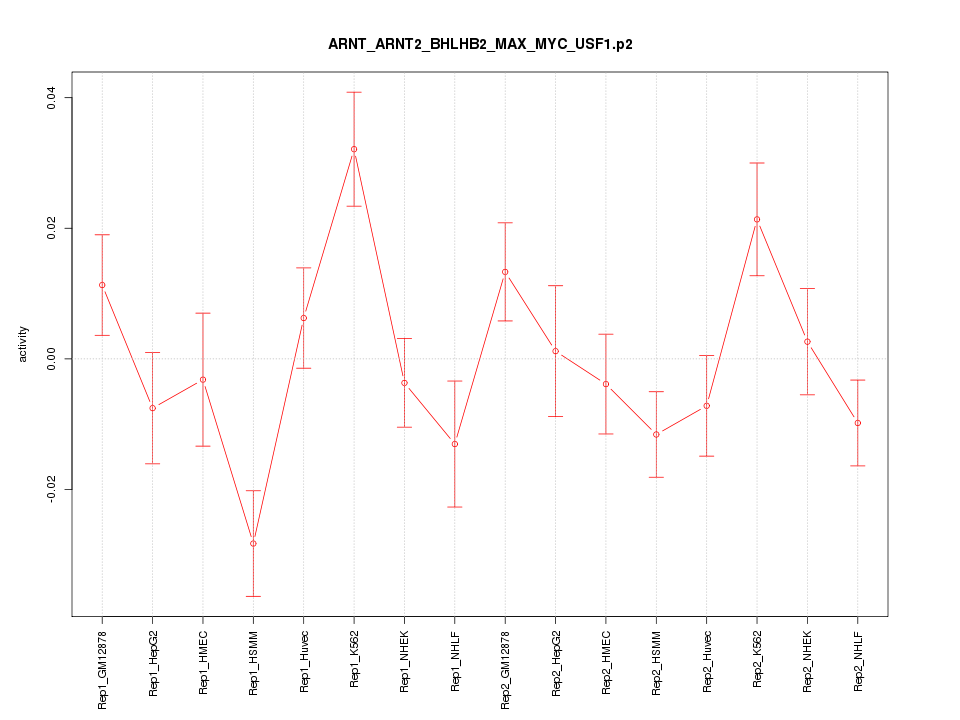
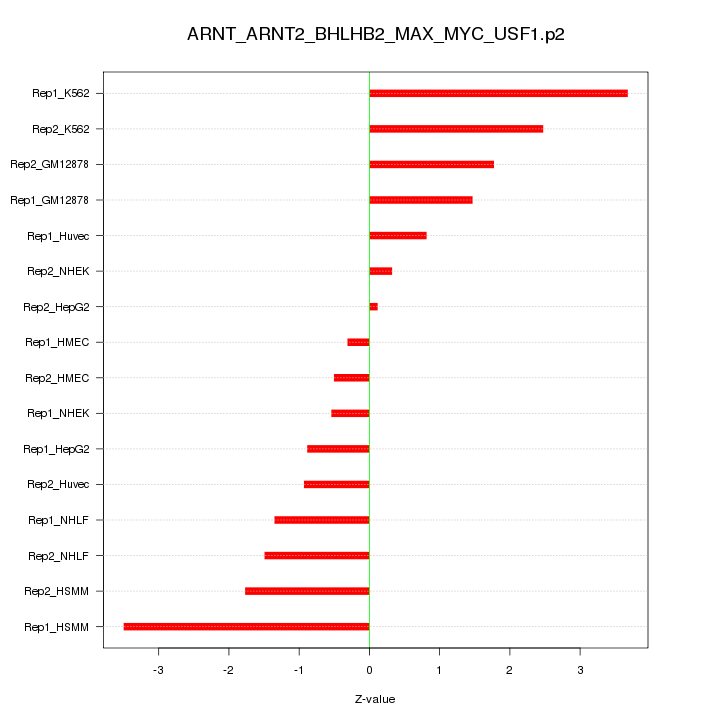
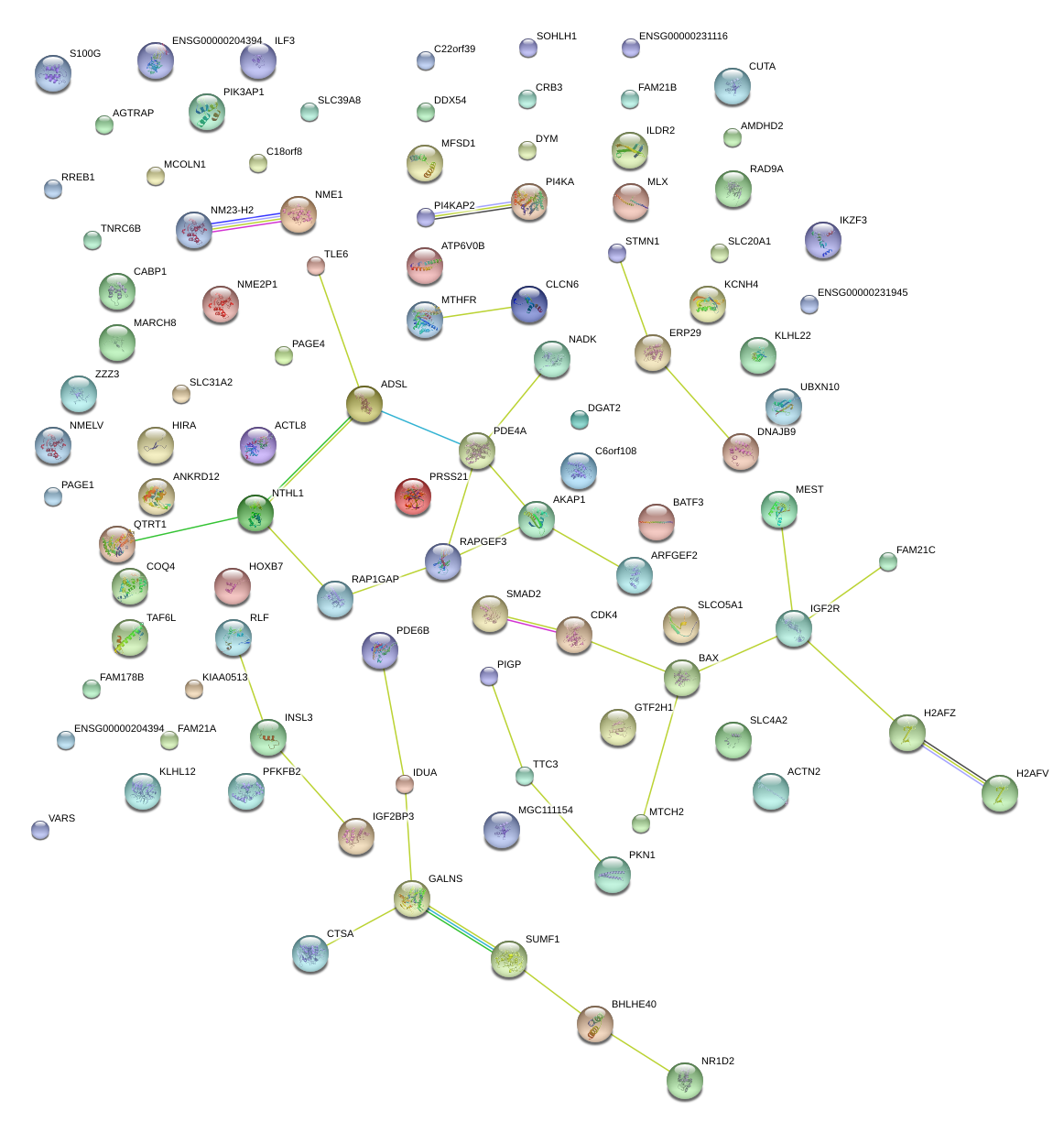
|
chr6_-_33493792
|
3.916
|
|
CUTA
|
cutA divalent cation tolerance homolog (E. coli)
|
|
chr16_+_2510323
|
3.871
|
NM_001145815
NM_015944
|
AMDHD2
|
amidohydrolase domain containing 2
|
|
chr6_-_33494042
|
3.543
|
NM_001014433
NM_001014837
NM_001014838
NM_001014840
NM_015921
|
CUTA
|
cutA divalent cation tolerance homolog (E. coli)
|
|
chr6_-_33493898
|
3.369
|
|
CUTA
|
cutA divalent cation tolerance homolog (E. coli)
|
|
chr2_-_97016027
|
3.217
|
NM_001122646
|
FAM178B
|
family with sequence similarity 178, member B
|
|
chr17_-_53764867
|
2.881
|
|
|
|
|
chr6_-_31871394
|
2.608
|
|
VARS
|
valyl-tRNA synthetase
|
|
chr1_-_1700088
|
2.377
|
NM_001198993
|
NADK
|
NAD kinase
|
|
chr19_+_10673105
|
2.318
|
NM_031209
|
QTRT1
|
queuine tRNA-ribosyltransferase 1
|
|
chr22_+_22703107
|
2.139
|
NM_001144931
|
LOC391322
|
D-dopachrome tautomerase-like
|
|
chr9_+_114953087
|
2.047
|
|
SLC31A2
|
solute carrier family 31 (copper transporters), member 2
|
|
chr12_-_56432292
|
1.923
|
|
CDK4
|
cyclin-dependent kinase 4
|
|
chr9_+_114953048
|
1.872
|
NM_001860
|
SLC31A2
|
solute carrier family 31 (copper transporters), member 2
|
|
chr1_-_1699723
|
1.871
|
NM_023018
|
NADK
|
NAD kinase
|
|
chr6_-_31871452
|
1.788
|
|
VARS
|
valyl-tRNA synthetase
|
|
chr8_+_142471290
|
1.752
|
|
|
|
|
chr18_+_9126742
|
1.747
|
NM_001083625
NM_015208
|
ANKRD12
|
ankyrin repeat domain 12
|
|
chr19_+_7493436
|
1.744
|
NM_020533
|
MCOLN1
|
mucolipin 1
|
|
chr12_-_56432377
|
1.737
|
|
CDK4
|
cyclin-dependent kinase 4
|
|
chr2_+_113119985
|
1.690
|
NM_005415
|
SLC20A1
|
solute carrier family 20 (phosphate transporter), member 1
|
|
chr12_-_56432375
|
1.682
|
|
CDK4
|
cyclin-dependent kinase 4
|
|
chr1_-_210939820
|
1.677
|
|
BATF3
|
basic leucine zipper transcription factor, ATF-like 3
|
|
chr1_-_210939929
|
1.665
|
NM_018664
|
BATF3
|
basic leucine zipper transcription factor, ATF-like 3
|
|
chr17_-_44043361
|
1.647
|
NM_004502
|
HOXB7
|
homeobox B7
|
|
chr21_+_37367432
|
1.638
|
NM_001001894
|
TTC3
|
tetratricopeptide repeat domain 3
|
|
chr1_-_21868380
|
1.610
|
NM_001145657
NM_002885
|
RAP1GAP
|
RAP1 GTPase activating protein
|
|
chr11_+_66915997
|
1.573
|
