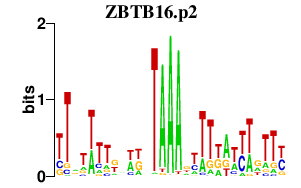
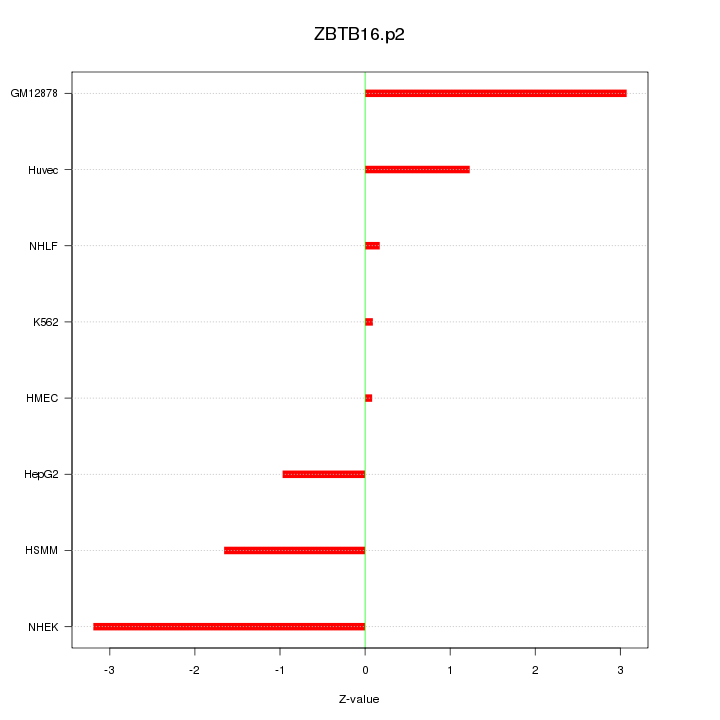
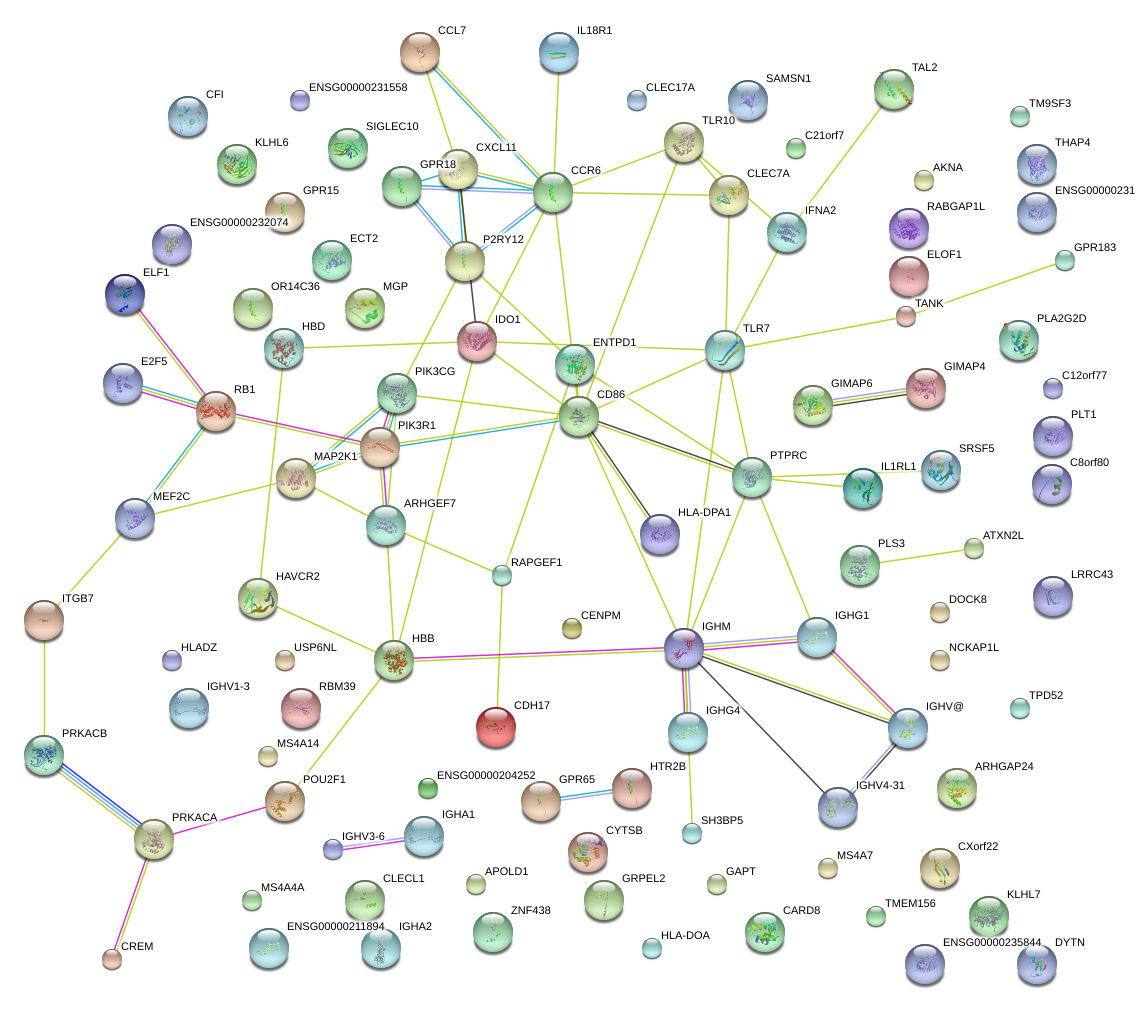
|
chr19_-_53444733
|
5.109
|
NM_001184901
NM_001184903
NM_014959
|
CARD8
|
caspase recruitment domain family, member 8
|
|
chr13_-_98708549
|
3.069
|
NM_001098200
NM_005292
|
GPR18
|
G protein-coupled receptor 18
|
|
chr2_+_161701711
|
3.039
|
NM_004180
NM_133484
|
TANK
|
TRAF family member-associated NFKB activator
|
|
chr12_-_51887318
|
2.894
|
|
ITGB7
|
integrin, beta 7
|
|
chr2_-_242205556
|
2.820
|
NM_001164356
|
THAP4
|
THAP domain containing 4
|
|
chr3_+_188197029
|
2.615
|
|
|
|
|
chr12_-_51887266
|
2.522
|
NM_000889
|
ITGB7
|
integrin, beta 7
|
|
chr4_-_38460983
|
2.176
|
NM_001017388
NM_001195106
NM_001195107
NM_001195108
NM_030956
|
TLR10
|
toll-like receptor 10
|
|
chr8_+_39890484
|
2.168
|
NM_002164
|
IDO1
|
indoleamine 2,3-dioxygenase 1
|
|
chr8_+_86287161
|
2.055
|
NM_001083589
|
E2F5
|
E2F transcription factor 5, p130-binding
|
|
chr12_+_121218218
|
1.889
|
NM_152759
|
LRRC43
|
leucine rich repeat containing 43
|
|
chr7_+_106293112
|
1.876
|
NM_002649
|
PIK3CG
|
phosphoinositide-3-kinase, catalytic, gamma polypeptide
|
|
chr19_+_14554895
|
1.874
|
NM_207390
|
CLEC17A
|
C-type lectin domain family 17, member A
|
|
chr13_-_40454360
|
1.853
|
NM_001145353
|
ELF1
|
E74-like factor 1 (ets domain transcription factor)
|
|
chr9_-_70780574
|
1.795
|
|
|
|
|
chr9_-_116196505
|
1.769
|
NM_030767
|
AKNA
|
AT-hook transcription factor
|
|
chr1_+_173111306
|
1.752
|
NM_001035230
|
RABGAP1L
|
RAB GTPase activating protein 1-like
|
|
chr2_-_231697999
|
1.751
|
NM_000867
|
HTR2B
|
5-hydroxytryptamine (serotonin) receptor 2B
|
|
chr10_-_11614279
|
1.660
|
NM_001080491
|
USP6NL
|
USP6 N-terminal like
|
|
chr21_+_29374743
|
1.652
|
NM_020152
|
C21orf7
|
chromosome 21 open reading frame 7
|
|
chr13_-_98757634
|
1.623
|
NM_004951
|
GPR183
|
G protein-coupled receptor 183
|
|
chr12_-_25041639
|
1.560
|
NM_001101339
|
C12orf77
|
chromosome 12 open reading frame 77
|
|
chr10_-_31328426
|
1.529
|
NM_001143769
|
ZNF438
|
zinc finger protein 438
|
|
chr3_+_173954940
|
1.524
|
