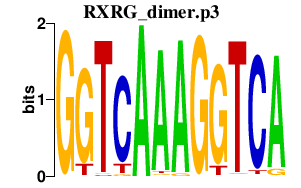
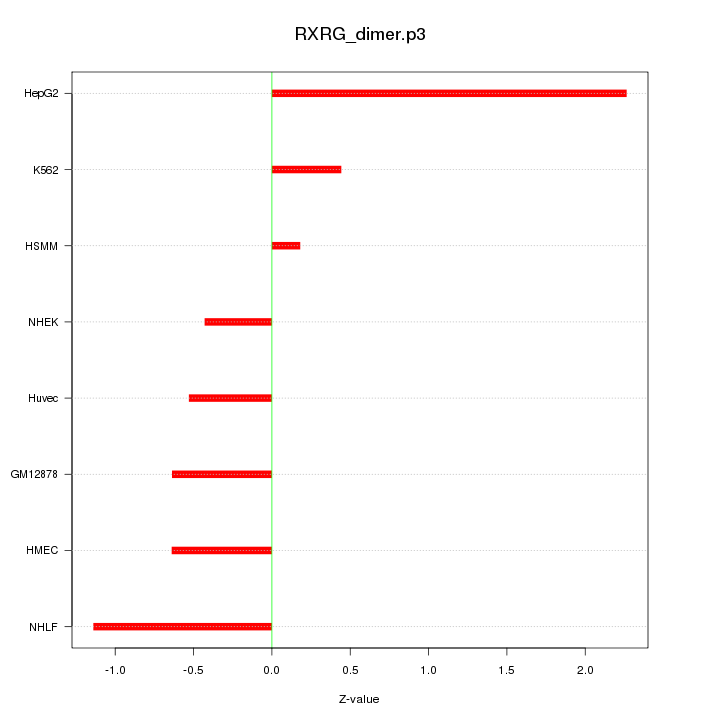
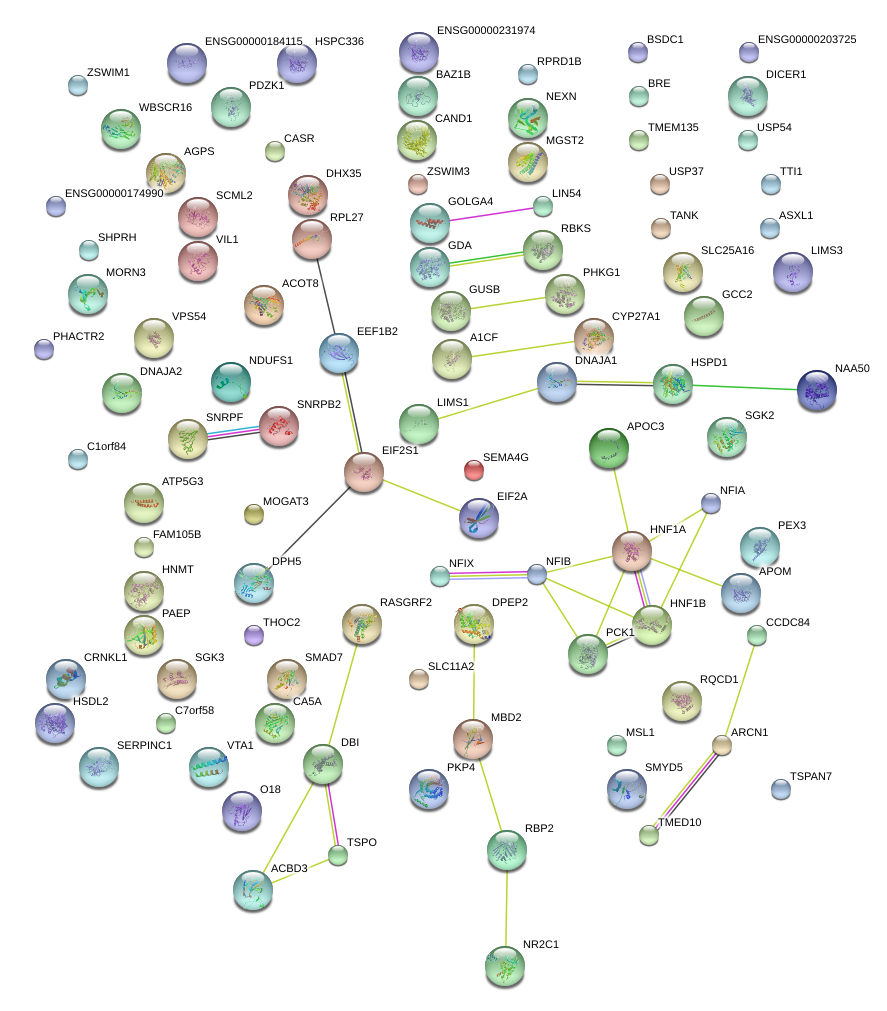
|
chr20_+_55569542
|
1.348
|
NM_002591
|
PCK1
|
phosphoenolpyruvate carboxykinase 1 (soluble)
|
|
chr2_+_177965734
|
1.255
|
|
AGPS
|
alkylglycerone phosphate synthase
|
|
chr2_-_177965626
|
1.104
|
|
LOC100130691
|
hypothetical LOC100130691
|
|
chr2_+_218992053
|
1.076
|
NM_007127
|
VIL1
|
villin 1
|
|
chr2_-_219141199
|
1.074
|
NM_020935
|
USP37
|
ubiquitin specific peptidase 37
|
|
chr2_-_64099935
|
1.032
|
|
VPS54
|
vacuolar protein sorting 54 homolog (S. cerevisiae)
|
|
chr6_+_30702586
|
1.006
|
NM_024909
NM_001031722
NM_001190724
|
ATAT1
|
alpha tubulin acetyltransferase 1
|
|
chr2_+_161725172
|
1.002
|
|
TANK
|
TRAF family member-associated NFKB activator
|
|
chr10_+_102722275
|
0.962
|
NM_017893
|
SEMA4G
|
sema domain, immunoglobulin domain (Ig), transmembrane domain (TM) and short cytoplasmic domain, (semaphorin) 4G
|
|
chr10_-_52315390
|
0.936
|
|
A1CF
|
APOBEC1 complementation factor
|
|
chr16_-_86527608
|
0.925
|
NM_001739
|
CA5A
|
carbonic anhydrase VA, mitochondrial
|
|
chr6_+_144040751
|
0.909
|
NM_001100164
NM_001100165
|
PHACTR2
|
phosphatase and actin regulator 2
|
|
chr1_+_61320213
|
0.909
|
NM_001134673
NM_005595
|
NFIA
|
nuclear factor I/A
|
|
chr11_+_116205830
|
0.906
|
NM_000040
|
APOC3
|
apolipoprotein C-III
|
|
chr2_+_206732859
|
0.904
|
|
EEF1B2
|
eukaryotic translation elongation factor 1 beta 2
|
|
chr3_+_37259684
|
0.902
|
NM_001172713
NM_002078
|
GOLGA4
|
golgin A4
|
|
chr2_+_108432008
|
0.893
|
NM_181453
|
GCC2
|
GRIP and coiled-coil domain containing 2
|
|
chr2_+_108432097
|
0.865
|
|
GCC2
|
GRIP and coiled-coil domain containing 2
|
|
chr3_+_142587973
|
0.854
|
|
|
|
|
chr10_-_52315436
|
0.853
|
NM_001198818
NM_001198819
NM_001198820
NM_014576
NM_138932
NM_138933
|
A1CF
|
APOBEC1 complementation factor
|
|
chr20_+_37024383
|
0.852
|
NM_001190809
NM_021931
|
DHX35
|
DEAH (Asp-Glu-Ala-His) box polypeptide 35
|
|
chr6_+_31731647
|
0.835
|
NM_019101
|
APOM
|
apolipoprotein M
|
|
chr2_-_206732348
|
0.832
|
NM_005006
|
NDUFS1
|
NADH dehydrogenase (ubiquinone) Fe-S protein 1, 75kDa (NADH-coenzyme Q reductase)
|
|
chr5_-_42847716
