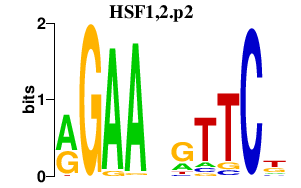
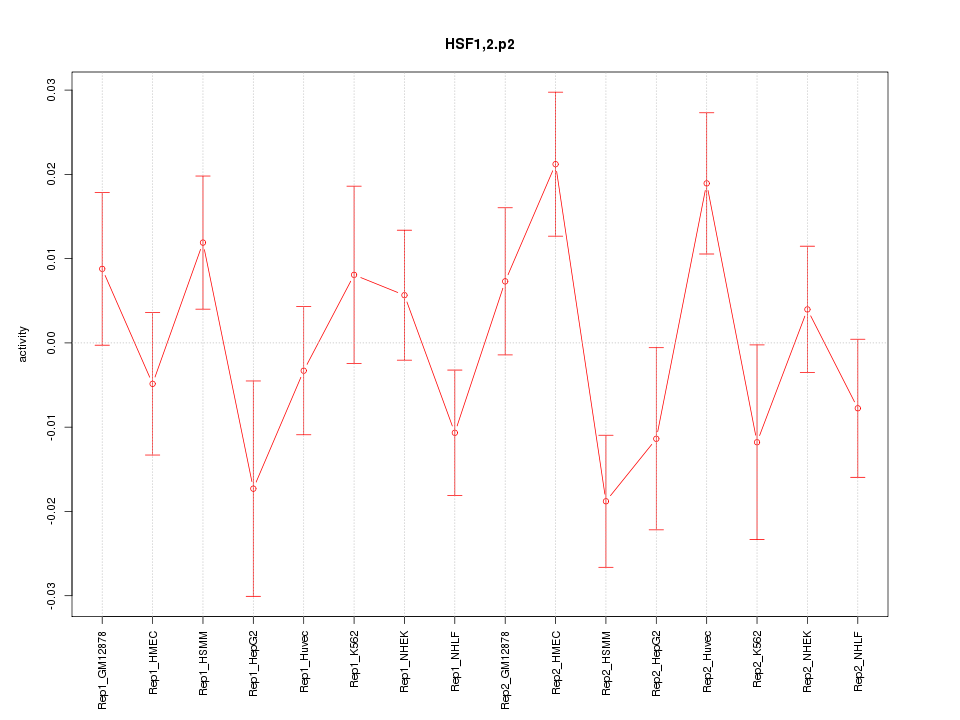
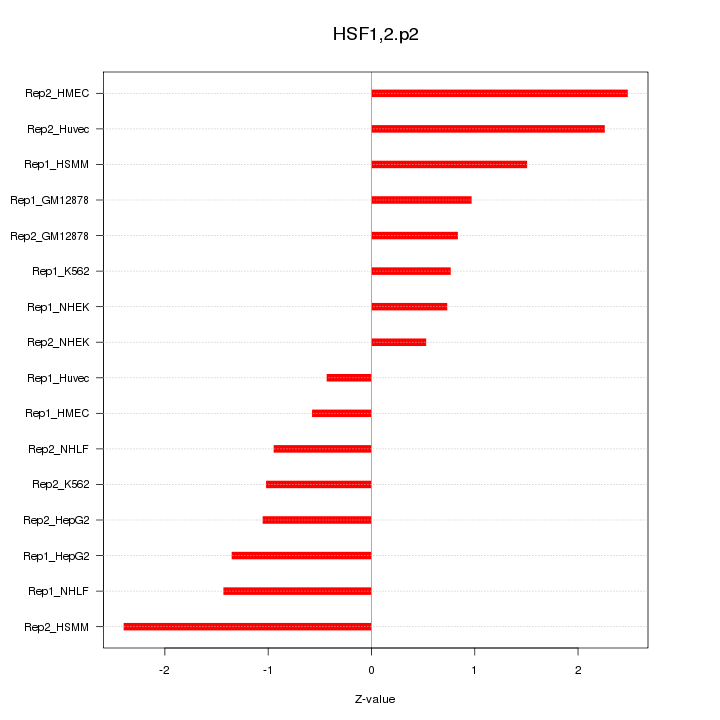
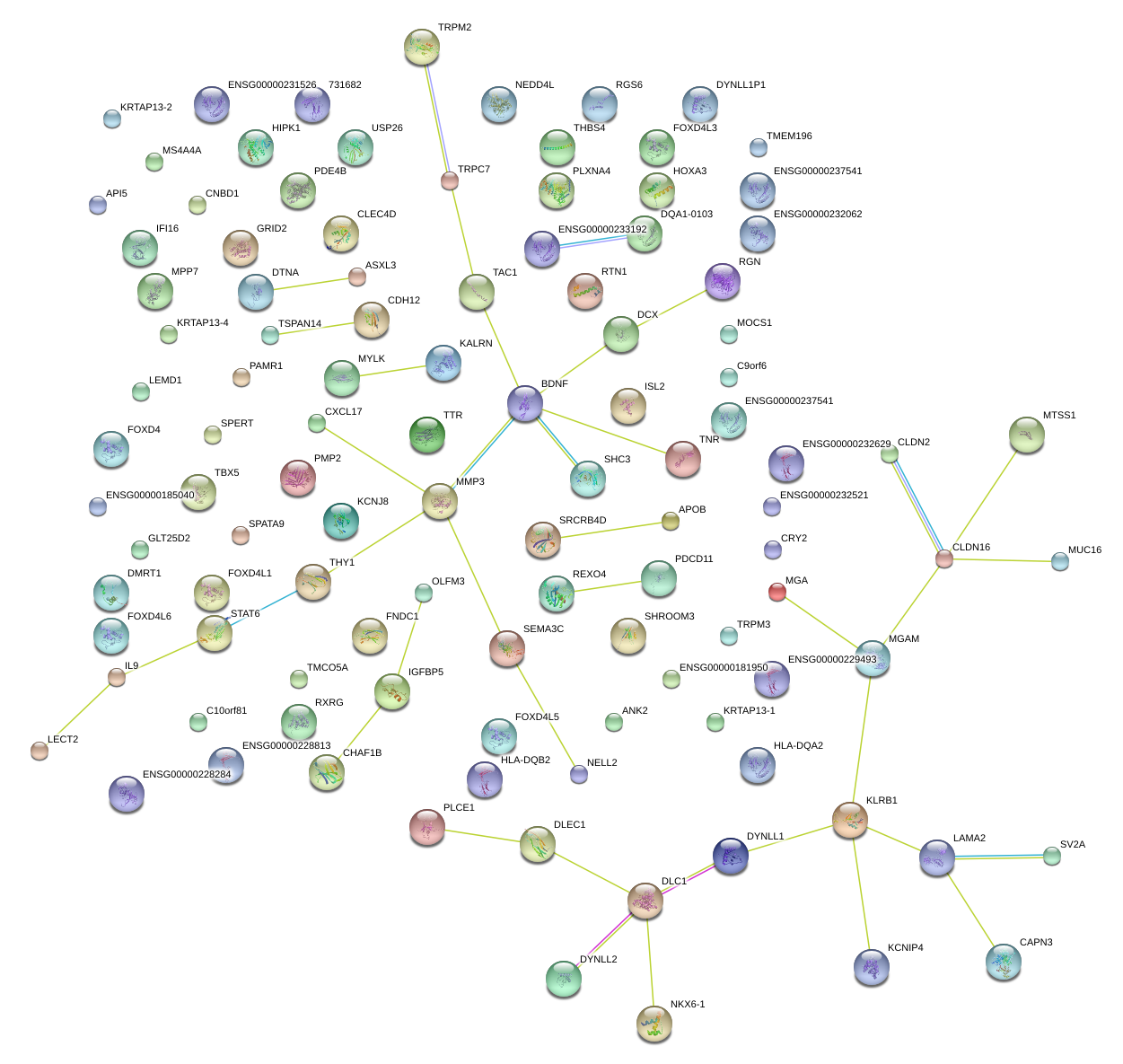
|
chr21_+_30724442
|
1.439
|
NM_181600
|
KRTAP13-4
|
keratin associated protein 13-4
|
|
chr11_-_118799452
|
1.250
|
NM_006288
|
THY1
|
Thy-1 cell surface antigen
|
|
chr6_+_129245978
|
1.219
|
NM_000426
NM_001079823
|
LAMA2
|
laminin, alpha 2
|
|
chr5_-_135318420
|
1.097
|
NM_002302
|
LECT2
|
leukocyte cell-derived chemotaxin 2
|
|
chr10_+_95838801
|
1.081
|
NM_001165979
|
PLCE1
|
phospholipase C, epsilon 1
|
|
chr4_+_77575276
|
1.002
|
NM_020859
|
SHROOM3
|
shroom family member 3
|
|
chr1_+_66570683
|
0.982
|
|
PDE4B
|
phosphodiesterase 4B, cAMP-specific
|
|
chr6_+_32817140
|
0.974
|
NM_020056
|
HLA-DQA2
|
major histocompatibility complex, class II, DQ alpha 2
|
|
chr1_-_102234959
|
0.961
|
|
OLFM3
|
olfactomedin 3
|
|
chr9_-_72926333
|
0.925
|
NM_001007471
|
TRPM3
|
transient receptor potential cation channel, subfamily M, member 3
|
|
chr12_+_8557402
|
0.898
|
NM_080387
|
CLEC4D
|
C-type lectin domain family 4, member D
|
|
chr12_-_21818881
|
0.894
|
NM_004982
|
KCNJ8
|
potassium inwardly-rectifying channel, subfamily J, member 8
|
|
chr19_-_47638890
|
0.881
|
NM_198477
|
CXCL17
|
chemokine (C-X-C motif) ligand 17
|
|
chr7_-_27133163
|
0.825
|
NM_153631
|
HOXA3
|
homeobox A3
|
|
chr1_-_148155986
|
0.822
|
NM_014849
|
SV2A
|
synaptic vesicle glycoprotein 2A
|
|
chr5_-_42211112
|
0.779
|
|
|
|
|
chr1_-_182273451
|
0.774
|
NM_015101
|
GLT25D2
|
glycosyltransferase 25 domain containing 2
|
|
chr13_+_45174446
|
0.766
|
NM_152719
|
SPERT
|
spermatid associated
|
|
chr6_-_32839244
|
0.728
|
NM_001198858
|
HLA-DQB2
|
major histocompatibility complex, class II, DQ beta 2
|
|
chr15_+_40438989
|
0.723
|
NM_000070
NM_024344
NM_173087
|
CAPN3
|
calpain 3, (p94)
|
|
chr7_-_75876830
|
0.694
|
NM_080744
|
SRCRB4D
|
scavenger receptor cysteine rich domain containing, group B (4 domains)
|
|
chr5_+_79366746
|
0.686
|
NM_003248
|
THBS4
|
thrombospondin 4
|
|
chrX_+_106050261
|
0.674
|
NM_020384
|
CLDN2
|
claudin 2
|
|
chr1_-_102235377
|
0.671
|
NM_058170
|
OLFM3
|
olfactomedin 3
|
|
chr3_+_191588354
|
0.669
|
NM_006580
|