NM_018098
|
ECT2
|
epithelial cell transforming sequence 2 oncogene
|
|
chr4_+_86918882
|
1.498
|
|
ARHGAP24
|
Rho GTPase activating protein 24
|
|
chr13_-_40454188
|
1.497
|
|
ELF1
|
E74-like factor 1 (ets domain transcription factor)
|
|
chr5_-_88155360
|
1.454
|
NM_001193348
NM_001193349
|
MEF2C
|
myocyte enhancer factor 2C
|
|
chr10_-_98315090
|
1.346
|
|
TM9SF3
|
transmembrane 9 superfamily member 3
|
|
chr19_-_46504831
|
1.338
|
|
|
|
|
chr3_-_152585226
|
1.330
|
NM_022788
|
P2RY12
|
purinergic receptor P2Y, G-protein coupled, 12
|
|
chr17_+_19917420
|
1.280
|
|
SPECC1
|
sperm antigen with calponin homology and coiled-coil domains 1
|
|
chr3_+_99733432
|
1.280
|
NM_005290
|
GPR15
|
G protein-coupled receptor 15
|
|
chr19_-_56612653
|
1.275
|
NM_001171156
NM_001171157
NM_001171158
NM_001171159
NM_001171160
NM_001171161
NM_033130
|
SIGLEC10
|
sialic acid binding Ig-like lectin 10
|
|
chr9_+_263025
|
1.273
|
NM_001190458
NM_001193536
|
DOCK8
|
dedicator of cytokinesis 8
|
|
chr1_+_165565006
|
1.216
|
|
POU2F1
|
POU class 2 homeobox 1
|
|
chr10_+_35466715
|
1.214
|
NM_183011
NM_183012
|
CREM
|
cAMP responsive element modulator
|
|
chr13_+_110637173
|
1.168
|
NM_001113513
|
ARHGEF7
|
Rho guanine nucleotide exchange factor (GEF) 7
|
|
chr11_+_59920062
|
1.161
|
NM_001079692
NM_032597
|
MS4A14
|
membrane-spanning 4-domains, subfamily A, member 14
|
|
chr3_-_184756101
|
1.158
|
NM_130446
|
KLHL6
|
kelch-like 6 (Drosophila)
|
|
chr12_+_53178834
|
1.149
|
NM_001184976
|
NCKAP1L
|
NCK-associated protein 1-like
|
|
chr10_+_97505398
|
1.136
|
NM_001164178
NM_001164181
NM_001164179
NM_001164182
NM_001164183
NM_001776
|
ENTPD1
|
ectonucleoside triphosphate diphosphohydrolase 1
|
|
chr7_+_149895390
|
1.134
|
NM_018326
|
GIMAP4
|
GTPase, IMAP family member 4
|
|
chr7_-_149960218
|
1.132
|
|
GIMAP6
|
GTPase, IMAP family member 6
|
|
chr21_-_14840506
|
1.112
|
NM_022136
|
SAMSN1
|
SAM domain, SH3 domain and nuclear localization signals 1
|
|
chr12_-_9777126
|
1.104
|
NM_172004
|
CLECL1
|
C-type lectin-like 1
|
|
chr20_-_33783409
|
1.069
|
|
RBM39
|
RNA binding motif protein 39
|
|
chr9_-_21375258
|