|
0.816
|
NM_001085486
NM_001093726
NM_005410
|
SEPP1
|
selenoprotein P, plasma, 1
|
|
chr12_+_94776836
|
0.805
|
NM_003095
|
SNRPF
|
small nuclear ribonucleoprotein polypeptide F
|
|
chr6_-_146326918
|
0.776
|
NM_001042683
NM_173082
|
SHPRH
|
SNF2 histone linker PHD RING helicase
|
|
chr2_-_175754366
|
0.772
|
NM_001002258
NM_001190329
NM_001689
|
ATP5G3
|
ATP synthase, H+ transporting, mitochondrial Fo complex, subunit C3 (subunit 9)
|
|
chr2_+_159021653
|
0.769
|
|
PKP4
|
plakophilin 4
|
|
chr7_-_100630912
|
0.749
|
NM_178176
|
MOGAT3
|
monoacylglycerol O-acyltransferase 3
|
|
chr16_-_45565068
|
0.747
|
NM_005880
|
DNAJA2
|
DnaJ (Hsp40) homolog, subfamily A, member 2
|
|
chr20_-_36095236
|
0.743
|
NM_014657
|
TTI1
|
Tel2 interacting protein 1 homolog (S. pombe)
|
|
chr6_+_142510108
|
0.741
|
|
VTA1
|
Vps20-associated 1 homolog (S. cerevisiae)
|
|
chr20_+_36095263
|
0.740
|
NM_021215
|
RPRD1B
|
regulation of nuclear pre-mRNA domain containing 1B
|
|
chr6_+_142510094
|
0.737
|
NM_016485
|
VTA1
|
Vps20-associated 1 homolog (S. cerevisiae)
|
|
chr2_+_177965617
|
0.736
|
NM_003659
|
AGPS
|
alkylglycerone phosphate synthase
|
|
chr9_+_114182216
|
0.708
|
|
HSDL2
|
hydroxysteroid dehydrogenase like 2
|
|
chr6_+_31731667
|
0.707
|
|
APOM
|
apolipoprotein M
|
|
chr18_-_50004458
|
0.700
|
|
MBD2
|
methyl-CpG binding domain protein 2
|
|
chr12_+_65949631
|
0.697
|
|
CAND1
|
cullin-associated and neddylation-dissociated 1
|
|
chr6_+_142510075
|
0.691
|
|
VTA1
|
Vps20-associated 1 homolog (S. cerevisiae)
|
|
chr3_-_114947330
|
0.679
|
|
NAA50
|
N(alpha)-acetyltransferase 50, NatE catalytic subunit
|
|
chr2_+_108604111
|
0.676
|
NM_001193484
|
LIMS1
|
LIM and senescent cell antigen-like domains 1
|
|
chr20_-_19981005
|
0.674
|
|
CRNKL1
|
crooked neck pre-mRNA splicing factor-like 1 (Drosophila)
|
|
chr12_-_93991235
|
0.669
|
NM_001032287
NM_001127362
NM_003297
|
NR2C1
|
nuclear receptor subfamily 2, group C, member 1
|
|
chr12_+_65949414
|
0.665
|
|
CAND1
|
cullin-associated and neddylation-dissociated 1
|
|
chr20_+_30409649
|
0.659
|
NM_001164603
NM_015338
|
ASXL1
|
additional sex combs like 1 (Drosophila)
|
|
chr9_+_114182009
|
0.651
|
NM_001195822
NM_032303
|
HSDL2
|
