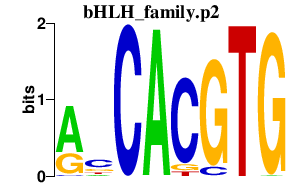
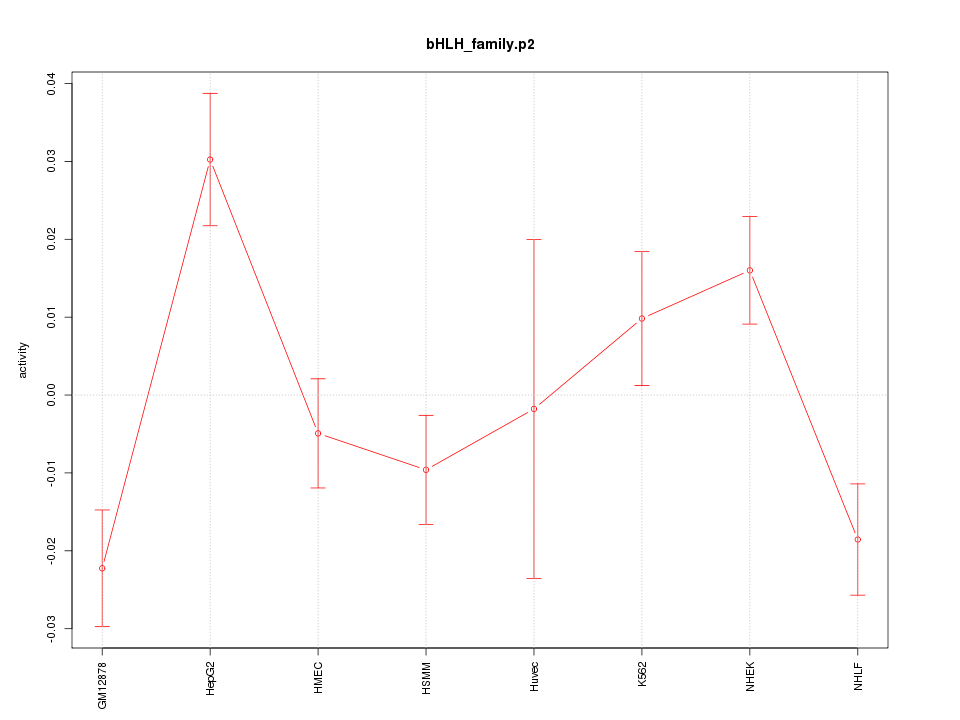
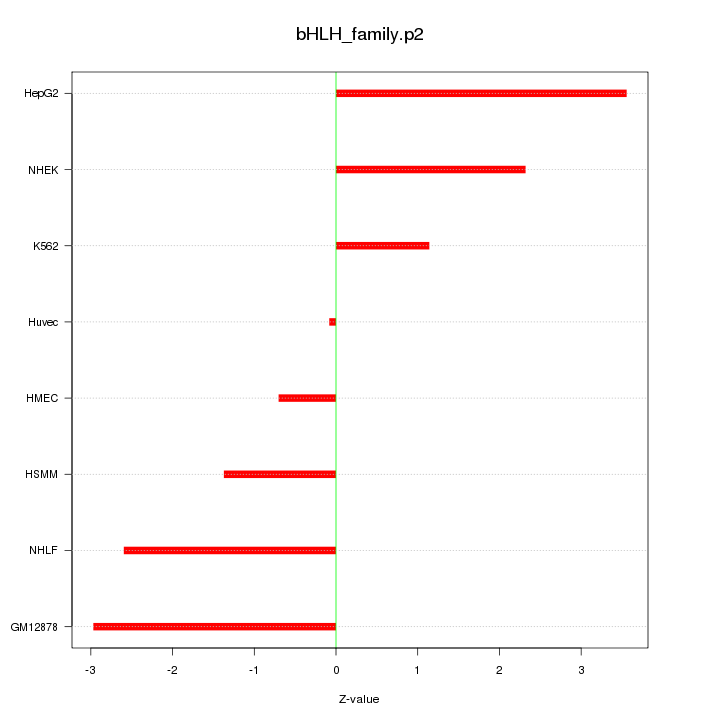
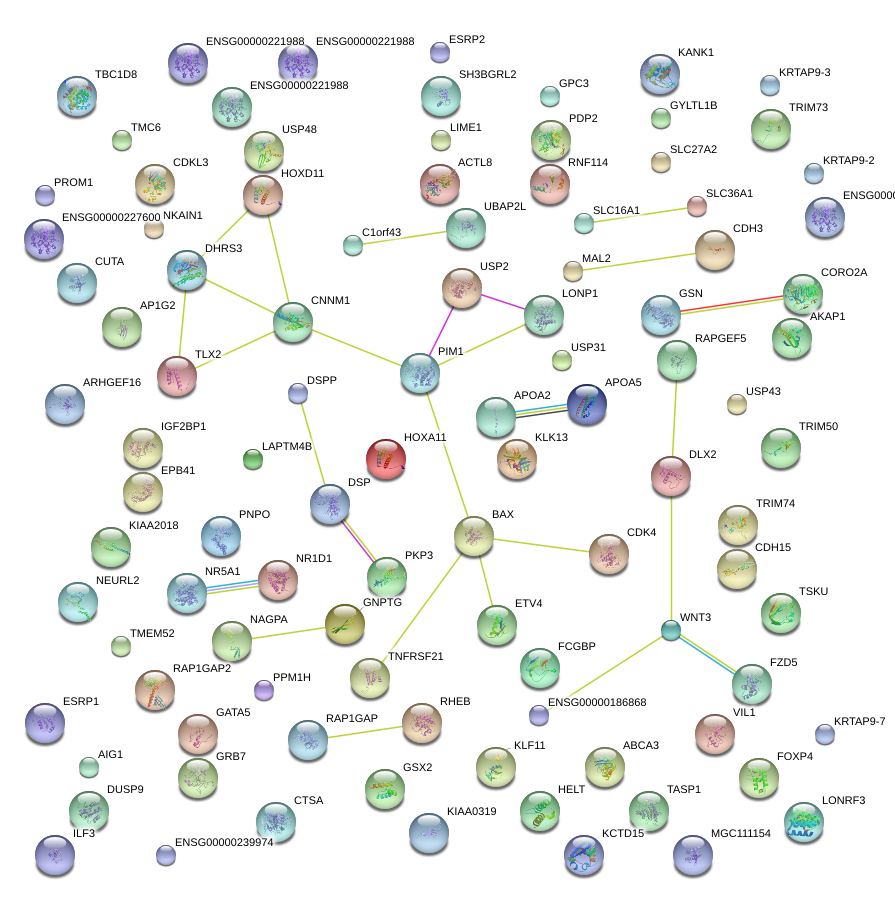
|
chr8_+_120289735
|
2.091
|
NM_052886
|
MAL2
|
mal, T-cell differentiation protein 2 (gene/pseudogene)
|
|
chr17_-_73636362
|
2.013
|
|
TMC6
|
transmembrane channel-like 6
|
|
chr6_-_47385313
|
1.553
|
|
TNFRSF21
|
tumor necrosis factor receptor superfamily, member 21
|
|
chr12_-_56432385
|
1.488
|
NM_000075
|
CDK4
|
cyclin-dependent kinase 4
|
|
chr1_-_12599934
|
1.463
|
NM_004753
|
DHRS3
|
dehydrogenase/reductase (SDR family) member 3
|
|
chr11_+_384199
|
1.435
|
NM_007183
|
PKP3
|
plakophilin 3
|
|
chr6_-_33493792
|
1.418
|
|
CUTA
|
cutA divalent cation tolerance homolog (E. coli)
|
|
chr19_+_10626106
|
1.344
|
|
ILF3
|
interleukin enhancer binding factor 3, 90kDa
|
|
chr2_+_218992053
|
1.309
|
NM_007127
|
VIL1
|
villin 1
|
|
chr12_-_61614930
|
1.305
|
NM_020700
|
PPM1H
|
protein phosphatase, Mg2+/Mn2+ dependent, 1H
|
|
chr11_+_384238
|
1.302
|
|
PKP3
|
plakophilin 3
|
|
chr5_-_133734616
|
1.269
|
|
CDKL3
|
cyclin-dependent kinase-like 3
|
|
chr16_-_5023934
|
1.266
|
NM_016256
|
NAGPA
|
N-acetylglucosamine-1-phosphodiester alpha-N-acetylglucosaminidase
|
|
chr2_-_208342296
|
1.266
|
NM_003468
|
FZD5
|
frizzled homolog 5 (Drosophila)
|
|
chr8_+_95722448
|
1.250
|
|
ESRP1
|
epithelial splicing regulatory protein 1
|
|
chr6_+_143423656
|
1.244
|
NM_016108
|
AIG1
|
androgen-induced 1
|
|
chrX_+_152562013
|
1.240
|
|
DUSP9
|
dual specificity phosphatase 9
|
|
chr12_-_61615124
|
1.221
|
|
PPM1H
|
protein phosphatase, Mg2+/Mn2+ dependent, 1H
|
|
chr11_+_76171780
|
1.206
|
NM_015516
|
TSKU
|
tsukushi small leucine rich proteoglycan homolog (Xenopus laevis)
|
|
chr1_+_3360880
|
1.187
|
NM_014448
|
ARHGEF16
|
Rho guanine nucleotide exchange factor (GEF) 16
|
|
chr17_+_35148084
|
1.172
|
NM_001030002
|
GRB7
|
growth factor receptor-bound protein 7
|
|
chr2_+_10101081
|
1.168
|
NM_003597
NM_001177716
|
KLF11
|
Kruppel-like factor 11
|
|
chr17_-_35510431
|
1.158
|
NM_021724
|
NR1D1
|
nuclear receptor subfamily 1, group D, member 1
|
|
chr17_+_77478858
|
1.147
|
|
LOC92659
|
