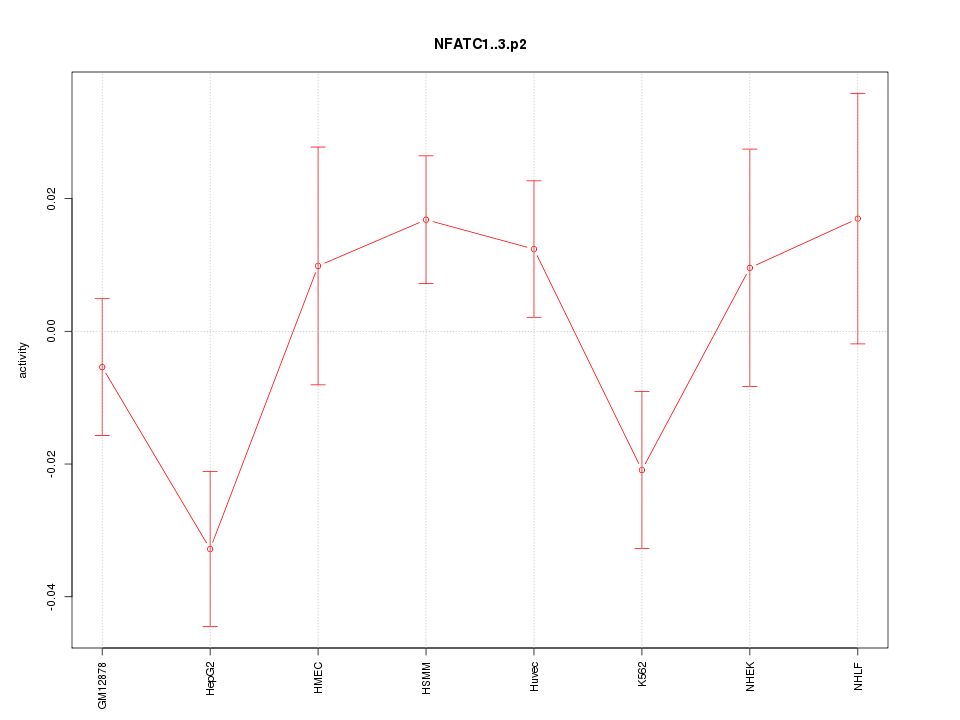
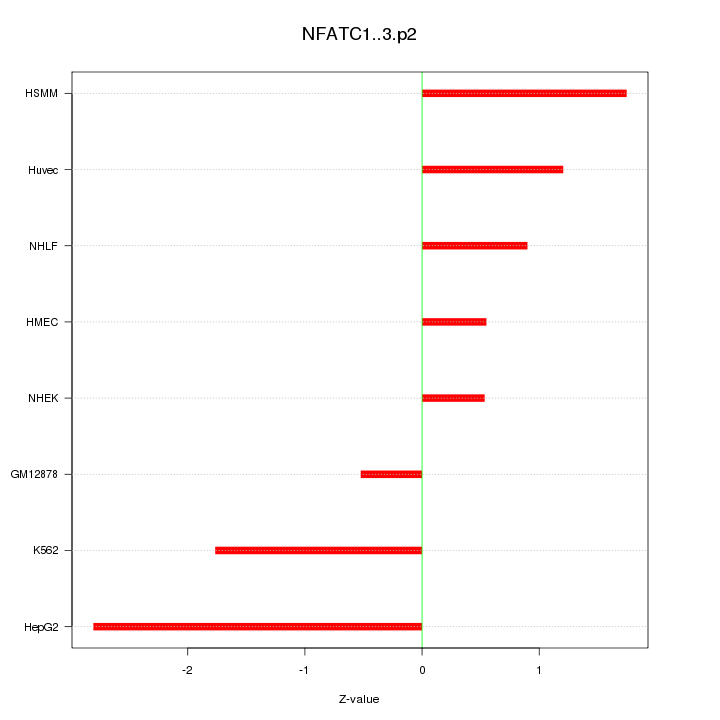
|
chr12_+_13240987
|
3.871
|
|
EMP1
|
epithelial membrane protein 1
|
|
chr12_+_13240868
|
3.506
|
NM_001423
|
EMP1
|
epithelial membrane protein 1
|
|
chr2_-_237987579
|
2.032
|
NM_004369
NM_057164
NM_057165
NM_057166
NM_057167
|
COL6A3
|
collagen, type VI, alpha 3
|
|
chr3_+_120799411
|
1.899
|
NM_015900
|
PLA1A
|
phospholipase A1 member A
|
|
chr1_+_157246305
|
1.888
|
NM_005531
|
IFI16
|
interferon, gamma-inducible protein 16
|
|
chr2_-_237987554
|
1.871
|
|
COL6A3
|
collagen, type VI, alpha 3
|
|
chr2_-_237987545
|
1.812
|
|
COL6A3
|
collagen, type VI, alpha 3
|
|
chr2_-_237987509
|
1.789
|
|
COL6A3
|
collagen, type VI, alpha 3
|
|
chr2_+_189547377
|
1.730
|
|
COL3A1
|
collagen, type III, alpha 1
|
|
chr2_+_189547358
|
1.649
|
|
COL3A1
|
collagen, type III, alpha 1
|
|
chr2_+_189547323
|
1.593
|
NM_000090
|
COL3A1
|
collagen, type III, alpha 1
|
|
chr9_+_74956494
|
1.548
|
NM_000700
|
ANXA1
|
annexin A1
|
|
chr4_+_183482130
|
1.418
|
NM_001080477
|
ODZ3
|
odz, odd Oz/ten-m homolog 3 (Drosophila)
|
|
chr1_+_100957883
|
1.394
|
NM_001078
NM_080682
|
VCAM1
|
vascular cell adhesion molecule 1
|
|
chr15_+_37330176
|
1.332
|
NM_207445
|
C15orf54
|
chromosome 15 open reading frame 54
|
|
chr4_-_149582948
|
1.319
|
NM_000901
NM_001166104
|
NR3C2
|
nuclear receptor subfamily 3, group C, member 2
|
|
chr2_-_237987553
|
1.254
|
|
COL6A3
|
collagen, type VI, alpha 3
|
|
chr6_+_106653429
|
1.199
|
NM_182907
|
PRDM1
|
PR domain containing 1, with ZNF domain
|
|
chr12_+_99274987
|
1.195
|
NM_001145288
NM_139319
|
SLC17A8
|
solute carrier family 17 (sodium-dependent inorganic phosphate cotransporter), member 8
|
|
chr4_-_149582754
|
1.189
|
|
NR3C2
|
nuclear receptor subfamily 3, group C, member 2
|
|
chr1_+_161305558
|
1.168
|
NM_001113381
|
RGS4
|
regulator of G-protein signaling 4
|
|
chr10_-_31328426
|
1.167
|
NM_001143769
|
ZNF438
|
zinc finger protein 438
|
|
chr11_-_85115105
|
1.144
|
NM_206927
NM_206928
|
SYTL2
|
synaptotagmin-like 2
|
|
chr4_-_100575313
|
1.137
|
NM_001166504
NM_000673
|
ADH7
|
alcohol dehydrogenase 7 (class IV), mu or sigma polypeptide
|
|
chr4_-_115120251
|
1.127
|
NM_024590
|
ARSJ
|
arylsulfatase family, member J
