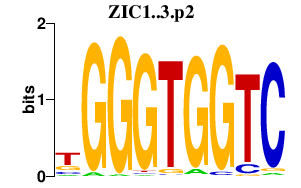
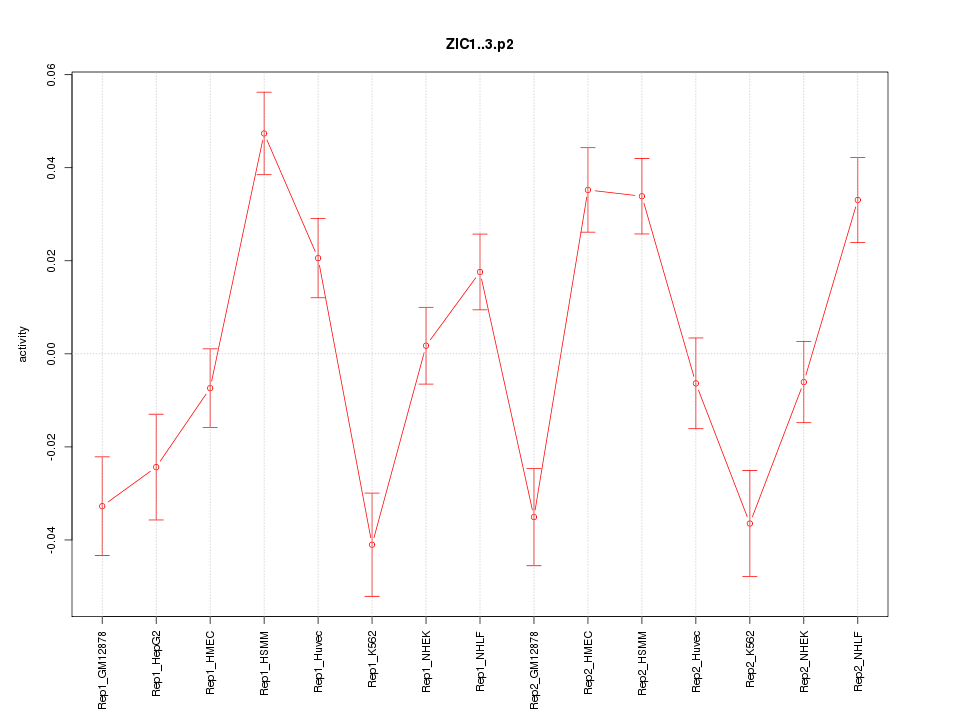
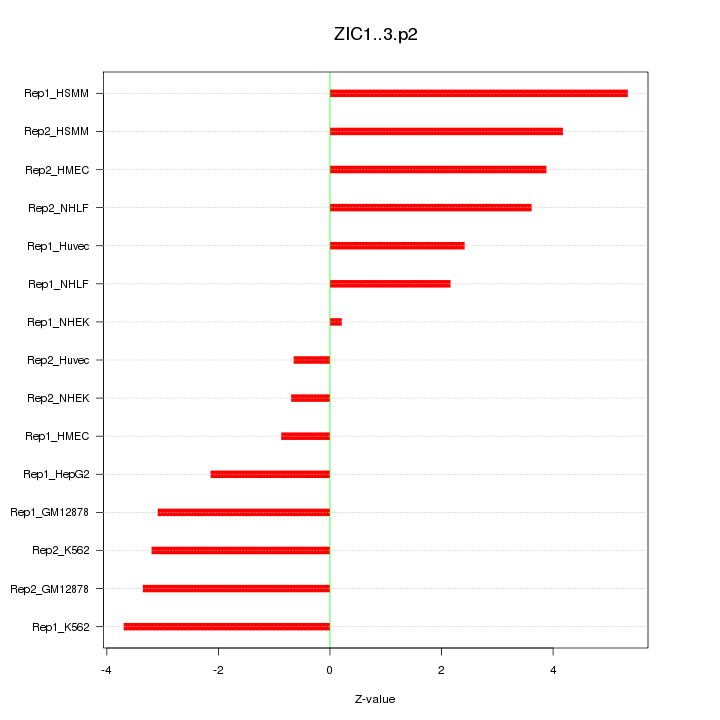
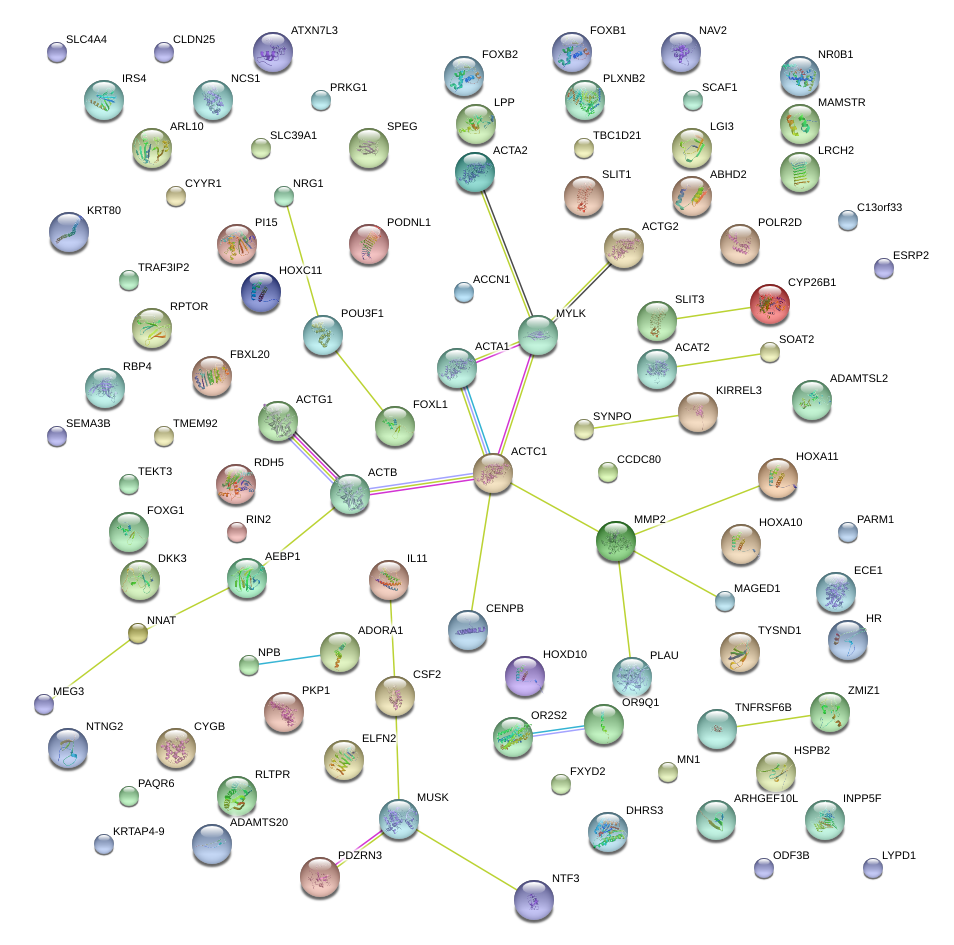
|
chr16_+_85169615
|
6.303
|
NM_005250
|
FOXL1
|
forkhead box L1
|
|
chr10_-_90702490
|
5.907
|
NM_001613
|
ACTA2
|
actin, alpha 2, smooth muscle, aorta
|
|
chr19_-_13910167
|
5.031
|
NM_024825
|
PODNL1
|
podocan-like 1
|
|
chr3_-_73756668
|
5.002
|
NM_015009
|
PDZRN3
|
PDZ domain containing ring finger 3
|
|
chr12_-_50871953
|
4.330
|
NM_001081492
NM_182507
|
KRT80
|
keratin 80
|
|
chr3_+_64645819
|
3.745
|
|
LOC100507098
|
hypothetical LOC100507098
|
|
chr17_-_72045218
|
3.182
|
NM_134268
|
CYGB
|
cytoglobin
|
|
chr16_+_66244314
|
3.172
|
|
RLTPR
|
RGD motif, leucine rich repeats, tropomodulin domain and proline-rich containing
|
|
chr21_-_26867138
|
3.164
|
NM_052954
|
CYYR1
|
cysteine/tyrosine-rich 1
|
|
chr7_-_27191287
|
2.932
|
NM_005523
|
HOXA11
|
homeobox A11
|
|
chr5_+_131437383
|
2.921
|
NM_000758
|
CSF2
|
colony stimulating factor 2 (granulocyte-macrophage)
|
|
chr22_-_36153449
|
2.897
|
NM_052906
|
ELFN2
|
extracellular leucine-rich repeat and fibronectin type III domain containing 2
|
|
chr7_-_27180398
|
2.887
|
NM_018951
|
HOXA10
|
homeobox A10
|
|
chr3_+_50281582
|
2.852
|
|
SEMA3B
|
sema domain, immunoglobulin domain (Ig), short basic domain, secreted, (semaphorin) 3B
|
|
chr5_-_168660293
|
2.832
|
NM_003062
|
SLIT3
|
slit homolog 3 (Drosophila)
|
|
chr6_-_112001590
|
2.817
|
NM_001164282
|
TRAF3IP2
|
TRAF3 interacting protein 2
|
|
chr10_+_75343352
|
2.719
|
|
PLAU
|
plasminogen activator, urokinase
|
|
chr1_-_21478571
|
2.711
|
NM_001113347
|
ECE1
|
endothelin converting enzyme 1
|
|
chr5_+_175725050
|
2.677
|
NM_173664
|
ARL10
|
ADP-ribosylation factor-like 10
|
|
chr22_-_49063088
|
2.620
|
|
PLXNB2
|
plexin B2
|
|
chr7_+_44110527
|
2.561
|
|
AEBP1
|
AE binding protein 1
|
|
chr19_-_53912025
|
2.519
|
NM_182574
|
MAMSTR
|
MEF2 activating motif and SAP domain containing transcriptional regulator
|
|
chr1_+_199519202
|
2.430
|
NM_000299
NM_001005337
|
PKP1
|
plakophilin 1 (ectodermal dysplasia/skin fragility syndrome)
|
|
