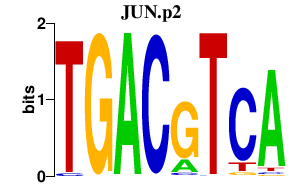
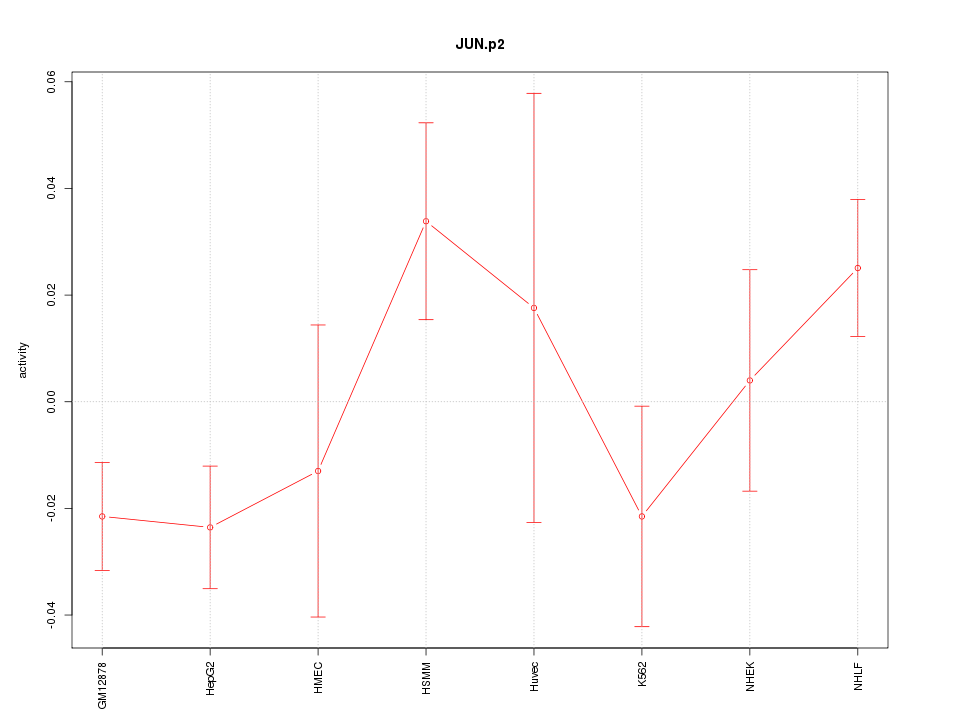
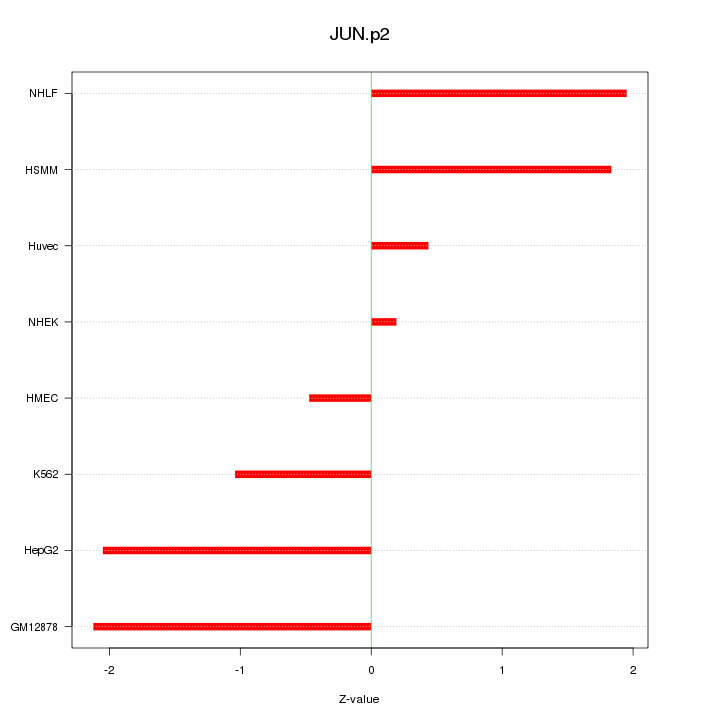
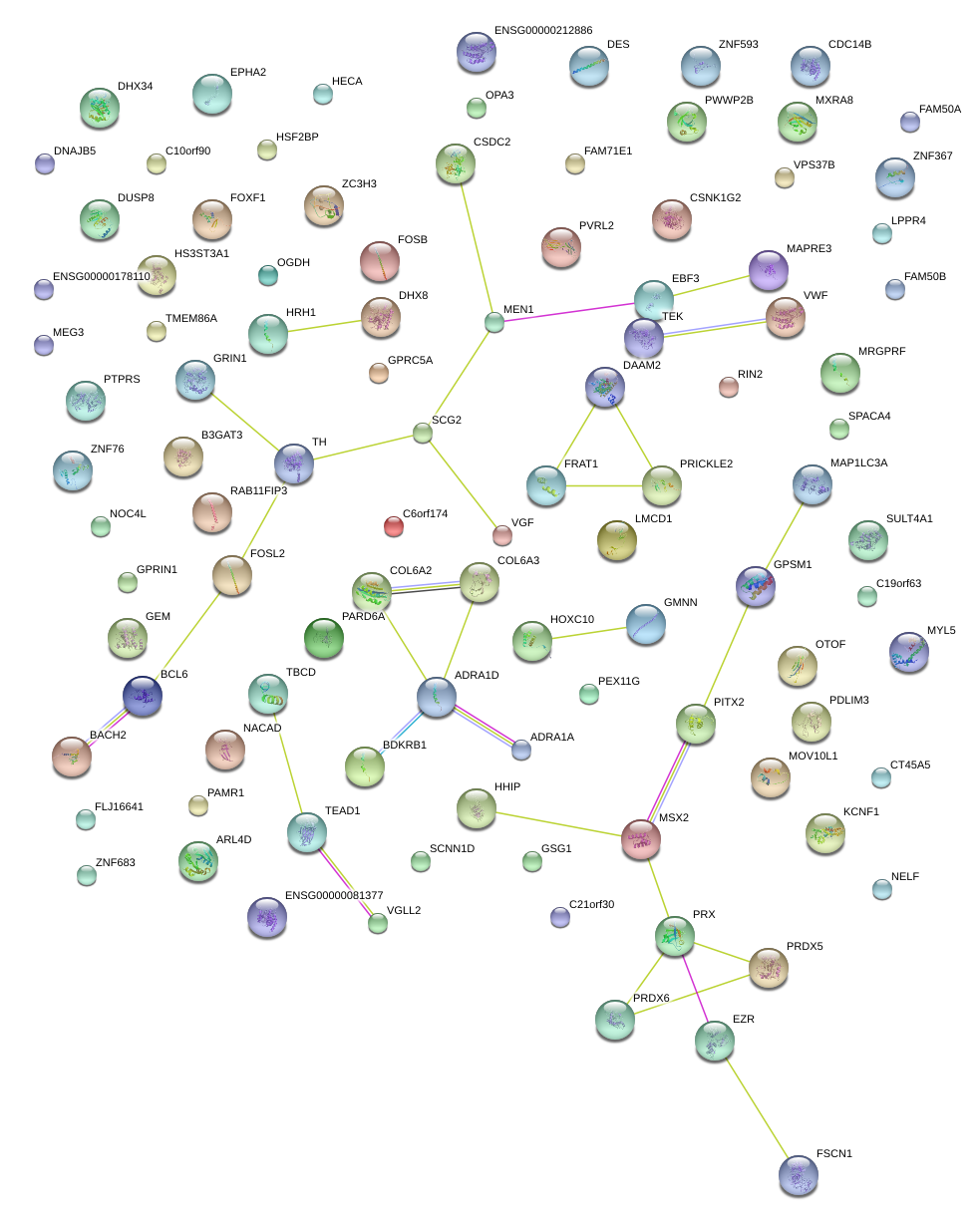
|
chr22_-_42589543
|
8.120
|
NM_014351
|
SULT4A1
|
sulfotransferase family 4A, member 1
|
|
chr12_-_121946581
|
4.736
|
|
VPS37B
|
vacuolar protein sorting 37 homolog B (S. cerevisiae)
|
|
chr12_-_121946635
|
4.337
|
NM_024667
|
VPS37B
|
vacuolar protein sorting 37 homolog B (S. cerevisiae)
|
|
chr1_+_26368969
|
3.469
|
NM_015871
|
ZNF593
|
zinc finger protein 593
|
|
chr6_+_139497933
|
3.404
|
NM_016217
|
HECA
|
headcase homolog (Drosophila)
|
|
chr11_-_1549719
|
2.906
|
NM_004420
|
DUSP8
|
dual specificity phosphatase 8
|
|
chr4_-_111763655
|
2.618
|
NM_000325
|
PITX2
|
paired-like homeodomain 2
|
|
chr20_+_32610161
|
2.429
|
NM_032514
|
MAP1LC3A
|
microtubule-associated protein 1 light chain 3 alpha
|
|
chr7_-_100595552
|
2.219
|
NM_003378
|
VGF
|
VGF nerve growth factor inducible
|
|
chr10_-_131652364
|
2.208
|
|
EBF3
|
early B-cell factor 3
|
|
chr12_+_131194967
|
2.202
|
|
NOC4L
|
nucleolar complex associated 4 homolog (S. cerevisiae)
|
|
chr10_+_134060681
|
2.197
|
NM_001098637
NM_138499
|
PWWP2B
|
PWWP domain containing 2B
|
|
chr19_+_55671600
|
2.184
|
|
C19orf63
|
chromosome 19 open reading frame 63
|
|
chr9_-_139473606
|
2.063
|
NM_001130969
NM_001130970
NM_001130971
NM_001178064
NM_015537
|
NELF
|
nasal embryonic LHRH factor
|
|
chr19_+_55671594
|
2.055
|
|
C19orf63
|
chromosome 19 open reading frame 63
|
|
chr12_+_131194931
|
2.043
|
NM_024078
|
NOC4L
|
nucleolar complex associated 4 homolog (S. cerevisiae)
|
|
chr19_+_55671584
|
2.041
|
|
C19orf63
|
chromosome 19 open reading frame 63
|
|
chr6_+_39868126
|
2.036
|
|
DAAM2
|
dishevelled associated activator of morphogenesis 2
|
|
chr12_-_121946561
|
2.031
|
|
|
|
|
chr19_+_50663089
|
2.029
|
NM_001114171
NM_006732
|
FOSB
|
FBJ murine osteosarcoma viral oncogene homolog B
|
|
chr7_-_45095017
|
1.997
|
NM_001146334
|
NACAD
|
NAC alpha domain containing
|
|
chr2_+_10969513
|
1.979
|
NM_002236
|
KCNF1
|
potassium voltage-gated channel, subfamily F, member 1
|
|
chr11_+_12652427
|
1.964
|