|
|
chr6_-_169395847
|
1.118
|
|
THBS2
|
thrombospondin 2
|
|
chr7_+_115952074
|
1.116
|
NM_001753
|
CAV1
|
caveolin 1, caveolae protein, 22kDa
|
|
chr6_-_169395972
|
1.105
|
NM_003247
|
THBS2
|
thrombospondin 2
|
|
chr4_+_169654742
|
1.082
|
NM_001166108
NM_016081
|
PALLD
|
palladin, cytoskeletal associated protein
|
|
chr3_+_161040439
|
1.080
|
|
SCHIP1
|
schwannomin interacting protein 1
|
|
chr2_-_37727472
|
1.051
|
|
CDC42EP3
|
CDC42 effector protein (Rho GTPase binding) 3
|
|
chr3_-_150533948
|
1.036
|
NM_001184723
NM_138786
|
TM4SF18
|
transmembrane 4 L six family member 18
|
|
chr18_+_64616296
|
1.033
|
NM_024781
|
CCDC102B
|
coiled-coil domain containing 102B
|
|
chrX_-_10504852
|
1.032
|
NM_001193277
|
MID1
|
midline 1 (Opitz/BBB syndrome)
|
|
chr2_-_188021117
|
1.031
|
NM_005795
|
CALCRL
|
calcitonin receptor-like
|
|
chr18_-_51219880
|
1.030
|
|
TCF4
|
transcription factor 4
|
|
chr12_+_13240983
|
1.022
|
|
EMP1
|
epithelial membrane protein 1
|
|
chr4_-_87989291
|
1.012
|
NM_197965
|
SLC10A6
|
solute carrier family 10 (sodium/bile acid cotransporter family), member 6
|
|
chr3_+_114733907
|
1.000
|
NM_017699
|
SIDT1
|
SID1 transmembrane family, member 1
|
|
chr3_-_46693902
|
0.994
|
NM_182775
|
ALS2CL
|
ALS2 C-terminal like
|
|
chr1_-_68470585
|
0.983
|
NM_001002292
NM_001193334
NM_024911
|
WLS
|
wntless homolog (Drosophila)
|
|
chr4_+_109033868
|
0.980
|
NM_001136257
NM_152621
|
SGMS2
|
sphingomyelin synthase 2
|
|
chr2_+_234245243
|
0.963
|
NM_021027
|
UGT1A9
|
UDP glucuronosyltransferase 1 family, polypeptide A9
|
|
chr15_+_67393795
|
0.952
|
NM_001104554
|
PAQR5
|
progestin and adipoQ receptor family member V
|
|
chr7_+_18501876
|
0.949
|
NM_014707
NM_178423
|
HDAC9
|
histone deacetylase 9
|
|
chr4_+_88790477
|
0.942
|
NM_001079911
NM_004407
|
DMP1
|
dentin matrix acidic phosphoprotein 1
|
|
chr21_-_26867138
|
0.913
|
NM_052954
|
CYYR1
|
cysteine/tyrosine-rich 1
|
|
chr21_-_27261276
|
0.912
|
NM_007038
|
ADAMTS5
|
ADAM metallopeptidase with thrombospondin type 1 motif, 5
|
|
chr1_-_109201228
|
0.907
|
NM_152763
|
AKNAD1
|
AKNA domain containing 1
|
|
chr19_+_46417117
|
0.893
|
|
AXL
|
AXL receptor tyrosine kinase
|
|
chr5_+_140848905
|
0.885
|
NM_018929
NM_032407
|
PCDHGC5
