |
USP4
|
ubiquitin specific peptidase 4 (proto-oncogene) |
| chr17_-_27503770 | 0.53 |
ENST00000533112.1
|
MYO18A
|
myosin XVIIIA |
| chr5_+_32788945 | 0.52 |
ENST00000326958.1
|
AC026703.1
|
AC026703.1 |
| chr18_-_43678241 | 0.49 |
ENST00000593152.2
ENST00000589252.1 ENST00000590665.1 ENST00000398752.6 |
ATP5A1
|
ATP synthase, H+ transporting, mitochondrial F1 complex, alpha subunit 1, cardiac muscle |
| chr16_+_56899114 | 0.47 |
ENST00000566786.1
ENST00000438926.2 ENST00000563236.1 ENST00000262502.5 |
SLC12A3
|
solute carrier family 12 (sodium/chloride transporter), member 3 |
| chr15_+_78441663 | 0.36 |
ENST00000299518.2
ENST00000558554.1 ENST00000557826.1 ENST00000561279.1 ENST00000559186.1 ENST00000560770.1 ENST00000559881.1 ENST00000559205.1 ENST00000441490.2 |
IDH3A
|
isocitrate dehydrogenase 3 (NAD+) alpha |
| chr6_+_44126545 | 0.28 |
ENST00000532171.1
ENST00000398776.1 ENST00000542245.1 |
CAPN11
|
calpain 11 |
| chr8_-_100905363 | 0.27 |
ENST00000524245.1
|
COX6C
|
cytochrome c oxidase subunit VIc |
| chr17_+_4855053 | 0.23 |
ENST00000518175.1
|
ENO3
|
enolase 3 (beta, muscle) |
| chr1_+_104293028 | 0.23 |
ENST00000370079.3
|
AMY1C
|
amylase, alpha 1C (salivary) |
| chr1_+_206858232 | 0.21 |
ENST00000294981.4
|
MAPKAPK2
|
mitogen-activated protein kinase-activated protein kinase 2 |
| chr11_+_64879317 | 0.18 |
ENST00000526809.1
ENST00000279263.7 ENST00000524986.1 ENST00000534371.1 ENST00000540748.1 ENST00000525385.1 ENST00000345348.5 ENST00000531321.1 ENST00000529414.1 ENST00000526085.1 ENST00000530750.1 |
TM7SF2
|
transmembrane 7 superfamily member 2 |
| chr11_-_67271723 | 0.16 |
ENST00000533391.1
ENST00000534749.1 ENST00000532703.1 |
PITPNM1
|
phosphatidylinositol transfer protein, membrane-associated 1 |
| chr2_+_134877740 | 0.07 |
ENST00000409645.1
|
MGAT5
|
mannosyl (alpha-1,6-)-glycoprotein beta-1,6-N-acetyl-glucosaminyltransferase |
Network of associatons between targets according to the STRING database.
First level regulatory network of RORA
Gene Ontology Analysis
Gene overrepresentation in biological_process category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 7.0 | 20.9 | GO:0007518 | myoblast fate determination(GO:0007518) |
| 6.1 | 42.4 | GO:1904044 | response to aldosterone(GO:1904044) |
| 6.0 | 18.1 | GO:0006172 | ADP biosynthetic process(GO:0006172) |
| 6.0 | 60.0 | GO:0007144 | female meiosis I(GO:0007144) |
| 4.9 | 19.4 | GO:0090296 | regulation of mitochondrial DNA replication(GO:0090296) |
| 4.8 | 14.4 | GO:2000276 | regulation of mitochondrial membrane permeability involved in programmed necrotic cell death(GO:1902445) negative regulation of oxidative phosphorylation uncoupler activity(GO:2000276) |
| 4.5 | 31.5 | GO:0070934 | CRD-mediated mRNA stabilization(GO:0070934) |
| 3.5 | 13.9 | GO:0043335 | protein unfolding(GO:0043335) |
| 3.3 | 10.0 | GO:0006097 | glyoxylate cycle(GO:0006097) |
| 2.9 | 11.4 | GO:0006627 | protein processing involved in protein targeting to mitochondrion(GO:0006627) |
| 2.6 | 12.8 | GO:0061088 | regulation of sequestering of zinc ion(GO:0061088) |
| 2.5 | 22.3 | GO:0045039 | protein import into mitochondrial inner membrane(GO:0045039) |
| 2.2 | 8.7 | GO:0019046 | release from viral latency(GO:0019046) |
| 1.9 | 5.7 | GO:0006121 | mitochondrial electron transport, succinate to ubiquinone(GO:0006121) |