chr2_-_133144094
|
2.414
|
|
LYPD1
|
LY6/PLAUR domain containing 1
|
|
chr17_+_36515166
|
2.396
|
NM_001146041
|
KRTAP4-9
|
keratin associated protein 4-9
|
|
chr8_-_22070208
|
2.379
|
NM_139278
|
LGI3
|
leucine-rich repeat LGI family, member 3
|
|
chr17_+_76549658
|
2.356
|
|
RPTOR
|
regulatory associated protein of MTOR, complex 1
|
|
chr3_+_50281309
|
2.309
|
|
SEMA3B
|
sema domain, immunoglobulin domain (Ig), short basic domain, secreted, (semaphorin) 3B
|
|
chr17_+_45703765
|
2.258
|
NM_001168215
|
TMEM92
|
transmembrane protein 92
|
|
chr6_-_169392771
|
2.204
|
|
|
|
|
chr3_-_124893875
|
2.198
|
|
MYLK
|
myosin light chain kinase
|
|
chr5_+_150008030
|
2.185
|
|
SYNPO
|
synaptopodin
|
|
chr3_+_189353502
|
2.170
|
|
LPP
|
LIM domain containing preferred translocation partner in lipoma
|
|
chr19_-_13808101
|
2.136
|
|
LOC284454
|
hypothetical LOC284454
|
|
chr20_+_19903778
|
2.136
|
|
RIN2
|
Ras and Rab interactor 2
|
|
chr11_+_57547928
|
2.126
|
NM_001005212
|
OR9Q1
|
olfactory receptor, family 9, subfamily Q, member 1
|
|
chr16_+_54070302
|
2.113
|
NM_004530
|
MMP2
|
matrix metallopeptidase 2 (gelatinase A, 72kDa gelatinase, 72kDa type IV collagenase)
|
|
chr8_+_31616809
|
2.103
|
NM_013962
|
NRG1
|
neuregulin 1
|
|
chr10_+_80672842
|
2.100
|
|
ZMIZ1
|
zinc finger, MIZ-type containing 1
|
|
chrX_+_51653348
|
2.053
|
NM_001005333
NM_006986
|
MAGED1
|
melanoma antigen family D, 1
|
|
chr8_+_75899326
|
2.011
|
NM_015886
|
PI15
|
peptidase inhibitor 15
|
|
chr12_+_54400417
|
2.007
|
NM_002905
|
RDH5
|
retinol dehydrogenase 5 (11-cis/9-cis)
|
|
chr6_+_160102978
|
2.003
|
NM_005891
|
ACAT2
|
acetyl-CoA acetyltransferase 2
|
|
chr19_-_60573552
|
1.993
|
NM_000641
|
IL11
|
interleukin 11
|
|
chr17_-_28644118
|
1.990
|
NM_183377
|
ACCN1
|
amiloride-sensitive cation channel 1, neuronal
|
|
chr16_-_66827449
|
1.986
|
NM_024939
|
ESRP2
|
epithelial splicing regulatory protein 2
|
|
chr22_-_26527469
|
1.967
|
NM_002430
|
MN1
|
meningioma (disrupted in balanced translocation) 1
|
|
chr11_+_111288686
|
1.958
|
|
HSPB2-C11ORF52
HSPB2
|
HSPB2-C11orf52 read-through transcript
heat shock 27kDa protein 2
|
|