NM_021961
|
TEAD1
|
TEA domain family member 1 (SV40 transcriptional enhancer factor)
|
|
chr20_+_19863716
|
1.960
|
|
RIN2
|
Ras and Rab interactor 2
|
|
chr16_+_85101605
|
1.953
|
NM_001451
|
FOXF1
|
forkhead box F1
|
|
chr2_+_28469172
|
1.877
|
|
FOSL2
|
FOS-like antigen 2
|
|
chr17_-_13445942
|
1.863
|
NM_006042
|
HS3ST3A1
|
heparan sulfate (glucosamine) 3-O-sulfotransferase 3A1
|
|
chr17_+_38831874
|
1.862
|
NM_001661
|
ARL4D
|
ADP-ribosylation factor-like 4D
|
|
chr14_+_100362204
|
1.861
|
|
MEG3
|
maternally expressed 3 (non-protein coding)
|
|
chr6_+_39868116
|
1.850
|
|
|
|
|
chr19_+_55671468
|
1.757
|
NM_175063
NM_206538
|
C19orf63
|
chromosome 19 open reading frame 63
|
|
chr19_+_53801811
|
1.750
|
NM_133498
|
SPACA4
|
sperm acrosome associated 4
|
|
chr11_-_68537243
|
1.688
|
NM_001098515
NM_145015
|
MRGPRF
|
MAS-related GPR, member F
|
|
chr19_-_7459867
|
1.674
|
NM_080662
|
PEX11G
|
peroxisomal biogenesis factor 11 gamma
|
|
chr12_+_52665196
|
1.661
|
NM_017409
|
HOXC10
|
homeobox C10
|
|
chr4_+_658250
|
1.632
|
|
MYL5
|
myosin, light chain 5, regulatory
|
|
chr19_-_55671814
|
1.632
|
NM_138411
|
FAM71E1
|
family with sequence similarity 71, member E1
|
|
chr7_+_5598964
|
1.621
|
|
FSCN1
|
fascin homolog 1, actin-bundling protein (Strongylocentrotus purpuratus)
|
|
chr2_-_237987509
|
1.616
|
|
COL6A3
|
collagen, type VI, alpha 3
|
|
chr6_+_135860631
|
1.586
|
|
NCRNA00271
|
non-protein coding RNA 271
|
|
chr19_-_50779861
|
1.583
|
|
OPA3
|
optic atrophy 3 (autosomal recessive, with chorea and spastic paraplegia)
|
|
chr3_+_50272648
|
1.569
|
|
|
|
|
chr2_+_28469275
|
1.565
|
NM_005253
|
FOSL2
|
FOS-like antigen 2
|
|
chr19_-_50779924
|
1.548
|
NM_001017989
NM_025136
|
OPA3
|
optic atrophy 3 (autosomal recessive, with chorea and spastic paraplegia)
|
|
chr7_+_5598979
|
1.544
|
NM_003088
|
FSCN1
|
fascin homolog 1, actin-bundling protein (Strongylocentrotus purpuratus)
|
|
chr2_-_237987554
|
1.526
|
|
COL6A3
|
collagen, type VI, alpha 3
|
|
chr18_+_52965290
|
1.525
|
|
BOD1P
|
biorientation of chromosomes in cell division 1 pseudogene
|
|
chr12_+_131194962
|
1.523
|
|
|
|
|
chr19_-_55671273
|
1.511
|
|