| 1.8 | 7.3 | GO:0021812 | neuronal-glial interaction involved in cerebral cortex radial glia guided migration(GO:0021812) |
| 1.8 | 21.3 | GO:0046051 | UTP biosynthetic process(GO:0006228) UTP metabolic process(GO:0046051) |
| 1.7 | 5.2 | GO:0016260 | selenocysteine biosynthetic process(GO:0016260) |
| 1.7 | 11.7 | GO:0034316 | negative regulation of Arp2/3 complex-mediated actin nucleation(GO:0034316) |
| 1.6 | 13.1 | GO:0070601 | centromeric sister chromatid cohesion(GO:0070601) |
| 1.4 | 19.5 | GO:0051599 | response to hydrostatic pressure(GO:0051599) |
| 1.2 | 4.7 | GO:0043456 | regulation of pentose-phosphate shunt(GO:0043456) |
| 1.1 | 3.2 | GO:0061394 | regulation of transcription from RNA polymerase II promoter in response to arsenic-containing substance(GO:0061394) |
| 1.1 | 16.0 | GO:0090344 | negative regulation of cell aging(GO:0090344) |
| 1.1 | 22.1 | GO:0033169 | histone H3-K9 demethylation(GO:0033169) |
| 1.0 | 8.3 | GO:0044351 | macropinocytosis(GO:0044351) |
| 1.0 | 17.3 | GO:0006044 | N-acetylglucosamine metabolic process(GO:0006044) |
| 1.0 | 3.8 | GO:0034982 | mitochondrial protein processing(GO:0034982) |
| 0.9 | 11.6 | GO:0006122 | mitochondrial electron transport, ubiquinol to cytochrome c(GO:0006122) |
| 0.8 | 2.4 | GO:0008050 | female courtship behavior(GO:0008050) |
| 0.8 | 4.0 | GO:0014886 | transition between slow and fast fiber(GO:0014886) |
| 0.6 | 24.5 | GO:0042407 | cristae formation(GO:0042407) |
| 0.6 | 39.7 | GO:0006120 | mitochondrial electron transport, NADH to ubiquinone(GO:0006120) |
| 0.6 | 1.7 | GO:0045082 | positive regulation of interleukin-10 biosynthetic process(GO:0045082) |
| 0.6 | 8.7 | GO:0016576 | histone dephosphorylation(GO:0016576) |
| 0.5 | 10.8 | GO:0051085 | chaperone mediated protein folding requiring cofactor(GO:0051085) |
| 0.5 | 5.4 | GO:0072321 | chaperone-mediated protein transport(GO:0072321) |
| 0.5 | 11.4 | GO:0097242 | beta-amyloid clearance(GO:0097242) |
| 0.4 | 6.2 | GO:0001553 | luteinization(GO:0001553) |
| 0.4 | 2.1 | GO:0032929 | negative regulation of superoxide anion generation(GO:0032929) |
| 0.4 | 20.2 | GO:0043486 | histone exchange(GO:0043486) |
| 0.4 | 2.7 | GO:0015760 | hexose phosphate transport(GO:0015712) glucose-6-phosphate transport(GO:0015760) |
| 0.4 | 2.6 | GO:0044245 | polysaccharide digestion(GO:0044245) |
| 0.4 | 1.1 | GO:0018894 | dibenzo-p-dioxin metabolic process(GO:0018894) cellular alkene metabolic process(GO:0043449) |
| 0.3 | 5.2 | GO:0006488 | dolichol-linked oligosaccharide biosynthetic process(GO:0006488) |
| 0.3 | 1.7 | GO:0030948 | negative regulation of vascular endothelial growth factor receptor signaling pathway(GO:0030948) regulation of receptor catabolic process(GO:2000644) |
| 0.2 | 5.6 | GO:0002230 | positive regulation of defense response to virus by host(GO:0002230) |
| 0.2 | 11.8 | GO:0006656 | phosphatidylcholine biosynthetic process(GO:0006656) |
| 0.2 | 10.1 | GO:0000083 | regulation of transcription involved in G1/S transition of mitotic cell cycle(GO:0000083) |
| 0.2 | 3.6 | GO:0090136 | epithelial cell-cell adhesion(GO:0090136) |
| 0.2 | 4.3 | GO:0036295 | cellular response to increased oxygen levels(GO:0036295) |
| 0.2 | 9.5 | GO:0006409 | tRNA export from nucleus(GO:0006409) tRNA-containing ribonucleoprotein complex export from nucleus(GO:0071431) |
| 0.2 | 7.2 | GO:0000462 | maturation of SSU-rRNA from tricistronic rRNA transcript (SSU-rRNA, 5.8S rRNA, LSU-rRNA)(GO:0000462) |
