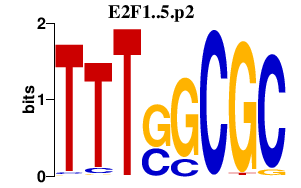
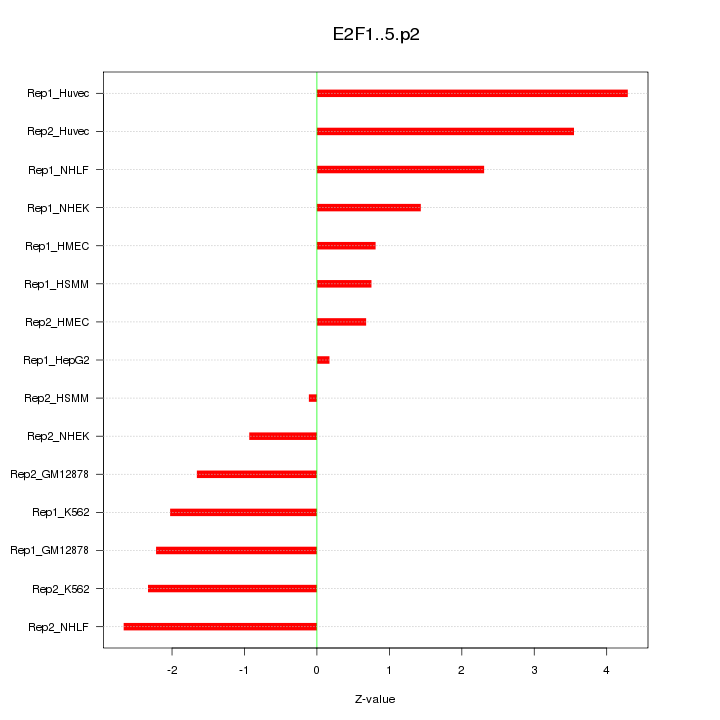
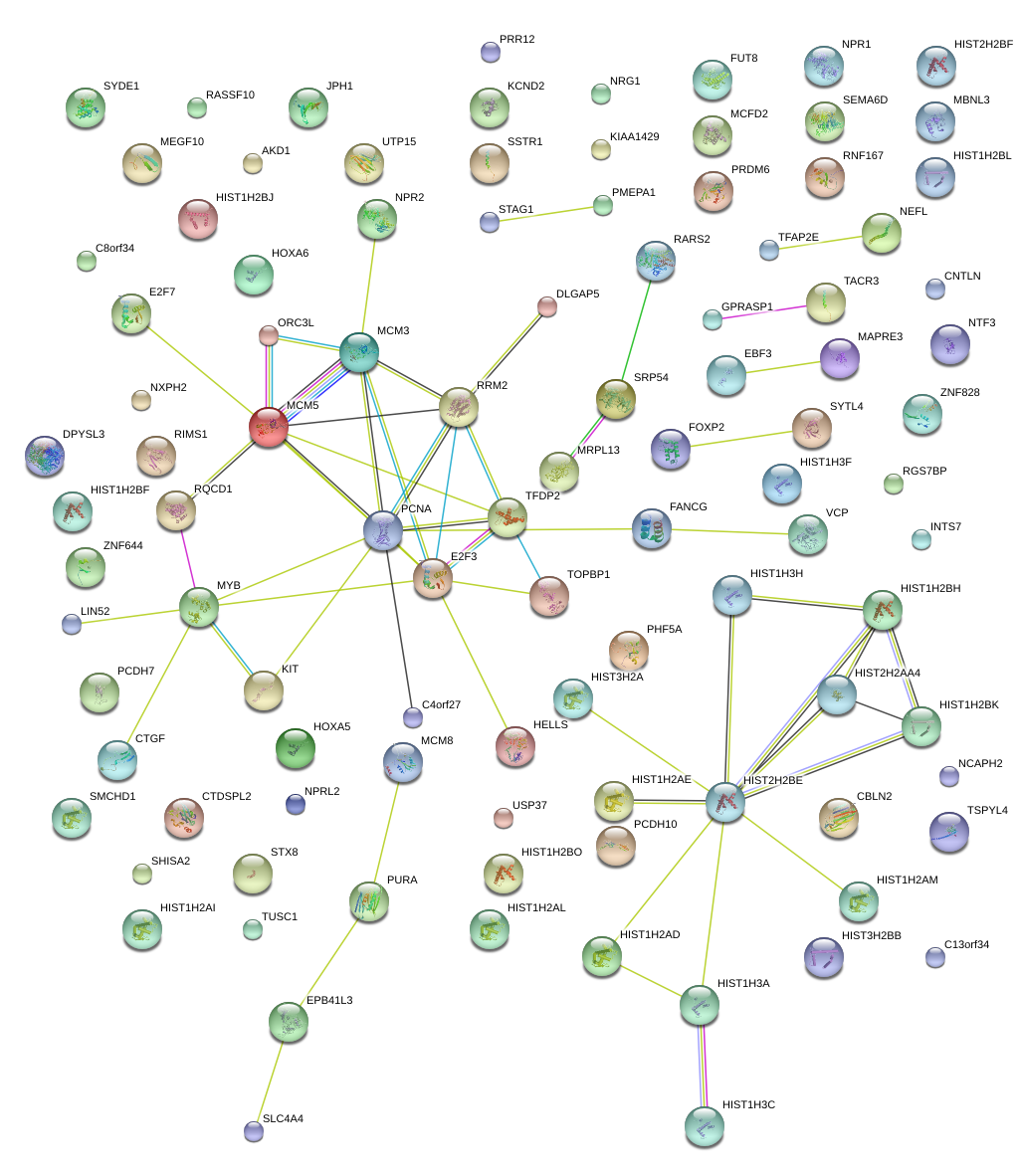
|
chr9_+_17124974
|
3.746
|
NM_001114395
NM_017738
|
CNTLN
|
centlein, centrosomal protein
|
|
chr6_+_72949092
|
2.173
|
|
RIMS1
|
regulating synaptic membrane exocytosis 1
|
|
chr6_-_52257500
|
1.845
|
NM_002388
|
MCM3
|
minichromosome maintenance complex component 3
|
|
chr6_-_52257475
|
1.834
|
|
MCM3
|
minichromosome maintenance complex component 3
|
|
chr5_-_146869613
|
1.737
|
|
DPYSL3
|
dihydropyrimidinase-like 3
|
|
chr6_-_52257431
|
1.698
|
|
MCM3
|
minichromosome maintenance complex component 3
|
|
chr15_+_45263517
|
1.616
|
NM_001198999
|
SEMA6D
|
sema domain, transmembrane domain (TM), and cytoplasmic domain, (semaphorin) 6D
|
|
chr10_+_96295511
|
1.606
|
NM_018063
|
HELLS
|
helicase, lymphoid-specific
|
|
chr7_-_27153892
|
1.573
|
NM_024014
|
HOXA6
|
homeobox A6
|
|
chr6_+_27941012
|
1.509
|
NM_003511
|
HIST1H2AI
HIST1H2AL
|
histone cluster 1, H2ai
histone cluster 1, H2al
|
|
chr5_-_146869811
|
1.464
|
NM_001197294
|
DPYSL3
|
dihydropyrimidinase-like 3
|
|
chr14_+_85069217
|
1.464
|
|
|
|
|
chr6_+_27208763
|
1.370
|
NM_021064
|
HIST1H2AG
|
histone cluster 1, H2ag
|
|
chr8_+_31616809
|
1.344
|
NM_013962
|
NRG1
|
neuregulin 1
|
|
chr10_-_131652007
|
1.313
|
NM_001005463
|
EBF3
|
early B-cell factor 3
|
|
chr20_-_5048443
|
1.280
|
NM_182649
|
PCNA
|
proliferating cell nuclear antigen
|
|
chr6_-_132314004
|
1.263
|
NM_001901
|
CTGF
|
connective tissue growth factor
|
|
chr8_-_24869945
|
1.240
|
NM_006158
|
NEFL
|
neurofilament, light polypeptide
|
|
chrX_-_131451675
|
1.232
|
NM_001170704
|
MBNL3
|
muscleblind-like 3 (Drosophila)
|
|
chr7_+_113513726
|
1.232
|
|
FOXP2
|
forkhead box P2
|
|
chr6_-_27208507
|
1.223
|
NM_021058
|
HIST1H2BJ
|
histone cluster 1, H2bj
|
|
chr15_+_42507123
|
1.223
|
|
CTDSPL2
|
CTD (carboxy-terminal domain, RNA polymerase II, polypeptide A) small phosphatase like 2
|
|
chr16_+_71256497
|
1.216
|
|
|
|
|
chr13_-_25523164
|
1.141
|
NM_001007538
|
SHISA2
|
shisa homolog 2 (Xenopus laevis)
|
|
chr7_+_113513828
|
1.132
|
|
FOXP2
|
forkhead box P2
|
|
chr12_-_75983329
|
1.116
|
NM_203394
|
E2F7
|
E2F transcription factor 7
|
|
