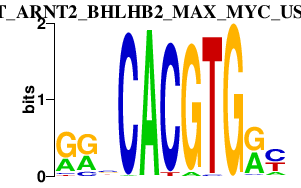
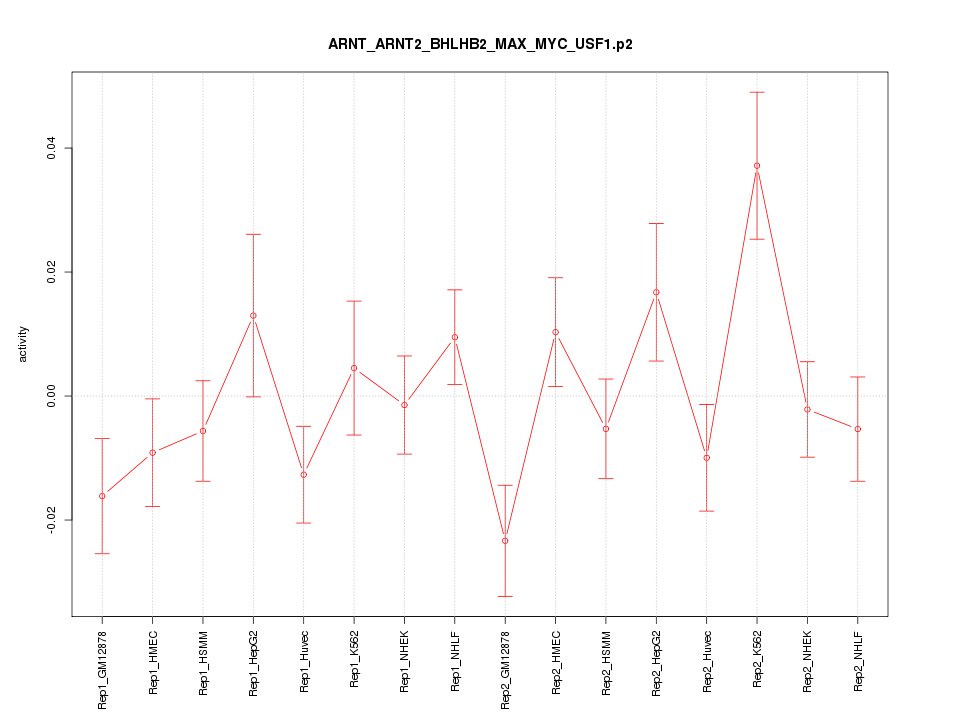
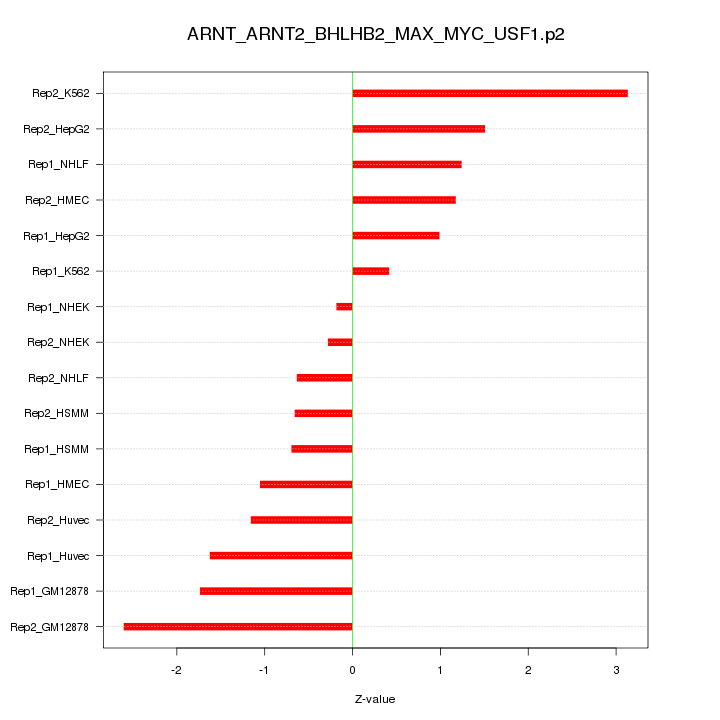
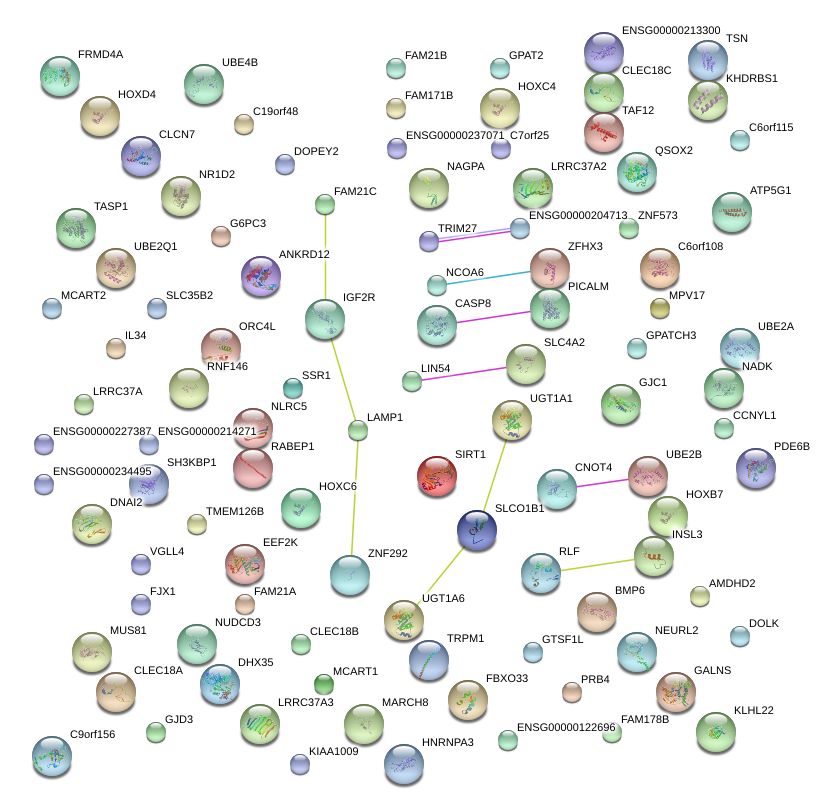
|
chr7_+_150390566
|
4.232
|
|
SLC4A2
|
solute carrier family 4, anion exchanger, member 2 (erythrocyte membrane protein band 3-like 1)
|
|
chr16_+_68542126
|
3.398
|
NM_001136214
|
CLEC18A
|
C-type lectin domain family 18, member A
|
|
chr2_-_97016027
|
3.223
|
NM_001122646
|
FAM178B
|
family with sequence similarity 178, member B
|
|
chr16_+_68542310
|
2.820
|
NM_182619
|
CLEC18C
CLEC18A
CLEC18B
|
C-type lectin domain family 18, member C
C-type lectin domain family 18, member A
C-type lectin domain family 18, member B
|
|
chr13_+_112999590
|
2.234
|
|
LAMP1
|
lysosomal-associated membrane protein 1
|
|
chr10_-_14090459
|
2.128
|
|
FRMD4A
|
FERM domain containing 4A
|
|
chr10_+_45542653
|
1.826
|
NM_001169106
NM_001169107
NM_015262
|
FAM21C
FAM21A
|
family with sequence similarity 21, member C
family with sequence similarity 21, member A
|
|
chr6_-_44333002
|
1.756
|
NM_178148
|
SLC35B2
|
solute carrier family 35, member B2
|
|
chr18_+_9126742
|
1.692
|
NM_001083625
NM_015208
|
ANKRD12
|
ankyrin repeat domain 12
|
|
chr4_+_609362
|
1.657
|
NM_000283
NM_001145291
|
PDE6B
|
phosphodiesterase 6B, cGMP-specific, rod, beta
|
|
chr16_+_2510323
|
1.618
|
NM_001145815
NM_015944
|
AMDHD2
|
amidohydrolase domain containing 2
|
|
chr17_+_44325127
|
1.612
|
NM_001002027
NM_005175
|
ATP5G1
|
ATP synthase, H+ transporting, mitochondrial Fo complex, subunit C1 (subunit 9)
|
|
chr2_+_187266899
|
1.602
|
NM_177454
|
FAM171B
|
family with sequence similarity 171, member B
|
|
chr17_-_41018291
|
1.540
|
|
|
|
|
chr1_-_152797760
|
1.520
|
|
UBE2Q1
|
ubiquitin-conjugating enzyme E2Q family member 1
|
|
chr1_-_152797820
|
1.513
|
|
UBE2Q1
|
ubiquitin-conjugating enzyme E2Q family member 1
|
|
chr8_+_142471290
|
1.465
|
|
|
|
|
chr10_-_45410197
|
1.459
|
NM_001002265
NM_145021
|
MARCH8
|
membrane-associated ring finger (C3HC4) 8
|
|
chr6_+_139391511
|
1.446
|
NM_021243
|
C6orf115
|
chromosome 6 open reading frame 115
|
|
chr2_+_177785667
|
1.442
|
NM_194247
|
HNRNPA3P1
HNRNPA3
|
heterogeneous nuclear ribonucleoprotein A3 pseudogene 1
heterogeneous nuclear ribonucleoprotein A3