NM_004584
|
RAD9A
|
RAD9 homolog A (S. pombe)
|
|
chr6_-_33493640
|
1.550
|
|
CUTA
|
cutA divalent cation tolerance homolog (E. coli)
|
|
chr7_+_150390566
|
1.525
|
|
SLC4A2
|
solute carrier family 4, anion exchanger, member 2 (erythrocyte membrane protein band 3-like 1)
|
|
chr1_-_210939727
|
1.514
|
|
|
|
|
chr10_-_98470164
|
1.449
|
NM_152309
|
PIK3AP1
|
phosphoinositide-3-kinase adaptor protein 1
|
|
chr1_+_20385164
|
1.438
|
NM_152376
|
UBXN10
|
UBX domain protein 10
|
|
chr10_-_45410197
|
1.432
|
NM_001002265
NM_145021
|
MARCH8
|
membrane-associated ring finger (C3HC4) 8
|
|
chr17_-_44043278
|
1.431
|
|
HOXB7
|
homeobox B7
|
|
chr1_+_40399618
|
1.421
|
NM_012421
|
RLF
|
rearranged L-myc fusion
|
|
chr1_+_40399642
|
1.419
|
|
RLF
|
rearranged L-myc fusion
|
|
chr8_-_70908130
|
1.415
|
NM_001146008
|
SLCO5A1
|
solute carrier organic anion transporter family, member 5A1
|
|
chr7_+_129913271
|
1.414
|
NM_177524
|
MEST
|
mesoderm specific transcript homolog (mouse)
|
|
chr12_-_56432385
|
1.405
|
NM_000075
|
CDK4
|
cyclin-dependent kinase 4
|
|
chr3_+_23961739
|
1.395
|
NM_005126
|
NR1D2
|
nuclear receptor subfamily 1, group D, member 2
|
|
chr19_+_54149963
|
1.385
|
|
BAX
|
BCL2-associated X protein
|
|
chr19_+_6415259
|
1.368
|
NM_139161
NM_174881
|
CRB3
|
crumbs homolog 3 (Drosophila)
|
|
chr19_+_2928480
|
1.350
|
NM_001143986
NM_024760
|
TLE6
|
transducin-like enhancer of split 6 (E(sp1) homolog, Drosophila)
|
|
chr1_+_11718715
|
1.341
|
NM_001040194
NM_001040195
NM_001040196
NM_001040197
NM_020350
|
AGTRAP
|
angiotensin II receptor-associated protein
|
|
chr1_-_201162938
|
1.324
|
|
KLHL12
|
kelch-like 12 (Drosophila)
|
|
chr22_-_19180075
|
1.311
|
NM_032775
|
KLHL22
|
kelch-like 22 (Drosophila)
|
|
chr1_-_201162991
|
1.291
|
NM_021633
|
KLHL12
|
kelch-like 12 (Drosophila)
|
|
chr18_-_43711452
|
1.283
|
NM_001003652
NM_001135937
|
SMAD2
|
SMAD family member 2
|
|
chr4_-_103485600
|
1.281
|
NM_001135146
|
SLC39A8
|
solute carrier family 39 (zinc transporter), member 8
|
|
chr20_+_43953606
|
1.277
|
|
CTSA
|
cathepsin A
|
|
chr1_+_11718764
|
1.241
|
|
AGTRAP
|
angiotensin II receptor-associated protein
|
|
chr17_+_52518098
|
1.229
|
|
AKAP1
|
A kinase (PRKA) anchor protein 1
|
|
chr19_+_10626106
|
1.211
|
|
ILF3
|
interleukin enhancer binding factor 3, 90kDa