chr1_-_226712140
|
1.114
|
NM_033445
|
HIST3H2A
|
histone cluster 3, H2a
|
|
chr2_+_10180304
|
1.091
|
NM_001034
|
RRM2
|
ribonucleotide reductase M2
|
|
chr19_+_54786723
|
1.073
|
NM_020719
|
PRR12
|
proline rich 12
|
|
chr15_+_42507057
|
1.071
|
|
CTDSPL2
|
CTD (carboxy-terminal domain, RNA polymerase II, polypeptide A) small phosphatase like 2
|
|
chr3_-_134863316
|
1.058
|
NM_007027
|
TOPBP1
|
topoisomerase (DNA) II binding protein 1
|
|
chr6_+_26153590
|
1.052
|
NM_003531
|
HIST1H3C
|
histone cluster 1, H3c
|
|
chr18_-_68361742
|
1.044
|
|
CBLN2
|
cerebellin 2 precursor
|
|
chr4_-_104860277
|
1.041
|
NM_001059
|
TACR3
|
tachykinin receptor 3
|
|
chr4_+_72271218
|
1.039
|
NM_001098484
NM_001134742
|
SLC4A4
|
solute carrier family 4, sodium bicarbonate cotransporter, member 4
|
|
chr4_+_134289893
|
1.026
|
NM_020815
NM_032961
|
PCDH10
|
protocadherin 10
|
|
chr18_-_5620629
|
1.020
|
|
EPB41L3
|
erythrocyte membrane protein band 4.1-like 3
|
|
chr22_+_34126154
|
1.011
|
|
MCM5
|
minichromosome maintenance complex component 5
|
|
chr6_+_88356503
|
0.995
|
NM_001197259
NM_012381
NM_181837
|
ORC3
|
origin recognition complex, subunit 3
|
|
chrX_+_101792949
|
0.987
|
NM_001099410
NM_001099411
NM_014710
NM_001184727
|
GPRASP1
|
G protein-coupled receptor associated sorting protein 1
|
|
chr2_-_139254273
|
0.979
|
NM_007226
|
NXPH2
|
neurexophilin 2
|
|
chr9_-_25668230
|
0.978
|
|
TUSC1
|
tumor suppressor candidate 1
|
|
chr3_-_143350898
|
0.978
|
NM_001178138
NM_001178139
|
TFDP2
|
transcription factor Dp-2 (E2F dimerization partner 2)
|
|
chr10_-_131652364
|
0.976
|
|
EBF3
|
early B-cell factor 3
|
|
chr22_+_34126104
|
0.976
|
NM_006739
|
MCM5
|
minichromosome maintenance complex component 5
|
|
chr6_+_135544214
|
0.963
|
|
MYB
|
v-myb myeloblastosis viral oncogene homolog (avian)
|
|
chr22_-_40194602
|
0.959
|
NM_032758
|
PHF5A
|
PHD finger protein 5A
|
|
chr17_+_4784524
|
0.955
|
|
RNF167
|
ring finger protein 167
|
|
chr9_+_35782405
|
0.955
|
NM_003995
|
NPR2
|
natriuretic peptide receptor B/guanylate cyclase B (atrionatriuretic peptide receptor B)
|
|
chr22_+_49293520
|
0.952
|
|
NCAPH2
|
non-SMC condensin II complex, subunit H2
|
|
chr14_+_37748345
|
0.950
|
|
SSTR1
|
