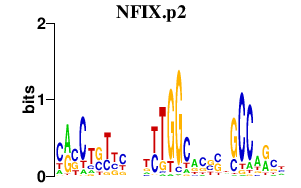
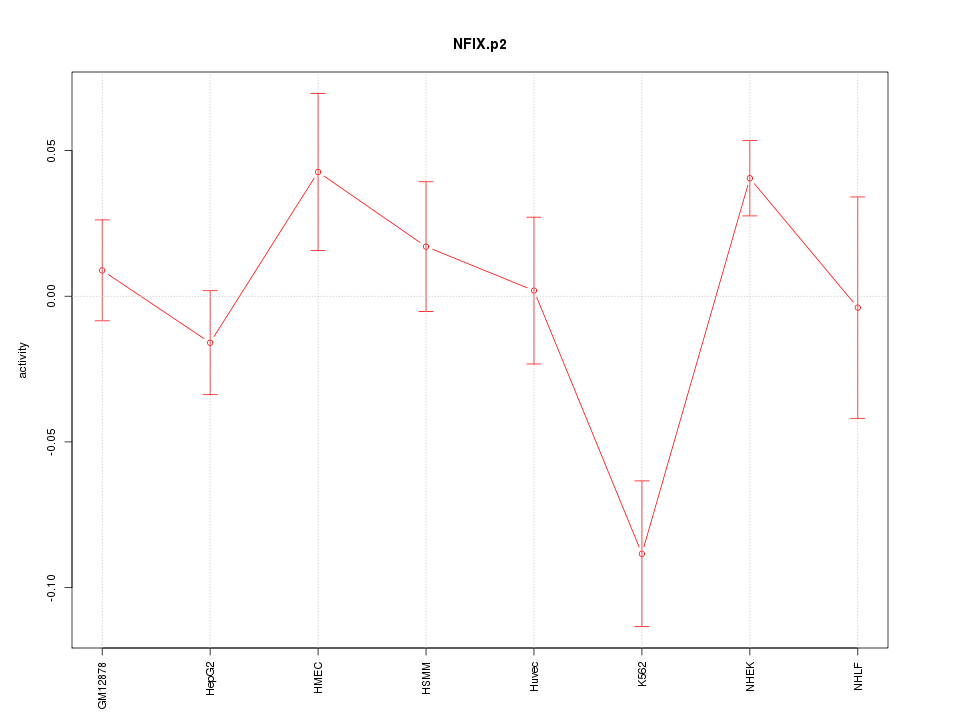
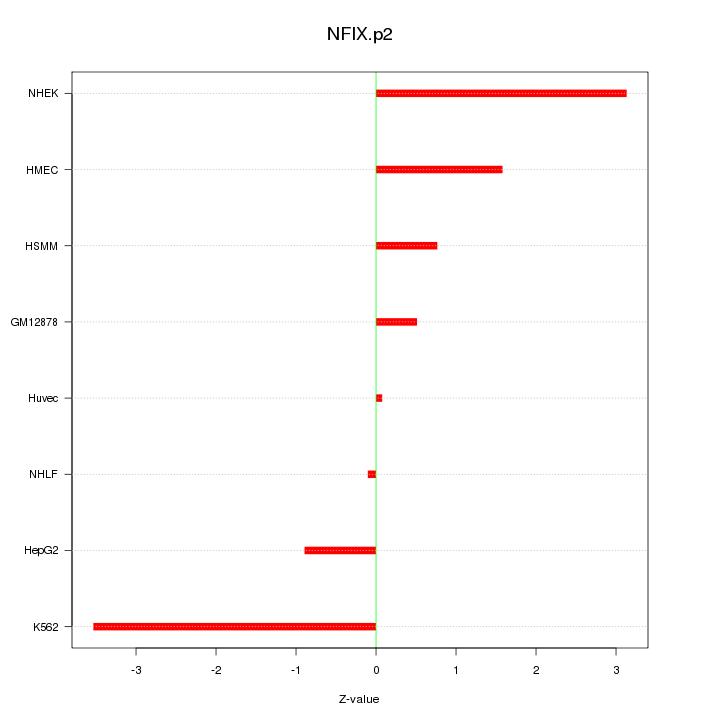
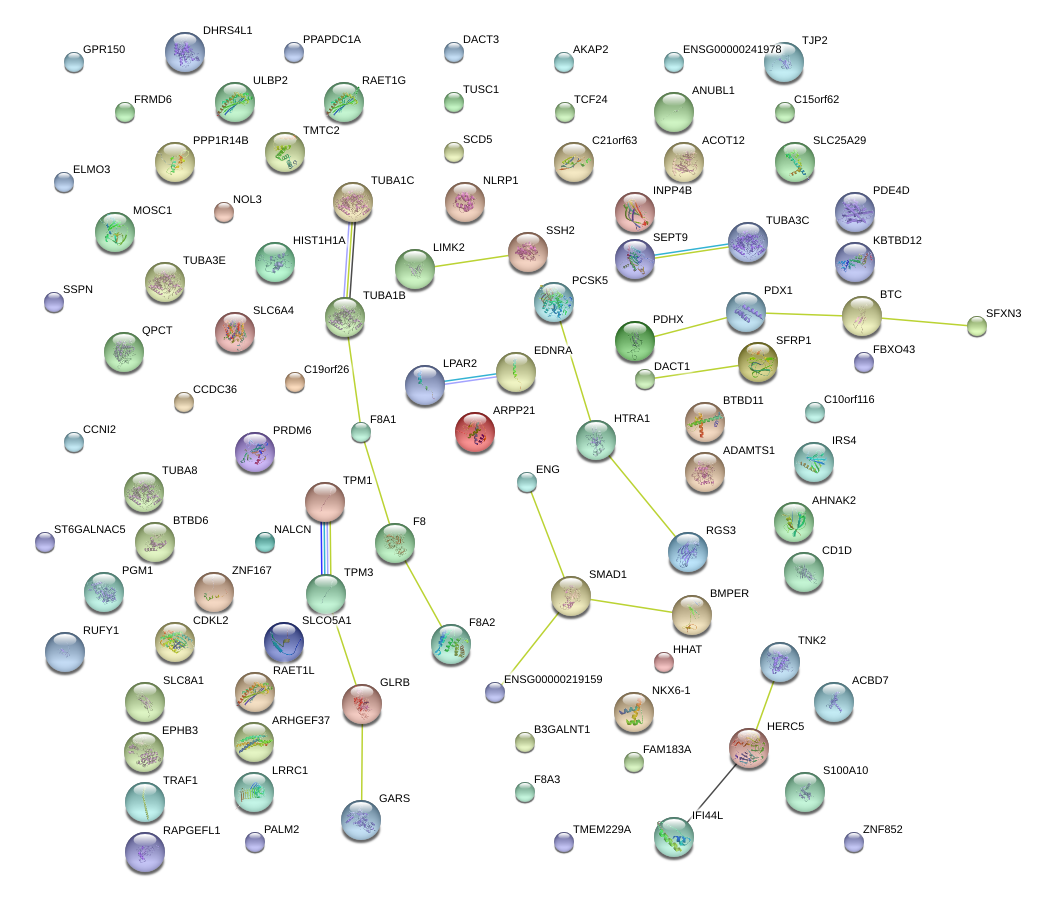
|
chr10_+_88718167
|
3.722
|
NM_006829
|
C10orf116
|
chromosome 10 open reading frame 116
|
|
chr17_-_25112393
|
3.554
|
|
SSH2
|
slingshot homolog 2 (Drosophila)
|
|
chr5_+_178919298
|
3.375
|
NM_001040451
NM_001040452
|
RUFY1
|
RUN and FYVE domain containing 1
|
|
chr8_-_101227203
|
3.018
|
NM_001029860
|
FBXO43
|
F-box protein 43
|
|
chr1_+_78858675
|
2.972
|
NM_006820
|
IFI44L
|
interferon-induced protein 44-like
|
|
chr1_+_156417302
|
2.878
|
|
CD1D
|
CD1d molecule
|
|
chr21_+_32706615
|
2.780
|
NM_058187
|
C21orf63
|
chromosome 21 open reading frame 63
|
|
chr14_+_51025604
|
2.754
|
NM_001042481
|
FRMD6
|
FERM domain containing 6
|
|
chr12_+_26239755
|
2.665
|
NM_005086
|
SSPN
|
sarcospan (Kras oncogene-associated gene)
|
|
chr5_+_148941228
|
2.636
|
NM_001001669
|
ARHGEF37
|
Rho guanine nucleotide exchange factor (GEF) 37
|
|
chr9_-_25668230
|
2.568
|
|
TUSC1
|
tumor suppressor candidate 1
|
|
chr4_-_85638410
|
2.461
|
NM_006168
|
NKX6-1
|
NK6 homeobox 1
|
|
chr17_-_5428491
|
2.450
|
NM_001033053
NM_014922
NM_033004
NM_033006
NM_033007
|
NLRP1
|
NLR family, pyrin domain containing 1
|
|
chr2_+_37425240
|
2.437
|
NM_012413
|
QPCT
|
glutaminyl-peptide cyclotransferase
|
|
chr2_-_56266408
|
2.399
|
|
LOC100129434
|
hypothetical LOC100129434
|
|
chr7_-_123460722
|
2.381
|
NM_001136002
|
TMEM229A
|
transmembrane protein 229A
|
|
chr4_+_158216631
|
2.347
|
NM_000824
NM_001166061
NM_001166060
|
GLRB
|
glycine receptor, beta
|
|
chr16_+_65765256
|
2.298
|
NM_003946
NM_001185057
|
NOL3
|
nucleolar protein 3 (apoptosis repressor with CARD domain)
|
|
chr6_-_26125939
|
2.160
|
NM_005325
|
HIST1H1A
|
histone cluster 1, H1a
|
|
chr6_+_53767251
|
2.120
|
|
LRRC1
|
leucine rich repeat containing 1
|
|
chr10_-_128884411
|
2.073
|
NM_001039762
|
FAM196A
|
family with sequence similarity 196, member A
|
|
chr22_+_44828338
|
2.018
|
|
|
|
|
chr17_+_35587767
|
1.983
|
NM_016339
|
RAPGEFL1
|
Rap guanine nucleotide exchange factor (GEF)-like 1
|
|
chr12_+_26239535
|
1.975
|
NM_001135823
|
SSPN
|
sarcospan (Kras oncogene-associated gene)
