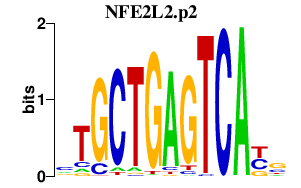
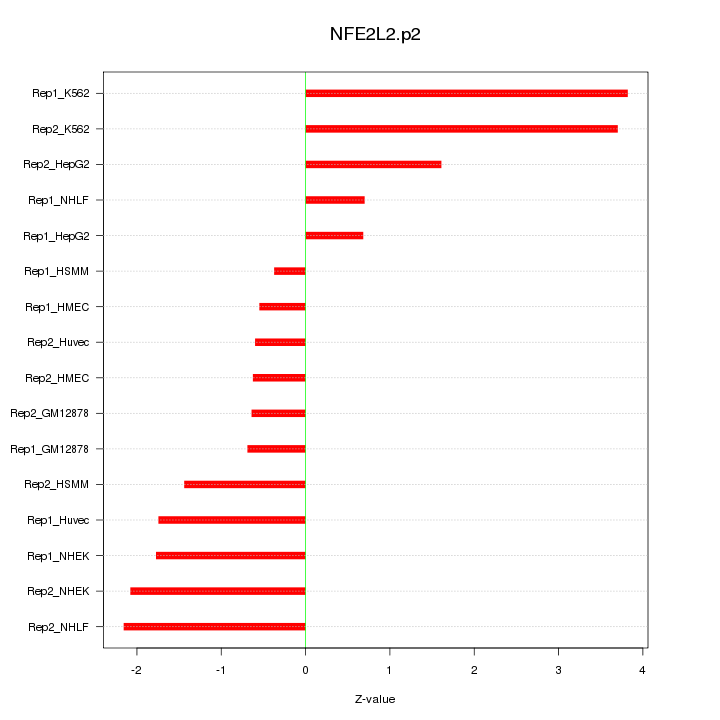
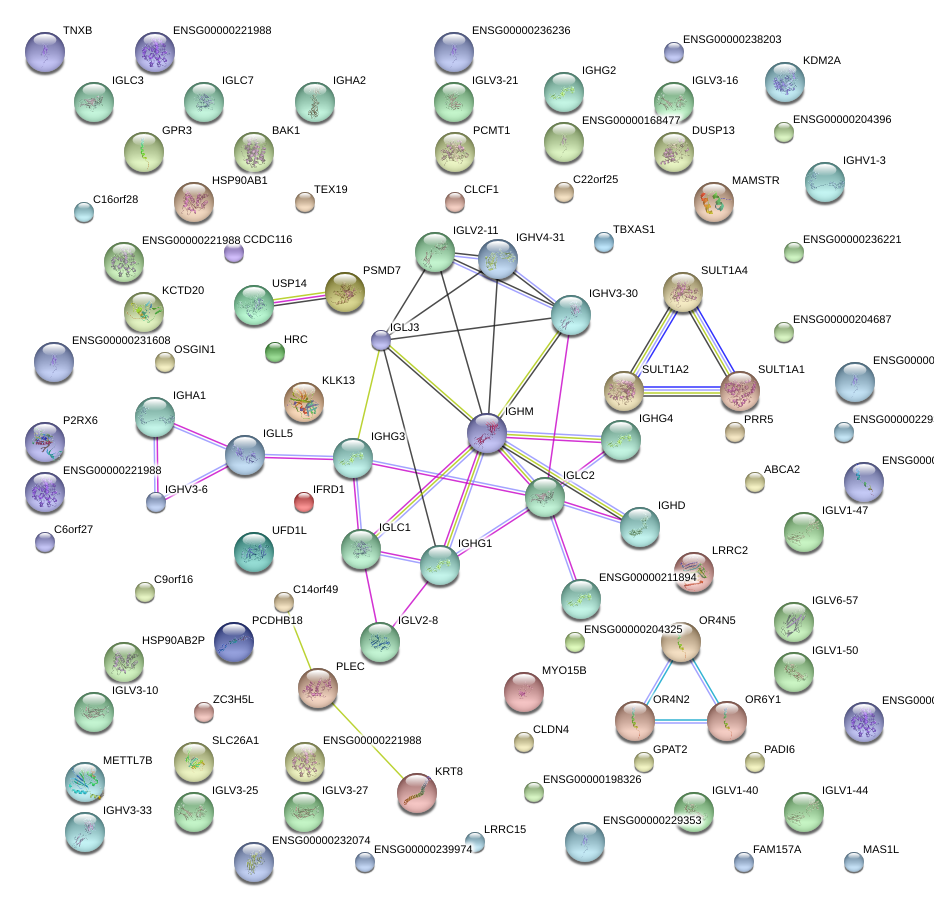
|
chr22_+_21484240
|
6.020
|
|
|
|
|
chr22_+_21340757
|
5.504
|
|
|
|
|
chr22_+_21094094
|
4.633
|
|
|
|
|
chr22_+_21393107
|
3.850
|
|
IGLJ3
|
immunoglobulin lambda joining 3
|
|
chr22_+_21079355
|
3.840
|
|
|
|
|
chr22_+_21359175
|
3.797
|
|
IGLV3-25
|
immunoglobulin lambda variable 3-25
|
|
chr22_+_21370394
|
3.556
|
|
IGL@
|
immunoglobulin lambda locus
|
|
chr22_+_21065202
|
3.135
|
|
|
|
|
chr22_+_21065207
|
3.098
|
|
IGLJ3
|
immunoglobulin lambda joining 3
|
|
chr22_+_21065134
|
3.074
|
|
LOC100290481
|
immunoglobulin lambda light chain-like
|
|
chr22_+_21053972
|
2.703
|
|
|
|
|
chr22_+_21407066
|
2.524
|
|
|
|
|
chr14_-_105712630
|
2.437
|
|
IGHV4-31
IGHA1
IGHA2
IGHG1
|
immunoglobulin heavy variable 4-31
immunoglobulin heavy constant alpha 1
immunoglobulin heavy constant alpha 2 (A2m marker)
immunoglobulin heavy constant gamma 1 (G1m marker)
|
|
chr22_+_20316968
|
2.368
|
NM_152612
|
CCDC116
|
coiled-coil domain containing 116
|
|
chr14_-_105804211
|
2.287
|
|
IGHG1
|
immunoglobulin heavy constant gamma 1 (G1m marker)
|
|
chr22_+_21042091
|
2.231
|
|
IGLV1-44
IGL@
|
immunoglobulin lambda variable 1-44
immunoglobulin lambda locus
|
|
chr10_-_76538859
|
2.183
|
NM_001007271
NM_001007272
NM_001007273
|
DUSP13
|
dual specificity phosphatase 13
|
|
chr22_+_20899155
|
2.180
|
|
|
|
|
chr4_-_977163
|
2.147
|
NM_022042
NM_134425
NM_213613
|
SLC26A1
|
solute carrier family 26 (sulfate transporter), member 1
|
|
chr12_+_54361685
|
2.086
|
|
METTL7B
|
methyltransferase like 7B
|
|
chr8_-_11242408
|
2.053
|
|
LOC100129129
|
hypothetical LOC100129129
|
|
chr12_+_54361596
|
2.047
|
NM_152637
|
METTL7B
|
methyltransferase like 7B
|
|
chr22_+_21116283
|
1.988
|
|
|
|
|
chr20_+_2744947
|
1.933
|
NM_001167670
|
LOC100288797
|
hypothetical protein LOC100288797
|
|
chr22_+_18388656
|
1.832
|
|
C22orf25
|
chromosome 22 open reading frame 25
|
|
chr6_-_32121784
|
1.823
|
NM_032470
|
TNXB
|
tenascin XB
|
|
chr1_+_17571277
|
1.782
|
NM_207421
|
PADI6
|
peptidyl arginine deiminase, type VI
|
|
chr7_+_139175420
|
1.734
|
NM_001061
NM_001166253
NM_030984
|
TBXAS1
|
thromboxane A synthase 1 (platelet)
|
|
chr14_+_19365447