|
protocadherin gamma subfamily C, 5
|
|
chr12_+_118100977
|
0.881
|
NM_014365
|
HSPB8
|
heat shock 22kDa protein 8
|
|
chr13_+_109932934
|
0.880
|
|
COL4A2
|
collagen, type IV, alpha 2
|
|
chr15_+_69220841
|
0.880
|
NM_024817
|
THSD4
|
thrombospondin, type I, domain containing 4
|
|
chr14_-_60818263
|
0.873
|
NM_001017970
|
TMEM30B
|
transmembrane protein 30B
|
|
chr1_+_169421011
|
0.865
|
NM_001460
|
FMO2
|
flavin containing monooxygenase 2 (non-functional)
|
|
chr4_-_177427252
|
0.864
|
NM_080874
|
ASB5
|
ankyrin repeat and SOCS box containing 5
|
|
chr15_+_71763674
|
0.862
|
NM_001024736
NM_025240
|
CD276
|
CD276 molecule
|
|
chr7_+_18502342
|
0.855
|
NM_058176
NM_178425
|
HDAC9
|
histone deacetylase 9
|
|
chr17_-_40339919
|
0.840
|
|
GFAP
|
glial fibrillary acidic protein
|
|
chr17_+_45617676
|
0.826
|
|
|
|
|
chr2_-_189363075
|
0.814
|
NM_052952
|
DIRC1
|
disrupted in renal carcinoma 1
|
|
chrX_-_131989963
|
0.811
|
NM_031907
|
USP26
|
ubiquitin specific peptidase 26
|
|
chr5_+_176494690
|
0.811
|
|
NSD1
|
nuclear receptor binding SET domain protein 1
|
|
chr5_-_54317102
|
0.810
|
NM_001135604
NM_007036
|
ESM1
|
endothelial cell-specific molecule 1
|
|
chr5_+_32824701
|
0.796
|
NM_024563
|
NPR3
|
natriuretic peptide receptor C/guanylate cyclase C (atrionatriuretic peptide receptor C)
|
|
chr8_+_32698892
|
0.796
|
NM_001159996
|
NRG1
|
neuregulin 1
|
|
chr7_+_28418668
|
0.794
|
NM_182898
|
CREB5
|
cAMP responsive element binding protein 5
|
|
chr1_-_79244948
|
0.792
|
NM_022159
|
ELTD1
|
EGF, latrophilin and seven transmembrane domain containing 1
|
|
chr16_-_80602546
|
0.791
|
NM_145168
|
SDR42E1
|
short chain dehydrogenase/reductase family 42E, member 1
|
|
chr7_-_131483925
|
0.790
|
|
PLXNA4
|
plexin A4
|
|
chr8_+_39561243
|
0.788
|
NM_001190956
NM_014237
|
ADAM18
|
ADAM metallopeptidase domain 18
|
|
chr1_-_149047329
|
0.786
|
NM_000396
|
CTSK
|
cathepsin K
|
|
chr3_-_113842652
|
0.781
|
NM_199511
NM_199512
|
CCDC80
|
coiled-coil domain containing 80
|
|
chr21_+_38550739
|
0.780
|
NM_170736
|
KCNJ15
|
potassium inwardly-rectifying channel, subfamily J, member 15
|
|
chr3_-_184756101
|
0.778
|
NM_130446
|
KLHL6
|
kelch-like 6 (Drosophila)
|
|
chr2_-_227952249
|
0.776
|
NM_024795
|
TM4SF20