|
|
chr9_+_70926205
|
1.961
|
|
TJP2
|
tight junction protein 2 (zona occludens 2)
|
|
chr15_+_38849450
|
1.955
|
NM_001130448
|
C15orf62
|
chromosome 15 open reading frame 62
|
|
chr15_+_61123327
|
1.949
|
|
TPM1
|
tropomyosin 1 (alpha)
|
|
chr3_-_197103848
|
1.943
|
|
TNK2
|
tyrosine kinase, non-receptor, 2
|
|
chr10_+_122206455
|
1.941
|
NM_001030059
|
PPAPDC1A
|
phosphatidic acid phosphatase type 2 domain containing 1A
|
|
chr4_+_89597257
|
1.921
|
NM_016323
|
HERC5
|
hect domain and RLD 5
|
|
chr1_+_43386180
|
1.895
|
NM_001101376
|
FAM183A
|
family with sequence similarity 183, member A
|
|
chr9_-_21549696
|
1.852
|
|
LOC554202
|
hypothetical LOC554202
|
|
chr19_-_19599973
|
1.808
|
NM_004720
|
LPAR2
|
lysophosphatidic acid receptor 2
|
|
chr1_+_208568811
|
1.805
|
NM_001170587
NM_001170588
NM_018194
|
HHAT
|
hedgehog acyltransferase
|
|
chr1_+_77105746
|
1.799
|
NM_030965
|
ST6GALNAC5
|
ST6 (alpha-N-acetyl-neuraminyl-2,3-beta-galactosyl-1,3)-N-acetylgalactosaminide alpha-2,6-sialyltransferase 5
|
|
chr3_+_129117007
|
1.794
|
|
KBTBD12
|
kelch repeat and BTB (POZ) domain containing 12
|
|
chr4_-_143987053
|
1.772
|
NM_001101669
NM_003866
|
INPP4B
|
inositol polyphosphate-4-phosphatase, type II, 105kDa
|
|
chr9_+_111582317
|
1.767
|
NM_053016
NM_007203
NM_147150
|
PALM2
PALM2-AKAP2
|
paralemmin 2
PALM2-AKAP2 readthrough
|
|
chr4_-_75938710
|
1.765
|
NM_001729
|
BTC
|
betacellulin
|
|
chr1_+_219026661
|
1.759
|
NM_022746
|
MOSC1
|
MOCO sulphurase C-terminal domain containing 1
|
|
chr1_+_208569262
|
1.731
|
NM_001170580
|
HHAT
|
hedgehog acyltransferase
|
|
chr4_+_148621518
|
1.718
|
NM_001166055
NM_001957
|
EDNRA
|
endothelin receptor type A
|
|
chr14_+_104785898
|
1.710
|
NM_033271
|
BTBD6
|
BTB (POZ) domain containing 6
|
|
chr1_+_77105709
|
1.706
|
|
ST6GALNAC5
|
ST6 (alpha-N-acetyl-neuraminyl-2,3-beta-galactosyl-1,3)-N-acetylgalactosaminide alpha-2,6-sialyltransferase 5
|
|
chr3_+_44571600
|
1.671
|
NM_018651
NM_025169
|
ZNF167
|
zinc finger protein 167
|
|
chr9_+_70926009
|
1.669
|
NM_001170414
|
TJP2
|
tight junction protein 2 (zona occludens 2)
|
|
chr17_-_25587079
|
1.663
|
NM_001045
|
SLC6A4
|
solute carrier family 6 (neurotransmitter transporter, serotonin), member 4