|
1.683
|
NM_001004723
|
OR4N2
|
olfactory receptor, family 4, subfamily N, member 2
|
|
chr22_+_21559959
|
1.609
|
NM_001178126
|
IGLL5
IGLC1
|
immunoglobulin lambda-like polypeptide 5
immunoglobulin lambda constant 1 (Mcg marker)
|
|
chr9_+_129962292
|
1.605
|
NM_024112
|
C9orf16
|
chromosome 9 open reading frame 16
|
|
chr6_-_31853027
|
1.573
|
NM_025258
|
C6orf27
|
chromosome 6 open reading frame 27
|
|
chr3_-_46582806
|
1.571
|
NM_024512
|
LRRC2
|
leucine rich repeat containing 2
|
|
chr22_+_21011653
|
1.567
|
|
|
|
|
chr22_+_21495273
|
1.519
|
|
|
|
|
chr22_+_21552897
|
1.515
|
|
LOC100290481
|
immunoglobulin lambda light chain-like
|
|
chr14_+_19681734
|
1.495
|
NM_001004724
|
OR4N5
|
olfactory receptor, family 4, subfamily N, member 5
|
|
chr22_+_19699441
|
1.459
|
NM_001159554
NM_005446
|
P2RX6
|
purinergic receptor P2X, ligand-gated ion channel, 6
|
|
chr6_+_32229599
|
1.438
|
NM_005155
|
PPT2-EGFL8
PPT2
|
PPT2-EGFL8 read-through transcript
palmitoyl-protein thioesterase 2
|
|
chr17_+_77910411
|
1.396
|
NM_207459
|
TEX19
|
testis expressed 19
|
|
chr6_+_44322831
|
1.394
|
|
HSP90AB1
|
heat shock protein 90kDa alpha (cytosolic), class B member 1
|
|
chr14_-_105862080
|
1.377
|
|
IGHV4-31
IGHA1
IGHD
IGHG1
IGHG2
IGHG3
IGHM
SCFV
|
immunoglobulin heavy variable 4-31
immunoglobulin heavy constant alpha 1
immunoglobulin heavy constant delta
immunoglobulin heavy constant gamma 1 (G1m marker)
immunoglobulin heavy constant gamma 2 (G2m marker)
immunoglobulin heavy constant gamma 3 (G3m marker)
immunoglobulin heavy constant mu
single-chain Fv fragment
|
|
chr19_-_54350457
|
1.377
|
NM_002152
|
HRC
|
histidine rich calcium binding protein
|
|
chr12_-_51629874
|
1.363
|
|
KRT8
|
keratin 8
|
|
chr22_+_20880159
|
1.324
|
|
IGLV6-57
|
immunoglobulin lambda variable 6-57
|
|
chr6_+_44322764
|
1.315
|
NM_007355
|
HSP90AB1
|
heat shock protein 90kDa alpha (cytosolic), class B member 1
|
|
chr6_+_32229278
|
1.305
|
NM_138717
|
PPT2
|
palmitoyl-protein thioesterase 2
|
|
chr5_+_140594121
|
1.303
|
|
PCDHB18
|
protocadherin beta 18 pseudogene
|
|
chr14_-_106084085
|
1.235
|
|
IGHV4-31
IGHG1
|
immunoglobulin heavy variable 4-31
immunoglobulin heavy constant gamma 1 (G1m marker)
|
|
chr16_+_82544327
|
1.235
|
NM_182981
|