hypothetical LOC92659
|
|
chr17_-_35510074
|
1.144
|
|
NR1D1
|
nuclear receptor subfamily 1, group D, member 1
|
|
chr17_+_44429748
|
1.143
|
NM_001160423
NM_006546
|
IGF2BP1
|
insulin-like growth factor 2 mRNA binding protein 1
|
|
chr6_-_47385599
|
1.138
|
|
TNFRSF21
|
tumor necrosis factor receptor superfamily, member 21
|
|
chr17_+_9489578
|
1.135
|
NM_153210
|
USP43
|
ubiquitin specific peptidase 43
|
|
chr4_-_15694407
|
1.127
|
|
PROM1
|
prominin 1
|
|
chr19_+_54149941
|
1.125
|
|
BAX
|
BCL2-associated X protein
|
|
chr6_+_37245860
|
1.123
|
NM_002648
|
PIM1
|
pim-1 oncogene
|
|
chr14_-_23106372
|
1.119
|
|
AP1G2
|
adaptor-related protein complex 1, gamma 2 subunit
|
|
chr6_-_47385205
|
1.099
|
|
TNFRSF21
|
tumor necrosis factor receptor superfamily, member 21
|
|
chr19_-_10625449
|
1.099
|
|
LOC147727
|
hypothetical LOC147727
|
|
chr6_-_47385604
|
1.091
|
|
TNFRSF21
|
tumor necrosis factor receptor superfamily, member 21
|
|
chr1_+_17954394
|
1.084
|
NM_030812
|
ACTL8
|
actin-like 8
|
|
chr20_-_43953036
|
1.071
|
NM_080749
|
NEURL2
|
neuralized homolog 2 (Drosophila)
|
|
chr11_-_118757477
|
1.071
|
|
USP2
|
ubiquitin specific peptidase 2
|
|
chr6_+_143423738
|
1.060
|
|
AIG1
|
androgen-induced 1
|
|
chr11_-_116168316
|
1.047
|
NM_001166598
NM_052968
|
APOA5
|
apolipoprotein A-V
|
|
chr17_-_73636386
|
1.031
|
NM_001127198
|
TMC6
|
transmembrane channel-like 6
|
|
chr16_+_65471881
|
1.031
|
NM_020786
|
PDP2
|
pyruvate dehyrogenase phosphatase catalytic subunit 2
|
|
chr2_-_101134146
|
1.028
|
NM_001102426
|
TBC1D8
|
TBC1 domain family, member 8 (with GRAM domain)
|
|
chr16_-_66827449
|
1.027
|
NM_024939
|
ESRP2
|
epithelial splicing regulatory protein 2
|
|
chr6_-_47385470
|
1.020
|
|
TNFRSF21
|
tumor necrosis factor receptor superfamily, member 21
|
|
chr16_-_5023914
|
1.019
|
|
NAGPA
|
N-acetylglucosamine-1-phosphodiester alpha-N-acetylglucosaminidase
|
|
chr6_-_47385605
|
1.018
|
NM_014452
|
TNFRSF21
|
tumor necrosis factor receptor superfamily, member 21
|
|
chr9_+_696744
|
1.015
|
NM_153186
|
KANK1
|
KN motif and ankyrin repeat domains 1
|
|
chr8_+_98857190
|
1.012
|
|
LAPTM4B
|
lysosomal protein transmembrane 4 beta
|
|