CLDN16
|
claudin 16
|
|
chr10_-_28611020
|
0.664
|
NM_173496
|
MPP7
|
membrane protein, palmitoylated 7 (MAGUK p55 subfamily member 7)
|
|
chr6_-_40003295
|
0.657
|
NM_005943
|
MOCS1
|
molybdenum cofactor synthesis 1
|
|
chr9_-_90983501
|
0.655
|
NM_016848
|
SHC3
|
SHC (Src homology 2 domain containing) transforming protein 3
|
|
chr1_-_163680941
|
0.650
|
NM_006917
|
RXRG
|
retinoid X receptor, gamma
|
|
chr8_-_125649926
|
0.646
|
|
MTSS1
|
metastasis suppressor 1
|
|
chr9_+_831689
|
0.645
|
NM_021951
|
DMRT1
|
doublesex and mab-3 related transcription factor 1
|
|
chr9_-_68492023
|
0.639
|
NM_001085476
|
FOXD4L6
|
forkhead box D4-like 6
|
|
chr18_+_29573152
|
0.631
|
|
ASXL3
|
additional sex combs like 3 (Drosophila)
|
|
chr11_+_59806661
|
0.629
|
NM_024021
|
MS4A4A
|
membrane-spanning 4-domains, subfamily A, member 4
|
|
chr8_+_87947788
|
0.626
|
NM_173538
|
CNBD1
|
cyclic nucleotide binding domain containing 1
|
|
chr2_-_21084911
|
0.621
|
|
APOB
|
apolipoprotein B (including Ag(x) antigen)
|
|
chr1_-_173979292
|
0.618
|
NM_003285
|
TNR
|
tenascin R (restrictin, janusin)
|
|
chr8_-_82522175
|
0.615
|
NM_002677
|
PMP2
|
peripheral myelin protein 2
|
|
chr9_-_90983195
|
0.613
|
|
SHC3
|
SHC (Src homology 2 domain containing) transforming protein 3
|
|
chr1_-_203657803
|
0.597
|
NM_001001552
NM_001199050
NM_001199051
NM_001199052
|
LEMD1
|
LEM domain containing 1
|
|
chr5_-_135729062
|
0.586
|
NM_001167576
NM_001167577
NM_020389
|
TRPC7
|
transient receptor potential cation channel, subfamily C, member 7
|
|
chr7_+_97199206
|
0.585
|
NM_003182
NM_013996
NM_013997
NM_013998
|
TAC1
|
tachykinin, precursor 1
|
|
chr15_+_39739864
|
0.582
|
NM_001080541
NM_001164273
|
MGA
|
MAX gene associated
|
|
chr2_-_39040981
|
0.580
|
|
LOC375196
|
hypothetical LOC375196
|
|
chr2_-_217268492
|
0.578
|
NM_000599
|
IGFBP5
|
insulin-like growth factor binding protein 5
|
|
chr15_+_74416129
|
0.573
|
|
ISL2
|
ISL LIM homeobox 2
|
|
chr5_-_135259403
|
0.572
|
NM_000590
|
IL9
|
interleukin 9
|
|
chr6_+_159510416
|
0.570
|
NM_032532
|
FNDC1
|
fibronectin type III domain containing 1
|
|
chr1_+_157246305
|
0.564
|
NM_005531
|
IFI16
|
interferon, gamma-inducible protein 16
|
|
chr3_+_125586241