|
|
chr6_+_139391570
|
1.439
|
|
C6orf115
|
chromosome 6 open reading frame 115
|
|
chr1_-_152797767
|
1.439
|
|
UBE2Q1
|
ubiquitin-conjugating enzyme E2Q family member 1
|
|
chr16_-_1465021
|
1.428
|
|
CLCN7
|
chloride channel 7
|
|
chr2_+_176724358
|
1.416
|
NM_014621
|
HOXD4
|
homeobox D4
|
|
chr16_+_55580910
|
1.403
|
NM_032206
|
NLRC5
|
NLR family, CARD domain containing 5
|
|
chr2_+_176724583
|
1.398
|
|
|
|
|
chr3_-_11736946
|
1.396
|
NM_014667
|
VGLL4
|
vestigial like 4 (Drosophila)
|
|
chr6_+_127629677
|
1.346
|
NM_030963
|
RNF146
|
ring finger protein 146
|
|
chr1_-_152797707
|
1.345
|
NM_017582
|
UBE2Q1
|
ubiquitin-conjugating enzyme E2Q family member 1
|
|
chr2_+_122229590
|
1.313
|
NM_004622
|
TSN
|
translin
|
|
chr9_-_138277467
|
1.312
|
NM_181701
|
QSOX2
|
quiescin Q6 sulfhydryl oxidase 2
|
|
chr16_-_71549149
|
1.300
|
|
ZFHX3
|
zinc finger homeobox 3
|
|
chr2_+_234333653
|
1.290
|
NM_000463
|
UGT1A1
|
UDP glucuronosyltransferase 1 family, polypeptide A1
|
|
chr11_-_85457469
|
1.283
|
|
PICALM
|
phosphatidylinositol binding clathrin assembly protein
|
|
chr9_-_130749604
|
1.282
|
NM_014908
|
DOLK
|
dolichol kinase
|
|
chr6_-_43305156
|
1.282
|
NM_006443
NM_199184
|
C6orf108
|
chromosome 6 open reading frame 108
|
|
chr1_-_1700088
|
1.215
|
NM_001198993
|
NADK
|
NAD kinase
|
|
chr20_-_32877029
|
1.205
|
NM_014071
|
NCOA6
|
nuclear receptor coactivator 6
|
|
chr17_-_40263124
|
1.188
|
NM_005497
|
GJC1
|
gap junction protein, gamma 1, 45kDa
|
|
chr20_-_41788975
|
1.180
|
NM_001008901
NM_176791
|
GTSF1L
|
gametocyte specific factor 1-like
|
|
chr6_-_28999388
|
1.178
|
|
TRIM27
|
tripartite motif containing 27
|
|
chr16_-_5023562
|
1.147
|
|
NAGPA
|
N-acetylglucosamine-1-phosphodiester alpha-N-acetylglucosaminidase
|
|
chr20_+_37024383
|
1.131
|
NM_001190809
NM_021931
|
DHX35
|
DEAH (Asp-Glu-Ala-His) box polypeptide 35
|
|
chr13_+_112999538
|
1.128
|
|
LAMP1
|
lysosomal-associated membrane protein 1
|
|
chr19_-_55999677
|
1.124
|
NM_199250
NM_199249
|
C19orf48
|
chromosome 19 open reading frame 48
|
|
chr6_-_7258351
|
1.113
|
NM_003144
|
SSR1
|
signal sequence receptor, alpha
|
|
chr6_+_87921980
|
1.110
|
NM_015021
|
ZNF292
|
zinc finger protein 292
|
|
chrX_-_19815586
|
1.110
|
NM_031892
|
SH3KBP1