|
transmembrane 4 L six family member 20
|
|
chr12_-_51153788
|
0.750
|
NM_173086
|
KRT6C
|
keratin 6C
|
|
chrX_+_2680092
|
0.749
|
NM_001141919
NM_001141920
NM_175569
|
XGPY2
XG
|
Xg pseudogene, Y-linked 2
Xg blood group
|
|
chr11_+_57714481
|
0.749
|
NM_001005283
|
OR9Q2
|
olfactory receptor, family 9, subfamily Q, member 2
|
|
chr9_+_5500544
|
0.739
|
NM_025239
|
PDCD1LG2
|
programmed cell death 1 ligand 2
|
|
chrX_+_100692071
|
0.733
|
NM_016608
|
ARMCX1
|
armadillo repeat containing, X-linked 1
|
|
chr4_-_153820535
|
0.730
|
NM_152680
|
TMEM154
|
transmembrane protein 154
|
|
chr19_-_46883935
|
0.730
|
NM_006890
|
CEACAM7
|
carcinoembryonic antigen-related cell adhesion molecule 7
|
|
chr12_-_90097384
|
0.727
|
NM_133503
|
DCN
|
decorin
|
|
chr9_+_88953378
|
0.721
|
NM_001001709
|
C9orf170
|
chromosome 9 open reading frame 170
|
|
chr16_-_63713382
|
0.718
|
NM_001797
|
CDH11
|
cadherin 11, type 2, OB-cadherin (osteoblast)
|
|
chrX_+_56772391
|
0.717
|
|
LOC550643
|
hypothetical LOC550643
|
|
chr5_+_140154627
|
0.717
|
NM_018905
NM_031495
|
PCDHA2
|
protocadherin alpha 2
|
|
chr8_+_55691179
|
0.717
|
NM_006269
|
RP1
|
retinitis pigmentosa 1 (autosomal dominant)
|
|
chr8_+_70567581
|
0.713
|
NM_001128205
NM_001128206
|
SULF1
|
sulfatase 1
|
|
chr6_+_116939500
|
0.706
|
NM_153711
|
FAM26E
|
family with sequence similarity 26, member E
|
|
chr7_+_93388951
|
0.706
|
NM_004126
|
GNG11
|
guanine nucleotide binding protein (G protein), gamma 11
|
|
chr6_+_116798791
|
0.705
|
NM_013352
|
DSE
|
dermatan sulfate epimerase
|
|
chr7_-_41709191
|
0.702
|
NM_002192
|
INHBA
|
inhibin, beta A
|
|
chr3_-_113842553
|
0.702
|
|
CCDC80
|
coiled-coil domain containing 80
|
|
chr1_+_66772393
|
0.701
|
NM_032291
|
SGIP1
|
SH3-domain GRB2-like (endophilin) interacting protein 1
|
|
chr8_+_70567634
|
0.694
|
|
SULF1
|
sulfatase 1
|
|
chr1_-_171286634
|
0.691
|
NM_005092
|
TNFSF18
|
tumor necrosis factor (ligand) superfamily, member 18
|
|
chr12_+_48652886
|
0.684
|
NM_001652
|
AQP6
|
aquaporin 6, kidney specific
|
|
chr11_-_35503720
|
0.679
|
NM_001001991
NM_015430
|
PAMR1
|
peptidase domain containing associated with muscle regeneration 1
|
|
chr11_-_68537243
|
0.677
|
NM_001098515
NM_145015
|
MRGPRF
|
MAS-related GPR, member F