|
0.563
|
|
KALRN
|
kalirin, RhoGEF kinase
|
|
chr18_+_54013575
|
0.558
|
NM_001144970
|
NEDD4L
|
neural precursor cell expressed, developmentally down-regulated 4-like
|
|
chr10_+_115501202
|
0.555
|
NM_001193434
NM_001193435
NM_024889
|
C10orf81
|
chromosome 10 open reading frame 81
|
|
chr15_+_39739933
|
0.555
|
|
MGA
|
MAX gene associated
|
|
chr11_-_35503720
|
0.547
|
NM_001001991
NM_015430
|
PAMR1
|
peptidase domain containing associated with muscle regeneration 1
|
|
chr7_-_80386258
|
0.547
|
|
SEMA3C
|
sema domain, immunoglobulin domain (Ig), short basic domain, secreted, (semaphorin) 3C
|
|
chr1_-_182273114
|
0.542
|
|
GLT25D2
|
glycosyltransferase 25 domain containing 2
|
|
chr14_+_71468902
|
0.537
|
|
RGS6
|
regulator of G-protein signaling 6
|
|
chr11_-_102219450
|
0.537
|
NM_002422
|
MMP3
|
matrix metallopeptidase 3 (stromelysin 1, progelatinase)
|
|
chr5_+_140762551
|
0.536
|
NM_018921
NM_032089
|
PCDHGA9
|
protocadherin gamma subfamily A, 9
|
|
chr1_+_114295289
|
0.536
|
NM_198269
|
HIPK1
|
homeodomain interacting protein kinase 1
|
|
chr7_-_131983986
|
0.527
|
NM_181775
|
PLXNA4
|
plexin A4
|
|
chrX_-_131989963
|
0.526
|
NM_031907
|
USP26
|
ubiquitin specific peptidase 26
|
|
chr4_-_20914626
|
0.526
|
NM_147183
|
KCNIP4
|
Kv channel interacting protein 4
|
|
chr18_+_30427279
|
0.526
|
NM_001128175
NM_001198938
NM_001198939
NM_001198940
|
DTNA
|
dystrobrevin, alpha
|
|
chr4_+_93444572
|
0.523
|
NM_001510
|
GRID2
|
glutamate receptor, ionotropic, delta 2
|
|
chr21_+_30690165
|
0.518
|
NM_181599
|
KRTAP13-1
|
keratin associated protein 13-1
|
|
chr12_-_113326085
|
0.508
|
NM_080718
|
TBX5
|
T-box 5
|
|
chr4_-_85638410
|
0.508
|
NM_006168
|
NKX6-1
|
NK6 homeobox 1
|
|
chr14_-_59167139
|
0.502
|
|
RTN1
|
reticulon 1
|
|
chr14_-_59167284
|
0.501
|
|
RTN1
|
reticulon 1
|
|
chr5_-_22889487
|
0.497
|
NM_004061
|
CDH12
|
cadherin 12, type 2 (N-cadherin 2)
|
|
chr12_-_43555607
|
0.491
|
NM_001145109
|
NELL2
|
NEL-like 2 (chicken)
|
|
chr2_-_21084077
|
0.491
|
|
APOB
|
apolipoprotein B (including Ag(x) antigen)
|
|
chrX_-_110542040
|
0.489
|
NM_178151
NM_001195553
NM_178152
NM_178153
|
DCX
|
doublecortin
|
|
chr15_+_36014748
|
0.488
|
NM_152453
|
TMCO5A
|