|
SH3-domain kinase binding protein 1
|
|
chr1_+_32251988
|
1.108
|
NM_006559
|
KHDRBS1
|
KH domain containing, RNA binding, signal transduction associated 1
|
|
chr17_+_63713171
|
1.078
|
|
|
|
|
chr16_-_72959492
|
1.068
|
|
LOC283922
|
pyruvate dehydrogenase phosphatase regulatory subunit pseudogene
|
|
chr6_+_160310118
|
1.067
|
NM_000876
|
IGF2R
|
insulin-like growth factor 2 receptor
|
|
chr16_-_87450734
|
1.061
|
|
GALNS
|
galactosamine (N-acetyl)-6-sulfate sulfatase
|
|
chrX_-_19815526
|
1.057
|
|
SH3KBP1
|
SH3-domain kinase binding protein 1
|
|
chr22_-_19180075
|
1.056
|
NM_032775
|
KLHL22
|
kelch-like 22 (Drosophila)
|
|
chr20_-_13567529
|
1.054
|
NM_017714
|
TASP1
|
taspase, threonine aspartase, 1
|
|
chr17_+_39503623
|
1.045
|
NM_138387
|
G6PC3
|
glucose 6 phosphatase, catalytic, 3
|
|
chr16_+_5023672
|
1.044
|
|
LOC100507589
|
hypothetical LOC100507589
|
|
chr16_-_1465007
|
1.021
|
|
CLCN7
|
chloride channel 7
|
|
chr4_-_84152985
|
1.021
|
NM_001115007
|
LIN54
|
lin-54 homolog (C. elegans)
|
|
chr5_+_133735004
|
1.002
|
|
UBE2B
|
ubiquitin-conjugating enzyme E2B (RAD6 homolog)
|
|
chr11_-_85457753
|
1.000
|
NM_001008660
NM_007166
|
PICALM
|
phosphatidylinositol binding clathrin assembly protein
|
|
chr17_-_44042924
|
0.992
|
|
HOXB7
|
homeobox B7
|
|
chr21_+_36458708
|
0.980
|
NM_005128
|
DOPEY2
|
dopey family member 2
|
|
chr1_+_10015736
|
0.971
|
|
UBE4B
|
ubiquitination factor E4B (UFD2 homolog, yeast)
|
|
chr6_+_7671267
|
0.961
|
|
BMP6
|
bone morphogenetic protein 6
|
|
chr2_-_27399405
|
0.941
|
NM_002437
|
MPV17
|
MpV17 mitochondrial inner membrane protein
|
|
chr1_+_40399642
|
0.932
|
|
RLF
|
rearranged L-myc fusion
|
|
chr16_+_22124870
|
0.928
|
NM_013302
|
EEF2K
|
eukaryotic elongation factor-2 kinase
|
|
chr10_+_51497660
|
0.916
|
NM_001005751
|
FAM21C
FAM21A
|
family with sequence similarity 21, member C
family with sequence similarity 21, member A
|
|
chr7_-_42918013
|
0.908
|
NM_001099858
NM_024054
|
C7orf25
|
chromosome 7 open reading frame 25
|
|
chr1_-_28842103
|
0.903
|
NM_001135218
NM_005644
|
TAF12
|
TAF12 RNA polymerase II, TATA box binding protein (TBP)-associated factor, 20kDa
|
|
chr12_+_52733927
|
0.901
|
NM_153633
|
HOXC4
HOXC6
|
homeobox C4
homeobox C6
|
|
chr3_+_23962518