| 0.2 | 2.5 | GO:0035970 | peptidyl-threonine dephosphorylation(GO:0035970) |
| 0.2 | 0.5 | GO:1903028 | positive regulation of opsonization(GO:1903028) |
| 0.2 | 8.7 | GO:0099601 | regulation of neurotransmitter receptor activity(GO:0099601) regulation of glutamate receptor signaling pathway(GO:1900449) |
| 0.2 | 6.7 | GO:0051894 | positive regulation of focal adhesion assembly(GO:0051894) |
| 0.1 | 2.9 | GO:0060117 | auditory receptor cell development(GO:0060117) |
| 0.1 | 2.6 | GO:0097031 | NADH dehydrogenase complex assembly(GO:0010257) mitochondrial respiratory chain complex I assembly(GO:0032981) mitochondrial respiratory chain complex I biogenesis(GO:0097031) |
| 0.1 | 6.6 | GO:0043276 | anoikis(GO:0043276) nucleotide-binding oligomerization domain containing signaling pathway(GO:0070423) |
| 0.1 | 2.9 | GO:0061418 | regulation of transcription from RNA polymerase II promoter in response to hypoxia(GO:0061418) |
| 0.1 | 3.0 | GO:1901379 | regulation of potassium ion transport(GO:0043266) regulation of potassium ion transmembrane transport(GO:1901379) |
| 0.1 | 7.2 | GO:0050909 | sensory perception of taste(GO:0050909) |
| 0.1 | 1.3 | GO:0030210 | heparin metabolic process(GO:0030202) heparin biosynthetic process(GO:0030210) |
| 0.1 | 1.7 | GO:0006123 | mitochondrial electron transport, cytochrome c to oxygen(GO:0006123) |
| 0.1 | 18.6 | GO:0043123 | positive regulation of I-kappaB kinase/NF-kappaB signaling(GO:0043123) |
| 0.1 | 0.9 | GO:0007168 | receptor guanylyl cyclase signaling pathway(GO:0007168) |
| 0.1 | 1.5 | GO:0035855 | megakaryocyte development(GO:0035855) |
| 0.1 | 6.8 | GO:0045333 | cellular respiration(GO:0045333) |
| 0.1 | 10.3 | GO:0043484 | regulation of RNA splicing(GO:0043484) |
| 0.1 | 1.5 | GO:0090162 | establishment of epithelial cell polarity(GO:0090162) |
| 0.1 | 13.5 | GO:0006457 | protein folding(GO:0006457) |
| 0.1 | 5.4 | GO:0035335 | peptidyl-tyrosine dephosphorylation(GO:0035335) |
| 0.1 | 7.0 | GO:0006414 | translational elongation(GO:0006414) |
| 0.0 | 0.5 | GO:0000244 | spliceosomal tri-snRNP complex assembly(GO:0000244) |
| 0.0 | 3.7 | GO:0038083 | peptidyl-tyrosine autophosphorylation(GO:0038083) |
| 0.0 | 1.1 | GO:0030728 | ovulation(GO:0030728) |
| 0.0 | 2.1 | GO:0007040 | lysosome organization(GO:0007040) lytic vacuole organization(GO:0080171) |
| 0.0 | 13.6 | GO:0042278 | purine nucleoside metabolic process(GO:0042278) |
| 0.0 | 2.5 | GO:0034113 | heterotypic cell-cell adhesion(GO:0034113) |
| 0.0 | 10.7 | GO:0000209 | protein polyubiquitination(GO:0000209) |
| 0.0 | 1.4 | GO:0031572 | G2 DNA damage checkpoint(GO:0031572) |
| 0.0 | 3.3 | GO:0051291 | protein heterooligomerization(GO:0051291) |
| 0.0 | 1.2 | GO:0001964 | startle response(GO:0001964) |
| 0.0 | 1.4 | GO:0050911 | detection of chemical stimulus involved in sensory perception of smell(GO:0050911) |
| 0.0 | 1.2 | GO:0006383 | transcription from RNA polymerase III promoter(GO:0006383) |
| 0.0 | 2.1 | GO:0006805 | xenobiotic metabolic process(GO:0006805) |
| 0.0 | 4.6 | GO:0043687 | post-translational protein modification(GO:0043687) |
Gene overrepresentation in cellular_component category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 5.6 | 22.3 | GO:0042721 | mitochondrial inner membrane protein insertion complex(GO:0042721) |
| 3.9 | 31.5 | GO:0071204 | histone pre-mRNA 3'end processing complex(GO:0071204) |