|
|
chr1_+_160868932
|
0.673
|
|
DDR2
|
discoidin domain receptor tyrosine kinase 2
|
|
chr5_+_140187746
|
0.671
|
NM_018909
NM_031848
NM_031849
|
PCDHA6
|
protocadherin alpha 6
|
|
chr11_-_127962662
|
0.668
|
NM_001143820
|
ETS1
|
v-ets erythroblastosis virus E26 oncogene homolog 1 (avian)
|
|
chr12_+_54611212
|
0.667
|
NM_001345
|
DGKA
|
diacylglycerol kinase, alpha 80kDa
|
|
chr8_-_15140162
|
0.666
|
NM_139167
|
SGCZ
|
sarcoglycan, zeta
|
|
chr1_+_160868822
|
0.665
|
NM_001014796
NM_006182
|
DDR2
|
discoidin domain receptor tyrosine kinase 2
|
|
chr20_+_9146036
|
0.660
|
NM_182797
|
PLCB4
|
phospholipase C, beta 4
|
|
chr18_+_27281729
|
0.656
|
NM_001944
|
DSG3
|
desmoglein 3
|
|
chr17_+_44642587
|
0.655
|
NM_001135186
NM_016428
|
ABI3
|
ABI family, member 3
|
|
chr1_-_207892420
|
0.652
|
NM_000228
|
LAMB3
|
laminin, beta 3
|
|
chr4_+_40953698
|
0.648
|
|
UCHL1
|
ubiquitin carboxyl-terminal esterase L1 (ubiquitin thiolesterase)
|
|
chr5_-_150928697
|
0.648
|
NM_001447
|
FAT2
|
FAT tumor suppressor homolog 2 (Drosophila)
|
|
chr16_-_63713466
|
0.646
|
|
CDH11
|
cadherin 11, type 2, OB-cadherin (osteoblast)
|
|
chr12_-_50871953
|
0.639
|
NM_001081492
NM_182507
|
KRT80
|
keratin 80
|
|
chr9_-_21192201
|
0.637
|
NM_021057
|
IFNA7
|
interferon, alpha 7
|
|
chr8_-_82606104
|
0.636
|
NM_001105281
|
FABP12
|
fatty acid binding protein 12
|
|
chr11_-_66898223
|
0.633
|
NM_001166212
|
CLCF1
|
cardiotrophin-like cytokine factor 1
|
|
chr11_-_18913121
|
0.632
|
NM_147199
|
MRGPRX1
|
MAS-related GPR, member X1
|
|
chr8_-_134141586
|
0.624
|
NM_006748
|
SLA
|
Src-like-adaptor
|
|
chr10_+_124310170
|
0.622
|
NM_004406
NM_007329
NM_017579
|
DMBT1
|
deleted in malignant brain tumors 1
|
|
chr5_-_88235624
|
0.622
|
NM_001131005
NM_001193347
|
MEF2C
|
myocyte enhancer factor 2C
|
|
chr3_-_57505107
|
0.619
|
NM_178504
NM_198564
|
DNAH12
|
dynein, axonemal, heavy chain 12
|
|
chr2_-_189752575
|
0.612
|
NM_000393
|
COL5A2
|
collagen, type V, alpha 2
|
|
chr3_+_161040343
|
0.610
|
NM_001197109
|
SCHIP1
|
schwannomin interacting protein 1
|
|
chr4_-_87593306
|
0.610
|
NM_002753
|
MAPK10
|
mitogen-activated protein kinase 10
|
|
chr4_-_77147625
|
0.605
|
NM_002416
|
CXCL9
|
chemokine (C-X-C motif) ligand 9
|