somatostatin receptor 1
|
|
chr14_+_73621408
|
0.932
|
NM_001024674
|
LIN52
|
lin-52 homolog (C. elegans)
|
|
chr8_+_69405510
|
0.932
|
NM_001195639
NM_052958
|
C8orf34
|
chromosome 8 open reading frame 34
|
|
chr20_+_5879297
|
0.924
|
NM_032485
NM_182802
|
MCM8
|
minichromosome maintenance complex component 8
|
|
chr2_+_219141761
|
0.915
|
NM_005444
|
RQCD1
|
RCD1 required for cell differentiation1 homolog (S. pombe)
|
|
chr1_+_226712430
|
0.908
|
NM_175055
|
HIST3H2BB
|
histone cluster 3, H2bb
|
|
chr2_-_47022406
|
0.898
|
NM_001171508
NM_001171511
|
MCFD2
|
multiple coagulation factor deficiency 2
|
|
chr8_-_95634845
|
0.890
|
NM_015496
NM_183009
|
KIAA1429
|
KIAA1429
|
|
chr6_+_135544138
|
0.890
|
NM_001130172
NM_001130173
NM_001161656
NM_001161657
NM_001161658
NM_001161659
NM_001161660
NM_005375
|
MYB
|
v-myb myeloblastosis viral oncogene homolog (avian)
|
|
chr5_+_63838183
|
0.887
|
NM_001029875
|
RGS7BP
|
regulator of G-protein signaling 7 binding protein
|
|
chr5_+_72897323
|
0.886
|
NM_032175
|
UTP15
|
UTP15, U3 small nucleolar ribonucleoprotein, homolog (S. cerevisiae)
|
|
chr7_+_113513600
|
0.885
|
|
FOXP2
|
forkhead box P2
|
|
chr9_-_35069520
|
0.867
|
|
FANCG
|
Fanconi anemia, complementation group G
|
|
chr5_+_122452739
|
0.840
|
NM_001136239
|
PRDM6
|
PR domain containing 6
|
|
chr6_+_27883877
|
0.839
|
NM_003509
|
HIST1H2AI
HIST1H3F
|
histone cluster 1, H2ai
histone cluster 1, H3f
|
|
chr4_-_147079410
|
0.825
|
|
|
|
|
chr6_+_20510019
|
0.820
|
NM_001949
|
E2F3
|
E2F transcription factor 3
|
|
chr7_-_27149750
|
0.812
|
NM_019102
|
HOXA5
|
homeobox A5
|
|
chr10_+_22580992
|
0.801
|
|
|
|
|
chr11_+_12987455
|
0.798
|
NM_001080521
|
RASSF10
|
Ras association (RalGDS/AF-6) domain family (N-terminal) member 10
|
|
chr8_-_75396016
|
0.792
|
NM_020647
|
JPH1
|
junctophilin 1
|
|
chr6_-_26358811
|
0.782
|
NM_021018
|
HIST1H3F
|
histone cluster 1, H3f
|
|
chr8_-_121526568
|
0.779
|
|
MRPL13
|
mitochondrial ribosomal protein L13
|
|
chr13_+_114098045
|
0.758
|
NM_001164144
NM_001164145
NM_032436
|
ZNF828
|
zinc finger protein 828
|
|
chr4_+_134292630
|
0.754
|
|
PCDH10
|
protocadherin 10
|
|
chr14_-_54727958
|
0.750
|
NM_001146015
NM_014750
|
DLGAP5
|