|
0.894
|
NM_001145425
|
NR1D2
|
nuclear receptor subfamily 1, group D, member 2
|
|
chr16_+_69171298
|
0.887
|
NM_001172771
NM_152456
|
IL34
|
interleukin 34
|
|
chr11_+_35596310
|
0.884
|
NM_014344
|
FJX1
|
four jointed box 1 (Drosophila)
|
|
chr1_-_27099515
|
0.883
|
|
GPATCH3
|
G patch domain containing 3
|
|
chr10_+_69314714
|
0.882
|
|
SIRT1
|
sirtuin 1
|
|
chr1_+_40399618
|
0.877
|
NM_012421
|
RLF
|
rearranged L-myc fusion
|
|
chr11_-_85457736
|
0.873
|
|
PICALM
|
phosphatidylinositol binding clathrin assembly protein
|
|
chr15_-_29181198
|
0.873
|
NM_002420
|
TRPM1
|
transient receptor potential cation channel, subfamily M, member 1
|
|
chr6_-_84994032
|
0.868
|
NM_014895
|
KIAA1009
|
KIAA1009
|
|
chr11_+_85017269
|
0.857
|
NM_001193537
NM_001193538
NM_018480
|
TMEM126B
|
transmembrane protein 126B
|
|
chr11_+_65384445
|
0.848
|
NM_025128
|
MUS81
|
MUS81 endonuclease homolog (S. cerevisiae)
|
|
chr10_+_51497716
|
0.842
|
|
FAM21A
|
family with sequence similarity 21, member A
|
|
chr14_-_38971370
|
0.840
|
NM_203301
|
FBXO33
|
F-box protein 33
|
|
chr9_-_37894098
|
0.828
|
|
MCART1
|
mitochondrial carrier triple repeat 1
|
|
chr16_-_1464927
|
0.827
|
|
CLCN7
|
chloride channel 7
|
|
chr7_-_44496703
|
0.825
|
NM_015332
|
NUDCD3
|
NudC domain containing 3
|
|
chr2_+_201806410
|
0.825
|
NM_001080124
NM_001228
NM_033355
NM_033358
|
CASP8
|
caspase 8, apoptosis-related cysteine peptidase
|
|
chr17_-_60346042
|
0.824
|
|
LRRC37A3
|
leucine rich repeat containing 37, member A3
|
|
chr9_-_99724559
|
0.824
|
NM_016481
|
C9orf156
|
chromosome 9 open reading frame 156
|
|
chr17_+_69781980
|
0.823
|
NM_001172810
NM_023036
|
DNAI2
|
dynein, axonemal, intermediate chain 2
|
|
chr16_-_1465041
|
0.823
|
NM_001114331
NM_001287
|
CLCN7
|
chloride channel 7
|
|
chr2_+_208284484
|
0.821
|
NM_001142300
NM_152523
|
CCNYL1
|
cyclin Y-like 1
|
|
chr7_-_134845363
|
0.818
|
NM_001008225
NM_001190847
NM_001190848
NM_001190849
NM_001190850
NM_013316
|
CNOT4
|
CCR4-NOT transcription complex, subunit 4
|
|
chr20_-_43953036
|
0.817
|
NM_080749
|
NEURL2
|
neuralized homolog 2 (Drosophila)
|
|
chr2_-_148494732
|
0.810
|
NM_001190882
NM_001190879
NM_002552
NM_001190881
NM_181741
|
ORC4
|
origin recognition complex, subunit 4
|
|