transmembrane and coiled-coil domains 5A
|
|
chr11_+_45825244
|
0.485
|
NM_001127457
|
CRY2
|
cryptochrome 2 (photolyase-like)
|
|
chr18_+_27425727
|
0.479
|
NM_000371
|
TTR
|
transthyretin
|
|
chr7_-_80386602
|
0.475
|
NM_006379
|
SEMA3C
|
sema domain, immunoglobulin domain (Ig), short basic domain, secreted, (semaphorin) 3C
|
|
chrX_+_46825179
|
0.471
|
|
RGN
|
regucalcin (senescence marker protein-30)
|
|
chr14_-_59407103
|
0.470
|
NM_021136
|
RTN1
|
reticulon 1
|
|
chr5_-_95044415
|
0.463
|
NM_031952
|
SPATA9
|
spermatogenesis associated 9
|
|
chr4_+_114433551
|
0.459
|
|
ANK2
|
ankyrin 2, neuronal
|
|
chr10_+_82268433
|
0.457
|
|
TSPAN14
|
tetraspanin 14
|
|
chr11_-_27700180
|
0.455
|
NM_170731
|
BDNF
|
brain-derived neurotrophic factor
|
|
chr2_+_113973130
|
0.454
|
NM_012184
|
FOXD4L1
|
forkhead box D4-like 1
|
|
chr7_-_19778928
|
0.454
|
NM_152774
|
TMEM196
|
transmembrane protein 196
|
|
chr3_-_124893875
|
0.453
|
|
MYLK
|
myosin light chain kinase
|
|
chr8_-_13018123
|
0.449
|
NM_001164271
|
DLC1
|
deleted in liver cancer 1
|
|
chr19_+_13066956
|
0.448
|
|
NFIX
|
nuclear factor I/X (CCAAT-binding transcription factor)
|
|
chr2_+_214984705
|
0.447
|
NM_001080500
|
VWC2L
|
von Willebrand factor C domain containing protein 2-like
|
|
chr14_-_56346849
|
0.441
|
NM_021728
|
OTX2
|
orthodenticle homeobox 2
|
|
chr8_-_93181091
|
0.439
|
NM_001198628
|
RUNX1T1
|
runt-related transcription factor 1; translocated to, 1 (cyclin D-related)
|
|
chr6_-_117193505
|
0.439
|
NM_001085480
|
FAM162B
|
family with sequence similarity 162, member B
|
|
chr13_+_23042722
|
0.436
|
NM_148957
|
TNFRSF19
|
tumor necrosis factor receptor superfamily, member 19
|
|
chr17_+_17339059
|
0.433
|
|
|
|
|
chr7_-_96489884
|
0.430
|
|
|
|
|
chr4_-_101658094
|
0.425
|
NM_001159694
NM_016242
|
EMCN
|
endomucin
|
|
chr3_+_149930656
|
0.417
|
NM_032049
|
AGTR1
|
angiotensin II receptor, type 1
|
|
chr4_+_22303645
|
0.415
|
NM_001128432
NM_020973
|
GBA3
|
glucosidase, beta, acid 3 (cytosolic)
|
|
chr7_-_4867963
|
0.414
|
NM_020144
|
PAPOLB
|
poly(A) polymerase beta (testis specific)
|
|
chr19_-_53941835
|
0.408
|
NM_182575
|
IZUMO1
|
izumo sperm-egg fusion 1
|
|
chr3_+_134978615
|
0.408
|
|
TF
|