chr16_+_67236699
|
0.989
|
|
CDH3
|
cadherin 3, type 1, P-cadherin (placental)
|
|
chr2_-_172675873
|
0.988
|
|
DLX2
|
distal-less homeobox 2
|
|
chr17_+_35147697
|
0.986
|
NM_005310
|
GRB7
|
growth factor receptor-bound protein 7
|
|
chr6_-_33493640
|
0.975
|
|
CUTA
|
cutA divalent cation tolerance homolog (E. coli)
|
|
chr16_-_23068010
|
0.975
|
NM_020718
|
USP31
|
ubiquitin specific peptidase 31
|
|
chr15_+_48261649
|
0.968
|
NM_001159629
NM_003645
|
SLC27A2
|
solute carrier family 27 (fatty acid transporter), member 2
|
|
chr1_-_159459999
|
0.967
|
NM_001643
|
APOA2
|
apolipoprotein A-II
|
|
chr17_-_38979178
|
0.961
|
NM_001079675
|
ETV4
|
ets variant 4
|
|
chr17_+_35147738
|
0.958
|
|
GRB7
|
growth factor receptor-bound protein 7
|
|
chr8_+_95722575
|
0.948
|
|
ESRP1
|
epithelial splicing regulatory protein 1
|
|
chr1_-_113300138
|
0.939
|
NM_001166496
|
SLC16A1
|
solute carrier family 16, member 1 (monocarboxylic acid transporter 1)
|
|
chr8_+_95722516
|
0.937
|
NM_001034915
NM_001122825
NM_001122826
NM_001122827
NM_017697
|
ESRP1
|
epithelial splicing regulatory protein 1
|
|
chr17_+_52518098
|
0.913
|
|
AKAP1
|
A kinase (PRKA) anchor protein 1
|
|
chr10_+_101079099
|
0.911
|
|
CNNM1
|
cyclin M1
|
|
chr6_+_32230172
|
0.890
|
|
PPT2
|
palmitoyl-protein thioesterase 2
|
|
chr20_+_43953428
|
0.889
|
|
CTSA
|
cathepsin A
|
|
chr1_-_21868380
|
0.885
|
NM_001145657
NM_002885
|
RAP1GAP
|
RAP1 GTPase activating protein
|
|
chr19_+_10625987
|
0.884
|
|
ILF3
|
interleukin enhancer binding factor 3, 90kDa
|
|
chr6_+_80397690
|
0.876
|
NM_031469
|
SH3BGRL2
|
SH3 domain binding glutamic acid-rich protein like 2
|
|
chrX_+_117992603
|
0.872
|
|
LONRF3
|
LON peptidase N-terminal domain and ring finger 3
|
|
chr20_-_13567529
|
0.869
|
NM_017714
|
TASP1
|
taspase, threonine aspartase, 1
|
|
chr22_-_39923412
|
0.867
|
|
|
|
|
chr1_+_29113589
|
0.853
|
NM_001166006
|
EPB41
|
erythrocyte membrane protein band 4.1 (elliptocytosis 1, RH-linked)
|
|
chr1_+_152459918
|
0.853
|
NM_014847
|
UBAP2L
|
ubiquitin associated protein 2-like
|
|
chr1_-_1840564
|
0.846
|
NM_178545
|
TMEM52
|
transmembrane protein 52
|
|
chr5_+_78568094
|
0.833
|
|
|
|
|
chr1_-_152459697