|
chr3_-_113334657
|
0.603
|
NM_001190259
NM_001190260
NM_152785
|
GCET2
|
germinal center expressed transcript 2
|
|
chr3_-_194118321
|
0.602
|
|
MB21D2
|
Mab-21 domain containing 2
|
|
chr1_-_224993498
|
0.596
|
NM_002221
|
ITPKB
|
inositol 1,4,5-trisphosphate 3-kinase B
|
|
chr16_-_63713485
|
0.593
|
|
CDH11
|
cadherin 11, type 2, OB-cadherin (osteoblast)
|
|
chr10_-_108448990
|
0.591
|
|
SORCS1
|
sortilin-related VPS10 domain containing receptor 1
|
|
chr12_-_10173947
|
0.591
|
NM_022570
NM_197947
NM_197948
NM_197949
NM_197950
NM_197954
|
CLEC7A
|
C-type lectin domain family 7, member A
|
|
chr20_+_19863716
|
0.587
|
|
RIN2
|
Ras and Rab interactor 2
|
|
chr5_+_140767953
|
0.584
|
NM_018926
NM_032100
|
PCDHGB6
|
protocadherin gamma subfamily B, 6
|
|
chr4_-_24590867
|
0.584
|
NM_173463
|
CCDC149
|
coiled-coil domain containing 149
|
|
chrX_-_62921937
|
0.580
|
NM_001173479
|
ARHGEF9
|
Cdc42 guanine nucleotide exchange factor (GEF) 9
|
|
chr10_+_105026773
|
0.577
|
NM_032727
|
INA
|
internexin neuronal intermediate filament protein, alpha
|
|
chr9_-_35679902
|
0.573
|
NM_003289
NM_213674
|
TPM2
|
tropomyosin 2 (beta)
|
|
chr6_-_112682459
|
0.572
|
NM_001105206
NM_001105207
NM_001105208
NM_001105209
NM_002290
|
LAMA4
|
laminin, alpha 4
|
|
chr1_+_246591505
|
0.570
|
NM_001004696
|
OR2T4
|
olfactory receptor, family 2, subfamily T, member 4
|
|
chr7_+_80145461
|
0.569
|
|
CD36
|
CD36 molecule (thrombospondin receptor)
|
|
chr1_-_167865973
|
0.568
|
NM_003005
|
SELP
|
selectin P (granule membrane protein 140kDa, antigen CD62)
|
|
chr6_+_152053323
|
0.566
|
NM_001122742
|
ESR1
|
estrogen receptor 1
|
|
chr17_-_44022595
|
0.566
|
|
HOXB3
|
homeobox B3
|
|
chr9_+_12764988
|
0.565
|
NM_203403
|
C9orf150
|
chromosome 9 open reading frame 150
|
|
chr9_+_22437353
|
0.562
|
|
DMRTA1
|
DMRT-like family A1
|
|
chr15_+_41672543
|
0.562
|
NM_020990
|
CKMT1B
CKMT1A
|
creatine kinase, mitochondrial 1B
creatine kinase, mitochondrial 1A
|
|
chr14_-_91268136
|
0.561
|
NM_024764
|
CATSPERB
|
cation channel, sperm-associated, beta
|
|
chr4_-_74704964
|
0.560
|
NM_177532
NM_201431
|
RASSF6
|
Ras association (RalGDS/AF-6) domain family member 6
|
|
chr1_-_94475840
|
0.558
|
NM_004815
|
ARHGAP29
|
Rho GTPase activating protein 29