transferrin
|
|
chr19_+_56049982
|
0.403
|
NM_001030047
NM_001030048
NM_001030050
NM_001648
|
KLK3
|
kallikrein-related peptidase 3
|
|
chr14_+_88130482
|
0.403
|
NM_207662
|
ZC3H14
|
zinc finger CCCH-type containing 14
|
|
chr15_+_94670160
|
0.403
|
NM_001145155
|
NR2F2
|
nuclear receptor subfamily 2, group F, member 2
|
|
chr3_+_12367993
|
0.401
|
NM_015869
|
PPARG
|
peroxisome proliferator-activated receptor gamma
|
|
chr14_-_59167193
|
0.399
|
NM_206852
|
RTN1
|
reticulon 1
|
|
chr5_-_22889192
|
0.396
|
|
CDH12
|
cadherin 12, type 2 (N-cadherin 2)
|
|
chrX_-_54841397
|
0.395
|
NM_198510
|
ITIH5L
|
inter-alpha (globulin) inhibitor H5-like
|
|
chr10_-_90601655
|
0.394
|
NM_144590
|
ANKRD22
|
ankyrin repeat domain 22
|
|
chr1_+_179164195
|
0.392
|
|
|
|
|
chr12_-_11230809
|
0.390
|
NM_181429
|
TAS2R42
|
taste receptor, type 2, member 42
|
|
chr12_-_24628348
|
0.390
|
|
C12orf67
|
chromosome 12 open reading frame 67
|
|
chr4_-_15686838
|
0.388
|
NM_001145849
NM_001145850
NM_001145851
NM_001145852
NM_006017
|
PROM1
|
prominin 1
|
|
chr19_-_60383280
|
0.387
|
NM_003180
|
SYT5
|
synaptotagmin V
|
|
chrX_-_52700674
|
0.387
|
NM_173358
|
SSX7
|
synovial sarcoma, X breakpoint 7
|
|
chr10_-_104587069
|
0.387
|
NM_000102
|
CYP17A1
|
cytochrome P450, family 17, subfamily A, polypeptide 1
|
|
chr11_-_113971607
|
0.387
|
|
FAM55D
|
family with sequence similarity 55, member D
|
|
chr3_-_48607596
|
0.384
|
NM_000094
|
COL7A1
|
collagen, type VII, alpha 1
|
|
chr1_+_26220852
|
0.384
|
NM_004455
|
EXTL1
|
exostoses (multiple)-like 1
|
|
chr12_+_1596482
|
0.383
|
NM_030775
|
WNT5B
|
wingless-type MMTV integration site family, member 5B
|
|
chr12_-_14612057
|
0.382
|
NM_024829
|
PLBD1
|
phospholipase B domain containing 1
|
|
chrX_-_31436335
|
0.382
|
NM_004014
|
DMD
|
dystrophin
|
|
chr6_+_29382381
|
0.376
|
NM_030946
|
OR14J1
|
olfactory receptor, family 14, subfamily J, member 1
|
|
chr7_-_11643113
|
0.375
|
|
THSD7A
|
thrombospondin, type I, domain containing 7A
|
|
chr9_-_69468634
|
0.372
|
NM_001126334
|
FOXD4L5
|
forkhead box D4-like 5
|
|
chr5_-_22248630
|
0.372
|
|
CDH12
|
cadherin 12, type 2 (N-cadherin 2)
|
|
chr6_-_127822227
|
0.372
|
NM_014702
|
KIAA0408
|
KIAA0408
|
|