OSGIN1
|
oxidative stress induced growth inhibitor 1
|
|
chr16_-_1369613
|
1.225
|
NM_001193389
NM_023076
|
UNKL
|
unkempt homolog (Drosophila)-like
|
|
chr16_-_28528780
|
1.224
|
NM_001055
NM_177529
|
SULT1A1
|
sulfotransferase family, cytosolic, 1A, phenol-preferring, member 1
|
|
chr3_-_195571748
|
1.215
|
NM_001135057
NM_130830
|
LRRC15
|
leucine rich repeat containing 15
|
|
chr17_+_71109734
|
1.197
|
|
MYO15B
|
myosin XVB pseudogene
|
|
chr4_+_22608252
|
1.171
|
|
|
|
|
chr14_-_95011790
|
1.168
|
NM_152592
|
C14orf49
|
chromosome 14 open reading frame 49
|
|
chr7_+_72883128
|
1.162
|
NM_001305
|
CLDN4
|
claudin 4
|
|
chr11_+_66764064
|
1.155
|
|
KDM2A
|
lysine (K)-specific demethylase 2A
|
|
chr14_-_106084519
|
1.141
|
|
IGHA1
IGHG1
|
immunoglobulin heavy constant alpha 1
immunoglobulin heavy constant gamma 1 (G1m marker)
|
|
chr18_+_148480
|
1.122
|
NM_001037334
NM_005151
|
USP14
|
ubiquitin specific peptidase 14 (tRNA-guanine transglycosylase)
|
|
chr8_-_145097031
|
1.118
|
NM_201380
|
PLEC
|
plectin
|
|
chr12_-_51585004
|
1.094
|
NM_002273
|
KRT8
|
keratin 8
|
|
chr6_-_29563657
|
1.091
|
NM_052967
|
MAS1L
|
MAS1 oncogene-like
|
|
chr1_+_27591738
|
1.078
|
NM_005281
|
GPR3
|
G protein-coupled receptor 3
|
|
chr7_+_111850367
|
1.074
|
NM_001007245
NM_001197080
|
IFRD1
|
interferon-related developmental regulator 1
|
|
chr9_-_139042526
|
1.061
|
NM_001606
|
ABCA2
|
ATP-binding cassette, sub-family A (ABC1), member 2
|
|
chr22_-_17846711
|
1.051
|
|
UFD1L
|
ubiquitin fusion degradation 1 like (yeast)
|
|
chr6_+_36518778
|
1.051
|
|
KCTD20
|
potassium channel tetramerisation domain containing 20
|
|
chr6_-_33655995
|
1.036
|
NM_001188
|
BAK1
|
BCL2-antagonist/killer 1
|
|
chr19_-_53914768
|
1.032
|
NM_001130915
|
MAMSTR
|
MEF2 activating motif and SAP domain containing transcriptional regulator
|
|
chr1_-_156784518
|
1.030
|
NM_001005189
|
OR6Y1
|
olfactory receptor, family 6, subfamily Y, member 1
|
|
chr22_+_43451600
|
1.028
|
|
PRR5
|
proline rich 5 (renal)
|
|
chr6_+_150112537
|
1.026
|
|
PCMT1
|
protein-L-isoaspartate (D-aspartate) O-methyltransferase
|
|
chr11_-_66898223
|
1.012
|
NM_001166212
|
CLCF1
|
cardiotrophin-like cytokine factor 1
|
|
chr6_-_33655811
|
1.011
|
|
BAK1
|
BCL2-antagonist/killer 1
|
|