hydroxysteroid dehydrogenase like 2
|
|
chr5_+_80292313
|
0.628
|
NM_006909
|
RASGRF2
|
Ras protein-specific guanine nucleotide-releasing factor 2
|
|
chr20_+_36095552
|
0.628
|
|
RPRD1B
|
regulation of nuclear pre-mRNA domain containing 1B
|
|
chr5_+_14717803
|
0.617
|
|
FAM105B
|
family with sequence similarity 105, member B
|
|
chr20_+_36095621
|
0.613
|
|
|
|
|
chr17_+_35532994
|
0.608
|
|
MSL1
|
male-specific lethal 1 homolog (Drosophila)
|
|
chr2_+_138438277
|
0.598
|
NM_001024074
NM_001024075
NM_006895
|
HNMT
|
histamine N-methyltransferase
|
|
chr17_+_38403935
|
0.592
|
NM_000988
|
RPL27
|
ribosomal protein L27
|
|
chr20_+_43943254
|
0.587
|
NM_080603
|
ZSWIM1
|
zinc finger, SWIM-type containing 1
|
|
chrX_-_122694569
|
0.583
|
NM_001081550
|
THOC2
|
THO complex 2
|
|
chr1_-_224441024
|
0.582
|
NM_022735
|
ACBD3
|
acyl-CoA binding domain containing 3
|
|
chr1_-_224441023
|
0.580
|
|
ACBD3
|
acyl-CoA binding domain containing 3
|
|
chr6_+_142510034
|
0.549
|
|
VTA1
|
Vps20-associated 1 homolog (S. cerevisiae)
|
|
chr12_+_94776901
|
0.547
|
|
SNRPF
|
small nuclear ribonucleoprotein polypeptide F
|
|
chr1_-_32632601
|
0.547
|
NM_001143888
NM_001143889
NM_001143890
NM_018045
|
BSDC1
|
BSD domain containing 1
|
|
chr2_+_206732562
|
0.546
|
NM_001037663
NM_001959
NM_021121
|
EEF1B2
|
eukaryotic translation elongation factor 1 beta 2
|
|
chr3_+_151747266
|
0.545
|
|
EIF2A
|
eukaryotic translation initiation factor 2A, 65kDa
|
|
chr1_+_144439069
|
0.543
|
NM_002614
|
PDZK1
|
PDZ domain containing 1
|
|
chr2_+_219354692
|
0.539
|
NM_000784
|
CYP27A1
|
cytochrome P450, family 27, subfamily A, polypeptide 1
|
|
chr7_-_72574383
|
0.532
|
|
BAZ1B
|
bromodomain adjacent to zinc finger domain, 1B
|
|
chr17_-_33179118
|
0.530
|
NM_000458
NM_001165923
|
HNF1B
|
HNF1 homeobox B
|
|
chr2_+_27967102
|
0.526
|
|
BRE
|
brain and reproductive organ-expressed (TNFRSF1A modulator)
|
|
chr1_-_101263867
|
0.526
|
NM_001077394
NM_001077395
NM_015958
|
DPH5
|
DPH5 homolog (S. cerevisiae)
|
|
chr2_+_219141761
|
0.518
|
NM_005444
|
RQCD1
|
RCD1 required for cell differentiation1 homolog (S. pombe)
|
|
chr14_-_94693359
|
0.515
|
|
DICER1
|
dicer 1, ribonuclease type III
|
|
chr2_+_206732575
|
0.514
|
|
EEF1B2
|