chrX_-_107866262
|
1.944
|
NM_003604
|
IRS4
|
insulin receptor substrate 4
|
|
chr8_-_22044463
|
1.929
|
NM_005144
NM_018411
|
HR
|
hairless homolog (mouse)
|
|
chr12_+_5411534
|
1.920
|
NM_001102654
|
NTF3
|
neurotrophin 3
|
|
chr10_+_52421120
|
1.902
|
|
PRKG1
|
protein kinase, cGMP-dependent, type I
|
|
chr6_+_101948102
|
1.901
|
|
|
|
|
chr20_+_35583020
|
1.895
|
NM_005386
NM_181689
|
NNAT
|
neuronatin
|
|
chr4_+_72271218
|
1.895
|
NM_001098484
NM_001134742
|
SLC4A4
|
solute carrier family 4, sodium bicarbonate cotransporter, member 4
|
|
chr17_-_15185623
|
1.885
|
NM_031898
|
TEKT3
|
tektin 3
|
|
chr11_-_126375864
|
1.868
|
|
KIRREL3
|
kin of IRRE like 3 (Drosophila)
|
|
chr11_-_11986704
|
1.858
|
NM_001018057
|
DKK3
|
dickkopf homolog 3 (Xenopus laevis)
|
|
chr9_-_35948150
|
1.852
|
NM_019897
|
OR2S2
|
olfactory receptor, family 2, subfamily S, member 2
|
|
chr20_+_61798512
|
1.847
|
|
TNFRSF6B
|
tumor necrosis factor receptor superfamily, member 6b, decoy
|
|
chr20_+_61798446
|
1.846
|
NM_003823
|
TNFRSF6B
|
tumor necrosis factor receptor superfamily, member 6b, decoy
|
|
chr1_+_201363458
|
1.839
|
NM_000674
NM_001048230
|
ADORA1
|
adenosine A1 receptor
|
|
chr19_+_54837181
|
1.834
|
NM_021228
|
SCAF1
|
SR-related CTD-associated factor 1
|
|
chr1_-_154484341
|
1.800
|
NM_198406
NM_024897
|
PAQR6
|
progestin and adipoQ receptor family member VI
|
|
chr17_-_40674883
|
1.789
|
|
|
|
|
chr15_+_87540238
|
1.789
|
|
ABHD2
|
abhydrolase domain containing 2
|
|
chr9_+_135389698
|
1.774
|
NM_014694
|
ADAMTSL2
|
ADAMTS-like 2
|
|
chr13_+_30378311
|
1.760
|
NM_032849
|
C13orf33
|
chromosome 13 open reading frame 33
|
|
chr11_+_111288669
|
1.747
|
NM_001541
|
HSPB2-C11ORF52
HSPB2
|
HSPB2-C11orf52 read-through transcript
heat shock 27kDa protein 2
|
|
chr6_+_160103052
|
1.737
|
|
ACAT2
|
acetyl-CoA acetyltransferase 2
|
|
chr12_-_42231990
|
1.713
|
NM_025003
|
ADAMTS20
|
ADAM metallopeptidase with thrombospondin type 1 motif, 20
|
|
chr9_+_112470871
|
1.702
|
NM_001166280
NM_001166281
NM_005592
|
MUSK
|
muscle, skeletal, receptor tyrosine kinase
|
|
chr5_-_92982927
|
1.689
|
|
|
|
|
chr11_+_19328846
|
1.671
|