|
|
chr10_+_102782151
|
1.646
|
|
SFXN3
|
sideroflexin 3
|
|
chr13_-_100866839
|
1.622
|
|
NALCN
|
sodium leak channel, non-selective
|
|
chr5_+_94981735
|
1.615
|
NM_199243
|
GPR150
|
G protein-coupled receptor 150
|
|
chrX_+_154264931
|
1.615
|
NM_001007523
NM_001007524
NM_012151
|
F8A2
F8A3
F8A1
|
coagulation factor VIII-associated (intronic transcript) 2
coagulation factor VIII-associated (intronic transcript) 3
coagulation factor VIII-associated (intronic transcript) 1
|
|
chr14_+_104785936
|
1.610
|
|
BTBD6
|
BTB (POZ) domain containing 6
|
|
chr14_+_58174489
|
1.600
|
NM_001079520
NM_016651
|
DACT1
|
dapper, antagonist of beta-catenin, homolog 1 (Xenopus laevis)
|
|
chr8_-_68037378
|
1.589
|
NM_001193502
|
TCF24
|
transcription factor 24
|
|
chr21_-_27139563
|
1.565
|
NM_006988
|
ADAMTS1
|
ADAM metallopeptidase with thrombospondin type 1 motif, 1
|
|
chr5_-_59819642
|
1.563
|
NM_001165899
|
PDE4D
|
phosphodiesterase 4D, cAMP-specific
|
|
chr19_-_1188840
|
1.530
|
NM_152769
|
C19orf26
|
chromosome 19 open reading frame 26
|
|
chr7_+_33911636
|
1.512
|
NM_133468
|
BMPER
|
BMP binding endothelial regulator
|
|
chr22_+_16973556
|
1.472
|
|
TUBA8
|
tubulin, alpha 8
|
|
chr19_+_45807909
|
1.469
|
|
|
|
|
chr15_-_30798417
|
1.459
|
|
|
|
|
chr6_-_150285838
|
1.437
|
NM_001001788
|
RAET1G
|
retinoic acid early transcript 1G
|
|
chr3_-_162305319
|
1.425
|
NM_033169
NM_001038628
NM_003781
NM_033167
NM_033168
|
B3GALNT1
|
beta-1,3-N-acetylgalactosaminyltransferase 1 (globoside blood group)
|
|
chr1_+_63832067
|
1.424
|
NM_001172819
|
PGM1
|
phosphoglucomutase 1
|
|
chr4_-_83938920
|
1.420
|
NM_001037582
NM_024906
|
SCD5
|
stearoyl-CoA desaturase 5
|
|
chr4_-_76774594
|
1.413
|
NM_003948
|
CDKL2
|
cyclin-dependent kinase-like 2 (CDC2-related kinase)
|
|
chr5_+_132111034
|
1.410
|
NM_001039780
|
CCNI2
|
cyclin I family, member 2
|
|
chr9_+_111850770
|
1.383
|
|
AKAP2
|
A kinase (PRKA) anchor protein 2
|
|
chr10_-_15170779
|
1.382
|
NM_001039844
|
ACBD7
|
acyl-CoA binding domain containing 7
|
|
chr9_-_129656413
|
1.372
|
|
ENG
|
endoglin
|
|
chr3_+_185762233
|
1.367
|
NM_004443
|
EPHB3
|
EPH receptor B3
|
|
chr16_+_65790514
|
1.341
|
NM_024712
|
ELMO3