discs, large (Drosophila) homolog-associated protein 5
|
|
chr6_-_27883662
|
0.738
|
NM_003519
|
HIST1H2BL
|
histone cluster 1, H2bl
|
|
chr19_+_15079141
|
0.734
|
NM_033025
|
SYDE1
|
synapse defective 1, Rho GTPase, homolog 1 (C. elegans)
|
|
chr6_-_116681863
|
0.733
|
NM_021648
|
TSPYL4
|
TSPY-like 4
|
|
chr5_+_139473873
|
0.727
|
NM_005859
|
PURA
|
purine-rich element binding protein A
|
|
chr3_-_137953876
|
0.726
|
NM_005862
|
STAG1
|
stromal antigen 1
|
|
chrX_-_99873152
|
0.726
|
NM_001174068
NM_080737
|
SYTL4
|
synaptotagmin-like 4
|
|
chr1_+_35811557
|
0.708
|
NM_178548
|
TFAP2E
|
transcription factor AP-2 epsilon (activating enhancer binding protein 2 epsilon)
|
|
chr22_+_34126294
|
0.706
|
|
MCM5
|
minichromosome maintenance complex component 5
|
|
chr13_+_72199981
|
0.706
|
NM_024808
|
C13orf34
|
chromosome 13 open reading frame 34
|
|
chr4_-_170915602
|
0.704
|
NM_017867
|
C4orf27
|
chromosome 4 open reading frame 27
|
|
chr12_+_5411534
|
0.699
|
NM_001102654
|
NTF3
|
neurotrophin 3
|
|
chr17_-_9419864
|
0.693
|
NM_004853
|
STX8
|
syntaxin 8
|
|
chr6_-_88356445
|
0.692
|
NM_020320
|
RARS2
|
arginyl-tRNA synthetase 2, mitochondrial
|
|
chr8_-_121526514
|
0.690
|
|
MRPL13
|
mitochondrial ribosomal protein L13
|
|
chr4_+_30331543
|
0.689
|
|
PCDH7
|
protocadherin 7
|
|
chr1_-_210275402
|
0.688
|
NM_015434
|
INTS7
|
integrator complex subunit 7
|
|
chr5_+_126654354
|
0.688
|
NM_032446
|
MEGF10
|
multiple EGF-like-domains 10
|
|
chr2_-_219141199
|
0.686
|
NM_020935
|
USP37
|
ubiquitin specific peptidase 37
|
|
chr4_+_55218663
|
0.684
|
NM_000222
NM_001093772
|
KIT
|
v-kit Hardy-Zuckerman 4 feline sarcoma viral oncogene homolog
|
|
chr7_+_119700924
|
0.682
|
NM_012281
|
KCND2
|
potassium voltage-gated channel, Shal-related subfamily, member 2
|
|
chr1_-_148050534
|
0.681
|
NM_001024599
NM_001161334
|
HIST2H2BF
|
histone cluster 2, H2bf
|
|
chr6_+_135544170
|
0.680
|
|
MYB
|
v-myb myeloblastosis viral oncogene homolog (avian)
|
|
chr1_-_91259601
|
0.677
|
NM_016620
|
ZNF644
|
zinc finger protein 644
|
|
chr14_+_34521945
|
0.677
|
|
SRP54
|
signal recognition particle 54kDa
|
|
chr14_+_64949447
|
0.675
|
|
FUT8
|
fucosyltransferase 8 (alpha (1,6) fucosyltransferase)
|
|
chrX_-_103386215
|
0.668
|