1.068
|
NM_000605
|
IFNA2
|
interferon, alpha 2
|
|
chr12_-_10173947
|
1.063
|
NM_022570
NM_197947
NM_197948
NM_197949
NM_197950
NM_197954
|
CLEC7A
|
C-type lectin domain family 7, member A
|
|
chr4_-_77176163
|
1.062
|
NM_005409
|
CXCL11
|
chemokine (C-X-C motif) ligand 11
|
|
chr5_+_67622242
|
1.054
|
NM_181504
|
PIK3R1
|
phosphoinositide-3-kinase, regulatory subunit 1 (alpha)
|
|
chrX_+_12795111
|
1.048
|
NM_016562
|
TLR7
|
toll-like receptor 7
|
|
chr14_+_69306019
|
1.041
|
|
SRSF5
|
serine/arginine-rich splicing factor 5
|
|
chr13_+_110653407
|
1.028
|
|
ARHGEF7
|
Rho guanine nucleotide exchange factor (GEF) 7
|
|
chr1_+_196874759
|
1.011
|
NM_002838
NM_080921
NM_080923
|
PTPRC
|
protein tyrosine phosphatase, receptor type, C
|
|
chr2_-_207291364
|
0.986
|
NM_001093730
|
DYTN
|
dystrotelin
|
|
chr3_-_15357878
|
0.978
|
NM_001018009
|
SH3BP5
|
SH3-domain binding protein 5 (BTK-associated)
|
|
chr8_-_81155564
|
0.972
|
NM_001025252
|
TPD52
|
tumor protein D52
|
|
chr9_+_107464558
|
0.965
|
NM_005421
|
TAL2
|
T-cell acute lymphocytic leukemia 2
|
|
chr8_-_95298706
|
0.963
|
NM_001144663
|
CDH17
|
cadherin 17, LI cadherin (liver-intestine)
|
|
chr6_-_33149244
|
0.963
|
NM_033554
|
HLA-DPA1
|
major histocompatibility complex, class II, DP alpha 1
|
|
chr9_-_133602745
|
0.960
|
NM_005312
|
RAPGEF1
|
Rap guanine nucleotide exchange factor (GEF) 1
|
|
chr11_-_5212168
|
0.956
|
NM_000519
|
HBD
|
hemoglobin, delta
|
|
chr2_+_102294393
|
0.951
|
NM_016232
|
IL18R1
IL1RL1
|
interleukin 18 receptor 1
interleukin 1 receptor-like 1
|
|
chr5_+_148711419
|
0.929
|
|
GRPEL2
|
GrpE-like 2, mitochondrial (E. coli)
|
|
chr11_+_59902598
|
0.928
|
|
MS4A7
|
membrane-spanning 4-domains, subfamily A, member 7
|
|
chr12_+_63510481
|
0.914
|
|
|
|
|
chr7_+_37689903
|
0.895
|
|
|
|
|
chr1_+_84382575
|
0.888
|
|
PRKACB
|
protein kinase, cAMP-dependent, catalytic, beta
|
|
chr5_+_57823017
|
0.887
|
NM_152687
|
GAPT
|
GRB2-binding adaptor protein, transmembrane
|
|
chr12_+_12829807
|
0.886
|
NM_030817
|
APOLD1
|
apolipoprotein L domain containing 1
|
|
chr15_+_64481270
|
0.883
|
|
MAP2K1
|
mitogen-activated protein kinase kinase 1