|
engulfment and cell motility 3
|
|
chr4_+_146622154
|
1.338
|
NM_005900
|
SMAD1
|
SMAD family member 1
|
|
chr22_+_29974368
|
1.335
|
|
LIMK2
|
LIM domain kinase 2
|
|
chr2_-_40532651
|
1.327
|
|
SLC8A1
|
solute carrier family 8 (sodium/calcium exchanger), member 1
|
|
chr5_+_122452739
|
1.325
|
NM_001136239
|
PRDM6
|
PR domain containing 6
|
|
chr14_-_104507862
|
1.319
|
|
AHNAK2
|
AHNAK nucleoprotein 2
|
|
chr10_-_105982052
|
1.315
|
NM_025145
|
WDR96
|
WD repeat domain 96
|
|
chr14_+_23575460
|
1.315
|
NM_001082488
|
DHRS4L1
|
dehydrogenase/reductase (SDR family) member 4 like 1
|
|
chr3_+_49211936
|
1.314
|
NM_001135197
|
CCDC36
|
coiled-coil domain containing 36
|
|
chr3_+_35656012
|
1.303
|
|
ARPP21
|
cAMP-regulated phosphoprotein, 21kDa
|
|
chr17_+_72795567
|
1.298
|
NM_001113492
|
SEPT9
|
septin 9
|
|
chr9_-_25668855
|
1.283
|
NM_001004125
|
TUSC1
|
tumor suppressor candidate 1
|
|
chr4_+_148621694
|
1.279
|
|
EDNRA
|
endothelin receptor type A
|
|
chr9_+_77695334
|
1.278
|
NM_001190482
NM_006200
|
PCSK5
|
proprotein convertase subtilisin/kexin type 5
|
|
chr10_+_124210980
|
1.272
|
NM_002775
|
HTRA1
|
HtrA serine peptidase 1
|
|
chr12_+_106236281
|
1.261
|
NM_001018072
|
BTBD11
|
BTB (POZ) domain containing 11
|
|
chr22_+_16973399
|
1.251
|
NM_001193414
NM_018943
|
TUBA8
|
tubulin, alpha 8
|
|
chr9_+_115382990
|
1.247
|
|
RGS3
|
regulator of G-protein signaling 3
|
|
chr1_-_150233337
|
1.240
|
NM_002966
|
S100A10
|
S100 calcium binding protein A10
|
|
chr13_+_27392136
|
1.239
|
NM_000209
|
PDX1
|
pancreatic and duodenal homeobox 1
|
|
chr12_+_81604752
|
1.227
|
|
TMTC2
|
transmembrane and tetratricopeptide repeat containing 2
|
|
chr9_+_111850698
|
1.216
|
NM_001004065
NM_001198656
|
AKAP2
|
A kinase (PRKA) anchor protein 2
|
|
chr8_-_70908130
|
1.213
|
NM_001146008
|
SLCO5A1
|
solute carrier organic anion transporter family, member 5A1
|
|
chr14_-_99842519
|
1.204
|
NM_001039355
|
SLC25A29
|
solute carrier family 25, member 29
|
|
chrX_-_107866262
|
1.202
|
NM_003604
|
IRS4
|
insulin receptor substrate 4
|
|
chr8_-_41286088
|
1.198
|
NM_003012
|
SFRP1
|
secreted frizzled-related protein 1
|
|
chr10_-_45488162
|
1.186
|
NM_001128324
NM_174890
|
ANUBL1
|