chr2_-_96064389
|
0.809
|
NM_207328
|
GPAT2
|
glycerol-3-phosphate acyltransferase 2, mitochondrial
|
|
chr17_+_5126275
|
0.807
|
NM_001083585
NM_004703
|
RABEP1
|
rabaptin, RAB GTPase binding effector protein 1
|
|
chr5_+_41940216
|
0.805
|
NM_175921
|
C5orf51
|
chromosome 5 open reading frame 51
|
|
chr21_-_42247007
|
0.805
|
NM_015500
|
C2CD2
|
C2 calcium-dependent domain containing 2
|
|
chr1_-_165211128
|
0.804
|
NM_199351
|
ILDR2
|
immunoglobulin-like domain containing receptor 2
|
|
chr17_-_43977273
|
0.802
|
NM_002145
|
HOXB2
|
homeobox B2
|
|
chr1_+_181871830
|
0.801
|
NM_015149
|
RGL1
|
ral guanine nucleotide dissociation stimulator-like 1
|
|
chr1_-_11788550
|
0.792
|
|
MTHFR
|
methylenetetrahydrofolate reductase (NAD(P)H)
|
|
chr5_-_158569067
|
0.788
|
|
RNF145
|
ring finger protein 145
|
|
chr8_-_42028634
|
0.781
|
NM_001099412
NM_001099413
NM_006766
|
MYST3
|
MYST histone acetyltransferase (monocytic leukemia) 3
|
|
chr10_-_61139328
|
0.780
|
NM_194298
|
LOC100129721
SLC16A9
|
hypothetical protein LOC100129721
solute carrier family 16, member 9 (monocarboxylic acid transporter 9)
|
|
chr16_+_24458349
|
0.777
|
NM_006910
NM_018703
NM_032626
|
RBBP6
|
retinoblastoma binding protein 6
|
|
chr6_+_35852369
|
0.775
|
NM_207409
|
C6orf126
|
chromosome 6 open reading frame 126
|
|
chr3_-_129024665
|
0.766
|
NM_001003794
|
MGLL
|
monoglyceride lipase
|
|
chr6_-_169392771
|
0.765
|
|
|
|
|
chr14_-_34253473
|
0.759
|
NM_138638
|
CFL2
|
cofilin 2 (muscle)
|
|
chr16_+_11669736
|
0.757
|
NM_003498
|
SNN
|
stannin
|
|
chr10_+_70609986
|
0.755
|
NM_003171
|
SUPV3L1
|
suppressor of var1, 3-like 1 (S. cerevisiae)
|
|
chr8_-_95343716
|
0.747
|
NM_005261
NM_181702
|
GEM
|
GTP binding protein overexpressed in skeletal muscle
|
|
chr17_-_40122542
|
0.738
|
NM_001099225
NM_144609
|
CCDC43
|
coiled-coil domain containing 43
|
|
chr2_-_176575171
|
0.738
|
NM_030650
|
KIAA1715
|
KIAA1715
|
|
chr17_-_71362971
|
0.737
|
NM_012478
|
WBP2
|
WW domain binding protein 2
|
|
chr6_-_31973418
|
0.737
|
|
EHMT2
|
euchromatic histone-lysine N-methyltransferase 2
|
|
chr9_+_130749812
|
0.735
|
|
NUP188
|
nucleoporin 188kDa
|
|
chr16_-_87244896
|
0.735
|
NM_000101
|
CYBA
|
cytochrome b-245, alpha polypeptide
|
|
chr2_+_68238490
|
0.734
|