|
|
chr8_-_27997306
|
0.881
|
NM_001010906
|
C8orf80
|
chromosome 8 open reading frame 80
|
|
chr1_+_196874867
|
0.865
|
|
PTPRC
|
protein tyrosine phosphatase, receptor type, C
|
|
chr12_-_14929991
|
0.862
|
NM_000900
NM_001190839
|
MGP
|
matrix Gla protein
|
|
chr1_+_196874839
|
0.854
|
|
PTPRC
|
protein tyrosine phosphatase, receptor type, C
|
|
chr13_+_47948586
|
0.851
|
|
RB1
|
retinoblastoma 1
|
|
chr1_-_20318578
|
0.839
|
NM_012400
|
PLA2G2D
|
phospholipase A2, group IID
|
|
chr14_+_87541220
|
0.838
|
NM_003608
|
GPR65
|
G protein-coupled receptor 65
|
|
chr4_-_110942576
|
0.829
|
|
CFI
|
complement factor I
|
|
chr5_-_156468614
|
0.822
|
NM_032782
|
HAVCR2
|
hepatitis A virus cellular receptor 2
|
|
chr17_+_29621352
|
0.821
|
NM_006273
|
CCL7
|
chemokine (C-C motif) ligand 7
|
|
chr22_-_40666168
|
0.821
|
NM_001110215
|
CENPM
|
centromere protein M
|
|
chr1_+_246578699
|
0.819
|
NM_001001918
|
OR14C36
|
olfactory receptor, family 14, subfamily C, member 36
|
|
chrX_+_114734074
|
0.818
|
NM_001136025
NM_001172335
|
PLS3
|
plastin 3
|
|
chr6_+_167445284
|
0.817
|
NM_004367
|
CCR6
|
chemokine (C-C motif) receptor 6
|
|
chr4_-_38710376
|
0.816
|
NM_024943
|
TMEM156
|
transmembrane protein 156
|
|
chr6_-_33085329
|
0.814
|
NM_002119
|
HLA-DOA
|
major histocompatibility complex, class II, DO alpha
|
|
chr3_+_123256902
|
0.806
|
NM_175862
|
CD86
|
CD86 molecule
|
|
chr5_-_156468571
|
0.802
|
|
HAVCR2
|
hepatitis A virus cellular receptor 2
|
|
chr14_-_105796665
|
0.797
|
|
IGHV4-31
IGHV3-23
IGHA1
IGHG1
IGHM
|
immunoglobulin heavy variable 4-31
immunoglobulin heavy variable 3-23
immunoglobulin heavy constant alpha 1
immunoglobulin heavy constant gamma 1 (G1m marker)
immunoglobulin heavy constant mu
|
|
chr7_-_107930170
|
0.780
|
|
PNPLA8
|
patatin-like phospholipase domain containing 8
|
|
chr3_+_123779122
|
0.772
|
NM_001113523
|
PARP15
|
poly (ADP-ribose) polymerase family, member 15
|
|
chr16_+_28993770
|
0.771
|
|
RRN3P2
|
RNA polymerase I transcription factor homolog (S. cerevisiae) pseudogene 2
|
|
chr11_-_104332631
|
0.769
|
NM_033306
|
CASP4
|