| 3.2 | 9.5 | GO:1990876 | cytoplasmic side of nuclear pore(GO:1990876) |
| 2.4 | 7.3 | GO:0005607 | laminin-2 complex(GO:0005607) |
| 2.3 | 11.4 | GO:0034688 | integrin alphaM-beta2 complex(GO:0034688) |
| 2.2 | 11.0 | GO:0045272 | plasma membrane respiratory chain complex I(GO:0045272) |
| 1.9 | 13.1 | GO:0031415 | NatA complex(GO:0031415) |
| 1.6 | 27.1 | GO:0043020 | NADPH oxidase complex(GO:0043020) |
| 1.6 | 11.0 | GO:0061617 | MICOS complex(GO:0061617) |
| 1.6 | 17.3 | GO:0033018 | sarcoplasmic reticulum lumen(GO:0033018) |
| 1.4 | 5.7 | GO:0045283 | mitochondrial respiratory chain complex II, succinate dehydrogenase complex (ubiquinone)(GO:0005749) succinate dehydrogenase complex (ubiquinone)(GO:0045257) respiratory chain complex II(GO:0045273) succinate dehydrogenase complex(GO:0045281) fumarate reductase complex(GO:0045283) |
| 1.2 | 8.7 | GO:1990635 | proximal dendrite(GO:1990635) |
| 1.1 | 3.2 | GO:1990622 | CHOP-ATF3 complex(GO:1990622) |
| 1.1 | 5.4 | GO:0042719 | mitochondrial intermembrane space protein transporter complex(GO:0042719) |
| 1.0 | 11.4 | GO:0045275 | mitochondrial respiratory chain complex III(GO:0005750) respiratory chain complex III(GO:0045275) |
| 1.0 | 20.2 | GO:0000812 | Swr1 complex(GO:0000812) |
| 0.9 | 27.9 | GO:0005753 | mitochondrial proton-transporting ATP synthase complex(GO:0005753) |
| 0.8 | 64.5 | GO:0005758 | mitochondrial intermembrane space(GO:0005758) |
| 0.7 | 7.2 | GO:0005853 | eukaryotic translation elongation factor 1 complex(GO:0005853) |
| 0.6 | 19.5 | GO:0016327 | apicolateral plasma membrane(GO:0016327) |
| 0.5 | 28.7 | GO:0005747 | mitochondrial respiratory chain complex I(GO:0005747) NADH dehydrogenase complex(GO:0030964) respiratory chain complex I(GO:0045271) |
| 0.5 | 6.6 | GO:0008385 | IkappaB kinase complex(GO:0008385) |
| 0.4 | 1.7 | GO:0033257 | Bcl3/NF-kappaB2 complex(GO:0033257) |
| 0.4 | 9.4 | GO:0071682 | endocytic vesicle lumen(GO:0071682) |
| 0.3 | 3.6 | GO:0005916 | fascia adherens(GO:0005916) |
| 0.3 | 4.3 | GO:0033180 | proton-transporting V-type ATPase, V1 domain(GO:0033180) |
| 0.3 | 58.0 | GO:0000922 | spindle pole(GO:0000922) |
| 0.3 | 7.2 | GO:0034706 | sodium channel complex(GO:0034706) |
| 0.2 | 1.7 | GO:0098554 | cytoplasmic side of endoplasmic reticulum membrane(GO:0098554) |
| 0.2 | 44.0 | GO:0101002 | ficolin-1-rich granule(GO:0101002) ficolin-1-rich granule lumen(GO:1904813) |
| 0.2 | 3.8 | GO:0031305 | intrinsic component of mitochondrial inner membrane(GO:0031304) integral component of mitochondrial inner membrane(GO:0031305) |
| 0.2 | 5.2 | GO:0005665 | DNA-directed RNA polymerase II, core complex(GO:0005665) |
| 0.2 | 12.3 | GO:0030315 | T-tubule(GO:0030315) |
| 0.1 | 0.6 | GO:1990393 | 3M complex(GO:1990393) |
| 0.1 | 2.0 | GO:0005862 | muscle thin filament tropomyosin(GO:0005862) |
| 0.1 | 7.2 | GO:0022627 | cytosolic small ribosomal subunit(GO:0022627) |
| 0.1 | 2.9 | GO:0032420 | stereocilium(GO:0032420) |
| 0.1 | 2.5 | GO:0008305 | integrin complex(GO:0008305) |
| 0.1 | 27.0 | GO:0005759 | mitochondrial matrix(GO:0005759) |
| 0.1 | 10.1 | GO:0090575 | RNA polymerase II transcription factor complex(GO:0090575) |
| 0.0 | 0.2 | GO:0000015 | phosphopyruvate hydratase complex(GO:0000015) |
| 0.0 | 0.9 | GO:0008074 | guanylate cyclase complex, soluble(GO:0008074) |