|
0.832
|
NM_001098616
NM_015449
NM_138740
|
C1orf43
|
chromosome 1 open reading frame 43
|
|
chr20_+_43953606
|
0.828
|
|
CTSA
|
cathepsin A
|
|
chr20_-_60484420
|
0.826
|
NM_080473
|
GATA5
|
GATA binding protein 5
|
|
chr17_+_43374021
|
0.821
|
|
PNPO
|
pyridoxamine 5'-phosphate oxidase
|
|
chr20_+_47986335
|
0.820
|
|
RNF114
|
ring finger protein 114
|
|
chr16_+_1341937
|
0.817
|
|
GNPTG
|
N-acetylglucosamine-1-phosphate transferase, gamma subunit
|
|
chr19_-_45132372
|
0.802
|
NM_003890
|
FCGBP
|
Fc fragment of IgG binding protein
|
|
chr17_+_36636404
|
0.800
|
NM_031961
|
KRTAP9-2
|
keratin associated protein 9-2
|
|
chr4_+_54660954
|
0.798
|
NM_133267
|
GSX2
|
GS homeobox 2
|
|
chr11_-_118757574
|
0.797
|
NM_004205
|
USP2
|
ubiquitin specific peptidase 2
|
|
chr9_-_99994704
|
0.795
|
NM_052820
|
CORO2A
|
coronin, actin binding protein, 2A
|
|
chr20_+_43953331
|
0.795
|
|
CTSA
|
cathepsin A
|
|
chr12_-_56432375
|
0.794
|
|
CDK4
|
cyclin-dependent kinase 4
|
|
chr7_-_22363036
|
0.788
|
NM_012294
|
RAPGEF5
|
Rap guanine nucleotide exchange factor (GEF) 5
|
|
chr16_-_5023562
|
0.786
|
|
NAGPA
|
N-acetylglucosamine-1-phosphodiester alpha-N-acetylglucosaminidase
|
|
chr6_+_32229771
|
0.786
|
|
PPT2-EGFL8
PPT2
|
PPT2-EGFL8 read-through transcript
palmitoyl-protein thioesterase 2
|
|
chr20_+_43953372
|
0.786
|
|
CTSA
|
cathepsin A
|
|
chr7_-_150847624
|
0.784
|
|
RHEB
|
Ras homolog enriched in brain
|
|
chr1_-_113300351
|
0.783
|
NM_003051
|
SLC16A1
|
solute carrier family 16, member 1 (monocarboxylic acid transporter 1)
|
|
chr9_-_126309510
|
0.782
|
NM_004959
|
NR5A1
|
nuclear receptor subfamily 5, group A, member 1
|
|
chr7_-_72380020
|
0.772
|
NM_178125
|
TRIM50
|
tripartite motif containing 50
|
|
chr2_+_176680251
|
0.762
|
NM_021192
|
HOXD11
|
homeobox D11
|
|
chr19_+_38979502
|
0.760
|
NM_001129994
NM_001129995
NM_024076
|
KCTD15
|
potassium channel tetramerisation domain containing 15
|
|
chr17_+_2627099
|
0.756
|
|
RAP1GAP2
|
RAP1 GTPase activating protein 2
|
|
chr1_-_152459667
|
0.750
|
|
C1orf43
|
chromosome 1 open reading frame 43
|
|
chr1_-_31485320
|
0.750
|
NM_024522
|
NKAIN1
|
Na+/K+ transporting ATPase interacting 1