NM_001111018
|
NAV2
|
neuron navigator 2
|
|
chr1_+_17817397
|
1.668
|
|
ARHGEF10L
|
Rho guanine nucleotide exchange factor (GEF) 10-like
|
|
chr2_-_72228427
|
1.665
|
NM_019885
|
CYP26B1
|
cytochrome P450, family 26, subfamily B, polypeptide 1
|
|
chr14_+_100362236
|
1.644
|
|
MEG3
|
maternally expressed 3 (non-protein coding)
|
|
chr7_+_27191534
|
1.640
|
|
HOXA11-AS1
|
HOXA11 antisense RNA 1 (non-protein coding)
|
|
chr11_-_126375895
|
1.634
|
NM_001161707
NM_032531
|
KIRREL3
|
kin of IRRE like 3 (Drosophila)
|
|
chr1_-_12599934
|
1.625
|
NM_004753
|
DHRS3
|
dehydrogenase/reductase (SDR family) member 3
|
|
chr22_+_35290271
|
1.624
|
|
|
|
|
chr10_-_71575953
|
1.620
|
|
TYSND1
|
trypsin domain containing 1
|
|
chr14_+_100362204
|
1.617
|
|
MEG3
|
maternally expressed 3 (non-protein coding)
|
|
chr15_+_71953020
|
1.610
|
NM_153356
|
TBC1D21
|
TBC1 domain family, member 21
|
|
chr14_+_28306633
|
1.606
|
|
FOXG1
|
forkhead box G1
|
|
chr17_-_39632845
|
1.600
|
|
ATXN7L3
|
ataxin 7-like 3
|
|
chr3_-_113841200
|
1.593
|
|
CCDC80
|
coiled-coil domain containing 80
|
|
chr1_-_152202574
|
1.592
|
|
SLC39A1
|
solute carrier family 39 (zinc transporter), member 1
|
|
chr9_+_132002692
|
1.585
|
NM_001128826
|
NCS1
|
neuronal calcium sensor 1
|
|
chr22_-_49317408
|
1.581
|
|
ODF3B
|
outer dense fiber of sperm tails 3B
|
|
chr11_+_113155727
|
1.580
|
NM_001101389
|
CLDN25
|
claudin 25
|
|
chr2_+_220033777
|
1.569
|
NM_001173476
|
SPEG
|
SPEG complex locus
|
|
chr11_-_117200618
|
1.560
|
NM_001127489
NM_001680
|
FXYD2
|
FXYD domain containing ion transport regulator 2
|
|
chr15_+_58083798
|
1.557
|
|
FOXB1
|
forkhead box B1
|
|
chr12_+_52653176
|
1.544
|
NM_014212
|
HOXC11
|
homeobox C11
|
|
chr19_-_13909877
|
1.542
|
|
PODNL1
|
podocan-like 1
|
|
chr14_+_104218487
|
1.522
|
|
|
|
|
chr9_+_78824390
|
1.508
|
NM_001013735
|
FOXB2
|
forkhead box B2
|
|
chr20_-_3713509
|
1.496
|
|
CENPB
|
centromere protein B, 80kDa
|
|
chr9_+_134026836
|
1.493
|
|
NTNG2
|
netrin G2
|
|
chr4_+_76077229
|
1.489
|
NM_015393
|
PARM1
|
prostate androgen-regulated mucin-like protein 1
|
|
chr1_-_38285033
|
1.464
|
NM_002699
|
POU3F1
|