FAM71E1
|
family with sequence similarity 71, member E1
|
|
chr2_-_237987579
|
1.509
|
NM_004369
NM_057164
NM_057165
NM_057166
NM_057167
|
COL6A3
|
collagen, type VI, alpha 3
|
|
chr16_+_66252296
|
1.495
|
NM_001037281
NM_016948
|
PARD6A
|
par-6 partitioning defective 6 homolog alpha (C. elegans)
|
|
chr3_+_158026769
|
1.486
|
NM_001004316
NM_001193283
|
LEKR1
|
leucine, glutamate and lysine rich 1
|
|
chr3_+_11242666
|
1.459
|
NM_001098211
|
HRH1
|
histamine receptor H1
|
|
chr11_-_2149591
|
1.453
|
NM_000360
NM_199292
NM_199293
|
TH
|
tyrosine hydroxylase
|
|
chr8_-_95343716
|
1.439
|
NM_005261
NM_181702
|
GEM
|
GTP binding protein overexpressed in skeletal muscle
|
|
chr8_-_95343696
|
1.437
|
|
GEM
|
GTP binding protein overexpressed in skeletal muscle
|
|
chr7_+_44612705
|
1.430
|
|
OGDH
|
oxoglutarate (alpha-ketoglutarate) dehydrogenase (lipoamide)
|
|
chr6_-_56515640
|
1.418
|
|
|
|
|
chr7_+_65596078
|
1.380
|
|
|
|
|
chr1_-_1283770
|
1.377
|
NM_032348
|
MXRA8
|
matrix-remodelling associated 8
|
|
chr14_-_49399317
|
1.377
|
|
RN7SL2
|
RNA, 7SL, cytoplasmic 2
|
|
chr2_-_26554413
|
1.356
|
NM_004802
NM_194322
NM_194323
|
OTOF
|
otoferlin
|
|
chr19_-_45611110
|
1.343
|
NM_020956
NM_181882
|
PRX
|
periaxin
|
|
chr3_-_64186151
|
1.337
|
NM_198859
|
PRICKLE2
|
prickle homolog 2 (Drosophila)
|
|
chr4_-_111777584
|
1.336
|
|
PITX2
|
paired-like homeodomain 2
|
|
chrX_+_153325684
|
1.334
|
|
FAM50A
|
family with sequence similarity 50, member A
|
|
chr11_+_18676913
|
1.333
|
NM_153347
|
TMEM86A
|
transmembrane protein 86A
|
|
chr14_+_100362236
|
1.311
|
|
MEG3
|
maternally expressed 3 (non-protein coding)
|
|
chr1_-_16355013
|
1.295
|
NM_004431
|
EPHA2
|
EPH receptor A2
|
|
chr4_-_186693574
|
1.288
|
NM_001114107
NM_014476
|
PDLIM3
|
PDZ and LIM domain 3
|
|
chr5_+_174084137
|
1.287
|
NM_002449
|
MSX2
|
msh homeobox 2
|
|
chr19_-_5291700
|
1.285
|
NM_002850
NM_130853
NM_130854
NM_130855
|
PTPRS
|
protein tyrosine phosphatase, receptor type, S
|
|
chr2_-_224175339
|
1.279
|
|
SCG2
|
secretogranin II
|
|
chr12_-_13147839
|
1.274
|
NM_001080554
NM_001080555
|
GSG1
|
germ cell associated 1
|
|
chr9_-_139473569
|
1.257