|
|
chr6_-_31871530
|
1.201
|
NM_006295
|
VARS
|
valyl-tRNA synthetase
|
|
chr22_+_38903825
|
1.198
|
NM_001162501
NM_015088
|
TNRC6B
|
trinucleotide repeat containing 6B
|
|
chr10_+_51497716
|
1.192
|
|
FAM21A
|
family with sequence similarity 21, member A
|
|
chr1_+_205293217
|
1.191
|
NM_001018053
NM_006212
|
PFKFB2
|
6-phosphofructo-2-kinase/fructose-2,6-biphosphatase 2
|
|
chr11_+_75157414
|
1.172
|
NM_032564
|
DGAT2
|
diacylglycerol O-acyltransferase 2
|
|
chr22_-_19543099
|
1.170
|
|
PI4KA
|
phosphatidylinositol 4-kinase, catalytic, alpha
|
|
chr18_-_45241014
|
1.159
|
NM_017653
|
DYM
|
dymeclin
|
|
chr12_+_119562737
|
1.159
|
NM_001033677
|
CABP1
|
calcium binding protein 1
|
|
chrX_-_49347279
|
1.150
|
NM_003785
|
PAGE1
|
P antigen family, member 1 (prostate associated)
|
|
chr9_-_137731139
|
1.142
|
NM_001012415
NM_001101677
|
SOHLH1
|
spermatogenesis and oogenesis specific basic helix-loop-helix 1
|
|
chr1_+_17954394
|
1.136
|
NM_030812
|
ACTL8
|
actin-like 8
|
|
chr12_-_46439127
|
1.135
|
NM_001098531
|
RAPGEF3
|
Rap guanine nucleotide exchange factor (GEF) 3
|
|
chr17_+_31964909
|
1.113
|
|
|
|
|
chr21_-_37367259
|
1.107
|
NM_153682
|
PIGP
|
phosphatidylinositol glycan anchor biosynthesis, class P
|
|
chr17_+_46599081
|
1.103
|
NM_001018138
|
NME2
|
non-metastatic cells 2, protein (NM23B) expressed in
|
|
chr4_+_970775
|
1.100
|
NM_000203
|
IDUA
|
iduronidase, alpha-L-
|
|
chr16_+_2807164
|
1.096
|
NM_006799
NM_144956
NM_144957
|
PRSS21
|
protease, serine, 21 (testisin)
|
|
chr1_+_44213260
|
1.081
|
|
ATP6V0B
|
ATPase, H+ transporting, lysosomal 21kDa, V0 subunit b
|
|
chr6_-_43305156
|
1.077
|
NM_006443
NM_199184
|
C6orf108
|
chromosome 6 open reading frame 108
|
|
chr12_+_110935618
|
1.074
|
|
ERP29
|
endoplasmic reticulum protein 29
|
|
chr3_+_160002605
|
1.073
|
|
MFSD1
|
major facilitator superfamily domain containing 1
|
|
chr7_+_107997527
|
1.065
|
|
DNAJB9
|
DnaJ (Hsp40) homolog, subfamily B, member 9
|
|
chr12_-_112107587
|
1.064
|
NM_001111322
NM_024072
|
DDX54
|
DEAD (Asp-Glu-Ala-Asp) box polypeptide 54
|
|
chr1_-_77920914
|
1.060
|
NM_015534
|
ZZZ3
|
zinc finger, ZZ-type containing 3
|
|
chr3_-_4483955
|
1.055
|
NM_001164674
NM_001164675
NM_182760
|
SUMF1
|
sulfatase modifying factor 1
|
|
chr7_-_23476497