POU class 3 homeobox 1
|
|
chrX_-_30237403
|
1.464
|
NM_000475
|
NR0B1
|
nuclear receptor subfamily 0, group B, member 1
|
|
chr10_-_95350949
|
1.464
|
NM_006744
|
RBP4
|
retinol binding protein 4, plasma
|
|
chr17_-_34811377
|
1.464
|
NM_001184906
NM_032875
|
FBXL20
|
F-box and leucine-rich repeat protein 20
|
|
chr7_+_44110470
|
1.461
|
NM_001129
|
AEBP1
|
AE binding protein 1
|
|
chrX_-_114374860
|
1.460
|
NM_020871
|
LRCH2
|
leucine-rich repeats and calponin homology (CH) domain containing 2
|
|
chr5_+_147775652
|
1.459
|
|
FBXO38
|
F-box protein 38
|
|
chr2_+_233605625
|
1.435
|
NM_005383
|
NEU2
|
sialidase 2 (cytosolic sialidase)
|
|
chr6_+_41714171
|
1.431
|
NM_005586
|
MDFI
|
MyoD family inhibitor
|
|
chr17_-_19231049
|
1.428
|
NM_001198695
NM_002404
|
MFAP4
|
microfibrillar-associated protein 4
|
|
chr7_+_47661366
|
1.427
|
NM_001123065
|
C7orf65
|
chromosome 7 open reading frame 65
|
|
chr17_-_45629401
|
1.427
|
|
|
|
|
chr13_-_98202884
|
1.422
|
NM_005073
|
SLC15A1
|
solute carrier family 15 (oligopeptide transporter), member 1
|
|
chr12_-_131697109
|
1.415
|
NM_001195520
|
LOC645277
|
hypothetical LOC645277
|
|
chr18_+_17076166
|
1.412
|
NM_001142966
|
GREB1L
|
growth regulation by estrogen in breast cancer-like
|
|
chr11_-_65082705
|
1.404
|
|
LTBP3
|
latent transforming growth factor beta binding protein 3
|
|
chr4_-_174686918
|
1.383
|
|
HAND2
|
heart and neural crest derivatives expressed 2
|
|
chr19_-_53808505
|
1.377
|
NM_017708
|
FAM83E
|
family with sequence similarity 83, member E
|
|
chr20_-_22978141
|
1.376
|
NM_000361
|
THBD
|
thrombomodulin
|
|
chr1_+_148788410
|
1.370
|
NM_019032
NM_025008
|
ADAMTSL4
|
ADAMTS-like 4
|
|
chr9_+_123101926
|
1.367
|
|
GSN
|
gelsolin
|
|
chr12_-_3732478
|
1.363
|
|
EFCAB4B
|
EF-hand calcium binding domain 4B
|
|
chr16_+_85158357
|
1.358
|
NM_005251
|
FOXC2
|
forkhead box C2 (MFH-1, mesenchyme forkhead 1)
|
|
chr11_-_27697768
|
1.354
|
NM_001143807
|
BDNF
|
brain-derived neurotrophic factor
|
|
chr15_+_98960277
|
1.351
|
NM_024708
NM_198243
|
ASB7
|
ankyrin repeat and SOCS box containing 7
|
|
chr12_-_2814410
|
1.350
|
NM_031474
|
NRIP2
|