|
|
NELF
|
nasal embryonic LHRH factor
|
|
chr2_-_224175293
|
1.257
|
|
SCG2
|
secretogranin II
|
|
chr10_-_128199986
|
1.249
|
NM_001004298
|
C10orf90
|
chromosome 10 open reading frame 90
|
|
chr1_+_99502655
|
1.240
|
|
LPPR4
|
lipid phosphate phosphatase-related protein type 4
|
|
chr2_+_219991342
|
1.235
|
NM_001927
|
DES
|
desmin
|
|
chr9_+_139152595
|
1.224
|
|
GRIN1
|
glutamate receptor, ionotropic, N-methyl D-aspartate 1
|
|
chr4_+_145786622
|
1.217
|
NM_022475
|
HHIP
|
hedgehog interacting protein
|
|
chr11_-_66367279
|
1.215
|
|
|
|
|
chr3_+_8518539
|
1.214
|
|
LMCD1
|
LIM and cysteine-rich domains 1
|
|
chr6_+_35335278
|
1.207
|
|
ZNF76
|
zinc finger protein 76
|
|
chr19_+_1892157
|
1.207
|
NM_001319
|
CSNK1G2
|
casein kinase 1, gamma 2
|
|
chr11_+_18676857
|
1.197
|
|
TMEM86A
|
transmembrane protein 86A
|
|
chr21_-_43903755
|
1.192
|
NM_007031
|
HSF2BP
|
heat shock transcription factor 2 binding protein
|
|
chr6_-_91063181
|
1.190
|
NM_021813
|
BACH2
|
BTB and CNC homology 1, basic leucine zipper transcription factor 2
|
|
chr19_+_1892184
|
1.189
|
|
CSNK1G2
|
casein kinase 1, gamma 2
|
|
chr22_+_40286957
|
1.189
|
NM_014460
|
CSDC2
|
cold shock domain containing C2, RNA binding
|
|
chr12_-_6103945
|
1.171
|
NM_000552
|
VWF
|
von Willebrand factor
|
|
chr2_-_224175386
|
1.167
|
NM_003469
|
SCG2
|
secretogranin II
|
|
chr6_+_117693413
|
1.166
|
NM_153453
NM_182645
|
VGLL2
|
vestigial like 2 (Drosophila)
|
|
chr16_+_85101734
|
1.165
|
|
FOXF1
|
forkhead box F1
|
|
chr11_-_62146144
|
1.164
|
|
B3GAT3
|
beta-1,3-glucuronyltransferase 3 (glucuronosyltransferase I)
|
|
chr19_+_50041387
|
1.163
|
|
PVRL2
|
poliovirus receptor-related 2 (herpesvirus entry mediator B)
|
|
chr1_+_99502435
|
1.163
|
NM_001166252
NM_014839
|
LPPR4
|
lipid phosphate phosphatase-related protein type 4
|
|
chr1_-_26571812
|
1.158
|
NM_001114759
NM_173574
|
ZNF683
|
zinc finger protein 683
|
|
chr9_-_98421894
|
1.153
|
NM_003671
NM_033331
|
CDC14B
|
CDC14 cell division cycle 14 homolog B (S. cerevisiae)
|
|
chr19_+_50041400
|
1.145
|
|
PVRL2
|
poliovirus receptor-related 2 (herpesvirus entry mediator B)
|
|
chr9_+_34980266
|
1.135