NM_020143
|
PNO1
|
partner of NOB1 homolog (S. cerevisiae)
|
|
chr2_+_46779553
|
0.734
|
NM_014011
|
SOCS5
|
suppressor of cytokine signaling 5
|
|
chr9_+_130749791
|
0.730
|
NM_015354
|
NUP188
|
nucleoporin 188kDa
|
|
chr16_+_80370426
|
0.729
|
NM_002661
|
PLCG2
|
phospholipase C, gamma 2 (phosphatidylinositol-specific)
|
|
chr6_-_8047731
|
0.728
|
NM_001135650
NM_004280
|
EEF1E1
|
eukaryotic translation elongation factor 1 epsilon 1
|
|
chr11_+_66915997
|
0.728
|
NM_004584
|
RAD9A
|
RAD9 homolog A (S. pombe)
|
|
chr1_-_52936573
|
0.727
|
|
C1orf163
|
chromosome 1 open reading frame 163
|
|
chr7_+_116380590
|
0.725
|
NM_018412
NM_021908
|
ST7
|
suppression of tumorigenicity 7
|
|
chr14_-_93512818
|
0.709
|
|
ASB2
|
ankyrin repeat and SOCS box containing 2
|
|
chr2_+_46779827
|
0.704
|
NM_144949
|
SOCS5
|
suppressor of cytokine signaling 5
|
|
chr1_+_109963950
|
0.696
|
NM_139156
|
AMPD2
|
adenosine monophosphate deaminase 2
|
|
chr1_-_1699723
|
0.692
|
NM_023018
|
NADK
|
NAD kinase
|
|
chr8_-_95343696
|
0.691
|
|
GEM
|
GTP binding protein overexpressed in skeletal muscle
|
|
chr16_-_5023934
|
0.690
|
NM_016256
|
NAGPA
|
N-acetylglucosamine-1-phosphodiester alpha-N-acetylglucosaminidase
|
|
chr6_-_31973434
|
0.686
|
NM_006709
NM_025256
|
EHMT2
|
euchromatic histone-lysine N-methyltransferase 2
|
|
chr16_-_5023914
|
0.684
|
|
NAGPA
|
N-acetylglucosamine-1-phosphodiester alpha-N-acetylglucosaminidase
|
|
chr3_+_51992339
|
0.681
|
NM_000666
NM_001198895
NM_001198896
NM_001198897
NM_001198898
|
ACY1
|
aminoacylase 1
|
|
chr3_-_196644864
|
0.681
|
NM_012287
|
ACAP2
|
ArfGAP with coiled-coil, ankyrin repeat and PH domains 2
|
|
chr6_+_151688327
|
0.677
|
NM_144497
|
AKAP12
|
A kinase (PRKA) anchor protein 12
|
|
chr8_-_70908130
|
0.672
|
NM_001146008
|
SLCO5A1
|
solute carrier organic anion transporter family, member 5A1
|
|
chr12_+_50981915
|
0.665
|
NM_002284
|
KRT86
|
keratin 86
|
|
chr2_+_241903395
|
0.658
|
NM_001008491
NM_006155
|
SEPT2
|
septin 2
|
|
chr17_+_37694007
|
0.655
|
|
STAT5A
|
signal transducer and activator of transcription 5A
|
|
chr12_-_119116848
|
0.653
|
NM_006836
|
GCN1L1
|
GCN1 general control of amino-acid synthesis 1-like 1 (yeast)
|
|
chr16_+_85169615
|
0.649
|
NM_005250
|
FOXL1
|
forkhead box L1