caspase 4, apoptosis-related cysteine peptidase
|
|
chr8_+_24297742
|
0.769
|
NM_001145271
NM_001145272
NM_014479
|
ADAMDEC1
|
ADAM-like, decysin 1
|
|
chr12_-_48902692
|
0.768
|
NM_001113547
|
LIMA1
|
LIM domain and actin binding 1
|
|
chr6_+_28032997
|
0.766
|
NM_012367
|
OR2B6
|
olfactory receptor, family 2, subfamily B, member 6
|
|
chr10_+_89257569
|
0.765
|
NM_001178118
|
MINPP1
|
multiple inositol-polyphosphate phosphatase 1
|
|
chr2_+_201833467
|
0.762
|
NM_033356
|
CASP8
|
caspase 8, apoptosis-related cysteine peptidase
|
|
chr7_-_143213296
|
0.749
|
|
FAM115A
|
family with sequence similarity 115, member A
|
|
chr5_+_61910318
|
0.742
|
NM_181506
|
LRRC70
|
leucine rich repeat containing 70
|
|
chr2_-_136590210
|
0.741
|
NM_001008540
|
CXCR4
|
chemokine (C-X-C motif) receptor 4
|
|
chr5_-_75044001
|
0.741
|
NM_152408
|
POC5
|
POC5 centriolar protein homolog (Chlamydomonas)
|
|
chr10_+_70517863
|
0.741
|
|
SRGN
|
serglycin
|
|
chr6_-_159386007
|
0.740
|
NM_054114
NM_138810
NM_152133
|
TAGAP
|
T-cell activation RhoGTPase activating protein
|
|
chr19_-_53450953
|
0.737
|
NM_001184902
NM_001184904
|
CARD8
|
caspase recruitment domain family, member 8
|
|
chr2_+_201755865
|
0.729
|
NM_001230
NM_032974
NM_032977
|
CASP10
|
caspase 10, apoptosis-related cysteine peptidase
|
|
chr17_+_63367052
|
0.726
|
|
BPTF
|
bromodomain PHD finger transcription factor
|
|
chr8_-_102005697
|
0.724
|
|
YWHAZ
|
tyrosine 3-monooxygenase/tryptophan 5-monooxygenase activation protein, zeta polypeptide
|
|
chr14_-_19867372
|
0.716
|
NM_182852
NM_182851
|
CCNB1IP1
|
cyclin B1 interacting protein 1, E3 ubiquitin protein ligase
|
|
chr13_-_48873735
|
0.715
|
NM_030925
|
CAB39L
|
calcium binding protein 39-like
|
|
chr4_-_47709717
|
0.714
|
NM_000087
|
CNGA1
|
cyclic nucleotide gated channel alpha 1
|
|
chr5_-_78318112
|
0.713
|
NM_000046
|
ARSB
|
arylsulfatase B
|
|
chr1_-_89437098
|
0.712
|
NM_052941
|
GBP4
|
guanylate binding protein 4
|
|
chr1_+_178232379
|
0.706
|
|
CEP350
|
centrosomal protein 350kDa
|
|
chr22_-_34886727
|
0.703
|
NM_145640
|
APOL3
|
apolipoprotein L, 3
|
|
chr2_+_228064531
|
0.701
|
|
AGFG1
|
ArfGAP with FG repeats 1