|
|
chr6_+_39868770
|
0.557
|
NM_015345
|
DAAM2
|
dishevelled associated activator of morphogenesis 2
|
|
chr4_+_154083584
|
0.557
|
NM_033393
|
FHDC1
|
FH2 domain containing 1
|
|
chr17_-_30783631
|
0.557
|
NM_018042
|
SLFN12
|
schlafen family member 12
|
|
chr1_+_24518501
|
0.556
|
NM_001195010
|
GRHL3
|
grainyhead-like 3 (Drosophila)
|
|
chr12_+_118515646
|
0.556
|
NM_001136534
|
TMEM233
|
transmembrane protein 233
|
|
chr2_+_234333653
|
0.555
|
NM_000463
|
UGT1A1
|
UDP glucuronosyltransferase 1 family, polypeptide A1
|
|
chr11_+_6898780
|
0.554
|
NM_001004684
|
OR2D3
|
olfactory receptor, family 2, subfamily D, member 3
|
|
chr15_+_69626594
|
0.554
|
|
THSD4
|
thrombospondin, type I, domain containing 4
|
|
chr5_-_64813459
|
0.552
|
NM_197941
|
ADAMTS6
|
ADAM metallopeptidase with thrombospondin type 1 motif, 6
|
|
chr11_+_55776251
|
0.552
|
NM_001004747
|
OR5T3
|
olfactory receptor, family 5, subfamily T, member 3
|
|
chrX_+_102471769
|
0.551
|
NM_152278
|
TCEAL7
|
transcription elongation factor A (SII)-like 7
|
|
chr1_+_24518398
|
0.549
|
NM_198173
NM_198174
|
GRHL3
|
grainyhead-like 3 (Drosophila)
|
|
chr2_-_40510923
|
0.547
|
NM_001112800
NM_001112801
NM_021097
|
SLC8A1
|
solute carrier family 8 (sodium/calcium exchanger), member 1
|
|
chr4_+_47181987
|
0.546
|
NM_020453
|
ATP10D
|
ATPase, class V, type 10D
|
|
chr8_-_95343696
|
0.545
|
|
GEM
|
GTP binding protein overexpressed in skeletal muscle
|
|
chr12_+_27740694
|
0.542
|
NM_001029874
|
REP15
|
RAB15 effector protein
|
|
chr8_+_42671718
|
0.541
|
NM_000749
|
CHRNB3
|
cholinergic receptor, nicotinic, beta 3
|
|
chr6_+_106640903
|
0.541
|
|
PRDM1
|
PR domain containing 1, with ZNF domain
|
|
chr3_+_189425886
|
0.539
|
NM_001167671
|
LPP
|
LIM domain containing preferred translocation partner in lipoma
|
|
chr14_+_62740878
|
0.538
|
NM_020663
|
RHOJ
|
ras homolog gene family, member J
|
|
chr13_-_48873735
|
0.534
|
NM_030925
|
CAB39L
|
calcium binding protein 39-like
|
|
chr3_+_11269384
|
0.533
|
NM_000861
|
HRH1
|
histamine receptor H1
|
|
chr19_-_48133089
|
0.533
|
NM_002783
|
PSG7
|
pregnancy specific beta-1-glycoprotein 7 (gene/pseudogene)
|
|
chr11_+_120478584
|
0.531
|
NM_005422
|
TECTA
|
tectorin alpha
|
|
chr13_-_66702433