eukaryotic translation elongation factor 1 beta 2
|
|
chr12_+_65949327
|
0.510
|
NM_018448
|
CAND1
|
cullin-associated and neddylation-dissociated 1
|
|
chr7_-_74127508
|
0.505
|
|
WBSCR16
|
Williams-Beuren syndrome chromosome region 16
|
|
chr7_-_72574528
|
0.505
|
NM_032408
|
BAZ1B
|
bromodomain adjacent to zinc finger domain, 1B
|
|
chr6_-_146326863
|
0.499
|
|
SHPRH
|
SNF2 histone linker PHD RING helicase
|
|
chr12_+_119900677
|
0.478
|
|
HNF1A
|
HNF1 homeobox A
|
|
chr6_-_146327238
|
0.477
|
|
SHPRH
|
SNF2 histone linker PHD RING helicase
|
|
chr2_-_27966715
|
0.472
|
NM_022128
|
RBKS
|
ribokinase
|
|
chr6_+_143813472
|
0.469
|
NM_003630
|
PEX3
|
peroxisomal biogenesis factor 3
|
|
chr7_-_72574474
|
0.467
|
|
BAZ1B
|
bromodomain adjacent to zinc finger domain, 1B
|
|
chr2_+_73294852
|
0.441
|
NM_006062
|
SMYD5
|
SMYD family member 5
|
|
chr11_+_86426518
|
0.438
|
NM_001168724
NM_022918
|
TMEM135
|
transmembrane protein 135
|
|
chrX_-_122694562
|
0.433
|
|
THOC2
|
THO complex 2
|
|
chr16_-_45564984
|
0.429
|
|
DNAJA2
|
DnaJ (Hsp40) homolog, subfamily A, member 2
|
|
chr20_+_41621079
|
0.428
|
NM_170693
|
SGK2
|
serum/glucocorticoid regulated kinase 2
|
|
chr12_-_49708279
|
0.428
|
NM_001174125
|
SLC11A2
|
solute carrier family 11 (proton-coupled divalent metal ion transporters), member 2
|
|
chr20_+_41621119
|
0.424
|
|
SGK2
|
serum/glucocorticoid regulated kinase 2
|
|
chr14_-_74713086
|
0.421
|
|
TMED10
|
transmembrane emp24-like trafficking protein 10 (yeast)
|
|
chr2_+_27967116
|
0.414
|
|
BRE
|
brain and reproductive organ-expressed (TNFRSF1A modulator)
|
|
chr2_+_119841611
|
0.393
|
NM_001079863
|
DBI
|
diazepam binding inhibitor (GABA receptor modulator, acyl-CoA binding protein)
|
|
chr17_-_33178987
|
0.391
|
|
HNF1B
|
HNF1 homeobox B
|
|
chr20_+_43919662
|
0.384
|
NM_080752
|
ZSWIM3
|
zinc finger, SWIM-type containing 3
|
|
chr11_+_117948301
|
0.384
|
NM_001142281
NM_001655
|
ARCN1
|
archain 1
|
|
chr11_+_117948350
|
0.384
|
|
ARCN1
|
archain 1
|
|
chr7_-_65084641
|
0.379
|
|
GUSB
|
glucuronidase, beta
|
|
chr4_+_4594619
|
0.377
|
|
LOC100507266
|
hypothetical LOC100507266
|
|
chrX_-_18282698
|
0.374
|
NM_006089
|
SCML2
|
sex comb on midleg-like 2 (Drosophila)
|
|