|
|
chr7_+_106292932
|
0.697
|
|
PIK3CG
|
phosphoinositide-3-kinase, catalytic, gamma polypeptide
|
|
chr13_-_32678142
|
0.691
|
NM_052851
|
STARD13
|
StAR-related lipid transfer (START) domain containing 13
|
|
chr1_-_204455269
|
0.688
|
NM_001007544
|
C1orf186
|
chromosome 1 open reading frame 186
|
|
chr1_+_157246305
|
0.687
|
NM_005531
|
IFI16
|
interferon, gamma-inducible protein 16
|
|
chr19_-_1601246
|
0.674
|
NM_001136139
|
TCF3
|
transcription factor 3 (E2A immunoglobulin enhancer binding factors E12/E47)
|
|
chr11_-_104411065
|
0.671
|
NM_001223
NM_033292
NM_033293
NM_033294
NM_033295
|
CASP1
|
caspase 1, apoptosis-related cysteine peptidase (interleukin 1, beta, convertase)
|
|
chr4_-_110942783
|
0.668
|
NM_000204
|
CFI
|
complement factor I
|
|
chr9_+_21430439
|
0.662
|
NM_024013
|
IFNA1
|
interferon, alpha 1
|
|
chr17_-_26672757
|
0.661
|
NM_001003927
NM_014210
|
EVI2A
|
ecotropic viral integration site 2A
|
|
chr12_+_15590811
|
0.658
|
|
PTPRO
|
protein tyrosine phosphatase, receptor type, O
|
|
chr2_-_44440358
|
0.658
|
NM_001042385
NM_001042386
|
PREPL
|
prolyl endopeptidase-like
|
|
chr5_-_150501393
|
0.657
|
NM_001193544
|
ANXA6
|
annexin A6
|
|
chr5_+_179065282
|
0.653
|
|
CANX
|
calnexin
|
|
chr8_-_110411283
|
0.650
|
NM_001128211
|
NUDCD1
|
NudC domain containing 1
|
|
chr10_+_70517823
|
0.649
|
NM_002727
|
SRGN
|
serglycin
|
|
chr3_+_39346200
|
0.642
|
NM_005201
|
CCR8
|
chemokine (C-C motif) receptor 8
|
|
chr2_-_97775300
|
0.641
|
|
TMEM131
|
transmembrane protein 131
|
|
chr12_+_9713472
|
0.638
|
NM_001197318
NM_013269
NM_001004419
NM_001197317
NM_001197319
|
CLEC2D
|
C-type lectin domain family 2, member D
|
|
chr20_-_51633042
|
0.638
|
NM_006526
|
ZNF217
|
zinc finger protein 217
|
|
chr22_+_20880112
|
0.637
|
|
|
|
|
chr16_-_45562840
|
0.626
|
|
DNAJA2
|
DnaJ (Hsp40) homolog, subfamily A, member 2
|
|
chr22_+_20880159
|
0.624
|
|
IGLV6-57
|
immunoglobulin lambda variable 6-57
|
|
chr1_+_185064654
|
0.624
|
NM_024420
|
PLA2G4A
|
phospholipase A2, group IVA (cytosolic, calcium-dependent)
|
|
chr3_+_32968069
|
0.621
|
NM_005508
|
CCR4
|
