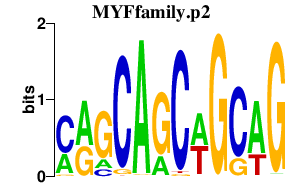
|
chr13_-_34948758
|
6.626
|
NM_005584
|
MAB21L1
|
mab-21-like 1 (C. elegans)
|
|
chr5_+_113725914
|
6.221
|
NM_021614
|
KCNN2
|
potassium intermediate/small conductance calcium-activated channel, subfamily N, member 2
|
|
chr5_+_113726203
|
5.945
|
|
KCNN2
|
potassium intermediate/small conductance calcium-activated channel, subfamily N, member 2
|
|
chr13_-_71338558
|
5.287
|
|
DACH1
|
dachshund homolog 1 (Drosophila)
|
|
chr13_+_34948885
|
5.194
|
|
NBEA
|
neurobeachin
|
|
chr8_+_56177567
|
4.970
|
NM_052898
|
XKR4
|
XK, Kell blood group complex subunit-related family, member 4
|
|
chr2_-_230287442
|
4.879
|
NM_139072
|
DNER
|
delta/notch-like EGF repeat containing
|
|
chr6_+_99389318
|
4.599
|
|
POU3F2
|
POU class 3 homeobox 2
|
|
chr13_-_71338144
|
4.547
|
|
DACH1
|
dachshund homolog 1 (Drosophila)
|
|
chr6_+_107917965
|
4.372
|
NM_018013
|
SOBP
|
sine oculis binding protein homolog (Drosophila)
|
|
chr5_-_59225377
|
4.171
|
NM_001104631
|
PDE4D
|
phosphodiesterase 4D, cAMP-specific
|
|
chr3_-_130807865
|
3.717
|
NM_015103
|
PLXND1
|
plexin D1
|
|
chr22_-_19122047
|
3.205
|
|
SCARF2
|
scavenger receptor class F, member 2
|
|
chr8_+_31616809
|
3.094
|
NM_013962
|
NRG1
|
neuregulin 1
|
|
chr22_+_28206180
|
3.045
|
NM_021076
|
NEFH
|
neurofilament, heavy polypeptide
|
|
chr12_+_101876151
|
3.016
|
|
ASCL1
|
achaete-scute complex homolog 1 (Drosophila)
|
|
chr2_+_104838177
|
2.884
|
NM_006236
|
POU3F3
|
POU class 3 homeobox 3
|
|
chr8_-_23768242
|
2.844
|
NM_003155
|
STC1
|
stanniocalcin 1
|
|
chrX_-_92815222
|
2.729
|
NM_004538
|
NAP1L3
|
nucleosome assembly protein 1-like 3
|
|
chr8_+_65656164
|
2.719
|
|
BHLHE22
|
basic helix-loop-helix family, member e22
|
|
chrX_-_99551926
|
2.618
|
NM_001105243
NM_001184880
NM_020766
|
PCDH19
|
protocadherin 19
|
|
chr17_+_14145051
|
2.481
|
NM_006041
|
HS3ST3B1
|
heparan sulfate (glucosamine) 3-O-sulfotransferase 3B1
|
|
chrX_-_74062005
|
2.439
|
NM_001008537
|
KIAA2022
|
KIAA2022
|
|
chr8_+_1937711
|
2.437
|
|
KBTBD11
|
kelch repeat and BTB (POZ) domain containing 11
|
|
chr5_-_178705036
|
2.410
|
NM_014244
NM_021599
|
ADAMTS2
|
ADAM metallopeptidase with thrombospondin type 1 motif, 2
|
|
chr4_+_61749540
|
2.406
|
|
LPHN3
|
latrophilin 3
|
|
chrX_-_107866262
|
2.388
|
NM_003604
|
IRS4
|
insulin receptor substrate 4
|
|
chr3_-_62835601
|
2.298
|
|
|
|
|
chrX_-_30237403
|
2.262
|
NM_000475
|
NR0B1
|
nuclear receptor subfamily 0, group B, member 1
|
|
chr9_-_139183675
|
2.260
|
|
|
|
|
chr5_-_146238336
|
2.248
|
NM_004576
NM_181675
NM_001127381
|
PPP2R2B
|
protein phosphatase 2, regulatory subunit B, beta
|
|
chr6_+_45498354
|
2.235
|
|
RUNX2
|
runt-related transcription factor 2
|
|
chr19_-_15204194
|
2.227
|
NM_024794
|
EPHX3
|
epoxide hydrolase 3
|
|
chr1_+_219119623
|
2.224