nuclear receptor interacting protein 2
|
|
chr9_-_121171400
|
1.347
|
NM_014618
|
DBC1
|
deleted in bladder cancer 1
|
|
chr2_+_15999599
|
1.344
|
|
MYCN
|
v-myc myelocytomatosis viral related oncogene, neuroblastoma derived (avian)
|
|
chr11_+_122806196
|
1.330
|
|
LOC100128242
|
hypothetical LOC100128242
|
|
chr3_+_45042754
|
1.325
|
NM_003278
|
CLEC3B
|
C-type lectin domain family 3, member B
|
|
chr8_-_93144366
|
1.310
|
NM_004349
|
RUNX1T1
|
runt-related transcription factor 1; translocated to, 1 (cyclin D-related)
|
|
chr4_-_174687104
|
1.295
|
|
HAND2
|
heart and neural crest derivatives expressed 2
|
|
chr9_+_111850698
|
1.290
|
NM_001004065
NM_001198656
|
AKAP2
|
A kinase (PRKA) anchor protein 2
|
|
chr21_-_27139143
|
1.281
|
|
ADAMTS1
|
ADAM metallopeptidase with thrombospondin type 1 motif, 1
|
|
chr11_-_66877519
|
1.279
|
NM_021173
|
POLD4
|
polymerase (DNA-directed), delta 4
|
|
chr9_-_72673753
|
1.272
|
NM_001007470
NM_020952
NM_024971
NM_206944
NM_206945
NM_206946
NM_206947
NM_206948
|
TRPM3
|
transient receptor potential cation channel, subfamily M, member 3
|
|
chr8_+_70541631
|
1.269
|
|
SULF1
|
sulfatase 1
|
|
chr16_+_65790514
|
1.267
|
NM_024712
|
ELMO3
|
engulfment and cell motility 3
|
|
chr11_+_131285747
|
1.266
|
NM_001144058
NM_001144059
NM_016522
|
NTM
|
neurotrimin
|
|
chr15_+_38520594
|
1.262
|
NM_014952
|
BAHD1
|
bromo adjacent homology domain containing 1
|
|
chr3_-_130761967
|
1.261
|
|
PLXND1
|
plexin D1
|
|
chr19_+_50004134
|
1.251
|
NM_001013257
NM_005581
|
BCAM
|
basal cell adhesion molecule (Lutheran blood group)
|
|
chr1_+_26986996
|
1.249
|
NM_017837
|
PIGV
|
phosphatidylinositol glycan anchor biosynthesis, class V
|
|
chr1_+_116181927
|
1.245
|
|
NHLH2
|
nescient helix loop helix 2
|
|
chr5_-_58607673
|
1.242
|
NM_001197220
|
PDE4D
|
phosphodiesterase 4D, cAMP-specific
|
|
chr11_-_111287657
|
1.231
|
NM_001885
|
CRYAB
|
crystallin, alpha B
|
|
chr11_-_132318741
|
1.226
|
|
OPCML
|
opioid binding protein/cell adhesion molecule-like
|
|
chr11_-_75057062
|
1.226
|
NM_033063
NM_207577
|
MAP6
|
microtubule-associated protein 6
|
|
chr5_+_167889123
|
1.222
|
|
|
|
|
chr17_+_7549138