chr9_+_73954174
|
0.368
|
|
GDA
|
guanine deaminase
|
|
chr1_-_172153087
|
0.362
|
NM_000488
|
SERPINC1
|
serpin peptidase inhibitor, clade C (antithrombin), member 1
|
|
chr6_-_146327155
|
0.361
|
|
SHPRH
|
SNF2 histone linker PHD RING helicase
|
|
chr20_-_43919360
|
0.361
|
NM_005469
|
ACOT8
|
acyl-CoA thioesterase 8
|
|
chr18_-_44730706
|
0.358
|
|
SMAD7
|
SMAD family member 7
|
|
chr4_-_84150885
|
0.358
|
|
LIN54
|
lin-54 homolog (C. elegans)
|
|
chr2_+_27967159
|
0.358
|
|
BRE
|
brain and reproductive organ-expressed (TNFRSF1A modulator)
|
|
chr5_+_14717764
|
0.357
|
NM_138348
|
FAM105B
|
family with sequence similarity 105, member B
|
|
chr18_-_44731078
|
0.347
|
NM_001190821
NM_005904
|
SMAD7
|
SMAD family member 7
|
|
chr7_-_74127572
|
0.346
|
|
WBSCR16
|
Williams-Beuren syndrome chromosome region 16
|
|
chr11_+_118374071
|
0.345
|
|
CCDC84
|
coiled-coil domain containing 84
|
|
chr7_+_120415958
|
0.343
|
NM_024913
|
C7orf58
|
chromosome 7 open reading frame 58
|
|
chr2_-_198072868
|
0.337
|
NM_002156
|
HSPD1
|
heat shock 60kDa protein 1 (chaperonin)
|
|
chrX_+_38305634
|
0.334
|
NM_004615
|
TSPAN7
|
tetraspanin 7
|
|
chr22_-_22389524
|
0.330
|
|
LOC91316
|
glucuronidase, beta/immunoglobulin lambda-like polypeptide 1 pseudogene
|
|
chr14_-_74713062
|
0.329
|
|
TMED10
|
transmembrane emp24-like trafficking protein 10 (yeast)
|
|
chr7_-_65084597
|
0.328
|
|
GUSB
|
glucuronidase, beta
|
|
chr1_+_43628136
|
0.327
|
NM_001012961
|
C1orf84
|
chromosome 1 open reading frame 84
|
|
chr1_+_78126787
|
0.324
|
NM_001172309
NM_144573
|
NEXN
|
nexilin (F actin binding protein)
|
|
chr3_-_114947681
|
0.318
|
NM_025146
|
NAA50
|
N(alpha)-acetyltransferase 50, NatE catalytic subunit
|
|
chr4_+_140806536
|
0.315
|
|
MGST2
|
microsomal glutathione S-transferase 2
|
|
chr7_-_106991941
|
0.312
|
NM_001161520
NM_006348
NM_181733
|
COG5
|
component of oligomeric golgi complex 5
|
|
chr11_+_117735530
|
0.312
|
|
UBE4A
|
ubiquitination factor E4A (UFD2 homolog, yeast)
|
|
chr1_-_15784048
|
0.305
|
NM_024758
|
AGMAT
|
agmatine ureohydrolase (agmatinase)
|
|
chr20_-_32363268
|
0.301
|
NM_001161766
|
AHCY
|
adenosylhomocysteinase
|
|
chr7_-_65084668
|
0.300
|
NM_000181
|
GUSB
|
glucuronidase, beta