|
|
HLX
|
H2.0-like homeobox
|
|
chr18_-_24010944
|
2.215
|
|
CDH2
|
cadherin 2, type 1, N-cadherin (neuronal)
|
|
chr10_-_132999798
|
2.205
|
|
TCERG1L
|
transcription elongation regulator 1-like
|
|
chr8_+_37773991
|
2.195
|
|
GPR124
|
G protein-coupled receptor 124
|
|
chr17_-_4636248
|
2.191
|
NM_001144939
NM_001144940
NM_001144941
NM_182566
|
VMO1
|
vitelline membrane outer layer 1 homolog (chicken)
|
|
chr7_-_137182125
|
2.159
|
NM_004717
|
DGKI
|
diacylglycerol kinase, iota
|
|
chr15_+_90738108
|
2.125
|
NM_006011
|
ST8SIA2
|
ST8 alpha-N-acetyl-neuraminide alpha-2,8-sialyltransferase 2
|
|
chr6_+_5030718
|
2.120
|
NM_001145115
|
PPP1R3G
|
protein phosphatase 1, regulatory (inhibitor) subunit 3G
|
|
chr2_-_27384617
|
2.091
|
NM_003353
|
UCN
|
urocortin
|
|
chr3_+_161426322
|
2.042
|
|
LOC401097
|
hypothetical protein LOC401097
|
|
chr19_+_1236766
|
2.031
|
NM_001405
|
EFNA2
|
ephrin-A2
|
|
chr10_-_132999973
|
2.027
|
NM_174937
|
TCERG1L
|
transcription elongation regulator 1-like
|
|
chr18_-_21185968
|
2.025
|
|
ZNF521
|
zinc finger protein 521
|
|
chr11_-_617172
|
1.982
|
NM_021920
|
SCT
|
secretin
|
|
chr8_-_140784416
|
1.968
|
NM_016601
|
KCNK9
|
potassium channel, subfamily K, member 9
|
|
chr5_+_137829077
|
1.925
|
NM_001964
|
EGR1
|
early growth response 1
|
|
chr4_-_155631861
|
1.920
|
|
DCHS2
|
dachsous 2 (Drosophila)
|
|
chr16_+_85101734
|
1.885
|
|
FOXF1
|
forkhead box F1
|
|
chr14_+_28306633
|
1.876
|
|
FOXG1
|
forkhead box G1
|
|
chr16_+_85101605
|
1.870
|
NM_001451
|
FOXF1
|
forkhead box F1
|
|
chrX_-_110400406
|
1.855
|
NM_014289
|
CAPN6
|
calpain 6
|
|
chr4_+_172971149
|
1.836
|
NM_001034845
|
GALNTL6
|
UDP-N-acetyl-alpha-D-galactosamine:polypeptide N-acetylgalactosaminyltransferase-like 6
|
|
chr8_-_133562185
|
1.833
|
NM_004519
|
KCNQ3
|
potassium voltage-gated channel, KQT-like subfamily, member 3
|
|
chr11_-_129803712
|
1.820
|
NM_007037
|
ADAMTS8
|
ADAM metallopeptidase with thrombospondin type 1 motif, 8
|
|
chr15_-_53822468
|
1.801
|
NM_173814
|
PRTG
|
protogenin
|
|
chr1_-_29321599
|
1.798
|
NM_001171868
|
TMEM200B
|
transmembrane protein 200B
|
|
chr13_-_78075683
|
1.798
|
NM_006237
|
POU4F1
|
POU class 4 homeobox 1
|
|
chrX_-_24941098
|
1.769
|
|
ARX
|
aristaless related homeobox
|
|
chr3_+_3816079
|
1.733
|
NM_020873
|
LRRN1
|
leucine rich repeat neuronal 1
|
|
chr2_+_114916368
|
1.720
|
NM_020868
|
DPP10
|
dipeptidyl-peptidase 10 (non-functional)
|
|
chr8_-_133561933
|
1.719
|
|
KCNQ3
|
potassium voltage-gated channel, KQT-like subfamily, member 3
|
|
chr12_+_6807831
|
1.715
|
NM_014262
|
LEPREL2
|
leprecan-like 2
|
|
chr4_+_147779492
|
1.705
|
NM_004575
|
POU4F2
|
POU class 4 homeobox 2
|
|
chr10_-_73518315
|
1.701
|
|
SPOCK2
|
sparc/osteonectin, cwcv and kazal-like domains proteoglycan (testican) 2
|
|
chr5_+_126654354