| 0.0 | 6.3 | GO:0031234 | extrinsic component of cytoplasmic side of plasma membrane(GO:0031234) |
| 0.0 | 1.3 | GO:0019005 | SCF ubiquitin ligase complex(GO:0019005) |
| 0.0 | 11.9 | GO:0000785 | chromatin(GO:0000785) |
| 0.0 | 0.5 | GO:0070469 | respiratory chain(GO:0070469) |
| 0.0 | 3.6 | GO:0005681 | spliceosomal complex(GO:0005681) |
| 0.0 | 6.4 | GO:0005911 | cell-cell junction(GO:0005911) |
| 0.0 | 19.1 | GO:0005615 | extracellular space(GO:0005615) |
| 0.0 | 4.8 | GO:0005813 | centrosome(GO:0005813) |
| 0.0 | 3.6 | GO:0005769 | early endosome(GO:0005769) |
Gene overrepresentation in molecular_function category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 5.3 | 15.9 | GO:0033867 | Fas-activated serine/threonine kinase activity(GO:0033867) |
| 3.9 | 11.6 | GO:0045155 | electron transporter, transferring electrons from CoQH2-cytochrome c reductase complex and cytochrome c oxidase complex activity(GO:0045155) |
| 3.3 | 10.0 | GO:0004450 | isocitrate dehydrogenase (NADP+) activity(GO:0004450) |
| 2.5 | 17.6 | GO:0004673 | protein histidine kinase activity(GO:0004673) |
| 2.4 | 11.8 | GO:0052740 | 1-acyl-2-lysophosphatidylserine acylhydrolase activity(GO:0052740) |
| 2.3 | 9.2 | GO:0004727 | prenylated protein tyrosine phosphatase activity(GO:0004727) |
| 2.2 | 17.3 | GO:0004563 | beta-N-acetylhexosaminidase activity(GO:0004563) |
| 2.2 | 19.4 | GO:1901612 | cardiolipin binding(GO:1901612) |
| 1.9 | 5.7 | GO:0008177 | succinate dehydrogenase (ubiquinone) activity(GO:0008177) |
| 1.7 | 5.2 | GO:0016781 | selenide, water dikinase activity(GO:0004756) phosphotransferase activity, paired acceptors(GO:0016781) |
| 1.7 | 5.2 | GO:0000026 | alpha-1,2-mannosyltransferase activity(GO:0000026) |
| 1.6 | 11.4 | GO:0030369 | ICAM-3 receptor activity(GO:0030369) |
| 1.5 | 18.1 | GO:0004017 | adenylate kinase activity(GO:0004017) |
| 1.5 | 11.7 | GO:0071933 | Arp2/3 complex binding(GO:0071933) |
| 1.4 | 13.9 | GO:0030911 | TPR domain binding(GO:0030911) |
| 1.4 | 27.1 | GO:0016175 | superoxide-generating NADPH oxidase activity(GO:0016175) |
| 1.2 | 4.7 | GO:0004082 | bisphosphoglycerate mutase activity(GO:0004082) phosphoglycerate mutase activity(GO:0004619) 2,3-bisphosphoglycerate-dependent phosphoglycerate mutase activity(GO:0046538) |
| 1.2 | 22.1 | GO:0032454 | histone demethylase activity (H3-K9 specific)(GO:0032454) |
| 1.0 | 14.4 | GO:0016018 | cyclosporin A binding(GO:0016018) |
| 1.0 | 13.1 | GO:0004596 | peptide alpha-N-acetyltransferase activity(GO:0004596) |
| 0.9 | 18.6 | GO:0050786 | RAGE receptor binding(GO:0050786) |
| 0.9 | 7.9 | GO:0016174 | NAD(P)H oxidase activity(GO:0016174) |
| 0.9 | 9.5 | GO:0005049 | nuclear export signal receptor activity(GO:0005049) |
| 0.7 | 39.7 | GO:0050136 | NADH dehydrogenase (ubiquinone) activity(GO:0008137) NADH dehydrogenase (quinone) activity(GO:0050136) |
| 0.7 | 2.2 | GO:0052596 | tryptamine:oxygen oxidoreductase (deaminating) activity(GO:0052593) aminoacetone:oxygen oxidoreductase(deaminating) activity(GO:0052594) aliphatic-amine oxidase activity(GO:0052595) phenethylamine:oxygen oxidoreductase (deaminating) activity(GO:0052596) |
| 0.7 | 2.7 | GO:0050309 | glucose-6-phosphatase activity(GO:0004346) sugar-terminal-phosphatase activity(GO:0050309) |
| 0.7 | 12.8 | GO:0005385 | zinc ion transmembrane transporter activity(GO:0005385) |
| 0.6 | 2.6 | GO:0016160 | amylase activity(GO:0016160) |