AN1, ubiquitin-like, homolog (Xenopus laevis)
|
|
chr5_-_80725704
|
1.171
|
NM_130767
|
ACOT12
|
acyl-CoA thioesterase 12
|
|
chr9_-_122731263
|
1.158
|
NM_001190945
|
TRAF1
|
TNF receptor-associated factor 1
|
|
chr14_-_99695684
|
1.148
|
NM_206918
|
DEGS2
|
degenerative spermatocyte homolog 2, lipid desaturase (Drosophila)
|
|
chr1_-_19487790
|
1.139
|
NM_012067
|
AKR7A3
|
aldo-keto reductase family 7, member A3 (aflatoxin aldehyde reductase)
|
|
chr9_-_129656581
|
1.134
|
|
ENG
|
endoglin
|
|
chr15_-_83002421
|
1.132
|
|
NMB
|
neuromedin B
|
|
chr1_+_148521852
|
1.128
|
NM_144697
|
C1orf51
|
chromosome 1 open reading frame 51
|
|
chrX_+_10084984
|
1.128
|
NM_001830
|
CLCN4
|
chloride channel 4
|
|
chr4_+_148621567
|
1.124
|
|
EDNRA
|
endothelin receptor type A
|
|
chr9_+_115382760
|
1.123
|
NM_021106
|
RGS3
|
regulator of G-protein signaling 3
|
|
chr1_+_199975674
|
1.115
|
|
NAV1
|
neuron navigator 1
|
|
chr5_+_96024434
|
1.106
|
|
CAST
|
calpastatin
|
|
chr10_+_71232281
|
1.081
|
|
COL13A1
|
collagen, type XIII, alpha 1
|
|
chr11_-_30564338
|
1.055
|
NM_001145399
|
MPPED2
|
metallophosphoesterase domain containing 2
|
|
chr4_+_166097548
|
1.052
|
NM_153027
|
C4orf39
|
chromosome 4 open reading frame 39
|
|
chr1_+_6008956
|
1.037
|
NM_003636
NM_172130
|
KCNAB2
|
potassium voltage-gated channel, shaker-related subfamily, beta member 2
|
|
chr15_+_61122957
|
1.035
|
|
TPM1
|
tropomyosin 1 (alpha)
|
|
chr3_-_50371896
|
1.035
|
|
TMEM115
|
transmembrane protein 115
|
|
chr1_+_226261374
|
1.022
|
NM_033131
|
WNT3A
|
wingless-type MMTV integration site family, member 3A
|
|
chr6_-_55551812
|
1.016
|
NM_001042406
NM_019036
|
HMGCLL1
|
3-hydroxymethyl-3-methylglutaryl-CoA lyase-like 1
|
|
chr5_+_76418545
|
1.013
|
|
LOC728723
|
hypothetical LOC728723
|
|
chr21_-_27139143
|
1.011
|
|
ADAMTS1
|
ADAM metallopeptidase with thrombospondin type 1 motif, 1
|
|
chr11_-_75057299
|
1.007
|
|
MAP6
|
microtubule-associated protein 6
|
|
chr4_+_156899575
|
0.983
|
NM_000857
|
GUCY1B3
|
guanylate cyclase 1, soluble, beta 3
|
|
chr10_+_89409332
|
0.977
|
NM_001015880
NM_004670
|
PAPSS2
|
3'-phosphoadenosine 5'-phosphosulfate synthase 2
|
|
chr1_+_199975563
|
0.976
|
NM_001167738
|
NAV1
|