chr22_-_17846725
|
1.008
|
NM_001035247
NM_005659
|
UFD1L
|
ubiquitin fusion degradation 1 like (yeast)
|
|
chr16_+_72888161
|
1.004
|
NM_002811
|
PSMD7
|
proteasome (prosome, macropain) 26S subunit, non-ATPase, 7
|
|
chr16_+_88696172
|
0.994
|
|
FAM157C
|
family with sequence similarity 157, member C
|
|
chr11_+_66764060
|
0.984
|
|
LOC100131150
|
hypothetical protein LOC100131150
|
|
chr22_-_17846621
|
0.980
|
|
UFD1L
|
ubiquitin fusion degradation 1 like (yeast)
|
|
chr19_-_56260135
|
0.969
|
NM_015596
|
KLK13
|
kallikrein-related peptidase 13
|
|
chr18_+_148586
|
0.961
|
|
USP14
|
ubiquitin specific peptidase 14 (tRNA-guanine transglycosylase)
|
|
chr19_-_40696393
|
0.942
|
NM_001126056
NM_001126057
NM_001126058
NM_001190347
NM_001190348
NM_001190349
NM_033317
|
DMKN
|
dermokine
|
|
chr2_+_161873031
|
0.940
|
NM_005805
|
PSMD14
|
proteasome (prosome, macropain) 26S subunit, non-ATPase, 14
|
|
chrX_-_100527799
|
0.939
|
NM_000061
|
BTK
|
Bruton agammaglobulinemia tyrosine kinase
|
|
chr1_-_6585268
|
0.925
|
|
KLHL21
|
kelch-like 21 (Drosophila)
|
|
chr1_-_199259447
|
0.917
|
NM_017596
|
KIF21B
|
kinesin family member 21B
|
|
chr9_-_139202855
|
0.914
|
NM_013366
|
ANAPC2
|
anaphase promoting complex subunit 2
|
|
chr16_+_30326864
|
0.903
|
|
ZNF771
|
zinc finger protein 771
|
|
chr1_+_3378030
|
0.895
|
|
ARHGEF16
|
Rho guanine nucleotide exchange factor (GEF) 16
|
|
chr9_+_139202874
|
0.884
|
NM_003731
|
SSNA1
|
Sjogren syndrome nuclear autoantigen 1
|
|
chr20_+_41621119
|
0.878
|
|
SGK2
|
serum/glucocorticoid regulated kinase 2
|
|
chr1_-_45760115
|
0.872
|
NM_002574
NM_181696
NM_181697
|
PRDX1
|
peroxiredoxin 1
|
|
chr19_-_43926953
|
0.869
|
NM_144691
|
CAPN12
|
calpain 12
|
|
chr16_-_30032330
|
0.866
|
NM_024307
|
GDPD3
|
glycerophosphodiester phosphodiesterase domain containing 3
|
|
chr9_-_139202804
|
0.857
|
|
ANAPC2
|
anaphase promoting complex subunit 2
|
|
chr9_-_139202831
|
0.852
|
|
ANAPC2
|
anaphase promoting complex subunit 2
|
|
chr15_-_62173041
|
0.848
|
NM_001014812
NM_032231
|
FAM96A
|
family with sequence similarity 96, member A
|
|
chr8_-_144887876
|
0.840
|
NM_198488
|
FAM83H
|
family with sequence similarity 83, member H
|
|
chr7_-_75280962
|
0.835
|
NM_002991
|
CCL24
|
chemokine (C-C motif) ligand 24