| 0.5 | 7.2 | GO:0015280 | ligand-gated sodium channel activity(GO:0015280) |
| 0.5 | 13.5 | GO:0046933 | proton-transporting ATP synthase activity, rotational mechanism(GO:0046933) |
| 0.5 | 6.6 | GO:0042975 | peroxisome proliferator activated receptor binding(GO:0042975) |
| 0.5 | 60.3 | GO:0004725 | protein tyrosine phosphatase activity(GO:0004725) |
| 0.5 | 3.7 | GO:0004550 | nucleoside diphosphate kinase activity(GO:0004550) |
| 0.4 | 8.3 | GO:0009931 | calcium-dependent protein serine/threonine kinase activity(GO:0009931) |
| 0.4 | 1.3 | GO:0030158 | protein xylosyltransferase activity(GO:0030158) |
| 0.3 | 3.6 | GO:0045294 | alpha-catenin binding(GO:0045294) |
| 0.3 | 40.1 | GO:0003697 | single-stranded DNA binding(GO:0003697) |
| 0.3 | 8.7 | GO:0001965 | G-protein alpha-subunit binding(GO:0001965) |
| 0.3 | 6.2 | GO:0016832 | aldehyde-lyase activity(GO:0016832) |
| 0.3 | 7.2 | GO:0003746 | translation elongation factor activity(GO:0003746) |
| 0.3 | 19.5 | GO:0019213 | deacetylase activity(GO:0019213) |
| 0.2 | 2.4 | GO:0070324 | thyroid hormone binding(GO:0070324) |
| 0.2 | 19.0 | GO:0097110 | scaffold protein binding(GO:0097110) |
| 0.2 | 1.1 | GO:0070576 | vitamin D 24-hydroxylase activity(GO:0070576) |
| 0.2 | 11.1 | GO:0019212 | phosphatase inhibitor activity(GO:0019212) |
| 0.2 | 4.3 | GO:0044769 | ATPase activity, coupled to transmembrane movement of ions, rotational mechanism(GO:0044769) proton-transporting ATPase activity, rotational mechanism(GO:0046961) |
| 0.2 | 9.2 | GO:0035255 | ionotropic glutamate receptor binding(GO:0035255) |
| 0.2 | 22.3 | GO:0008565 | protein transporter activity(GO:0008565) |
| 0.2 | 0.5 | GO:0031685 | adenosine receptor binding(GO:0031685) |
| 0.2 | 2.1 | GO:0003993 | acid phosphatase activity(GO:0003993) |
| 0.2 | 2.9 | GO:0008510 | sodium:bicarbonate symporter activity(GO:0008510) |
| 0.1 | 15.2 | GO:0004222 | metalloendopeptidase activity(GO:0004222) |
| 0.1 | 1.2 | GO:0004972 | NMDA glutamate receptor activity(GO:0004972) |
| 0.1 | 0.4 | GO:0004449 | isocitrate dehydrogenase (NAD+) activity(GO:0004449) |
| 0.1 | 1.4 | GO:0004993 | G-protein coupled serotonin receptor activity(GO:0004993) |
| 0.1 | 10.8 | GO:0051082 | unfolded protein binding(GO:0051082) |
| 0.1 | 6.7 | GO:0005201 | extracellular matrix structural constituent(GO:0005201) |
| 0.1 | 1.4 | GO:0031434 | mitogen-activated protein kinase kinase binding(GO:0031434) |
| 0.1 | 0.2 | GO:0050613 | delta14-sterol reductase activity(GO:0050613) |
| 0.1 | 1.7 | GO:0015002 | cytochrome-c oxidase activity(GO:0004129) heme-copper terminal oxidase activity(GO:0015002) oxidoreductase activity, acting on a heme group of donors, oxygen as acceptor(GO:0016676) |
| 0.1 | 3.3 | GO:0043425 | bHLH transcription factor binding(GO:0043425) |
| 0.1 | 13.1 | GO:0061630 | ubiquitin protein ligase activity(GO:0061630) |
| 0.1 | 0.9 | GO:0019992 | diacylglycerol binding(GO:0019992) |
| 0.0 | 3.0 | GO:0015459 | potassium channel regulator activity(GO:0015459) |
| 0.0 | 3.7 | GO:0004715 | non-membrane spanning protein tyrosine kinase activity(GO:0004715) |
| 0.0 | 32.9 | GO:0008270 | zinc ion binding(GO:0008270) |
| 0.0 | 1.5 | GO:0000980 | RNA polymerase II distal enhancer sequence-specific DNA binding(GO:0000980) |
| 0.0 | 0.2 | GO:0004634 | phosphopyruvate hydratase activity(GO:0004634) |
| 0.0 | 0.9 | GO:0004383 | guanylate cyclase activity(GO:0004383) |