|
|
chr11_+_117735558
|
0.299
|
|
UBE4A
|
ubiquitination factor E4A (UFD2 homolog, yeast)
|
|
chr16_+_30617962
|
0.298
|
NM_006662
|
SRCAP
|
Snf2-related CREBBP activator protein
|
|
chr4_+_109761170
|
0.294
|
NM_000995
NM_033625
|
RPL34
|
ribosomal protein L34
|
|
chr12_-_117294933
|
0.294
|
|
TAOK3
|
TAO kinase 3
|
|
chr11_-_116213511
|
0.294
|
NM_000039
|
APOA1
|
apolipoprotein A-I
|
|
chr2_+_26422495
|
0.287
|
|
EPT1
|
ethanolaminephosphotransferase 1 (CDP-ethanolamine-specific)
|
|
chr1_-_149404954
|
0.282
|
NM_212551
NM_001136543
|
LYSMD1
|
LysM, putative peptidoglycan-binding, domain containing 1
|
|
chr14_-_74713039
|
0.281
|
|
TMED10
|
transmembrane emp24-like trafficking protein 10 (yeast)
|
|
chr9_+_73954237
|
0.280
|
|
GDA
|
guanine deaminase
|
|
chr5_+_1854493
|
0.271
|
NM_004553
|
NDUFS6
|
NADH dehydrogenase (ubiquinone) Fe-S protein 6, 13kDa (NADH-coenzyme Q reductase)
|
|
chr1_-_154147787
|
0.267
|
NM_006912
|
RIT1
|
Ras-like without CAAX 1
|
|
chr7_-_65084622
|
0.266
|
|
GUSB
|
glucuronidase, beta
|
|
chr4_-_7120656
|
0.263
|
NM_025196
|
GRPEL1
|
GrpE-like 1, mitochondrial (E. coli)
|
|
chr14_-_74713088
|
0.262
|
NM_006827
|
TMED10
|
transmembrane emp24-like trafficking protein 10 (yeast)
|
|
chr8_+_96106362
|
0.257
|
NM_152416
|
C8orf38
|
chromosome 8 open reading frame 38
|
|
chr20_+_54400979
|
0.250
|
NM_001324
|
CSTF1
|
cleavage stimulation factor, 3' pre-RNA, subunit 1, 50kDa
|
|
chr3_+_151747204
|
0.247
|
NM_032025
|
EIF2A
|
eukaryotic translation initiation factor 2A, 65kDa
|
|
chr16_+_79829794
|
0.246
|
NM_017429
|
BCMO1
|
beta-carotene 15,15'-monooxygenase 1
|
|
chr8_-_124355670
|
0.240
|
NM_001017926
NM_007222
|
ZHX1
|
zinc fingers and homeoboxes 1
|
|
chr17_+_71508581
|
0.235
|
NM_001258
|
CDK3
|
cyclin-dependent kinase 3
|
|
chr19_+_10223912
|
0.234
|
|
MRPL4
|
mitochondrial ribosomal protein L4
|
|
chr14_-_74713012
|
0.233
|
|
TMED10
|
transmembrane emp24-like trafficking protein 10 (yeast)
|
|
chr19_+_54708312
|
0.228
|
|
FCGRT
|
Fc fragment of IgG, receptor, transporter, alpha
|
|
chr1_-_172153024
|
0.228
|
|
SERPINC1
|
serpin peptidase inhibitor, clade C (antithrombin), member 1
|
|
chr11_+_117735529
|
0.228
|
|
UBE4A
|