|
1.052
|
NM_006547
|
IGF2BP3
|
insulin-like growth factor 2 mRNA binding protein 3
|
|
chr16_-_87450734
|
1.041
|
|
GALNS
|
galactosamine (N-acetyl)-6-sulfate sulfatase
|
|
chr19_+_54149941
|
1.023
|
|
BAX
|
BCL2-associated X protein
|
|
chr16_+_2807228
|
1.019
|
|
PRSS21
|
protease, serine, 21 (testisin)
|
|
chr1_+_111544469
|
1.013
|
|
|
|
|
chr17_-_37586716
|
1.006
|
NM_012285
|
KCNH4
|
potassium voltage-gated channel, subfamily H (eag-related), member 4
|
|
chr22_-_19543012
|
1.004
|
|
PI4KA
|
phosphatidylinositol 4-kinase, catalytic, alpha
|
|
chr16_+_29373365
|
1.001
|
NM_001014999
NM_001015000
NM_024044
NM_178044
|
SLX1A-SULT1A3
SLX1A
SLX1B
|
SLX1A-SULT1A3 read-through transcript
SLX1 structure-specific endonuclease subunit homolog A (S. cerevisiae)
SLX1 structure-specific endonuclease subunit homolog B (S. cerevisiae)
|
|
chr3_-_4483952
|
0.999
|
|
SUMF1
|
sulfatase modifying factor 1
|
|
chr1_-_11788550
|
0.996
|
|
MTHFR
|
methylenetetrahydrofolate reductase (NAD(P)H)
|
|
chr1_+_11718798
|
0.991
|
|
AGTRAP
|
angiotensin II receptor-associated protein
|
|
chr1_-_165211128
|
0.990
|
NM_199351
|
ILDR2
|
immunoglobulin-like domain containing receptor 2
|
|
chr17_-_35273947
|
0.986
|
NM_012481
NM_183228
NM_183229
NM_183230
NM_183231
NM_183232
|
IKZF3
|
IKAROS family zinc finger 3 (Aiolos)
|
|
chrX_+_49480643
|
0.983
|
NM_007003
|
PAGE4
|
P antigen family, member 4 (prostate associated)
|
|
chr1_+_44213229
|
0.983
|
|
ATP6V0B
|
ATPase, H+ transporting, lysosomal 21kDa, V0 subunit b
|
|
chr10_+_51497660
|
0.983
|
NM_001005751
|
FAM21C
FAM21A
|
family with sequence similarity 21, member C
family with sequence similarity 21, member A
|
|
chr18_+_19337437
|
0.975
|
NM_013326
|
C18orf8
|
chromosome 18 open reading frame 8
|
|
chr19_+_14405100
|
0.965
|
NM_002741
|
PKN1
|
protein kinase N1
|
|
chr11_+_75157450
|
0.961
|
|
DGAT2
|
diacylglycerol O-acyltransferase 2
|
|
chr16_-_2037819
|
0.953
|
NM_002528
|
NTHL1
|
nth endonuclease III-like 1 (E. coli)
|
|
chr3_+_160002582
|
0.952
|
|
MFSD1
|
major facilitator superfamily domain containing 1
|
|
chr1_+_234916376
|
0.949
|
NM_001103
|
ACTN2
|
actinin, alpha 2
|
|
chr1_-_26105954
|
0.944
|
NM_203399
|
STMN1
|
stathmin 1
|
|
chr9_+_130124566
|
0.942
|
NM_016035
|
COQ4
|
coenzyme Q4 homolog (S. cerevisiae)