|
NM_001135005
|
DNAJB5
|
DnaJ (Hsp40) homolog, subfamily B, member 5
|
|
chr12_+_12935619
|
1.127
|
|
GPRC5A
|
G protein-coupled receptor, family C, group 5, member A
|
|
chr3_-_188946168
|
1.122
|
NM_001706
|
BCL6
|
B-cell CLL/lymphoma 6
|
|
chr1_+_1207351
|
1.116
|
NM_001130413
NM_002978
|
SCNN1D
|
sodium channel, nonvoltage-gated 1, delta
|
|
chr9_+_27099138
|
1.116
|
NM_000459
|
TEK
|
TEK tyrosine kinase, endothelial
|
|
chr8_-_144694722
|
1.116
|
|
ZC3H3
|
zinc finger CCCH-type containing 3
|
|
chr22_+_48870434
|
1.111
|
NM_001164104
NM_018995
|
MOV10L1
|
Mov10l1, Moloney leukemia virus 10-like 1, homolog (mouse)
|
|
chr11_-_35503720
|
1.098
|
NM_001001991
NM_015430
|
PAMR1
|
peptidase domain containing associated with muscle regeneration 1
|
|
chr21_+_46369606
|
1.098
|
|
COL6A2
|
collagen, type VI, alpha 2
|
|
chr6_-_127822227
|
1.096
|
NM_014702
|
KIAA0408
|
KIAA0408
|
|
chr10_+_99069007
|
1.090
|
NM_005479
|
FRAT1
|
frequently rearranged in advanced T-cell lymphomas
|
|
chr16_+_415630
|
1.089
|
NM_014700
|
RAB11FIP3
|
RAB11 family interacting protein 3 (class II)
|
|
chr2_-_237987545
|
1.077
|
|
COL6A3
|
collagen, type VI, alpha 3
|
|
chr8_-_26780665
|
1.068
|
|
ADRA1A
|
adrenergic, alpha-1A-, receptor
|
|
chr4_-_111777651
|
1.062
|
|
PITX2
|
paired-like homeodomain 2
|
|
chr2_+_28469199
|
1.061
|
|
FOSL2
|
FOS-like antigen 2
|
|
chr2_-_237987553
|
1.060
|
|
COL6A3
|
collagen, type VI, alpha 3
|
|
chr9_-_139473433
|
1.059
|
|
NELF
|
nasal embryonic LHRH factor
|
|
chr9_+_138341752
|
1.058
|
NM_001145638
NM_015597
|
GPSM1
|
G-protein signaling modulator 1
|
|
chr14_+_75521806
|
1.056
|
NM_001102564
NM_052873
|
C14orf179
|
chromosome 14 open reading frame 179
|
|
chr7_+_28415495
|
1.054
|
|
LOC401317
|
hypothetical LOC401317
|
|
chr3_-_8518327
|
1.048
|
|
LOC100288428
|
hypothetical LOC100288428
|
|
chr6_-_91063301
|
1.041
|
NM_001170794
|
BACH2
|
BTB and CNC homology 1, basic leucine zipper transcription factor 2
|
|
chr21_+_46369911
|
1.040
|
|
COL6A2
|
collagen, type VI, alpha 2
|
|
chrX_-_83644054
|
1.040
|
NM_001177478
NM_001177479
NM_144657
|
HDX
|
highly divergent homeobox
|
|
chr19_+_40811774
|