|
1.697
|
NM_032446
|
MEGF10
|
multiple EGF-like-domains 10
|
|
chr2_-_192767844
|
1.669
|
NM_016192
|
TMEFF2
|
transmembrane protein with EGF-like and two follistatin-like domains 2
|
|
chr3_+_142252890
|
1.667
|
|
SPSB4
|
splA/ryanodine receptor domain and SOCS box containing 4
|
|
chr3_+_26639300
|
1.654
|
NM_052953
|
LRRC3B
|
leucine rich repeat containing 3B
|
|
chr4_+_134292630
|
1.645
|
|
PCDH10
|
protocadherin 10
|
|
chr2_-_39040981
|
1.643
|
|
LOC375196
|
hypothetical LOC375196
|
|
chrX_-_110400321
|
1.640
|
|
CAPN6
|
calpain 6
|
|
chr5_-_178704934
|
1.634
|
|
ADAMTS2
|
ADAM metallopeptidase with thrombospondin type 1 motif, 2
|
|
chr5_+_92946348
|
1.623
|
|
NR2F1
|
nuclear receptor subfamily 2, group F, member 1
|
|
chr7_+_137182230
|
1.617
|
|
|
|
|
chr8_+_85258056
|
1.607
|
NM_173848
|
RALYL
|
RALY RNA binding protein-like
|
|
chr15_-_50869379
|
1.604
|
NM_004498
|
ONECUT1
|
one cut homeobox 1
|
|
chr19_-_13478037
|
1.588
|
NM_000068
NM_001127221
NM_001127222
NM_001174080
NM_023035
|
CACNA1A
|
calcium channel, voltage-dependent, P/Q type, alpha 1A subunit
|
|
chr8_+_37773915
|
1.586
|
|
GPR124
|
G protein-coupled receptor 124
|
|
chr1_+_100777044
|
1.571
|
|
GPR88
|
G protein-coupled receptor 88
|
|
chr6_+_129245978
|
1.550
|
NM_000426
NM_001079823
|
LAMA2
|
laminin, alpha 2
|
|
chr5_-_147142457
|
1.537
|
|
JAKMIP2
|
janus kinase and microtubule interacting protein 2
|
|
chr11_-_129803515
|
1.534
|
|
ADAMTS8
|
ADAM metallopeptidase with thrombospondin type 1 motif, 8
|
|
chr5_+_170779264
|
1.531
|
NM_003862
|
FGF18
|
fibroblast growth factor 18
|
|
chr4_-_168392307
|
1.528
|
NM_001040159
NM_016950
|
SPOCK3
|
sparc/osteonectin, cwcv and kazal-like domains proteoglycan (testican) 3
|
|
chr3_+_141136896
|
1.527
|
|
CLSTN2
|
calsyntenin 2
|
|
chr8_+_85258007
|
1.508
|
NM_001100392
NM_001100393
|
RALYL
|
RALY RNA binding protein-like
|
|
chr3_+_161425946
|
1.508
|
NM_001168214
|
LOC401097
|
hypothetical protein LOC401097
|
|
chr12_+_6808025
|
1.498
|
|
LEPREL2
|
leprecan-like 2
|
|
chr14_+_58174489
|
1.486
|
NM_001079520
NM_016651
|
DACT1
|
dapper, antagonist of beta-catenin, homolog 1 (Xenopus laevis)
|
|
chr19_+_18584666
|
1.482
|
NM_012109
|
TMEM59L
|
transmembrane protein 59-like
|
|
chr6_+_45498262
|
1.481
|
|
RUNX2
|
runt-related transcription factor 2
|
|
chr12_+_130878870
|
1.476
|
NM_016155
|
MMP17
|
matrix metallopeptidase 17 (membrane-inserted)
|
|
chr4_-_13155135
|
1.453
|
NM_001189
|
NKX3-2
|
NK3 homeobox 2
|
|
chr11_+_627268
|
1.446
|
NM_000797
|
DRD4
|
dopamine receptor D4
|
|
chr9_-_119217147
|
1.442
|
|
ASTN2
|
astrotactin 2
|
|
chr4_-_168392266
|
1.432
|
|
SPOCK3
|
sparc/osteonectin, cwcv and kazal-like domains proteoglycan (testican) 3
|
|
chr10_+_83625049
|
1.431
|
NM_001010848
NM_001165972
|
NRG3
|
neuregulin 3
|
|
chrX_+_53095230