|
|
chr3_+_160002408
|
0.937
|
NM_001167903
NM_022736
|
MFSD1
|
major facilitator superfamily domain containing 1
|
|
chr1_+_44213182
|
0.937
|
NM_001039457
NM_004047
|
ATP6V0B
|
ATPase, H+ transporting, lysosomal 21kDa, V0 subunit b
|
|
chr4_+_609362
|
0.934
|
NM_000283
NM_001145291
|
PDE6B
|
phosphodiesterase 6B, cGMP-specific, rod, beta
|
|
chr16_+_83618804
|
0.933
|
NM_014732
|
KIAA0513
|
KIAA0513
|
|
chr22_-_17798998
|
0.933
|
|
HIRA
|
HIR histone cell cycle regulation defective homolog A (S. cerevisiae)
|
|
chr1_+_44213200
|
0.926
|
|
ATP6V0B
|
ATPase, H+ transporting, lysosomal 21kDa, V0 subunit b
|
|
chr19_+_10392128
|
0.925
|
NM_001111307
|
PDE4A
|
phosphodiesterase 4A, cAMP-specific
|
|
chr17_+_37972590
|
0.922
|
NM_170607
NM_198204
NM_198205
|
MLX
|
MAX-like protein X
|
|
chr11_-_47620590
|
0.912
|
NM_014342
|
MTCH2
|
mitochondrial carrier homolog 2 (C. elegans)
|
|
chr11_+_18300689
|
0.903
|
|
GTF2H1
|
general transcription factor IIH, polypeptide 1, 62kDa
|
|
chr17_-_41018291
|
0.898
|
|
|
|
|
chr22_-_17799177
|
0.892
|
|
HIRA
|
HIR histone cell cycle regulation defective homolog A (S. cerevisiae)
|
|
chr1_+_11788889
|
0.878
|
|
CLCN6
|
chloride channel 6
|
|
chr22_-_17799215
|
0.878
|
NM_003325
|
HIRA
|
HIR histone cell cycle regulation defective homolog A (S. cerevisiae)
|
|
chr11_+_62295412
|
0.873
|
NM_006473
|
TAF6L
|
TAF6-like RNA polymerase II, p300/CBP-associated factor (PCAF)-associated factor, 65kDa
|
|
chr1_+_44213210
|
0.854
|
|
ATP6V0B
|
ATPase, H+ transporting, lysosomal 21kDa, V0 subunit b
|
|
chr3_-_4483928
|
0.847
|
|
SUMF1
|
sulfatase modifying factor 1
|
|
chr20_+_46971618
|
0.846
|
NM_006420
|
ARFGEF2
|
ADP-ribosylation factor guanine nucleotide-exchange factor 2 (brefeldin A-inhibited)
|
|
chr6_+_160310118
|
0.841
|
NM_000876
|
IGF2R
|
insulin-like growth factor 2 receptor
|
|
chr21_-_37366684
|
0.836
|
NM_153681
|
PIGP
|
phosphatidylinositol glycan anchor biosynthesis, class P
|
|
chr6_+_7052985
|
0.835
|
NM_001003698
NM_001003699
NM_001003700
|
RREB1
|
ras responsive element binding protein 1
|
|
chr3_+_4996096
|
0.832
|
NM_003670
|
BHLHE40
|
basic helix-loop-helix family, member e40
|
|
chr4_-_101090431
|
0.830
|
|
H2AFZ
|
H2A histone family, member Z
|
|
chr19_-_14748623
|
0.828
|
|
EMR2
|