|
|
chr7_+_101860958
|
0.649
|
NM_001126340
NM_032831
|
ORAI2
|
ORAI calcium release-activated calcium modulator 2
|
|
chr7_-_135083906
|
0.649
|
NM_205855
|
FAM180A
|
family with sequence similarity 180, member A
|
|
chr1_-_201162991
|
0.644
|
NM_021633
|
KLHL12
|
kelch-like 12 (Drosophila)
|
|
chr13_+_48582389
|
0.643
|
NM_014923
|
FNDC3A
|
fibronectin type III domain containing 3A
|
|
chr19_+_14405216
|
0.641
|
|
PKN1
|
protein kinase N1
|
|
chr2_-_68238079
|
0.639
|
NM_138458
|
WDR92
|
WD repeat domain 92
|
|
chr12_+_102982365
|
0.637
|
NM_013320
|
HCFC2
|
host cell factor C2
|
|
chr20_+_43953372
|
0.635
|
|
CTSA
|
cathepsin A
|
|
chr5_+_132237253
|
0.634
|
NM_052971
|
LEAP2
|
liver expressed antimicrobial peptide 2
|
|
chr1_-_77921681
|
0.631
|
|
ZZZ3
|
zinc finger, ZZ-type containing 3
|
|
chr17_+_63252704
|
0.630
|
|
BPTF
|
bromodomain PHD finger transcription factor
|
|
chr1_-_153477673
|
0.626
|
NM_001171812
NM_000157
|
GBA
|
glucosidase, beta, acid
|
|
chr1_+_234372454
|
0.625
|
NM_003272
|
GPR137B
|
G protein-coupled receptor 137B
|
|
chr19_-_13088212
|
0.625
|
NM_001142554
NM_017722
NM_001136035
|
TRMT1
|
TRM1 tRNA methyltransferase 1 homolog (S. cerevisiae)
|
|
chr6_-_35804337
|
0.624
|
NM_001145775
|
FKBP5
|
FK506 binding protein 5
|
|
chr11_-_57054756
|
0.624
|
NM_012456
|
TIMM10
|
translocase of inner mitochondrial membrane 10 homolog (yeast)
|
|
chr2_-_61618165
|
0.622
|
|
XPO1
|
exportin 1 (CRM1 homolog, yeast)
|
|
chr17_-_73636362
|
0.620
|
|
TMC6
|
transmembrane channel-like 6
|
|
chr19_+_10626106
|
0.619
|
|
ILF3
|
interleukin enhancer binding factor 3, 90kDa
|
|
chr17_+_74542066
|
0.616
|
NM_198593
|
C1QTNF1
|
C1q and tumor necrosis factor related protein 1
|
|
chr5_+_154115046
|
0.614
|
|
LARP1
|
La ribonucleoprotein domain family, member 1
|
|
chr17_-_33043514
|
0.612
|
NM_001163544
NM_001163545
NM_001163546
NM_001163547
NM_007247
NM_080550
NM_198882
|
SYNRG
|
synergin, gamma
|
|
chr17_-_44010543
|
0.612
|
NM_024015
|
HOXB4
|
homeobox B4
|
|
chr1_-_27099524
|
0.611
|
NM_022078
|
GPATCH3
|
G patch domain containing 3
|
|
chr12_-_62902279
|
0.610
|
NM_152440
|
C12orf66
|
chromosome 12 open reading frame 66
|
|
chr4_-_100069265
|
0.610
|
NM_001130678
|
EIF4E
|
eukaryotic translation initiation factor 4E