neuron navigator 1
|
|
chr11_-_63014986
|
0.973
|
NM_001146728
NM_001146729
NM_054108
|
HRASLS5
|
HRAS-like suppressor family, member 5
|
|
chr5_+_76418214
|
0.956
|
|
LOC728723
|
hypothetical LOC728723
|
|
chr3_-_46710137
|
0.953
|
NM_001190707
NM_147129
|
ALS2CL
|
ALS2 C-terminal like
|
|
chr9_-_112840801
|
0.950
|
|
LPAR1
|
lysophosphatidic acid receptor 1
|
|
chr2_-_127694123
|
0.949
|
|
CYP27C1
|
cytochrome P450, family 27, subfamily C, polypeptide 1
|
|
chr3_-_50371934
|
0.948
|
NM_007024
|
TMEM115
|
transmembrane protein 115
|
|
chr1_-_234295293
|
0.945
|
|
NID1
|
nidogen 1
|
|
chr9_-_111300350
|
0.943
|
NM_001145368
NM_002829
|
PTPN3
|
protein tyrosine phosphatase, non-receptor type 3
|
|
chr1_-_68470585
|
0.942
|
NM_001002292
NM_001193334
NM_024911
|
WLS
|
wntless homolog (Drosophila)
|
|
chr6_+_86216020
|
0.936
|
NM_002526
|
NT5E
|
5'-nucleotidase, ecto (CD73)
|
|
chr20_-_3696344
|
0.936
|
NM_001039140
|
C20orf27
|
chromosome 20 open reading frame 27
|
|
chrX_-_20044670
|
0.934
|
NM_001168467
|
MAP7D2
|
MAP7 domain containing 2
|
|
chr1_+_26744929
|
0.930
|
NM_001006665
|
RPS6KA1
|
ribosomal protein S6 kinase, 90kDa, polypeptide 1
|
|
chr10_-_123346148
|
0.924
|
NM_001144915
|
FGFR2
|
fibroblast growth factor receptor 2
|
|
chr6_-_119512035
|
0.922
|
NM_001100411
|
FAM184A
|
family with sequence similarity 184, member A
|
|
chr11_+_122258600
|
0.919
|
NM_024806
NM_199124
|
C11orf63
|
chromosome 11 open reading frame 63
|
|
chr13_-_100866706
|
0.914
|
NM_052867
|
NALCN
|
sodium leak channel, non-selective
|
|
chr11_+_10428799
|
0.913
|
NM_000480
|
AMPD3
|
adenosine monophosphate deaminase 3
|
|
chr2_-_96174502
|
0.912
|
|
DUSP2
|
dual specificity phosphatase 2
|
|
chr5_-_169340321
|
0.909
|
NM_001129891
|
FAM196B
|
family with sequence similarity 196, member B
|
|
chr9_-_129656778
|
0.909
|
|
ENG
|
endoglin
|
|
chr21_+_25856311
|
0.908
|
|
MIR155HG
|
MIR155 host gene (non-protein coding)
|
|
chr1_-_159306327
|
0.902
|
|
ARHGAP30
|
Rho GTPase activating protein 30
|
|
chr1_-_72520735
|
0.902
|
NM_173808
|
NEGR1
|
neuronal growth regulator 1
|
|
chr22_+_29974314
|
0.899
|
NM_001031801
NM_016733
|
LIMK2
|
LIM domain kinase 2
|
|
chr9_-_83493415
|
0.897
|
NM_005077
|