chr11_-_123685871
|
0.371
|
NM_001002917
|
OR8D1
|
olfactory receptor, family 8, subfamily D, member 1
|
|
chr2_+_143603368
|
0.368
|
NM_018460
|
ARHGAP15
|
Rho GTPase activating protein 15
|
|
chr15_+_90815910
|
0.367
|
NM_153040
|
C15orf32
|
chromosome 15 open reading frame 32
|
|
chr18_-_28606839
|
0.367
|
NM_020805
|
KLHL14
|
kelch-like 14 (Drosophila)
|
|
chr7_+_30927439
|
0.364
|
NM_001185060
NM_001185061
NM_001185062
|
AQP1
|
aquaporin 1 (Colton blood group)
|
|
chr17_+_11726943
|
0.362
|
NM_004662
|
DNAH9
|
dynein, axonemal, heavy chain 9
|
|
chr18_-_30057512
|
0.360
|
NM_001198546
NM_001198548
NM_003787
|
NOL4
|
nucleolar protein 4
|
|
chr12_-_14611820
|
0.360
|
|
PLBD1
|
phospholipase B domain containing 1
|
|
chr4_-_122304925
|
0.359
|
NM_024873
|
TNIP3
|
TNFAIP3 interacting protein 3
|
|
chr6_-_127882192
|
0.357
|
NM_001012279
|
C6orf174
|
chromosome 6 open reading frame 174
|
|
chr11_+_123405506
|
0.355
|
NM_001004464
|
OR10G8
|
olfactory receptor, family 10, subfamily G, member 8
|
|
chr17_-_25685175
|
0.353
|
NM_206832
|
TMIGD1
|
transmembrane and immunoglobulin domain containing 1
|
|
chr4_-_123092358
|
0.352
|
NM_001130698
|
TRPC3
|
transient receptor potential cation channel, subfamily C, member 3
|
|
chr11_-_113971681
|
0.351
|
NM_001077639
NM_017678
|
FAM55D
|
family with sequence similarity 55, member D
|
|
chr1_+_117098579
|
0.350
|
NM_001767
|
CD2
|
CD2 molecule
|
|
chr7_+_18296569
|
0.349
|
|
HDAC9
|
histone deacetylase 9
|
|
chr3_-_152144712
|
0.347
|
NM_052995
|
CLRN1
|
clarin 1
|
|
chr4_+_96980261
|
0.346
|
NM_005390
|
PDHA2
|
pyruvate dehydrogenase (lipoamide) alpha 2
|
|
chr3_+_160269734
|
0.342
|
NM_001197113
NM_001197114
NM_001042705
NM_001042706
NM_001197100
|
IQCJ-SCHIP1
IQCJ
|
IQCJ-SCHIP1 readthrough
IQ motif containing J
|
|
chr1_+_153232685
|
0.341
|
NM_018655
|
LENEP
|
lens epithelial protein
|
|
chr2_+_143603439
|
0.341
|
|
ARHGAP15
|
Rho GTPase activating protein 15
|
|
chr14_-_69616616
|
0.340
|
NM_182936
NM_001130417
|
SLC8A3
|
solute carrier family 8 (sodium/calcium exchanger), member 3
|
|
chr8_-_23768242
|
0.339
|
NM_003155
|
STC1
|
stanniocalcin 1
|
|
chr15_+_60838053
|
0.339
|
|
TLN2
|
talin 2
|
|
chr15_-_72804763
|
0.338
|