ubiquitination factor E4A (UFD2 homolog, yeast)
|
|
chr22_+_22445035
|
0.227
|
NM_005940
|
MMP11
|
matrix metallopeptidase 11 (stromelysin 3)
|
|
chr12_-_6949877
|
0.227
|
|
PHB2
|
prohibitin 2
|
|
chr11_+_117735502
|
0.224
|
NM_004788
|
UBE4A
|
ubiquitination factor E4A (UFD2 homolog, yeast)
|
|
chr15_-_50316996
|
0.223
|
|
MYO5C
|
myosin VC
|
|
chr2_+_219141546
|
0.221
|
|
RQCD1
|
RCD1 required for cell differentiation1 homolog (S. pombe)
|
|
chr19_+_54708302
|
0.219
|
|
FCGRT
|
Fc fragment of IgG, receptor, transporter, alpha
|
|
chr11_+_85690900
|
0.219
|
NM_016401
|
C11orf73
|
chromosome 11 open reading frame 73
|
|
chr6_+_33280396
|
0.218
|
NM_014234
|
HSD17B8
|
hydroxysteroid (17-beta) dehydrogenase 8
|
|
chr2_+_86521781
|
0.218
|
NM_018433
|
KDM3A
|
lysine (K)-specific demethylase 3A
|
|
chr10_+_103995297
|
0.217
|
NM_004193
|
GBF1
|
golgi brefeldin A resistant guanine nucleotide exchange factor 1
|
|
chr2_+_198072965
|
0.217
|
NM_002157
|
HSPE1
|
heat shock 10kDa protein 1 (chaperonin 10)
|
|
chr12_+_119900931
|
0.215
|
NM_000545
|
HNF1A
|
HNF1 homeobox A
|
|
chr14_-_30927931
|
0.215
|
NM_015473
|
HEATR5A
|
HEAT repeat containing 5A
|
|
chr2_+_119841298
|
0.210
|
NM_001178017
|
DBI
|
diazepam binding inhibitor (GABA receptor modulator, acyl-CoA binding protein)
|
|
chr17_+_7063970
|
0.209
|
|
ACADVL
|
acyl-CoA dehydrogenase, very long chain
|
|
chr4_-_84150997
|
0.209
|
NM_001115008
NM_194282
|
LIN54
|
lin-54 homolog (C. elegans)
|
|
chr5_+_92944680
|
0.208
|
NM_005654
|
NR2F1
|
nuclear receptor subfamily 2, group F, member 1
|
|
chr19_+_6323805
|
0.207
|
|
ALKBH7
|
alkB, alkylation repair homolog 7 (E. coli)
|
|
chr4_-_4594591
|
0.207
|
|
STX18
|
syntaxin 18
|
|
chr17_+_38403991
|
0.204
|
|
RPL27
|
ribosomal protein L27
|
|
chr19_-_40317829
|
0.202
|
NM_139284
|
LGI4
|
leucine-rich repeat LGI family, member 4
|
|
chr2_+_201691664
|
0.196
|
|
CFLAR
|
CASP8 and FADD-like apoptosis regulator
|
|
chr3_-_151746970
|
0.195
|
NM_014445
|
SERP1
|
stress-associated endoplasmic reticulum protein 1
|
|
chr7_-_74127622
|
0.194
|
NM_030798
|
WBSCR16
|
Williams-Beuren syndrome chromosome region 16
|
|
chr9_+_103335983
|
0.192
|
|
RNF20
|
ring finger protein 20
|
|
chr12_-_117295095
|
0.192
|
NM_016281
|