NM_153448
|
ESX1
|
ESX homeobox 1
|
|
chr15_-_81743198
|
0.661
|
|
BNC1
|
basonuclin 1
|
|
chr3_-_123036597
|
0.659
|
NM_001023570
NM_001023571
|
IQCB1
|
IQ motif containing B1
|
|
chr9_-_4289574
|
0.659
|
|
GLIS3
|
GLIS family zinc finger 3
|
|
chr12_-_47396724
|
0.651
|
NM_001240
|
CCNT1
|
cyclin T1
|
|
chr1_-_200946038
|
0.644
|
NM_177402
|
SYT2
|
synaptotagmin II
|
|
chr2_+_15998090
|
0.640
|
NM_005378
|
MYCN
|
v-myc myelocytomatosis viral related oncogene, neuroblastoma derived (avian)
|
|
chr21_-_27139143
|
0.637
|
|
ADAMTS1
|
ADAM metallopeptidase with thrombospondin type 1 motif, 1
|
|
chr12_-_107479294
|
0.634
|
NM_014706
|
SART3
|
squamous cell carcinoma antigen recognized by T cells 3
|
|
chr2_+_10180145
|
0.633
|
NM_001165931
|
RRM2
|
ribonucleotide reductase M2
|
|
chr2_-_160627366
|
0.631
|
NM_001007267
NM_001195641
NM_007366
|
PLA2R1
|
phospholipase A2 receptor 1, 180kDa
|
|
chr10_-_105604942
|
0.631
|
NM_014631
|
SH3PXD2A
|
SH3 and PX domains 2A
|
|
chr6_-_88356411
|
0.630
|
|
RARS2
|
arginyl-tRNA synthetase 2, mitochondrial
|
|
chr6_-_27222556
|
0.629
|
NM_080593
|
HIST1H2BK
|
histone cluster 1, H2bk
|
|
chr6_+_90596348
|
0.629
|
|
CASP8AP2
|
caspase 8 associated protein 2
|
|
chr2_+_15998010
|
0.628
|
|
MYCN
|
v-myc myelocytomatosis viral related oncogene, neuroblastoma derived (avian)
|
|
chr22_+_49293493
|
0.626
|
NM_001185011
NM_014551
NM_152299
|
NCAPH2
|
non-SMC condensin II complex, subunit H2
|
|
chr3_+_64645819
|
0.624
|
|
LOC100507098
|
hypothetical LOC100507098
|
|
chr7_+_120378052
|
0.624
|
NM_019071
NM_198267
|
ING3
|
inhibitor of growth family, member 3
|
|
chr9_-_35069751
|
0.619
|
NM_004629
|
FANCG
|
Fanconi anemia, complementation group G
|
|
chr2_-_10747553
|
0.617
|
NM_024894
|
NOL10
|
nucleolar protein 10
|
|
chr3_+_135996990
|
0.615
|
|
EPHB1
|
EPH receptor B1
|
|
chr6_+_43036442
|
0.614
|
NM_018960
|
GNMT
|
glycine N-methyltransferase
|
|
chr18_+_52469559
|
0.613
|
NM_015285
NM_052834
|
WDR7
|
WD repeat domain 7
|
|
chr18_-_53621277
|
0.613
|
NM_005603
|
ATP8B1
|
ATPase, aminophospholipid transporter, class I, type 8B, member 1
|
|
chr3_-_10003512
|
0.611
|
NM_018447
|
TMEM111
|
transmembrane protein 111
|
|
chr14_+_35365266
|
0.611
|