|
1.426
|
NM_018969
|
GPR173
|
G protein-coupled receptor 173
|
|
chr3_-_144164867
|
1.426
|
NM_198504
|
PAQR9
|
progestin and adipoQ receptor family member IX
|
|
chr15_-_39593285
|
1.426
|
NM_001135685
NM_002344
NM_206961
|
LTK
|
leukocyte receptor tyrosine kinase
|
|
chr7_+_153380673
|
1.419
|
NM_130797
|
DPP6
|
dipeptidyl-peptidase 6
|
|
chr15_-_63457378
|
1.419
|
NM_004884
|
IGDCC3
|
immunoglobulin superfamily, DCC subclass, member 3
|
|
chr7_-_72822471
|
1.418
|
NM_001306
|
CLDN3
|
claudin 3
|
|
chr4_-_142273881
|
1.410
|
NM_020724
|
RNF150
|
ring finger protein 150
|
|
chrX_+_12066505
|
1.385
|
NM_014728
|
FRMPD4
|
FERM and PDZ domain containing 4
|
|
chr10_-_73518269
|
1.384
|
|
SPOCK2
|
sparc/osteonectin, cwcv and kazal-like domains proteoglycan (testican) 2
|
|
chr1_-_11841476
|
1.384
|
NM_002521
|
NPPB
|
natriuretic peptide B
|
|
chr11_-_2113297
|
1.378
|
|
IGF2
|
insulin-like growth factor 2 (somatomedin A)
|
|
chr10_+_23521464
|
1.376
|
NM_178161
|
PTF1A
|
pancreas specific transcription factor, 1a
|
|
chr5_+_115326035
|
1.366
|
NM_173800
|
AQPEP
|
laeverin
|
|
chr6_+_43151922
|
1.365
|
NM_002821
NM_152880
NM_152881
NM_152882
|
PTK7
|
PTK7 protein tyrosine kinase 7
|
|
chr10_+_25504295
|
1.355
|
NM_020752
|
GPR158
|
G protein-coupled receptor 158
|
|
chr9_-_103540038
|
1.352
|
|
GRIN3A
|
glutamate receptor, ionotropic, N-methyl-D-aspartate 3A
|
|
chr10_+_115793795
|
1.346
|
NM_000684
|
ADRB1
|
adrenergic, beta-1-, receptor
|
|
chr3_+_112273464
|
1.333
|
NM_015480
|
PVRL3
|
poliovirus receptor-related 3
|
|
chr13_+_34948977
|
1.333
|
|
NBEA
|
neurobeachin
|
|
chr11_+_14621972
|
1.332
|
|
PDE3B
|
phosphodiesterase 3B, cGMP-inhibited
|
|
chr16_-_47872880
|
1.326
|
|
|
|
|
chr8_+_99508425
|
1.321
|
NM_020697
|
KCNS2
|
potassium voltage-gated channel, delayed-rectifier, subfamily S, member 2
|
|
chr22_-_19122107
|
1.315
|
NM_153334
NM_182895
|
SCARF2
|
scavenger receptor class F, member 2
|
|
chr16_+_2009741
|
1.300
|
|
NPW
|
neuropeptide W
|
|
chr11_-_132907499
|
1.296
|
NM_001012393
|
OPCML
|
opioid binding protein/cell adhesion molecule-like
|
|
chr3_+_195889845
|
1.285
|
|
FAM43A
|
family with sequence similarity 43, member A
|
|
chr22_+_19699441
|
1.280
|
NM_001159554
NM_005446
|
P2RX6
|
purinergic receptor P2X, ligand-gated ion channel, 6
|
|
chr1_+_211190631
|
1.277
|
|
VASH2
|
vasohibin 2
|
|
chr9_-_139184247
|
1.272
|
NM_001013653
|
LRRC26
|
leucine rich repeat containing 26
|
|
chr6_+_72653126
|
1.235
|
|
RIMS1
|
regulating synaptic membrane exocytosis 1
|
|
chr10_-_73518284
|
1.228
|
|
SPOCK2
|
sparc/osteonectin, cwcv and kazal-like domains proteoglycan (testican) 2
|
|
chr3_+_112273352
|
1.226
|
|
PVRL3
|
poliovirus receptor-related 3
|
|
chrX_-_53367246
|
1.224
|
NM_001111125
|
IQSEC2
|
IQ motif and Sec7 domain 2
|
|
chr4_+_15313850
|