1.033
|
|
RBM42
|
RNA binding motif protein 42
|
|
chr18_+_75540786
|
1.030
|
NM_004715
NM_048368
|
CTDP1
|
CTD (carboxy-terminal domain, RNA polymerase II, polypeptide A) phosphatase, subunit 1
|
|
chr16_+_415606
|
1.029
|
|
RAB11FIP3
|
RAB11 family interacting protein 3 (class II)
|
|
chr4_+_658312
|
1.026
|
|
MYL5
|
myosin, light chain 5, regulatory
|
|
chr5_+_99898922
|
1.026
|
NM_198507
|
FAM174A
|
family with sequence similarity 174, member A
|
|
chr19_+_40716153
|
1.026
|
NM_014364
|
GAPDHS
|
glyceraldehyde-3-phosphate dehydrogenase, spermatogenic
|
|
chr18_+_75540831
|
1.026
|
|
CTDP1
|
CTD (carboxy-terminal domain, RNA polymerase II, polypeptide A) phosphatase, subunit 1
|
|
chr17_+_30472710
|
1.025
|
NM_017559
|
FNDC8
|
fibronectin type III domain containing 8
|
|
chrX_+_153325643
|
1.019
|
NM_004699
|
FAM50A
|
family with sequence similarity 50, member A
|
|
chr5_-_95794416
|
1.018
|
NM_000439
|
PCSK1
|
proprotein convertase subtilisin/kexin type 1
|
|
chr17_+_4590067
|
0.998
|
NM_001136046
NM_032265
|
ZMYND15
|
zinc finger, MYND-type containing 15
|
|
chr19_-_52308848
|
0.993
|
NM_015168
|
ZC3H4
|
zinc finger CCCH-type containing 4
|
|
chr9_-_34387786
|
0.985
|
NM_032596
|
C9orf24
|
chromosome 9 open reading frame 24
|
|
chr11_+_124539814
|
0.983
|
|
PKNOX2
|
PBX/knotted 1 homeobox 2
|
|
chr2_-_96899396
|
0.980
|
NM_017789
|
SEMA4C
|
sema domain, immunoglobulin domain (Ig), transmembrane domain (TM) and short cytoplasmic domain, (semaphorin) 4C
|
|
chr22_+_17539198
|
0.980
|
|
|
|
|
chr10_-_105208634
|
0.980
|
NM_001001412
|
CALHM1
|
calcium homeostasis modulator 1
|
|
chr17_-_76064842
|
0.970
|
NM_002522
|
NPTX1
|
neuronal pentraxin I
|
|
chr9_-_128924819
|
0.968
|
NM_012098
|
ANGPTL2
|
angiopoietin-like 2
|
|
chr19_+_53814359
|
0.957
|
NM_020126
|
SPHK2
|
sphingosine kinase 2
|
|
chr22_+_36927884
|
0.954
|
NM_001161572
NM_001161574
NM_012323
|
MAFF
|
v-maf musculoaponeurotic fibrosarcoma oncogene homolog F (avian)
|
|
chr1_-_226670928
|
0.951
|
|
TRIM17
|
tripartite motif containing 17
|
|
chr2_-_241408296
|
0.951
|
NM_004321
|
KIF1A
|
kinesin family member 1A
|
|
chr3_-_48489474
|
0.947
|
|
SHISA5