|
1.220
|
NM_001406
|
EFNB3
|
ephrin-B3
|
|
chr11_+_1897374
|
1.204
|
NM_001042780
NM_001042781
NM_001042782
NM_006757
|
TNNT3
|
troponin T type 3 (skeletal, fast)
|
|
chr5_+_170779664
|
1.200
|
|
FGF18
|
fibroblast growth factor 18
|
|
chr6_+_142664490
|
1.198
|
NM_001032394
NM_001032395
NM_020455
NM_198569
|
GPR126
|
G protein-coupled receptor 126
|
|
chr17_-_77474597
|
1.190
|
NM_032711
|
MAFG
|
v-maf musculoaponeurotic fibrosarcoma oncogene homolog G (avian)
|
|
chr9_-_103289154
|
1.187
|
NM_032342
|
C9orf125
|
chromosome 9 open reading frame 125
|
|
chr15_+_58083737
|
1.187
|
|
|
|
|
chr8_-_93144039
|
1.185
|
|
|
|
|
chr2_-_236837726
|
1.185
|
NM_212556
|
ASB18
|
ankyrin repeat and SOCS box containing 18
|
|
chr17_+_4784524
|
1.177
|
|
RNF167
|
ring finger protein 167
|
|
chrX_+_101887562
|
1.172
|
NM_001142529
NM_001142530
|
BHLHB9
|
basic helix-loop-helix domain containing, class B, 9
|
|
chr19_-_12747263
|
1.171
|
|
HOOK2
|
hook homolog 2 (Drosophila)
|
|
chr12_-_3732614
|
1.168
|
NM_001144958
NM_001144959
NM_032680
|
EFCAB4B
|
EF-hand calcium binding domain 4B
|
|
chr14_-_50631407
|
1.168
|
|
TRIM9
|
tripartite motif containing 9
|
|
chr12_-_80677223
|
1.167
|
NM_003625
|
PPFIA2
|
protein tyrosine phosphatase, receptor type, f polypeptide (PTPRF), interacting protein (liprin), alpha 2
|
|
chr21_-_27139563
|
1.167
|
NM_006988
|
ADAMTS1
|
ADAM metallopeptidase with thrombospondin type 1 motif, 1
|
|
chr2_+_217206450
|
1.165
|
|
IGFBP2
|
insulin-like growth factor binding protein 2, 36kDa
|
|
chr8_-_17599438
|
1.164
|
NM_020749
|
MTUS1
|
microtubule associated tumor suppressor 1
|
|
chrX_-_43717864
|
1.160
|
NM_000266
|
NDP
|
Norrie disease (pseudoglioma)
|
|
chr12_+_52733927
|
1.160
|
NM_153633
|
HOXC4
HOXC6
|
homeobox C4
homeobox C6
|
|
chr11_-_110917632
|
1.158
|
|
|
|
|
chr9_+_4480193
|
1.156
|
NM_004170
|
SLC1A1
|
solute carrier family 1 (neuronal/epithelial high affinity glutamate transporter, system Xag), member 1
|
|
chr16_+_51722285
|
1.150
|
|
CHD9
|
chromodomain helicase DNA binding protein 9
|
|
chr12_-_51734434
|
1.146
|
|
LOC283335
|
hypothetical LOC283335
|
|
chr1_-_203657803