TLE1
|
transducin-like enhancer of split 1 (E(sp1) homolog, Drosophila)
|
|
chr3_+_127170717
|
0.895
|
NM_001012337
|
ROPN1B
|
rhophilin associated tail protein 1B
|
|
chr3_-_50371960
|
0.893
|
|
TMEM115
|
transmembrane protein 115
|
|
chr1_-_159306273
|
0.890
|
|
ARHGAP30
|
Rho GTPase activating protein 30
|
|
chr8_-_70907957
|
0.888
|
NM_001146009
|
SLCO5A1
|
solute carrier organic anion transporter family, member 5A1
|
|
chr8_+_24827173
|
0.887
|
NM_005382
|
NEFM
|
neurofilament, medium polypeptide
|
|
chr1_-_115433570
|
0.886
|
NM_005725
|
TSPAN2
|
tetraspanin 2
|
|
chr17_+_18069593
|
0.886
|
NM_004140
|
LLGL1
|
lethal giant larvae homolog 1 (Drosophila)
|
|
chr17_-_68100537
|
0.883
|
|
LOC100499467
|
hypothetical LOC100499467
|
|
chr4_+_166097579
|
0.882
|
|
C4orf39
|
chromosome 4 open reading frame 39
|
|
chr1_+_20788027
|
0.880
|
NM_001785
|
CDA
|
cytidine deaminase
|
|
chr1_+_46632505
|
0.878
|
NM_001441
|
FAAH
|
fatty acid amide hydrolase
|
|
chr17_-_53335748
|
0.876
|
NM_017949
|
CUEDC1
|
CUE domain containing 1
|
|
chr9_-_129656830
|
0.873
|
NM_000118
NM_001114753
|
ENG
|
endoglin
|
|
chr12_+_81605032
|
0.872
|
NM_152588
|
TMTC2
|
transmembrane and tetratricopeptide repeat containing 2
|
|
chr4_+_3264509
|
0.865
|
|
RGS12
|
regulator of G-protein signaling 12
|
|
chr3_+_191714533
|
0.863
|
NM_001167928
NM_001167929
NM_001167930
NM_001167931
NM_002182
NM_134470
|
IL1RAP
|
interleukin 1 receptor accessory protein
|
|
chr10_-_91001496
|
0.858
|
|
LIPA
|
lipase A, lysosomal acid, cholesterol esterase
|
|
chr15_+_39739864
|
0.849
|
NM_001080541
NM_001164273
|
MGA
|
MAX gene associated
|
|
chr18_-_24010944
|
0.845
|
|
CDH2
|
cadherin 2, type 1, N-cadherin (neuronal)
|
|
chr5_-_157011949
|
0.845
|
NM_007017
NM_178424
|
SOX30
|
SRY (sex determining region Y)-box 30
|
|
chr11_-_11986704
|
0.840
|
NM_001018057
|
DKK3
|
dickkopf homolog 3 (Xenopus laevis)
|
|
chr11_+_34599227
|
0.840
|
|
EHF
|
ets homologous factor
|
|
chr4_+_107036042
|
0.837
|
NM_001033047
NM_001184690
NM_001184691
NM_001184692
NM_001184693
|
NPNT
|
nephronectin
|
|
chr1_-_234295117
|
0.835
|
|
NID1
|
nidogen 1
|
|
chr10_-_123343320
|
0.834
|
NM_001144913
NM_001144914
|
FGFR2
|
fibroblast growth factor receptor 2