NM_000499
|
CYP1A1
|
cytochrome P450, family 1, subfamily A, polypeptide 1
|
|
chr10_+_90474280
|
0.338
|
NM_001080518
|
LIPK
|
lipase, family member K
|
|
chr2_+_207512522
|
0.338
|
NM_173077
|
CPO
|
carboxypeptidase O
|
|
chr3_-_13032130
|
0.337
|
|
|
|
|
chr16_+_68516401
|
0.336
|
NM_199424
|
WWP2
|
WW domain containing E3 ubiquitin protein ligase 2
|
|
chr9_-_108416
|
0.336
|
NM_207305
|
FOXD4
|
forkhead box D4
|
|
chrX_-_134798808
|
0.332
|
NM_001017438
|
CT45A3
CT45A4
CT45A5
CT45A6
CT45A1
|
cancer/testis antigen family 45, member A3
cancer/testis antigen family 45, member A4
cancer/testis antigen family 45, member A5
cancer/testis antigen family 45, member A6
cancer/testis antigen family 45, member A1
|
|
chr12_+_54106304
|
0.331
|
NM_001005183
|
OR6C76
|
olfactory receptor, family 6, subfamily C, member 76
|
|
chr22_+_44059491
|
0.328
|
NM_001167574
NM_006953
|
UPK3A
|
uroplakin 3A
|
|
chr10_-_62002672
|
0.328
|
|
ANK3
|
ankyrin 3, node of Ranvier (ankyrin G)
|
|
chr4_+_5577783
|
0.328
|
NM_005750
|
C4orf6
|
chromosome 4 open reading frame 6
|
|
chr14_-_80934523
|
0.328
|
NM_033104
|
STON2
|
stonin 2
|
|
chr3_-_9895471
|
0.327
|
|
CIDEC
|
cell death-inducing DFFA-like effector c
|
|
chr6_+_106640903
|
0.326
|
|
PRDM1
|
PR domain containing 1, with ZNF domain
|
|
chr15_+_45263517
|
0.326
|
NM_001198999
|
SEMA6D
|
sema domain, transmembrane domain (TM), and cytoplasmic domain, (semaphorin) 6D
|
|
chr7_+_137411717
|
0.325
|
NM_001190906
NM_001190907
NM_005989
|
AKR1D1
|
aldo-keto reductase family 1, member D1 (delta 4-3-ketosteroid-5-beta-reductase)
|
|
chr15_+_87540238
|
0.324
|
|
ABHD2
|
abhydrolase domain containing 2
|
|
chr14_-_59166976
|
0.324
|
|
RTN1
|
reticulon 1
|
|
chr5_+_127012611
|
0.322
|
NM_001048252
|
CTXN3
|
cortexin 3
|
|
chr12_-_91346054
|
0.322
|
NM_001025232
|
CLLU1OS
|
chronic lymphocytic leukemia up-regulated 1 opposite strand
|
|
chr13_+_52500830
|
0.321
|
NM_006418
|
OLFM4
|
olfactomedin 4
|
|
chr19_-_295778
|
0.320
|
NM_017550
|
MIER2
|
mesoderm induction early response 1, family member 2
|
|
chr1_+_109450116
|
0.319
|
|
|
|
|
chr15_-_79403494
|
0.317
|
NM_181900
|
STARD5
|
StAR-related lipid transfer (START) domain containing 5
|
|