chemokine (C-C motif) receptor 4
|
|
chr14_+_64947592
|
0.618
|
NM_178155
NM_178156
|
FUT8
|
fucosyltransferase 8 (alpha (1,6) fucosyltransferase)
|
|
chr4_+_74825138
|
0.613
|
NM_000584
|
IL8
|
interleukin 8
|
|
chr10_+_127419021
|
0.612
|
|
C10orf137
|
chromosome 10 open reading frame 137
|
|
chr14_-_105549177
|
0.612
|
|
IGHV4-31
IGHD
IGHG3
IGHM
|
immunoglobulin heavy variable 4-31
immunoglobulin heavy constant delta
immunoglobulin heavy constant gamma 3 (G3m marker)
immunoglobulin heavy constant mu
|
|
chr18_-_62422195
|
0.608
|
NM_021153
|
CDH19
|
cadherin 19, type 2
|
|
chr14_-_105804211
|
0.608
|
|
IGHG1
|
immunoglobulin heavy constant gamma 1 (G1m marker)
|
|
chr6_+_134800533
|
0.602
|
|
LOC154092
|
hypothetical LOC154092
|
|
chr11_+_59027624
|
0.600
|
NM_001004706
|
OR4D11
|
olfactory receptor, family 4, subfamily D, member 11
|
|
chr9_+_104797413
|
0.599
|
NM_001340
|
CYLC2
|
cylicin, basic protein of sperm head cytoskeleton 2
|
|
chr12_+_10351683
|
0.599
|
NM_002262
NM_007334
|
KLRD1
|
killer cell lectin-like receptor subfamily D, member 1
|
|
chr2_-_45648608
|
0.593
|
|
SRBD1
|
S1 RNA binding domain 1
|
|
chr2_-_232034638
|
0.581
|
|
NCL
|
nucleolin
|
|
chr8_+_42671718
|
0.578
|
NM_000749
|
CHRNB3
|
cholinergic receptor, nicotinic, beta 3
|
|
chr8_+_104900591
|
0.576
|
NM_014677
|
RIMS2
|
regulating synaptic membrane exocytosis 2
|
|
chr4_+_86918870
|
0.573
|
NM_001042669
|
ARHGAP24
|
Rho GTPase activating protein 24
|
|
chr3_-_52688768
|
0.571
|
NM_018165
NM_181042
|
PBRM1
|
polybromo 1
|
|
chr12_+_25096755
|
0.571
|
|
LRMP
|
lymphoid-restricted membrane protein
|
|
chr11_-_104421240
|
0.569
|
NM_001017534
NM_052889
|
CARD16
CASP1
|
caspase recruitment domain family, member 16
caspase 1, apoptosis-related cysteine peptidase (interleukin 1, beta, convertase)
|
|
chr19_-_44518516
|
0.567
|
NM_004877
|
GMFG
|
glia maturation factor, gamma
|
|
chr2_+_183290912
|
0.566
|
|
DNAJC10
|
DnaJ (Hsp40) homolog, subfamily C, member 10
|
|
chr4_-_17415918
|
0.564
|
|
DCAF16
|
DDB1 and CUL4 associated factor 16
|
|
chr9_-_21229975
|
0.560
|
NM_002172
|
IFNA14
|
interferon, alpha 14
|
|
chr13_-_45654292