NM_032352
|
BRMS1L
|
breast cancer metastasis-suppressor 1-like
|
|
chr3_-_51950919
|
0.610
|
NM_004704
|
RRP9
|
ribosomal RNA processing 9, small subunit (SSU) processome component, homolog (yeast)
|
|
chr17_-_9419849
|
0.607
|
|
STX8
|
syntaxin 8
|
|
chr5_-_139760838
|
0.607
|
|
|
|
|
chr9_-_112840064
|
0.605
|
NM_001401
NM_057159
|
LPAR1
|
lysophosphatidic acid receptor 1
|
|
chr5_+_1854493
|
0.605
|
NM_004553
|
NDUFS6
|
NADH dehydrogenase (ubiquinone) Fe-S protein 6, 13kDa (NADH-coenzyme Q reductase)
|
|
chr18_+_11841388
|
0.603
|
NM_020412
|
CHMP1B
|
chromatin modifying protein 1B
|
|
chr9_-_21984414
|
0.598
|
NM_058195
|
CDKN2A
|
cyclin-dependent kinase inhibitor 2A (melanoma, p16, inhibits CDK4)
|
|
chr2_-_164300758
|
0.597
|
NM_018086
|
FIGN
|
fidgetin
|
|
chr11_-_62379788
|
0.595
|
|
SNHG1
|
small nucleolar RNA host gene 1 (non-protein coding)
|
|
chr8_-_120937215
|
0.592
|
NM_024094
|
DSCC1
|
defective in sister chromatid cohesion 1 homolog (S. cerevisiae)
|
|
chr16_+_66153810
|
0.592
|
NM_001191022
NM_006565
|
CTCF
|
CCCTC-binding factor (zinc finger protein)
|
|
chr3_-_10003344
|
0.591
|
|
TMEM111
|
transmembrane protein 111
|
|
chr1_-_100370994
|
0.588
|
NM_194292
|
SASS6
|
spindle assembly 6 homolog (C. elegans)
|
|
chr8_-_121526806
|
0.587
|
NM_014078
|
MRPL13
|
mitochondrial ribosomal protein L13
|
|
chr13_+_33290254
|
0.583
|
|
RFC3
|
replication factor C (activator 1) 3, 38kDa
|
|
chr2_+_219141546
|
0.581
|
|
RQCD1
|
RCD1 required for cell differentiation1 homolog (S. pombe)
|
|
chr4_-_130233855
|
0.579
|
NM_144643
|
SCLT1
|
sodium channel and clathrin linker 1
|
|
chr11_-_19219000
|
0.571
|
NM_024680
|
E2F8
|
E2F transcription factor 8
|
|
chr5_+_50714714
|
0.569
|
NM_002202
|
ISL1
|
ISL LIM homeobox 1
|
|
chr2_+_131966970
|
0.567
|
|
LOC150776
|
sphingomyelin phosphodiesterase 4, neutral membrane pseudogene
|
|
chr1_-_26058434
|
0.567
|
NM_024037
|
C1orf135
|
chromosome 1 open reading frame 135
|
|
chr22_+_22639025
|
0.566
|
NM_001084393
|
DDTL
|
D-dopachrome tautomerase-like
|
|
chr5_+_94981735
|
0.565
|
NM_199243
|
GPR150
|
G protein-coupled receptor 150
|
|
chr6_-_27914095
|
0.563
|
NM_003510
|
HIST1H2AK
|
histone cluster 1, H2ak
|
|
chr1_-_68470585
|
0.559