|
0.530
|
NM_020403
NM_203487
|
PCDH9
|
protocadherin 9
|
|
chr8_-_95343716
|
0.527
|
NM_005261
NM_181702
|
GEM
|
GTP binding protein overexpressed in skeletal muscle
|
|
chr8_+_76059262
|
0.525
|
NM_031461
|
CRISPLD1
|
cysteine-rich secretory protein LCCL domain containing 1
|
|
chr5_-_146813343
|
0.525
|
NM_001387
|
DPYSL3
|
dihydropyrimidinase-like 3
|
|
chr4_-_90978469
|
0.515
|
NM_001146055
|
SNCA
|
synuclein, alpha (non A4 component of amyloid precursor)
|
|
chr19_-_48465422
|
0.512
|
NM_002784
|
PSG9
|
pregnancy specific beta-1-glycoprotein 9
|
|
chr10_+_74323344
|
0.512
|
NM_152635
|
OIT3
|
oncoprotein induced transcript 3
|
|
chr4_+_15038726
|
0.509
|
NM_031911
|
C1QTNF7
|
C1q and tumor necrosis factor related protein 7
|
|
chr10_+_96688399
|
0.509
|
NM_000771
|
CYP2C9
|
cytochrome P450, family 2, subfamily C, polypeptide 9
|
|
chr8_-_18585721
|
0.509
|
|
PSD3
|
pleckstrin and Sec7 domain containing 3
|
|
chr18_+_57308754
|
0.509
|
NM_031891
|
CDH20
|
cadherin 20, type 2
|
|
chr7_+_28692122
|
0.506
|
NM_001011666
|
CREB5
|
cAMP responsive element binding protein 5
|
|
chr9_+_34979674
|
0.506
|
NM_001135004
NM_012266
|
DNAJB5
|
DnaJ (Hsp40) homolog, subfamily B, member 5
|
|
chr1_+_154816142
|
0.506
|
NM_001105669
|
TTC24
|
tetratricopeptide repeat domain 24
|
|
chr12_+_39372456
|
0.505
|
NM_001843
NM_175038
|
CNTN1
|
contactin 1
|
|
chr6_-_112682401
|
0.500
|
|
LAMA4
|
laminin, alpha 4
|
|
chr18_+_59571256
|
0.499
|
NM_001040147
|
SERPINB7
|
serpin peptidase inhibitor, clade B (ovalbumin), member 7
|
|
chr5_+_132037271
|
0.499
|
NM_000589
NM_172348
|
IL4
|
interleukin 4
|
|
chr18_-_68683913
|
0.499
|
NM_138999
NM_153181
|
NETO1
|
neuropilin (NRP) and tolloid (TLL)-like 1
|
|
chr11_-_64327197
|
0.497
|
NM_004579
|
MAP4K2
|
mitogen-activated protein kinase kinase kinase kinase 2
|
|
chr11_+_123491313
|
0.496
|
NM_001130142
NM_014622
NM_198315
|
VWA5A
|
von Willebrand factor A domain containing 5A
|
|
chr14_+_85066265
|
0.495
|
|
FLRT2
|
fibronectin leucine rich transmembrane protein 2
|
|
chr1_-_6502655
|
0.495
|
NM_198681
|
PLEKHG5
|
pleckstrin homology domain containing, family G (with RhoGef domain) member 5
|
|
chr1_+_78888064
|
0.494
|
NM_006417
|
IFI44
|
interferon-induced protein 44
|