|
|
chr6_+_31782618
|
0.829
|
NM_001003693
|
LY6G6F
|
lymphocyte antigen 6 complex, locus G6F
|
|
chr20_+_41621079
|
0.827
|
NM_170693
|
SGK2
|
serum/glucocorticoid regulated kinase 2
|
|
chr17_-_43254132
|
0.815
|
NM_145798
|
OSBPL7
|
oxysterol binding protein-like 7
|
|
chr6_+_34314545
|
0.815
|
NM_145905
|
HMGA1
|
high mobility group AT-hook 1
|
|
chr16_-_28528149
|
0.813
|
NM_177530
NM_177534
|
SULT1A1
|
sulfotransferase family, cytosolic, 1A, phenol-preferring, member 1
|
|
chr17_-_43254117
|
0.810
|
|
OSBPL7
|
oxysterol binding protein-like 7
|
|
chr10_-_73280964
|
0.805
|
|
PSAP
|
prosaposin
|
|
chr10_-_73281047
|
0.805
|
|
PSAP
|
prosaposin
|
|
chr2_+_161873190
|
0.801
|
|
PSMD14
|
proteasome (prosome, macropain) 26S subunit, non-ATPase, 14
|
|
chr1_-_199259133
|
0.794
|
|
KIF21B
|
kinesin family member 21B
|
|
chr16_+_72888231
|
0.793
|
|
PSMD7
|
proteasome (prosome, macropain) 26S subunit, non-ATPase, 7
|
|
chr6_-_170704251
|
0.793
|
|
PSMB1
|
proteasome (prosome, macropain) subunit, beta type, 1
|
|
chr9_-_139042416
|
0.775
|
|
ABCA2
|
ATP-binding cassette, sub-family A (ABC1), member 2
|
|
chr21_-_35181956
|
0.774
|
|
RUNX1
|
runt-related transcription factor 1
|
|
chr22_+_21006814
|
0.769
|
|
|
|
|
chr6_-_28999621
|
0.756
|
|
TRIM27
|
tripartite motif containing 27
|
|
chr16_+_30326077
|
0.755
|
NM_001142305
NM_016643
|
ZNF771
|
zinc finger protein 771
|
|
chr6_+_44302973
|
0.754
|
|
SLC29A1
|
solute carrier family 29 (nucleoside transporters), member 1
|
|
chr6_+_150112511
|
0.748
|
NM_005389
|
PCMT1
|
protein-L-isoaspartate (D-aspartate) O-methyltransferase
|
|
chr3_+_122794859
|
0.747
|
NM_016298
|
FBXO40
|
F-box protein 40
|
|
chr19_-_51820133
|
0.744
|
NM_000960
|
PTGIR
|
prostaglandin I2 (prostacyclin) receptor (IP)
|
|
chr10_+_134751398
|
0.738
|
NM_001083909
|
GPR123
|
G protein-coupled receptor 123
|
|
chr22_+_20880112
|
0.732
|
|
|
|
|
chr16_+_65869558
|
0.729
|
NM_001129727
|
PLEKHG4
|
pleckstrin homology domain containing, family G (with RhoGef domain) member 4
|
|
chr16_+_30876115
|
0.726
|
NM_014712
|
SETD1A
|
SET domain containing 1A
|
|
chr15_-_38417564
|
0.717
|
|
C15orf52
|
chromosome 15 open reading frame 52
|
|
chr3_+_46894239
|
0.716
|
NM_000316
|
PTH1R
|
parathyroid hormone 1 receptor