egf-like module containing, mucin-like, hormone receptor-like 2
|
|
chr6_-_44333002
|
0.827
|
NM_178148
|
SLC35B2
|
solute carrier family 35, member B2
|
|
chr3_-_20202622
|
0.826
|
NM_001012409
NM_001012410
NM_001012411
NM_001012412
NM_001012413
NM_138484
|
SGOL1
|
shugoshin-like 1 (S. pombe)
|
|
chr11_+_120399966
|
0.821
|
NM_001130047
NM_152715
|
TBCEL
|
tubulin folding cofactor E-like
|
|
chr6_+_36670117
|
0.819
|
|
SRSF3
|
serine/arginine-rich splicing factor 3
|
|
chr10_+_45542653
|
0.817
|
NM_001169106
NM_001169107
NM_015262
|
FAM21C
FAM21A
|
family with sequence similarity 21, member C
family with sequence similarity 21, member A
|
|
chr19_-_46883935
|
0.814
|
NM_006890
|
CEACAM7
|
carcinoembryonic antigen-related cell adhesion molecule 7
|
|
chr15_+_86983038
|
0.814
|
NM_002201
|
ISG20
|
interferon stimulated exonuclease gene 20kDa
|
|
chr17_-_34157962
|
0.812
|
NM_007144
|
PCGF2
|
polycomb group ring finger 2
|
|
chr16_+_65840275
|
0.807
|
NM_004594
|
SLC9A5
|
solute carrier family 9 (sodium/hydrogen exchanger), member 5
|
|
chr13_-_35686665
|
0.803
|
NM_017826
|
SOHLH2
|
spermatogenesis and oogenesis specific basic helix-loop-helix 2
|
|
chr1_+_32251988
|
0.802
|
NM_006559
|
KHDRBS1
|
KH domain containing, RNA binding, signal transduction associated 1
|
|
chr1_+_1157485
|
0.802
|
NM_080605
|
B3GALT6
|
UDP-Gal:betaGal beta 1,3-galactosyltransferase polypeptide 6
|
|
chr1_+_26368969
|
0.801
|
NM_015871
|
ZNF593
|
zinc finger protein 593
|
|
chr3_+_160002571
|
0.800
|
|
MFSD1
|
major facilitator superfamily domain containing 1
|
|
chr3_+_160002619
|
0.789
|
|
MFSD1
|
major facilitator superfamily domain containing 1
|
|
chr12_+_110935644
|
0.785
|
|
ERP29
|
endoplasmic reticulum protein 29
|
|
chr16_+_24458349
|
0.784
|
NM_006910
NM_018703
NM_032626
|
RBBP6
|
retinoblastoma binding protein 6
|
|
chr12_+_51932136
|
0.782
|
NM_032889
|
MFSD5
|
major facilitator superfamily domain containing 5
|
|
chr16_-_1465007
|
0.782
|
|
CLCN7
|
chloride channel 7
|
|
chr10_-_43223226
|
0.777
|
NM_001098206
|
HNRNPF
|
heterogeneous nuclear ribonucleoprotein F
|
|
chr16_+_2037986
|
0.777
|
NM_000548
NM_001077183
NM_001114382
|
TSC2
|
tuberous sclerosis 2
|
|
chr4_+_186176988
|
0.776
|
NM_001029887
|
HELT
|
HES/HEY-like transcription factor
|
|
chr2_+_26840626
|
0.774