|
|
chr17_-_42250961
|
0.747
|
NM_030753
|
WNT3
|
wingless-type MMTV integration site family, member 3
|
|
chr2_+_74594093
|
0.743
|
|
TLX2
|
T-cell leukemia homeobox 2
|
|
chr6_+_7486836
|
0.733
|
|
DSP
|
desmoplakin
|
|
chrX_-_132947141
|
0.729
|
|
GPC3
|
glypican 3
|
|
chr5_+_150807380
|
0.727
|
|
SLC36A1
|
solute carrier family 36 (proton/amino acid symporter), member 1
|
|
chr11_+_45900731
|
0.726
|
|
GYLTL1B
|
glycosyltransferase-like 1B
|
|
chr6_+_7486862
|
0.725
|
NM_001008844
NM_004415
|
DSP
|
desmoplakin
|
|
chr10_+_101078784
|
0.723
|
NM_020348
|
CNNM1
|
cyclin M1
|
|
chr4_+_186176988
|
0.717
|
NM_001029887
|
HELT
|
HES/HEY-like transcription factor
|
|
chr6_+_41622267
|
0.715
|
|
FOXP4
|
forkhead box P4
|
|
chr11_-_71492019
|
0.714
|
NM_017907
|
LAMTOR1
|
late endosomal/lysosomal adaptor, MAPK and MTOR activator 1
|
|
chr12_+_94861201
|
0.714
|
NM_152435
|
AMDHD1
|
amidohydrolase domain containing 1
|
|
chr6_+_36346214
|
0.709
|
NM_001145717
|
PNPLA1
|
patatin-like phospholipase domain containing 1
|
|
chr17_-_36507900
|
0.705
|
NM_031960
|
KRTAP4-8
|
keratin associated protein 4-8
|
|
chr3_-_188945921
|
0.701
|
|
BCL6
|
B-cell CLL/lymphoma 6
|
|
chr2_+_219751207
|
0.697
|
|
FAM134A
|
family with sequence similarity 134, member A
|
|
chr12_+_54760211
|
0.697
|
|
ERBB3
|
v-erb-b2 erythroblastic leukemia viral oncogene homolog 3 (avian)
|
|
chr4_-_15694691
|
0.696
|
NM_001145847
NM_001145848
|
PROM1
|
prominin 1
|
|
chr6_+_160310118
|
0.693
|
NM_000876
|
IGF2R
|
insulin-like growth factor 2 receptor
|
|
chr14_-_54438913
|
0.692
|
NM_000161
NM_001024024
NM_001024070
NM_001024071
|
GCH1
|
GTP cyclohydrolase 1
|
|
chr20_-_2769120
|
0.692
|
|
FAM113A
|
family with sequence similarity 113, member A
|
|
chr11_-_66482366
|
0.688
|
NM_000920
NM_001040716
|
PC
|
pyruvate carboxylase
|
|
chr6_+_138230045
|
0.688
|
NM_006290
|
TNFAIP3
|
tumor necrosis factor, alpha-induced protein 3
|
|
chr1_+_61320213
|
0.685
|
NM_001134673
NM_005595
|
NFIA
|
nuclear factor I/A
|
|
chr2_-_239988269
|
0.683
|
|
HDAC4
|
histone deacetylase 4
|
|
chr19_-_2734324
|
0.681
|
|
SGTA
|
small glutamine-rich tetratricopeptide repeat (TPR)-containing, alpha