|
|
chr7_+_130445394
|
0.712
|
NM_001145354
|
MKLN1
|
muskelin 1, intracellular mediator containing kelch motifs
|
|
chr17_-_59258762
|
0.709
|
NM_017647
|
FTSJ3
|
FtsJ homolog 3 (E. coli)
|
|
chr9_+_138760432
|
0.700
|
|
LOC100128593
|
hypothetical LOC100128593
|
|
chr10_-_73281056
|
0.700
|
NM_001042465
NM_001042466
NM_002778
|
PSAP
|
prosaposin
|
|
chr15_+_39638445
|
0.699
|
NM_006293
|
TYRO3
|
TYRO3 protein tyrosine kinase
|
|
chr6_-_30793436
|
0.697
|
NM_014641
|
MDC1
|
mediator of DNA-damage checkpoint 1
|
|
chr2_-_70901186
|
0.694
|
NM_173535
|
CLEC4F
|
C-type lectin domain family 4, member F
|
|
chr16_+_30326386
|
0.690
|
|
ZNF771
|
zinc finger protein 771
|
|
chr6_+_36518521
|
0.689
|
NM_173562
|
KCTD20
|
potassium channel tetramerisation domain containing 20
|
|
chr10_-_73281011
|
0.681
|
|
PSAP
|
prosaposin
|
|
chr22_-_25344037
|
0.678
|
|
CRYBB1
|
crystallin, beta B1
|
|
chr6_-_36518643
|
0.677
|
NM_152990
|
PXT1
|
peroxisomal, testis specific 1
|
|
chr7_-_150352484
|
0.671
|
NM_173681
|
ATG9B
|
ATG9 autophagy related 9 homolog B (S. cerevisiae)
|
|
chr1_-_6585485
|
0.666
|
NM_014851
|
KLHL21
|
kelch-like 21 (Drosophila)
|
|
chr2_+_161873239
|
0.665
|
|
PSMD14
|
proteasome (prosome, macropain) 26S subunit, non-ATPase, 14
|
|
chr1_-_93030538
|
0.661
|
NM_005665
|
EVI5
|
ecotropic viral integration site 5
|
|
chr17_-_53764867
|
0.661
|
|
|
|
|
chr16_+_88511787
|
0.653
|
NM_002386
|
MC1R
|
melanocortin 1 receptor (alpha melanocyte stimulating hormone receptor)
|
|
chr6_-_28999388
|
0.652
|
|
TRIM27
|
tripartite motif containing 27
|
|
chr3_+_142569988
|
0.649
|
|
ZBTB38
|
zinc finger and BTB domain containing 38
|
|
chr9_-_85512937
|
0.649
|
NM_013438
NM_053067
|
UBQLN1
|
ubiquilin 1
|
|
chr19_+_45564549
|
0.647
|
|
PLD3
|
phospholipase D family, member 3
|
|
chr9_-_115880372
|
0.640
|
|
AMBP
|
alpha-1-microglobulin/bikunin precursor
|
|
chr11_+_123600553
|
0.635
|
NM_001007249
|
OR8G2
|
olfactory receptor, family 8, subfamily G, member 2
|
|
chr16_+_85983499
|
0.629
|
|
MAP1LC3B
|
microtubule-associated protein 1 light chain 3 beta
|
|
chr9_-_129381086
|
0.626
|
NM_001035534
|
FAM129B
|
family with sequence similarity 129, member B
|
|
chr22_+_43451351
|
0.625
|
NM_001017530
NM_001198721
NM_015366