|
NM_017877
|
CENPA
C2orf18
|
centromere protein A
chromosome 2 open reading frame 18
|
|
chr7_+_106472243
|
0.774
|
NM_002736
|
PRKAR2B
|
protein kinase, cAMP-dependent, regulatory, type II, beta
|
|
chr11_-_71492019
|
0.773
|
NM_017907
|
LAMTOR1
|
late endosomal/lysosomal adaptor, MAPK and MTOR activator 1
|
|
chr4_-_103485419
|
0.772
|
NM_001135147
|
SLC39A8
|
solute carrier family 39 (zinc transporter), member 8
|
|
chr6_+_36670067
|
0.768
|
NM_003017
|
SRSF3
|
serine/arginine-rich splicing factor 3
|
|
chr12_-_54995923
|
0.767
|
|
CNPY2
|
canopy 2 homolog (zebrafish)
|
|
chr7_+_107997544
|
0.766
|
NM_012328
|
DNAJB9
|
DnaJ (Hsp40) homolog, subfamily B, member 9
|
|
chr5_-_176869943
|
0.764
|
NM_001144875
NM_001144876
|
DOK3
|
docking protein 3
|
|
chr6_+_36670113
|
0.760
|
|
SRSF3
|
serine/arginine-rich splicing factor 3
|
|
chr12_-_121316907
|
0.759
|
|
VPS33A
|
vacuolar protein sorting 33 homolog A (S. cerevisiae)
|
|
chr1_+_10015577
|
0.759
|
NM_001105562
NM_006048
|
UBE4B
|
ubiquitination factor E4B (UFD2 homolog, yeast)
|
|
chr17_-_31964723
|
0.756
|
|
MYO19
|
myosin XIX
|
|
chr12_+_70519753
|
0.756
|
NM_001146213
NM_001146214
NM_022771
|
TBC1D15
|
TBC1 domain family, member 15
|
|
chr10_+_70150936
|
0.755
|
NM_018237
|
CCAR1
|
cell division cycle and apoptosis regulator 1
|
|
chr1_+_32992131
|
0.753
|
|
|
|
|
chr7_+_882704
|
0.753
|
NM_015949
|
GET4
|
golgi to ER traffic protein 4 homolog (S. cerevisiae)
|
|
chr5_+_154115046
|
0.750
|
|
LARP1
|
La ribonucleoprotein domain family, member 1
|
|
chr7_-_100331417
|
0.747
|
NM_000665
NM_015831
|
ACHE
|
acetylcholinesterase
|
|
chr19_+_1226510
|
0.743
|
NM_017914
|
C19orf24
|
chromosome 19 open reading frame 24
|
|
chr3_-_110319598
|
0.739
|
NM_014429
|
MORC1
|
MORC family CW-type zinc finger 1
|
|
chr12_-_54996044
|
0.737
|
|
CNPY2
|
canopy 2 homolog (zebrafish)
|
|
chr3_+_4996141
|
0.737
|
|
BHLHE40
|
basic helix-loop-helix family, member e40
|
|
chr12_+_51932173
|
0.737
|
|
MFSD5
|
major facilitator superfamily domain containing 5
|
|
chr18_-_19271814
|
0.736
|
NM_032933
|
C18orf45
|
chromosome 18 open reading frame 45
|
|
chr1_-_227828339
|
0.735
|
|
TAF5L
|
TAF5-like RNA polymerase II, p300/CBP-associated factor (PCAF)-associated factor, 65kDa
|
|
chr7_+_101861007
|
0.734
|
|
ORAI2
|