|
1.145
|
NM_001001552
NM_001199050
NM_001199051
NM_001199052
|
LEMD1
|
LEM domain containing 1
|
|
chr2_+_176695658
|
1.142
|
NM_014213
|
HOXD9
|
homeobox D9
|
|
chr3_+_50272648
|
1.139
|
|
|
|
|
chr10_+_119292699
|
1.138
|
|
|
|
|
chr9_+_134027154
|
1.137
|
NM_032536
|
NTNG2
|
netrin G2
|
|
chr1_+_113038202
|
1.136
|
|
MOV10
|
Mov10, Moloney leukemia virus 10, homolog (mouse)
|
|
chr3_-_166397142
|
1.135
|
NM_014926
|
SLITRK3
|
SLIT and NTRK-like family, member 3
|
|
chr1_+_176748834
|
1.134
|
NM_001170722
NM_001170723
NM_001170724
NM_032126
|
C1orf49
|
chromosome 1 open reading frame 49
|
|
chr21_+_39117064
|
1.129
|
|
ETS2
|
v-ets erythroblastosis virus E26 oncogene homolog 2 (avian)
|
|
chr17_-_36507900
|
1.129
|
NM_031960
|
KRTAP4-8
|
keratin associated protein 4-8
|
|
chr14_+_102662416
|
1.116
|
NM_006291
|
TNFAIP2
|
tumor necrosis factor, alpha-induced protein 2
|
|
chr17_+_56884560
|
1.112
|
|
TBX4
|
T-box 4
|
|
chr9_+_123101701
|
1.111
|
NM_000177
|
GSN
|
gelsolin
|
|
chr11_+_7491789
|
1.110
|
|
PPFIBP2
|
PTPRF interacting protein, binding protein 2 (liprin beta 2)
|
|
chr19_+_54786723
|
1.107
|
NM_020719
|
PRR12
|
proline rich 12
|
|
chr2_+_217206327
|
1.101
|
NM_000597
|
IGFBP2
|
insulin-like growth factor binding protein 2, 36kDa
|
|
chr7_-_112513379
|
1.096
|
NM_001146266
NM_001146267
|
GPR85
|
G protein-coupled receptor 85
|
|
chr3_+_185580554
|
1.096
|
NM_003741
|
CHRD
|
chordin
|
|
chr15_+_72253940
|
1.093
|
NM_201526
|
ISLR
|
immunoglobulin superfamily containing leucine-rich repeat
|
|
chr22_-_42589543
|
1.086
|
NM_014351
|
SULT4A1
|
sulfotransferase family 4A, member 1
|
|
chr12_-_80676562
|
1.085
|
|
PPFIA2
|
protein tyrosine phosphatase, receptor type, f polypeptide (PTPRF), interacting protein (liprin), alpha 2
|
|
chr5_+_139473873
|
1.084
|
NM_005859
|
PURA
|
purine-rich element binding protein A
|
|
chr2_+_235525355
|
1.078
|
NM_014521
|
SH3BP4
|
SH3-domain binding protein 4
|
|
chr7_-_27120015
|
1.075
|
|
HOXA3
|
homeobox A3
|
|
chr8_+_39084206
|
1.073
|
NM_145004
|
ADAM32
|
ADAM metallopeptidase domain 32
|
|
chr11_-_122571046
|
1.072
|
|
CLMP
|
CXADR-like membrane protein
|