|
|
chr19_-_5291700
|
0.681
|
NM_002850
NM_130853
NM_130854
NM_130855
|
PTPRS
|
protein tyrosine phosphatase, receptor type, S
|
|
chr14_-_93919305
|
0.679
|
|
SERPINA1
|
serpin peptidase inhibitor, clade A (alpha-1 antiproteinase, antitrypsin), member 1
|
|
chr10_-_82039158
|
0.678
|
NM_000429
|
MAT1A
|
methionine adenosyltransferase I, alpha
|
|
chr2_-_220116508
|
0.678
|
NM_024536
|
CHPF
|
chondroitin polymerizing factor
|
|
chr10_-_120504445
|
0.677
|
|
C10orf46
|
chromosome 10 open reading frame 46
|
|
chr11_+_18300370
|
0.673
|
NM_001142307
NM_005316
|
GTF2H1
|
general transcription factor IIH, polypeptide 1, 62kDa
|
|
chr6_+_143423723
|
0.672
|
|
AIG1
|
androgen-induced 1
|
|
chr20_+_13924263
|
0.672
|
|
MACROD2
|
MACRO domain containing 2
|
|
chr1_+_61315516
|
0.669
|
NM_001145511
|
NFIA
|
nuclear factor I/A
|
|
chr9_-_130749604
|
0.663
|
NM_014908
|
DOLK
|
dolichol kinase
|
|
chr11_-_71491942
|
0.660
|
|
LAMTOR1
|
late endosomal/lysosomal adaptor, MAPK and MTOR activator 1
|
|
chrX_+_130985316
|
0.659
|
NM_001042452
|
MST4
|
serine/threonine protein kinase MST4
|
|
chr6_+_31903374
|
0.657
|
NM_005346
|
HSPA1A
HSPA1B
|
heat shock 70kDa protein 1A
heat shock 70kDa protein 1B
|
|
chrX_-_107866262
|
0.654
|
NM_003604
|
IRS4
|
insulin receptor substrate 4
|
|
chr20_+_48781457
|
0.654
|
NM_032521
|
PARD6B
|
par-6 partitioning defective 6 homolog beta (C. elegans)
|
|
chr20_+_37024383
|
0.652
|
NM_001190809
NM_021931
|
DHX35
|
DEAH (Asp-Glu-Ala-His) box polypeptide 35
|
|
chr17_+_73676219
|
0.650
|
NM_004710
|
SYNGR2
|
synaptogyrin 2
|
|
chr17_+_75689955
|
0.648
|
|
GAA
|
glucosidase, alpha; acid
|
|
chr16_+_56616782
|
0.647
|
NM_002428
|
MMP15
|
matrix metallopeptidase 15 (membrane-inserted)
|
|
chr15_+_57696200
|
0.647
|
|
GCNT3
|
glucosaminyl (N-acetyl) transferase 3, mucin type
|
|
chr3_-_188946168
|
0.645
|
NM_001706
|
BCL6
|
B-cell CLL/lymphoma 6
|
|
chr2_-_172675480
|
0.643
|
NM_004405
|
DLX2
|
distal-less homeobox 2
|
|
chr2_-_39859689
|
0.640
|
|
THUMPD2
|
THUMP domain containing 2
|
|
chr21_+_32167175
|
0.637
|
NM_014586
|
HUNK
|
hormonally up-regulated Neu-associated kinase
|
|
chr4_+_164484540
|
0.634
|
NM_006174