TAOK3
|
TAO kinase 3
|
|
chr10_-_62431203
|
0.191
|
|
RHOBTB1
|
Rho-related BTB domain containing 1
|
|
chr19_+_6323720
|
0.190
|
|
ALKBH7
|
alkB, alkylation repair homolog 7 (E. coli)
|
|
chr9_-_138277467
|
0.187
|
NM_181701
|
QSOX2
|
quiescin Q6 sulfhydryl oxidase 2
|
|
chr12_-_49706364
|
0.187
|
NM_000617
NM_001174126
NM_001174127
NM_001174128
|
SLC11A2
|
solute carrier family 11 (proton-coupled divalent metal ion transporters), member 2
|
|
chr19_-_60272708
|
0.186
|
NM_138412
|
RDH13
|
retinol dehydrogenase 13 (all-trans/9-cis)
|
|
chr10_+_103995248
|
0.186
|
|
GBF1
|
golgi brefeldin A resistant guanine nucleotide exchange factor 1
|
|
chr3_-_52287297
|
0.180
|
|
WDR82
|
WD repeat domain 82
|
|
chr16_+_31105579
|
0.179
|
|
|
|
|
chr2_+_27966985
|
0.179
|
NM_199193
NM_199194
NM_004899
NM_199191
NM_199192
|
BRE
|
brain and reproductive organ-expressed (TNFRSF1A modulator)
|
|
chr3_-_151746954
|
0.176
|
|
SERP1
|
stress-associated endoplasmic reticulum protein 1
|
|
chr9_+_103335964
|
0.174
|
|
RNF20
|
ring finger protein 20
|
|
chr6_-_2785682
|
0.173
|
|
SERPINB1
|
serpin peptidase inhibitor, clade B (ovalbumin), member 1
|
|
chr5_+_172416959
|
0.173
|
|
C5orf41
|
chromosome 5 open reading frame 41
|
|
chr2_-_191107660
|
0.172
|
|
TMEM194B
|
transmembrane protein 194B
|
|
chr3_-_12856948
|
0.165
|
NM_001007074
|
RPL32
|
ribosomal protein L32
|
|
chr4_+_140806371
|
0.165
|
NM_002413
|
MGST2
|
microsomal glutathione S-transferase 2
|
|
chr12_+_67290881
|
0.164
|
NM_001010942
NM_015646
|
RAP1B
|
RAP1B, member of RAS oncogene family
|
|
chr14_-_94693418
|
0.163
|
NM_177438
|
DICER1
|
dicer 1, ribonuclease type III
|
|
chr19_-_62759536
|
0.163
|
NM_001039654
|
ZNF550
|
zinc finger protein 550
|
|
chr19_+_54708389
|
0.159
|
|
FCGRT
|
Fc fragment of IgG, receptor, transporter, alpha
|
|
chr18_-_50004770
|
0.159
|
|
MBD2
|
methyl-CpG binding domain protein 2
|
|
chr1_-_209915443
|
0.157
|
NM_002497
|
NEK2
|
NIMA (never in mitosis gene a)-related kinase 2
|
|
chr12_+_67489063
|
0.153
|
|
MDM2
|
Mdm2 p53 binding protein homolog (mouse)
|
|
chr18_-_45594206
|
0.150
|
NM_006111
|
ACAA2
|
acetyl-CoA acyltransferase 2
|
|
chrX_-_130792297
|
0.150
|
|
LOC286467
|
hypothetical LOC286467
|