|
|
chr1_+_891739
|
0.832
|
NM_001160184
NM_032129
|
PLEKHN1
|
pleckstrin homology domain containing, family N member 1
|
|
chr15_-_83002797
|
0.830
|
NM_021077
NM_205858
|
NMB
|
neuromedin B
|
|
chr21_+_33364316
|
0.825
|
NM_138983
|
OLIG1
|
oligodendrocyte transcription factor 1
|
|
chr12_+_50271283
|
0.820
|
NM_014191
|
SCN8A
|
sodium channel, voltage gated, type VIII, alpha subunit
|
|
chr6_+_150326802
|
0.816
|
NM_025218
|
ULBP1
|
UL16 binding protein 1
|
|
chr2_-_96174865
|
0.813
|
NM_004418
|
DUSP2
|
dual specificity phosphatase 2
|
|
chr11_+_125262164
|
0.810
|
NM_001134793
|
HYLS1
|
hydrolethalus syndrome 1
|
|
chrX_-_100070553
|
0.797
|
NM_212559
|
XKRX
|
XK, Kell blood group complex subunit-related, X-linked
|
|
chr11_+_63815348
|
0.797
|
NM_033310
|
KCNK4
|
potassium channel, subfamily K, member 4
|
|
chr17_+_6495694
|
0.796
|
NM_001105520
|
C17orf100
|
chromosome 17 open reading frame 100
|
|
chr16_-_674357
|
0.793
|
NM_001005920
|
JMJD8
|
jumonji domain containing 8
|
|
chr14_-_103248193
|
0.784
|
|
XRCC3
|
X-ray repair complementing defective repair in Chinese hamster cells 3
|
|
chr7_-_76883642
|
0.783
|
NM_017439
|
PION
|
pigeon homolog (Drosophila)
|
|
chr2_-_225615556
|
0.780
|
NM_014689
|
DOCK10
|
dedicator of cytokinesis 10
|
|
chr4_-_99798563
|
0.777
|
NM_005723
|
TSPAN5
|
tetraspanin 5
|
|
chr12_+_122812994
|
0.774
|
NM_207437
|
DNAH10
|
dynein, axonemal, heavy chain 10
|
|
chr17_-_78249750
|
0.769
|
NM_006822
|
RAB40B
|
RAB40B, member RAS oncogene family
|
|
chr12_+_74160779
|
0.766
|
NM_006851
|
GLIPR1
|
GLI pathogenesis-related 1
|
|
chr3_-_126414252
|
0.761
|
NM_024628
|
SLC12A8
|
solute carrier family 12 (potassium/chloride transporters), member 8
|
|
chr17_+_74531873
|
0.760
|
|
C1QTNF1
|
C1q and tumor necrosis factor related protein 1
|
|
chr22_+_38720881
|
0.759
|
NM_138435
|
FAM83F
|
family with sequence similarity 83, member F
|
|
chr4_-_154900466
|
0.753
|
|
|
|
|
chr11_+_124438407
|
0.752
|
|
SLC37A2
|
solute carrier family 37 (glycerol-3-phosphate transporter), member 2
|
|
chr3_+_32408346
|
0.751
|
|
CMTM7
|
CKLF-like MARVEL transmembrane domain containing 7
|
|
chr9_+_5440455
|
0.746
|
NM_014143
|
CD274
|
CD274 molecule
|