|
PRR5
|
proline rich 5 (renal)
|
|
chr17_-_70401299
|
0.619
|
NM_178128
|
FADS6
|
fatty acid desaturase domain family, member 6
|
|
chr6_+_150112549
|
0.619
|
|
PCMT1
|
protein-L-isoaspartate (D-aspartate) O-methyltransferase
|
|
chr20_+_48240772
|
0.610
|
NM_005194
|
CEBPB
|
CCAAT/enhancer binding protein (C/EBP), beta
|
|
chr16_+_88515917
|
0.607
|
NM_001197181
|
TUBB3
|
tubulin, beta 3
|
|
chr11_-_62539857
|
0.606
|
NM_001184732
NM_001184733
NM_001184736
NM_004254
|
SLC22A8
|
solute carrier family 22 (organic anion transporter), member 8
|
|
chr6_-_33374275
|
0.606
|
|
RGL2
|
ral guanine nucleotide dissociation stimulator-like 2
|
|
chr7_+_44045202
|
0.594
|
|
FLJ35390
|
hypothetical LOC255031
|
|
chr6_+_56927665
|
0.593
|
NM_152731
|
BEND6
|
BEN domain containing 6
|
|
chr11_-_14498443
|
0.587
|
|
PSMA1
|
proteasome (prosome, macropain) subunit, alpha type, 1
|
|
chr7_-_100667750
|
0.587
|
NM_014343
|
CLDN15
|
claudin 15
|
|
chr1_+_153232685
|
0.586
|
NM_018655
|
LENEP
|
lens epithelial protein
|
|
chr4_-_84152985
|
0.581
|
NM_001115007
|
LIN54
|
lin-54 homolog (C. elegans)
|
|
chr20_+_34206070
|
0.576
|
NM_012156
|
EPB41L1
|
erythrocyte membrane protein band 4.1-like 1
|
|
chr5_+_10406980
|
0.575
|
|
|
|
|
chr15_+_86983038
|
0.575
|
NM_002201
|
ISG20
|
interferon stimulated exonuclease gene 20kDa
|
|
chr22_-_18687511
|
0.566
|
NM_033257
|
DGCR6L
|
DiGeorge syndrome critical region gene 6-like
|
|
chr7_-_75826251
|
0.566
|
NM_012479
|
YWHAG
|
tyrosine 3-monooxygenase/tryptophan 5-monooxygenase activation protein, gamma polypeptide
|
|
chr16_+_85983375
|
0.565
|
|
MAP1LC3B
|
microtubule-associated protein 1 light chain 3 beta
|
|
chr22_+_18090118
|
0.563
|
|
SEPT5-GP1BB
|
SEPT5-GP1BB read-through transcript
|
|
chr8_+_144888275
|
0.562
|
|
LOC100128338
|
hypothetical LOC100128338
|
|
chr22_-_39189369
|
0.559
|
|
MKL1
|
megakaryoblastic leukemia (translocation) 1
|
|
chr19_-_4510713
|
0.558
|
|
SEMA6B
|
sema domain, transmembrane domain (TM), and cytoplasmic domain, (semaphorin) 6B
|
|
chrX_-_154216929
|
0.554
|
NM_001289
|
CLIC2
|
chloride intracellular channel 2
|
|
chr20_+_48240867
|
0.551
|
|
CEBPB
|
CCAAT/enhancer binding protein (C/EBP), beta
|
|
chr14_-_105589467
|
0.544
|
|
LOC100126583
IGHA1
IGHG1
IGHG2