|
NPY5R
|
neuropeptide Y receptor Y5
|
|
chr2_-_11727705
|
0.633
|
NM_012344
|
NTSR2
|
neurotensin receptor 2
|
|
chr1_-_109742020
|
0.631
|
NM_002959
|
SORT1
|
sortilin 1
|
|
chr2_-_85683268
|
0.628
|
NM_001031738
|
TMEM150A
|
transmembrane protein 150A
|
|
chr4_-_100069303
|
0.625
|
|
EIF4E
|
eukaryotic translation initiation factor 4E
|
|
chr17_+_77528782
|
0.625
|
|
ASPSCR1
|
alveolar soft part sarcoma chromosome region, candidate 1
|
|
chr7_-_103417198
|
0.620
|
NM_005045
NM_173054
|
RELN
|
reelin
|
|
chrX_-_131919896
|
0.620
|
|
HS6ST2
|
heparan sulfate 6-O-sulfotransferase 2
|
|
chr6_+_105511615
|
0.620
|
NM_001004317
|
LIN28B
|
lin-28 homolog B (C. elegans)
|
|
chr1_-_152459506
|
0.620
|
|
C1orf43
|
chromosome 1 open reading frame 43
|
|
chr16_-_69277355
|
0.614
|
NM_138383
|
MTSS1L
|
metastasis suppressor 1-like
|
|
chr22_-_35233032
|
0.611
|
NM_024955
NM_001102371
|
FOXRED2
|
FAD-dependent oxidoreductase domain containing 2
|
|
chr6_+_41622111
|
0.609
|
NM_001012426
NM_001012427
NM_138457
|
FOXP4
|
forkhead box P4
|
|
chr6_-_44333002
|
0.607
|
NM_178148
|
SLC35B2
|
solute carrier family 35, member B2
|
|
chr8_-_145609737
|
0.606
|
|
SLC39A4
|
solute carrier family 39 (zinc transporter), member 4
|
|
chr17_-_35140307
|
0.606
|
NM_032339
|
C17orf37
|
chromosome 17 open reading frame 37
|
|
chr18_+_9698299
|
0.605
|
|
RAB31
|
RAB31, member RAS oncogene family
|
|
chrX_+_117992623
|
0.603
|
NM_001031855
NM_024778
|
LONRF3
|
LON peptidase N-terminal domain and ring finger 3
|
|
chr19_-_8292273
|
0.600
|
NM_005001
|
NDUFA7
|
NADH dehydrogenase (ubiquinone) 1 alpha subcomplex, 7, 14.5kDa
|
|
chr22_-_17798998
|
0.598
|
|
HIRA
|
HIR histone cell cycle regulation defective homolog A (S. cerevisiae)
|
|
chr19_+_38376980
|
0.597
|
|
LRP3
|
low density lipoprotein receptor-related protein 3
|
|
chr16_-_23429295
|
0.597
|
|
GGA2
|
golgi-associated, gamma adaptin ear containing, ARF binding protein 2
|
|
chr11_-_16992524
|
0.595
|
NM_175058
|
PLEKHA7
|
pleckstrin homology domain containing, family A member 7
|
|
chr1_-_24307514
|
0.590
|
|
MYOM3
|
myomesin family, member 3
|
|
chr6_-_151814861
|
0.590
|
NM_017909
|
RMND1
|