chr2_+_222871109
|
0.316
|
NM_153038
|
CCDC140
|
coiled-coil domain containing 140
|
|
chr1_+_92456160
|
0.315
|
NM_001012425
|
C1orf146
|
chromosome 1 open reading frame 146
|
|
chr12_+_5473558
|
0.315
|
NM_002527
|
NTF3
|
neurotrophin 3
|
|
chr9_+_42707229
|
0.314
|
NM_001099279
NM_199244
|
FOXD4L2
FOXD4L4
|
forkhead box D4-like 2
forkhead box D4-like 4
|
|
chr11_-_65070625
|
0.313
|
|
LTBP3
|
latent transforming growth factor beta binding protein 3
|
|
chr4_-_122357104
|
0.312
|
NM_001128843
|
TNIP3
|
TNFAIP3 interacting protein 3
|
|
chr1_+_144004589
|
0.310
|
NM_001039703
|
NBPF10
NBPF8
|
neuroblastoma breakpoint family, member 10
neuroblastoma breakpoint family, member 8
|
|
chr9_+_110664366
|
0.309
|
NM_006687
|
ACTL7A
|
actin-like 7A
|
|
chr1_+_66570363
|
0.309
|
NM_001037339
|
PDE4B
|
phosphodiesterase 4B, cAMP-specific
|
|
chr3_+_45042754
|
0.308
|
NM_003278
|
CLEC3B
|
C-type lectin domain family 3, member B
|
|
chr4_-_108176901
|
0.308
|
NM_014421
|
DKK2
|
dickkopf homolog 2 (Xenopus laevis)
|
|
chr3_-_44464722
|
0.305
|
|
|
|
|
chr2_+_108604111
|
0.305
|
NM_001193484
|
LIMS1
|
LIM and senescent cell antigen-like domains 1
|
|
chr19_-_2036227
|
0.304
|
|
MOBKL2A
|
MOB1, Mps One Binder kinase activator-like 2A (yeast)
|
|
chr10_+_18280773
|
0.303
|
NM_001145195
NM_152725
|
SLC39A12
|
solute carrier family 39 (zinc transporter), member 12
|
|
chr2_+_166137091
|
0.302
|
NM_024969
|
CSRNP3
|
cysteine-serine-rich nuclear protein 3
|
|
chr9_+_70107602
|
0.297
|
NM_199135
|
FOXD4L3
FOXD4L6
|
forkhead box D4-like 3
forkhead box D4-like 6
|
|
chr8_+_24297742
|
0.295
|
NM_001145271
NM_001145272
NM_014479
|
ADAMDEC1
|
ADAM-like, decysin 1
|
|
chr3_-_42427026
|
0.295
|
NM_144634
|
LYZL4
|
lysozyme-like 4
|
|
chr3_+_46386636
|
0.294
|
NM_000579
NM_001100168
|
CCR5
|
chemokine (C-C motif) receptor 5
|
|
chrX_-_48156287
|
0.293
|
NM_001034832
NM_001040612
NM_005636
NM_175729
|
SSX4B
SSX4
|
synovial sarcoma, X breakpoint 4B
synovial sarcoma, X breakpoint 4
|
|
chr12_-_67643286
|
0.293
|
NM_001874
|
CPM
|
carboxypeptidase M
|
|
chr7_-_7541974
|
0.293
|
NM_001037763
|
COL28A1
|
collagen, type XXVIII, alpha 1
|
|
chr10_-_75080784
|
0.292
|
NM_024875
|
SYNPO2L
|
synaptopodin 2-like
|