1.219
|
|
BST1
|
bone marrow stromal cell antigen 1
|
|
chr6_-_33268177
|
1.214
|
NM_001163771
NM_080679
NM_080680
NM_080681
|
COL11A2
|
collagen, type XI, alpha 2
|
|
chr17_-_15104768
|
1.212
|
NM_153322
|
PMP22
|
peripheral myelin protein 22
|
|
chr10_-_118022965
|
1.195
|
NM_005264
|
GFRA1
|
GDNF family receptor alpha 1
|
|
chr10_+_52504239
|
1.176
|
NM_006258
|
PRKG1
|
protein kinase, cGMP-dependent, type I
|
|
chr5_-_147142249
|
1.176
|
|
JAKMIP2
|
janus kinase and microtubule interacting protein 2
|
|
chr16_-_2186465
|
1.175
|
NM_020764
|
CASKIN1
|
CASK interacting protein 1
|
|
chr8_+_143542378
|
1.175
|
NM_001702
|
BAI1
|
brain-specific angiogenesis inhibitor 1
|
|
chr12_+_93066370
|
1.174
|
NM_005761
|
PLXNC1
|
plexin C1
|
|
chr7_+_98084531
|
1.171
|
NM_002523
|
NPTX2
|
neuronal pentraxin II
|
|
chr20_-_30534848
|
1.168
|
NM_080616
|
C20orf112
|
chromosome 20 open reading frame 112
|
|
chr8_+_106400322
|
1.167
|
NM_012082
|
ZFPM2
|
zinc finger protein, multitype 2
|
|
chrX_+_149280486
|
1.167
|
|
MAMLD1
|
mastermind-like domain containing 1
|
|
chr3_-_124650081
|
1.158
|
NM_183357
|
ADCY5
|
adenylate cyclase 5
|
|
chr5_+_75414994
|
1.153
|
NM_014979
|
SV2C
|
synaptic vesicle glycoprotein 2C
|
|
chr16_+_55180767
|
1.152
|
NM_005954
|
MT3
|
metallothionein 3
|
|
chr10_-_30356344
|
1.151
|
|
KIAA1462
|
KIAA1462
|
|
chr8_-_121893468
|
1.151
|
NM_021021
|
SNTB1
|
syntrophin, beta 1 (dystrophin-associated protein A1, 59kDa, basic component 1)
|
|
chr12_+_79634820
|
1.150
|
NM_005593
|
MYF5
|
myogenic factor 5
|
|
chr18_-_21186107
|
1.149
|
NM_015461
|
ZNF521
|
zinc finger protein 521
|
|
chr4_-_168392151
|
1.147
|
|
SPOCK3
|
sparc/osteonectin, cwcv and kazal-like domains proteoglycan (testican) 3
|
|
chr8_+_37773564
|
1.144
|
|
GPR124
|
G protein-coupled receptor 124
|
|
chr9_-_38414422
|
1.137
|
NM_001007563
|
IGFBPL1
|
insulin-like growth factor binding protein-like 1
|
|
chr9_+_86474500
|
1.125
|
|
NTRK2
|
neurotrophic tyrosine kinase, receptor, type 2
|
|
chr19_+_47509274
|
1.122
|
NM_173633
|
TMEM145
|
transmembrane protein 145
|
|
chr1_-_102235377
|
1.118
|
NM_058170
|
OLFM3
|
olfactomedin 3
|
|
chrX_+_117992623
|
1.104
|
NM_001031855
NM_024778
|
LONRF3
|
LON peptidase N-terminal domain and ring finger 3
|
|
chr3_+_141136691
|
1.103
|
NM_022131
|
CLSTN2
|
calsyntenin 2
|
|
chr1_-_102234959
|
1.099
|
|
OLFM3
|
olfactomedin 3
|
|
chr10_+_42892506
|
1.097
|
NM_020630
NM_020975
|
RET
|
ret proto-oncogene
|
|
chr8_+_37773553
|
1.096
|
NM_032777
|
GPR124
|
G protein-coupled receptor 124
|
|
chr16_-_47872973
|
1.096
|
|
CBLN1
|
cerebellin 1 precursor
|
|
chr15_+_97462675
|
1.091
|
NM_015286
NM_145728
|
SYNM
|
synemin, intermediate filament protein
|
|
chr19_+_56319948
|
1.089
|
NM_001198558
NM_014441
|
SIGLEC9
|
sialic acid binding Ig-like lectin 9
|
|