required for meiotic nuclear division 1 homolog (S. cerevisiae)
|
|
chr7_+_55054416
|
0.588
|
|
EGFR
|
epidermal growth factor receptor
|
|
chr19_-_8292245
|
0.585
|
|
NDUFA7
|
NADH dehydrogenase (ubiquinone) 1 alpha subcomplex, 7, 14.5kDa
|
|
chr1_+_160306093
|
0.581
|
NM_001164757
NM_014697
|
NOS1AP
|
nitric oxide synthase 1 (neuronal) adaptor protein
|
|
chr8_+_123863169
|
0.579
|
|
ZHX2
|
zinc fingers and homeoboxes 2
|
|
chr2_-_39859912
|
0.578
|
NM_025264
|
THUMPD2
|
THUMP domain containing 2
|
|
chr8_+_123863075
|
0.578
|
NM_014943
|
ZHX2
|
zinc fingers and homeoboxes 2
|
|
chr1_-_26105954
|
0.575
|
NM_203399
|
STMN1
|
stathmin 1
|
|
chr12_-_101835183
|
0.570
|
|
PAH
|
phenylalanine hydroxylase
|
|
chr1_-_27113018
|
0.568
|
NM_021969
|
NR0B2
|
nuclear receptor subfamily 0, group B, member 2
|
|
chr14_-_23981493
|
0.566
|
NM_020195
|
SDR39U1
|
short chain dehydrogenase/reductase family 39U, member 1
|
|
chr6_-_43305156
|
0.562
|
NM_006443
NM_199184
|
C6orf108
|
chromosome 6 open reading frame 108
|
|
chr11_+_118443738
|
0.559
|
|
VPS11
|
vacuolar protein sorting 11 homolog (S. cerevisiae)
|
|
chr17_-_18848713
|
0.559
|
NM_001039999
|
FAM83G
|
family with sequence similarity 83, member G
|
|
chr17_-_37586716
|
0.557
|
NM_012285
|
KCNH4
|
potassium voltage-gated channel, subfamily H (eag-related), member 4
|
|
chr3_-_4483952
|
0.555
|
|
SUMF1
|
sulfatase modifying factor 1
|
|
chr14_-_93859394
|
0.553
|
NM_001756
|
SERPINA6
|
serpin peptidase inhibitor, clade A (alpha-1 antiproteinase, antitrypsin), member 6
|
|
chr16_+_29726195
|
0.553
|
|
MAZ
|
MYC-associated zinc finger protein (purine-binding transcription factor)
|
|
chr8_+_11599119
|
0.552
|
NM_002052
|
GATA4
|
GATA binding protein 4
|
|
chr16_+_23597691
|
0.551
|
|
PLK1
|
polo-like kinase 1
|
|
chr1_+_237616457
|
0.551
|
|
CHRM3
|
cholinergic receptor, muscarinic 3
|
|
chr20_-_2769245
|
0.550
|
NM_022760
|
FAM113A
|
family with sequence similarity 113, member A
|
|
chr12_-_115660227
|
0.550
|
|
C12orf49
|
chromosome 12 open reading frame 49
|
|
chr7_+_1050704
|
0.549
|
|
|
|
|
chr17_-_74487527
|
0.549
|
NM_005567
|
LGALS3BP
|
lectin, galactoside-binding, soluble, 3 binding protein
|