ORAI calcium release-activated calcium modulator 2
|
|
chr11_+_63463009
|
0.730
|
NM_024771
|
NAA40
|
N(alpha)-acetyltransferase 40, NatD catalytic subunit, homolog (S. cerevisiae)
|
|
chr8_-_103494073
|
0.730
|
|
UBR5
|
ubiquitin protein ligase E3 component n-recognin 5
|
|
chr1_+_55044323
|
0.729
|
NM_001110533
NM_152607
|
C1orf177
|
chromosome 1 open reading frame 177
|
|
chr8_-_133140808
|
0.721
|
NM_001080399
|
OC90
|
otoconin 90
|
|
chr12_+_112107747
|
0.718
|
NM_032848
|
C12orf52
|
chromosome 12 open reading frame 52
|
|
chr1_+_10015736
|
0.717
|
|
UBE4B
|
ubiquitination factor E4B (UFD2 homolog, yeast)
|
|
chr8_-_42028634
|
0.715
|
NM_001099412
NM_001099413
NM_006766
|
MYST3
|
MYST histone acetyltransferase (monocytic leukemia) 3
|
|
chr21_-_42247007
|
0.714
|
NM_015500
|
C2CD2
|
C2 calcium-dependent domain containing 2
|
|
chr12_-_121316997
|
0.714
|
NM_022916
|
VPS33A
|
vacuolar protein sorting 33 homolog A (S. cerevisiae)
|
|
chr12_+_49918921
|
0.712
|
|
DAZAP2
|
DAZ associated protein 2
|
|
chrX_-_19815526
|
0.711
|
|
SH3KBP1
|
SH3-domain kinase binding protein 1
|
|
chr16_-_47872973
|
0.709
|
|
CBLN1
|
cerebellin 1 precursor
|
|
chrX_-_47394937
|
0.709
|
NM_001114123
NM_005229
|
ELK1
|
ELK1, member of ETS oncogene family
|
|
chr9_+_130749791
|
0.706
|
NM_015354
|
NUP188
|
nucleoporin 188kDa
|
|
chr3_-_4483939
|
0.705
|
|
SUMF1
|
sulfatase modifying factor 1
|
|
chrX_+_130984912
|
0.705
|
NM_001042453
NM_016542
|
MST4
|
serine/threonine protein kinase MST4
|
|
chr1_-_11788666
|
0.705
|
NM_005957
|
MTHFR
|
methylenetetrahydrofolate reductase (NAD(P)H)
|
|
chr1_-_27099515
|
0.704
|
|
GPATCH3
|
G patch domain containing 3
|
|
chr11_-_59334706
|
0.693
|
NM_017840
|
MRPL16
|
mitochondrial ribosomal protein L16
|
|
chr2_+_176724583
|
0.691
|
|
|
|
|
chr2_-_175741096
|
0.689
|
NM_001880
|
ATF2
|
activating transcription factor 2
|
|
chr1_+_11788827
|
0.684
|
|
CLCN6
|
chloride channel 6
|
|
chr11_-_3620060
|
0.684
|
NM_053017
NM_001079536
|
ART5
|
ADP-ribosyltransferase 5
|
|
chr7_-_107997132
|
0.684
|
NM_182529
NM_001130475
|
THAP5
|
THAP domain containing 5
|
|
chr9_-_139316523
|
0.683
|
NM_001004354
|
NRARP
|
NOTCH-regulated ankyrin repeat protein
|
|
chr12_+_51932252
|
0.681
|
|
MFSD5
|
major facilitator superfamily domain containing 5
|