| 0.0 | 2.1 | GO:0046983 | protein dimerization activity(GO:0046983) |
| 0.0 | 0.5 | GO:0015377 | cation:chloride symporter activity(GO:0015377) |
| 0.0 | 7.2 | GO:0003735 | structural constituent of ribosome(GO:0003735) |
| 0.0 | 4.0 | GO:0003705 | transcription factor activity, RNA polymerase II distal enhancer sequence-specific binding(GO:0003705) |
| 0.0 | 0.2 | GO:0008526 | phosphatidylinositol transporter activity(GO:0008526) |
| 0.0 | 0.5 | GO:0043531 | ADP binding(GO:0043531) |
| 0.0 | 5.4 | GO:0046982 | protein heterodimerization activity(GO:0046982) |
Gene overrepresentation in C2:CP category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 4.3 | 60.0 | SA G2 AND M PHASES | Cdc25 activates the cdc2/cyclin B complex to induce the G2/M transition. |
| 0.6 | 17.6 | PID ARF6 DOWNSTREAM PATHWAY | Arf6 downstream pathway |
| 0.5 | 7.3 | PID INTEGRIN4 PATHWAY | Alpha6 beta4 integrin-ligand interactions |
| 0.4 | 13.9 | PID ERBB NETWORK PATHWAY | ErbB receptor signaling network |
| 0.4 | 9.2 | PID PRL SIGNALING EVENTS PATHWAY | Signaling events mediated by PRL |
| 0.4 | 27.7 | PID P38 ALPHA BETA DOWNSTREAM PATHWAY | Signaling mediated by p38-alpha and p38-beta |
| 0.4 | 27.1 | PID RAC1 PATHWAY | RAC1 signaling pathway |
| 0.4 | 26.9 | PID CERAMIDE PATHWAY | Ceramide signaling pathway |
| 0.3 | 33.1 | PID PTP1B PATHWAY | Signaling events mediated by PTP1B |
| 0.3 | 14.0 | PID INTEGRIN CS PATHWAY | Integrin family cell surface interactions |
| 0.3 | 9.5 | PID SMAD2 3PATHWAY | Regulation of cytoplasmic and nuclear SMAD2/3 signaling |
| 0.2 | 6.2 | PID S1P META PATHWAY | Sphingosine 1-phosphate (S1P) pathway |
| 0.2 | 6.6 | PID TOLL ENDOGENOUS PATHWAY | Endogenous TLR signaling |
| 0.2 | 1.7 | PID NFKAPPAB ATYPICAL PATHWAY | Atypical NF-kappaB pathway |
| 0.1 | 3.6 | PID RHOA PATHWAY | RhoA signaling pathway |
| 0.1 | 5.7 | PID ATF2 PATHWAY | ATF-2 transcription factor network |
| 0.1 | 6.7 | PID ILK PATHWAY | Integrin-linked kinase signaling |
| 0.1 | 10.1 | PID E2F PATHWAY | E2F transcription factor network |
| 0.1 | 8.7 | PID REG GR PATHWAY | Glucocorticoid receptor regulatory network |
| 0.1 | 2.4 | PID RXR VDR PATHWAY | RXR and RAR heterodimerization with other nuclear receptor |
| 0.1 | 1.4 | PID P38 MKK3 6PATHWAY | p38 MAPK signaling pathway |
| 0.0 | 14.7 | NABA SECRETED FACTORS | Genes encoding secreted soluble factors |
| 0.0 | 0.9 | PID CONE PATHWAY | Visual signal transduction: Cones |
| 0.0 | 0.9 | PID RAS PATHWAY | Regulation of Ras family activation |
| 0.0 | 3.0 | PID PDGFRB PATHWAY | PDGFR-beta signaling pathway |
Gene overrepresentation in C2:CP:REACTOME category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 2.2 | 60.0 | REACTOME CYCLIN A B1 ASSOCIATED EVENTS DURING G2 M TRANSITION | Genes involved in Cyclin A/B1 associated events during G2/M transition |
| 1.1 | 35.7 | REACTOME SYNTHESIS AND INTERCONVERSION OF NUCLEOTIDE DI AND TRIPHOSPHATES | Genes involved in Synthesis and interconversion of nucleotide di- and triphosphates |
| 0.8 | 70.1 | REACTOME RESPIRATORY ELECTRON TRANSPORT | Genes involved in Respiratory electron transport |
| 0.8 | 12.8 | REACTOME ZINC TRANSPORTERS | Genes involved in Zinc transporters |
| 0.7 | 31.3 | REACTOME LATENT INFECTION OF HOMO SAPIENS WITH MYCOBACTERIUM TUBERCULOSIS | Genes involved in Latent infection of Homo sapiens with Mycobacterium tuberculosis |