chr13_-_109236897
|
1.085
|
NM_003749
|
IRS2
|
insulin receptor substrate 2
|
|
chr12_+_70119785
|
1.084
|
|
LGR5
|
leucine-rich repeat containing G protein-coupled receptor 5
|
|
chr3_-_18441763
|
1.078
|
NM_001195470
NM_002971
|
SATB1
|
SATB homeobox 1
|
|
chr1_-_32002222
|
1.076
|
NM_001703
|
BAI2
|
brain-specific angiogenesis inhibitor 2
|
|
chr1_+_22909916
|
1.071
|
NM_004442
NM_017449
|
EPHB2
|
EPH receptor B2
|
|
chr6_+_150326802
|
1.069
|
NM_025218
|
ULBP1
|
UL16 binding protein 1
|
|
chr8_+_24827181
|
1.062
|
|
NEFM
|
neurofilament, medium polypeptide
|
|
chr9_-_100510657
|
1.062
|
NM_005458
|
GABBR2
|
gamma-aminobutyric acid (GABA) B receptor, 2
|
|
chr19_+_55985483
|
1.061
|
NM_033068
|
ACPT
|
acid phosphatase, testicular
|
|
chr21_+_33364420
|
1.060
|
|
OLIG1
|
oligodendrocyte transcription factor 1
|
|
chr9_-_119217128
|
1.057
|
NM_014010
|
ASTN2
|
astrotactin 2
|
|
chr11_+_359723
|
1.054
|
NM_178537
|
B4GALNT4
|
beta-1,4-N-acetyl-galactosaminyl transferase 4
|
|
chr22_+_22445035
|
1.051
|
NM_005940
|
MMP11
|
matrix metallopeptidase 11 (stromelysin 3)
|
|
chr8_+_24827173
|
1.049
|
NM_005382
|
NEFM
|
neurofilament, medium polypeptide
|
|
chr4_+_7245218
|
1.047
|
NM_020777
|
SORCS2
|
sortilin-related VPS10 domain containing receptor 2
|
|
chr14_-_50631407
|
1.043
|
|
TRIM9
|
tripartite motif containing 9
|
|
chr12_+_50342814
|
1.041
|
NM_001177984
|
SCN8A
|
sodium channel, voltage gated, type VIII, alpha subunit
|
|
chr7_+_96473225
|
1.027
|
NM_005222
|
DLX6
|
distal-less homeobox 6
|
|
chr9_+_86474457
|
1.023
|
|
NTRK2
|
neurotrophic tyrosine kinase, receptor, type 2
|
|
chr6_+_99389300
|
1.019
|
NM_005604
|
POU3F2
|
POU class 3 homeobox 2
|
|
chr8_+_104452961
|
1.018
|
NM_138455
|
CTHRC1
|
collagen triple helix repeat containing 1
|
|
chr11_-_86343737
|
1.017
|
|
FZD4
|
frizzled homolog 4 (Drosophila)
|
|
chr19_-_51829778
|
1.012
|
NM_033258
|
GNG8
|
guanine nucleotide binding protein (G protein), gamma 8
|
|
chr3_+_142253102
|
1.010
|
|
SPSB4
|
splA/ryanodine receptor domain and SOCS box containing 4
|
|
chr13_+_34414390
|
1.009
|
NM_015678
|
NBEA
|
neurobeachin
|
|
chr9_+_86474553
|
1.007
|
|
NTRK2
|
neurotrophic tyrosine kinase, receptor, type 2
|
|
chr5_-_139992621
|
1.006
|
NM_000591
|
CD14
|
CD14 molecule
|
|
chr16_-_47873183
|
1.006
|
NM_004352
|
CBLN1
|
cerebellin 1 precursor
|
|
chr5_+_137829065
|
1.005
|
|
EGR1
|
early growth response 1
|
|
chrX_+_23262905
|
1.004
|
NM_173495
|
PTCHD1
|
patched domain containing 1
|
|
chr13_+_30378311
|
0.993
|
NM_032849
|
C13orf33
|
chromosome 13 open reading frame 33
|
|
chr10_+_134893766
|
0.987
|
NM_003577
|
UTF1
|
undifferentiated embryonic cell transcription factor 1
|
|
chr1_+_109594163
|
0.983
|
NM_001408
|
CELSR2
|
cadherin, EGF LAG seven-pass G-type receptor 2 (flamingo homolog, Drosophila)
|