|
NM_001002292
NM_001193334
NM_024911
|
WLS
|
wntless homolog (Drosophila)
|
|
chr14_-_64040072
|
0.554
|
NM_006977
|
ZBTB25
|
zinc finger and BTB domain containing 25
|
|
chr1_-_147665783
|
0.553
|
|
HIST2H2BF
|
histone cluster 2, H2bf
|
|
chr14_+_92743381
|
0.553
|
|
UBR7
|
ubiquitin protein ligase E3 component n-recognin 7 (putative)
|
|
chr12_-_47532171
|
0.551
|
NM_004818
|
DDX23
|
DEAD (Asp-Glu-Ala-Asp) box polypeptide 23
|
|
chr4_+_30330963
|
0.545
|
NM_001173523
NM_002589
NM_032456
NM_032457
|
PCDH7
|
protocadherin 7
|
|
chr11_+_62380042
|
0.544
|
NM_001012662
NM_001012664
NM_002394
NM_001012661
NM_001012663
|
SLC3A2
|
solute carrier family 3 (activators of dibasic and neutral amino acid transport), member 2
|
|
chr3_-_62334229
|
0.543
|
NM_018008
|
FEZF2
|
FEZ family zinc finger 2
|
|
chr6_+_90596290
|
0.541
|
NM_001137667
NM_001137668
NM_012115
|
CASP8AP2
|
caspase 8 associated protein 2
|
|
chr19_-_57082907
|
0.541
|
NM_001135590
NM_032679
|
ZNF577
|
zinc finger protein 577
|
|
chr2_-_10747438
|
0.540
|
|
NOL10
|
nucleolar protein 10
|
|
chr13_+_102047286
|
0.540
|
NM_003291
|
TPP2
|
tripeptidyl peptidase II
|
|
chr8_+_121526823
|
0.536
|
NM_022045
|
MTBP
|
Mdm2, transformed 3T3 cell double minute 2, p53 binding protein (mouse) binding protein, 104kDa
|
|
chr7_+_121300446
|
0.529
|
|
PTPRZ1
|
protein tyrosine phosphatase, receptor-type, Z polypeptide 1
|
|
chrX_-_62891066
|
0.523
|
|
ARHGEF9
|
Cdc42 guanine nucleotide exchange factor (GEF) 9
|
|
chr20_-_5879036
|
0.522
|
NM_015939
|
TRMT6
|
tRNA methyltransferase 6 homolog (S. cerevisiae)
|
|
chr19_-_10305186
|
0.522
|
NM_133452
|
RAVER1
|
ribonucleoprotein, PTB-binding 1
|
|
chr1_-_119333701
|
0.521
|
NM_152380
|
TBX15
|
T-box 15
|
|
chr12_-_55013987
|
0.521
|
NM_001166279
NM_014871
|
PAN2
|
PAN2 poly(A) specific ribonuclease subunit homolog (S. cerevisiae)
|
|
chr11_+_118374071
|
0.518
|
|
CCDC84
|
coiled-coil domain containing 84
|
|
chr9_-_23811808
|
0.516
|
|
ELAVL2
|
ELAV (embryonic lethal, abnormal vision, Drosophila)-like 2 (Hu antigen B)
|
|
chr4_-_74154272
|
0.515
|
NM_173827
|
COX18
|
COX18 cytochrome c oxidase assembly homolog (S. cerevisiae)
|
|
chr2_-_667367
|
0.513
|
NM_152834
|
TMEM18
|
transmembrane protein 18