| 0.7 | 13.9 | REACTOME TETRAHYDROBIOPTERIN BH4 SYNTHESIS RECYCLING SALVAGE AND REGULATION | Genes involved in Tetrahydrobiopterin (BH4) synthesis, recycling, salvage and regulation |
| 0.6 | 13.5 | REACTOME FORMATION OF ATP BY CHEMIOSMOTIC COUPLING | Genes involved in Formation of ATP by chemiosmotic coupling |
| 0.6 | 6.6 | REACTOME IRAK1 RECRUITS IKK COMPLEX | Genes involved in IRAK1 recruits IKK complex |
| 0.5 | 8.3 | REACTOME DESTABILIZATION OF MRNA BY BRF1 | Genes involved in Destabilization of mRNA by Butyrate Response Factor 1 (BRF1) |
| 0.5 | 27.7 | REACTOME MITOCHONDRIAL PROTEIN IMPORT | Genes involved in Mitochondrial Protein Import |
| 0.3 | 2.8 | REACTOME DIGESTION OF DIETARY CARBOHYDRATE | Genes involved in Digestion of dietary carbohydrate |
| 0.3 | 9.5 | REACTOME NEP NS2 INTERACTS WITH THE CELLULAR EXPORT MACHINERY | Genes involved in NEP/NS2 Interacts with the Cellular Export Machinery |
| 0.2 | 6.7 | REACTOME CELL EXTRACELLULAR MATRIX INTERACTIONS | Genes involved in Cell-extracellular matrix interactions |
| 0.2 | 10.4 | REACTOME PYRUVATE METABOLISM AND CITRIC ACID TCA CYCLE | Genes involved in Pyruvate metabolism and Citric Acid (TCA) cycle |
| 0.2 | 5.2 | REACTOME BIOSYNTHESIS OF THE N GLYCAN PRECURSOR DOLICHOL LIPID LINKED OLIGOSACCHARIDE LLO AND TRANSFER TO A NASCENT PROTEIN | Genes involved in Biosynthesis of the N-glycan precursor (dolichol lipid-linked oligosaccharide, LLO) and transfer to a nascent protein |
| 0.2 | 28.7 | REACTOME MRNA SPLICING | Genes involved in mRNA Splicing |
| 0.2 | 23.9 | REACTOME INTEGRIN CELL SURFACE INTERACTIONS | Genes involved in Integrin cell surface interactions |
| 0.2 | 2.7 | REACTOME GLUCOSE TRANSPORT | Genes involved in Glucose transport |
| 0.1 | 3.2 | REACTOME ACTIVATION OF GENES BY ATF4 | Genes involved in Activation of Genes by ATF4 |
| 0.1 | 5.0 | REACTOME GLYCOLYSIS | Genes involved in Glycolysis |
| 0.1 | 18.8 | REACTOME FACTORS INVOLVED IN MEGAKARYOCYTE DEVELOPMENT AND PLATELET PRODUCTION | Genes involved in Factors involved in megakaryocyte development and platelet production |
| 0.1 | 14.4 | REACTOME RESPONSE TO ELEVATED PLATELET CYTOSOLIC CA2 | Genes involved in Response to elevated platelet cytosolic Ca2+ |
| 0.1 | 7.2 | REACTOME FORMATION OF THE TERNARY COMPLEX AND SUBSEQUENTLY THE 43S COMPLEX | Genes involved in Formation of the ternary complex, and subsequently, the 43S complex |
| 0.1 | 1.2 | REACTOME RAS ACTIVATION UOPN CA2 INFUX THROUGH NMDA RECEPTOR | Genes involved in Ras activation uopn Ca2+ infux through NMDA receptor |
| 0.1 | 1.1 | REACTOME XENOBIOTICS | Genes involved in Xenobiotics |
| 0.0 | 13.6 | REACTOME ANTIGEN PROCESSING UBIQUITINATION PROTEASOME DEGRADATION | Genes involved in Antigen processing: Ubiquitination & Proteasome degradation |
| 0.0 | 7.2 | REACTOME TRANSLATION | Genes involved in Translation |
| 0.0 | 2.1 | REACTOME METABOLISM OF VITAMINS AND COFACTORS | Genes involved in Metabolism of vitamins and cofactors |
| 0.0 | 1.4 | REACTOME OLFACTORY SIGNALING PATHWAY | Genes involved in Olfactory Signaling Pathway |
| 0.0 | 2.9 | REACTOME TRANSPORT OF INORGANIC CATIONS ANIONS AND AMINO ACIDS OLIGOPEPTIDES | Genes involved in Transport of inorganic cations/anions and amino acids/oligopeptides |
| 0.0 | 0.5 | REACTOME SIGNALING BY FGFR1 FUSION MUTANTS | Genes involved in Signaling by FGFR1 fusion mutants |