|
shisa homolog 5 (Xenopus laevis)
|
|
chr3_+_129690735
|
0.947
|
|
|
|
|
chr1_+_29435534
|
0.942
|
NM_001195001
NM_005704
NM_133177
NM_133178
|
PTPRU
|
protein tyrosine phosphatase, receptor type, U
|
|
chr8_-_98359347
|
0.937
|
NM_033512
|
TSPYL5
|
TSPY-like 5
|
|
chr9_-_19023183
|
0.936
|
NM_153707
|
FAM154A
|
family with sequence similarity 154, member A
|
|
chr19_-_48791967
|
0.935
|
NM_001007561
|
IRGQ
|
immunity-related GTPase family, Q
|
|
chr11_-_118740069
|
0.935
|
NM_171997
|
USP2
|
ubiquitin specific peptidase 2
|
|
chr14_+_95792299
|
0.932
|
NM_000710
|
BDKRB1
|
bradykinin receptor B1
|
|
chr10_-_31360778
|
0.931
|
NM_001143766
NM_001143767
NM_001143768
NM_001143770
NM_001143771
NM_182755
|
ZNF438
|
zinc finger protein 438
|
|
chr16_-_78192111
|
0.931
|
NM_001031804
NM_005360
|
MAF
|
v-maf musculoaponeurotic fibrosarcoma oncogene homolog (avian)
|
|
chr11_-_65550425
|
0.930
|
NM_053054
|
CATSPER1
|
cation channel, sperm associated 1
|
|
chr19_+_50041424
|
0.925
|
|
PVRL2
|
poliovirus receptor-related 2 (herpesvirus entry mediator B)
|
|
chr22_+_36928011
|
0.924
|
|
MAFF
|
v-maf musculoaponeurotic fibrosarcoma oncogene homolog F (avian)
|
|
chr1_+_156229686
|
0.923
|
NM_018240
|
KIRREL
|
kin of IRRE like (Drosophila)
|
|
chr9_+_36159384
|
0.922
|
NM_005893
|
CCIN
|
calicin
|
|
chr11_+_65945050
|
0.919
|
NM_178864
|
NPAS4
|
neuronal PAS domain protein 4
|
|
chr11_+_110916629
|
0.919
|
|
LAYN
|
layilin
|
|
chr20_-_43853945
|
0.917
|
NM_080614
|
WFDC3
|
WAP four-disulfide core domain 3
|
|
chr2_+_5750229
|
0.917
|
NM_003108
|
SOX11
|
SRY (sex determining region Y)-box 11
|
|
chr4_+_119732338
|
0.914
|
|
LOC729218
|
hypothetical LOC729218
|
|
chr7_-_41709191
|
0.911
|
NM_002192
|
INHBA
|
inhibin, beta A
|
|
chr20_+_33667160
|
0.910
|
NM_003116
|
SPAG4
|
sperm associated antigen 4
|
|
chr3_+_141136896
|
0.904
|
|
CLSTN2
|
calsyntenin 2
|
|
chr17_+_32368611
|
0.898
|
NM_005568
|
LHX1
|
LIM homeobox 1
|
|
chr9_+_129518163
|
0.896
|
NM_144965
|
TTC16
|
tetratricopeptide repeat domain 16
|
|
chr9_+_34979674
|
0.893
|
NM_001135004
NM_012266
|
DNAJB5
|
DnaJ (Hsp40) homolog, subfamily B, member 5
|
|
chr1_+_165330710