IGHM
|
hypothetical LOC100126583
immunoglobulin heavy constant alpha 1
immunoglobulin heavy constant gamma 1 (G1m marker)
immunoglobulin heavy constant gamma 2 (G2m marker)
immunoglobulin heavy constant mu
|
|
chr11_+_72660894
|
0.542
|
NM_004154
|
P2RY6
|
pyrimidinergic receptor P2Y, G-protein coupled, 6
|
|
chr2_+_27358800
|
0.540
|
NM_032546
NM_187841
|
TRIM54
|
tripartite motif containing 54
|
|
chr14_-_106193403
|
0.538
|
|
IGKV3-20
|
immunoglobulin kappa variable 3-20
|
|
chr11_-_14498462
|
0.536
|
NM_001143937
NM_002786
|
PSMA1
|
proteasome (prosome, macropain) subunit, alpha type, 1
|
|
chr11_+_62405736
|
0.535
|
|
SLC3A2
|
solute carrier family 3 (activators of dibasic and neutral amino acid transport), member 2
|
|
chr16_-_66075216
|
0.527
|
NM_001138
|
AGRP
|
agouti related protein homolog (mouse)
|
|
chr16_+_29984647
|
0.527
|
|
ALDOA
|
aldolase A, fructose-bisphosphate
|
|
chr6_+_32003380
|
0.523
|
|
C2
|
complement component 2
|
|
chr1_-_161105174
|
0.522
|
NM_178550
|
C1orf110
|
chromosome 1 open reading frame 110
|
|
chr7_-_96177010
|
0.522
|
|
SHFM1
|
split hand/foot malformation (ectrodactyly) type 1
|
|
chr1_+_46578976
|
0.512
|
NM_199044
|
NSUN4
|
NOP2/Sun domain family, member 4
|
|
chr11_+_60309396
|
0.510
|
NM_206893
|
MS4A10
|
membrane-spanning 4-domains, subfamily A, member 10
|
|
chr4_-_26100986
|
0.507
|
NM_000730
|
CCKAR
|
cholecystokinin A receptor
|
|
chr19_+_56337368
|
0.507
|
NM_014385
NM_016543
|
SIGLEC7
|
sialic acid binding Ig-like lectin 7
|
|
chr15_-_71830868
|
0.502
|
NM_001039614
|
C15orf59
|
chromosome 15 open reading frame 59
|
|
chr20_+_48240705
|
0.500
|
|
CEBPB
|
CCAAT/enhancer binding protein (C/EBP), beta
|
|
chr19_-_59438411
|
0.495
|
NM_024318
|
LILRB3
LILRA6
|
leukocyte immunoglobulin-like receptor, subfamily B (with TM and ITIM domains), member 3
leukocyte immunoglobulin-like receptor, subfamily A (with TM domain), member 6
|
|
chr14_-_105785639
|
0.491
|
|
|
|
|
chrX_-_15421279
|
0.486
|
NM_001018109
NM_003662
|
PIR
|
pirin (iron-binding nuclear protein)
|
|
chr9_-_126309510
|
0.482
|
NM_004959
|
NR5A1
|
nuclear receptor subfamily 5, group A, member 1
|
|
chr11_+_74548472
|
0.481
|
NM_001145211
|
SLCO2B1
|
solute carrier organic anion transporter family, member 2B1
|