|
|
chr5_+_96024434
|
0.513
|
|
CAST
|
calpastatin
|
|
chr2_-_47022374
|
0.511
|
|
MCFD2
|
multiple coagulation factor deficiency 2
|
|
chrX_+_130984912
|
0.510
|
NM_001042453
NM_016542
|
MST4
|
serine/threonine protein kinase MST4
|
|
chr6_-_27968906
|
0.506
|
NM_003514
|
HIST1H2AM
|
histone cluster 1, H2am
|
|
chr10_+_112317401
|
0.506
|
NM_005445
|
SMC3
|
structural maintenance of chromosomes 3
|
|
chr8_-_132122016
|
0.505
|
NM_001115
|
ADCY8
|
adenylate cyclase 8 (brain)
|
|
chr8_+_69026895
|
0.502
|
|
PREX2
|
phosphatidylinositol-3,4,5-trisphosphate-dependent Rac exchange factor 2
|
|
chr9_-_23811842
|
0.499
|
NM_001171195
|
ELAVL2
|
ELAV (embryonic lethal, abnormal vision, Drosophila)-like 2 (Hu antigen B)
|
|
chr9_+_85785456
|
0.499
|
NM_024945
|
RMI1
|
RMI1, RecQ mediated genome instability 1, homolog (S. cerevisiae)
|
|
chr3_-_49868889
|
0.498
|
NM_005879
|
TRAIP
|
TRAF interacting protein
|
|
chr9_-_21984379
|
0.497
|
|
CDKN2A
|
cyclin-dependent kinase inhibitor 2A (melanoma, p16, inhibits CDK4)
|
|
chr9_-_78710698
|
0.495
|
NM_015225
|
PRUNE2
|
prune homolog 2 (Drosophila)
|
|
chr3_+_148609841
|
0.490
|
NM_003412
|
ZIC1
|
Zic family member 1 (odd-paired homolog, Drosophila)
|
|
chr6_+_26333361
|
0.488
|
NM_003532
|
HIST1H3E
|
histone cluster 1, H3e
|
|
chr10_+_75580914
|
0.487
|
NM_006721
|
ADK
|
adenosine kinase
|
|
chr8_+_69027166
|
0.486
|
|
PREX2
|
phosphatidylinositol-3,4,5-trisphosphate-dependent Rac exchange factor 2
|
|
chrX_+_101887562
|
0.485
|
NM_001142529
NM_001142530
|
BHLHB9
|
basic helix-loop-helix domain containing, class B, 9
|
|
chr10_-_123724251
|
0.483
|
|
NSMCE4A
|
non-SMC element 4 homolog A (S. cerevisiae)
|
|
chr13_+_44592647
|
0.481
|
|
GTF2F2
|
general transcription factor IIF, polypeptide 2, 30kDa
|
|
chr1_-_29321599
|
0.479
|
NM_001171868
|
TMEM200B
|
transmembrane protein 200B
|
|
chr3_+_10003579
|
0.478
|
|
LOC442075
|
hypothetical LOC442075
|
|
chr2_+_188865796
|
0.478
|
|
GULP1
|
GULP, engulfment adaptor PTB domain containing 1
|
|
chr13_+_97426713
|
0.476
|
|
IPO5
|
importin 5
|
|
chr17_-_34860840
|
0.475
|
NM_004774
|
MED1
|
mediator complex subunit 1
|
|
chr5_-_37406643
|
0.474
|
NM_004298
|
NUP155
|
nucleoporin 155kDa
|