|
0.889
|
NM_001080426
|
DUSP27
|
dual specificity phosphatase 27 (putative)
|
|
chr2_+_241023703
|
0.885
|
NM_002081
|
GPC1
|
glypican 1
|
|
chr8_-_29263714
|
0.884
|
NM_001394
|
DUSP4
|
dual specificity phosphatase 4
|
|
chr9_-_129517740
|
0.883
|
NM_001002913
|
PTRH1
|
peptidyl-tRNA hydrolase 1 homolog (S. cerevisiae)
|
|
chr15_+_79080341
|
0.881
|
NM_022566
|
MESDC1
|
mesoderm development candidate 1
|
|
chr10_-_90957028
|
0.881
|
NM_003956
|
CH25H
|
cholesterol 25-hydroxylase
|
|
chr8_-_98359248
|
0.880
|
|
TSPYL5
|
TSPY-like 5
|
|
chr19_-_50779914
|
0.879
|
|
OPA3
|
optic atrophy 3 (autosomal recessive, with chorea and spastic paraplegia)
|
|
chr12_-_51734434
|
0.878
|
|
LOC283335
|
hypothetical LOC283335
|
|
chr11_-_62145952
|
0.875
|
NM_012200
|
B3GAT3
|
beta-1,3-glucuronyltransferase 3 (glucuronosyltransferase I)
|
|
chr4_-_171161486
|
0.873
|
NM_001009554
|
MFAP3L
|
microfibrillar-associated protein 3-like
|
|
chr19_-_13978096
|
0.873
|
NM_002918
|
RFX1
|
regulatory factor X, 1 (influences HLA class II expression)
|
|
chr2_-_27967458
|
0.871
|
|
LOC100302650
|
hypothetical LOC100302650
|
|
chr2_-_96899326
|
0.870
|
|
SEMA4C
|
sema domain, immunoglobulin domain (Ig), transmembrane domain (TM) and short cytoplasmic domain, (semaphorin) 4C
|
|
chr3_-_188945921
|
0.867
|
|
BCL6
|
B-cell CLL/lymphoma 6
|
|
chr1_-_160613203
|
0.866
|
NM_182581
|
C1orf111
|
chromosome 1 open reading frame 111
|
|
chr17_-_40923864
|
0.866
|
|
PLEKHM1
|
pleckstrin homology domain containing, family M (with RUN domain) member 1
|
|
chr7_+_44612745
|
0.866
|
|
OGDH
|
oxoglutarate (alpha-ketoglutarate) dehydrogenase (lipoamide)
|
|
chr3_-_10042989
|
0.865
|
|
CIDECP
|
cell death-inducing DFFA-like effector c pseudogene
|
|
chr5_-_99898788
|
0.862
|
|
|
|
|
chr20_+_55718619
|
0.862
|
|
|
|
|
chr20_-_33489383
|
0.862
|
NM_000557
|
GDF5
|
growth differentiation factor 5
|
|
chr10_+_102782151
|
0.861
|
|
SFXN3
|
sideroflexin 3
|
|
chr14_+_60517682
|
0.860
|
|
SLC38A6
|
solute carrier family 38, member 6
|
|
chr17_+_56884560
|
0.858
|
|
TBX4
|
T-box 4
|
|
chr19_+_10388448
|
0.854
|
|
PDE4A
|
phosphodiesterase 4A, cAMP-specific
|