|
|
chr11_+_65384513
|
0.607
|
|
MUS81
|
MUS81 endonuclease homolog (S. cerevisiae)
|
|
chr16_+_29819647
|
0.606
|
NM_181718
|
ASPHD1
|
aspartate beta-hydroxylase domain containing 1
|
|
chr13_+_112999462
|
0.606
|
NM_005561
|
LAMP1
|
lysosomal-associated membrane protein 1
|
|
chr16_-_15644465
|
0.605
|
NM_001184998
NM_001184999
NM_014647
|
KIAA0430
|
KIAA0430
|
|
chrX_-_47394808
|
0.605
|
|
ELK1
|
ELK1, member of ETS oncogene family
|
|
chr17_-_35510431
|
0.602
|
NM_021724
|
NR1D1
|
nuclear receptor subfamily 1, group D, member 1
|
|
chr2_-_175741096
|
0.593
|
NM_001880
|
ATF2
|
activating transcription factor 2
|
|
chr7_+_1050704
|
0.592
|
|
|
|
|
chr20_+_43953428
|
0.584
|
|
CTSA
|
cathepsin A
|
|
chr18_-_45241014
|
0.582
|
NM_017653
|
DYM
|
dymeclin
|
|
chr17_-_34263487
|
0.581
|
NM_000978
|
SNORA21
RPL23
|
small nucleolar RNA, H/ACA box 21
ribosomal protein L23
|
|
chrX_-_47394937
|
0.580
|
NM_001114123
NM_005229
|
ELK1
|
ELK1, member of ETS oncogene family
|
|
chr11_-_8849485
|
0.578
|
|
ST5
|
suppression of tumorigenicity 5
|
|
chr10_+_70150936
|
0.578
|
NM_018237
|
CCAR1
|
cell division cycle and apoptosis regulator 1
|
|
chr9_-_139128777
|
0.577
|
|
DPP7
|
dipeptidyl-peptidase 7
|
|
chr2_-_131567458
|
0.577
|
NM_001009993
|
FAM168B
|
family with sequence similarity 168, member B
|
|
chrX_+_53139936
|
0.575
|
|
LOC100132984
|
hypothetical protein LOC100132984
|
|
chr12_-_119826073
|
0.575
|
|
SPPL3
|
signal peptide peptidase 3
|
|
chr22_+_38903825
|
0.573
|
NM_001162501
NM_015088
|
TNRC6B
|
trinucleotide repeat containing 6B
|
|
chr6_+_30797393
|
0.571
|
|
TUBB
|
tubulin, beta
|
|
chr16_-_87450849
|
0.570
|
NM_000512
|
GALNS
|
galactosamine (N-acetyl)-6-sulfate sulfatase
|
|
chr3_-_20202622
|
0.569
|
NM_001012409
NM_001012410
NM_001012411
NM_001012412
NM_001012413
NM_138484
|
SGOL1
|
shugoshin-like 1 (S. pombe)
|
|
chr15_+_41871809
|
0.569
|
|
SERF2-C15ORF63
SERF2
|
SERF2-C15orf63 read-through transcript
small EDRK-rich factor 2
|
|
chr6_+_10803141
|
0.562
|
NM_017906
|
PAK1IP1
|
PAK1 interacting protein 1
|
|
chr19_+_12709281
|
0.557
|
NM_004317
|
ASNA1
|
arsA arsenite transporter, ATP-binding, homolog 1 (bacterial)
|
|
chr4_-_101090431
|
0.556
|
|
H2AFZ
|
H2A histone family, member Z
|