|
0.560
|
|
LCP1
|
lymphocyte cytosolic protein 1 (L-plastin)
|
|
chr12_+_14429499
|
0.554
|
|
ATF7IP
|
activating transcription factor 7 interacting protein
|
|
chr12_-_10953427
|
0.553
|
NM_023920
|
TAS2R13
|
taste receptor, type 2, member 13
|
|
chr12_+_15590552
|
0.553
|
NM_030668
NM_030669
NM_030670
NM_030671
|
PTPRO
|
protein tyrosine phosphatase, receptor type, O
|
|
chr19_-_14750347
|
0.553
|
NM_013447
NM_152916
NM_152917
NM_152918
NM_152919
NM_152920
NM_152921
|
EMR2
|
egf-like module containing, mucin-like, hormone receptor-like 2
|
|
chr1_+_67404756
|
0.549
|
NM_144701
|
IL23R
|
interleukin 23 receptor
|
|
chr8_+_11388918
|
0.543
|
NM_001715
|
BLK
|
B lymphoid tyrosine kinase
|
|
chr11_-_58032153
|
0.542
|
NM_001005218
|
OR5B21
|
olfactory receptor, family 5, subfamily B, member 21
|
|
chr1_+_51340487
|
0.535
|
NM_001136508
|
C1orf185
|
chromosome 1 open reading frame 185
|
|
chr19_-_16111695
|
0.533
|
|
|
|
|
chr17_+_65009986
|
0.532
|
|
MAP2K6
|
mitogen-activated protein kinase kinase 6
|
|
chr11_-_117600918
|
0.532
|
NM_001098526
|
AMICA1
|
adhesion molecule, interacts with CXADR antigen 1
|
|
chr1_-_111544820
|
0.526
|
|
DENND2D
|
DENN/MADD domain containing 2D
|
|
chr12_+_48443588
|
0.525
|
|
TMBIM6
|
transmembrane BAX inhibitor motif containing 6
|
|
chr1_+_84382529
|
0.524
|
NM_182948
|
PRKACB
|
protein kinase, cAMP-dependent, catalytic, beta
|
|
chr7_-_131983986
|
0.521
|
NM_181775
|
PLXNA4
|
plexin A4
|
|
chr1_-_157551072
|
0.521
|
NM_001004467
|
OR10J3
|
olfactory receptor, family 10, subfamily J, member 3
|
|
chr7_-_33047042
|
0.517
|
NM_001166118
NM_016489
|
NT5C3
|
5'-nucleotidase, cytosolic III
|
|
chr4_-_83566045
|
0.510
|
|
|
|
|
chr12_-_16652202
|
0.509
|
NM_001001395
|
LMO3
|
LIM domain only 3 (rhombotin-like 2)
|
|
chrX_+_48427137
|
0.508
|
|
WAS
|
Wiskott-Aldrich syndrome (eczema-thrombocytopenia)
|
|
chrX_-_138742050
|
0.507
|
NM_001010986
NM_173694
|
ATP11C
|
ATPase, class VI, type 11C
|
|
chr7_+_90177082
|
0.507
|
|
CDK14
|
cyclin-dependent kinase 14
|
|
chr8_+_24207524
|
0.507
|
NM_014265
NM_021777
|
ADAM28
|
ADAM metallopeptidase domain 28
|
|
chr10_+_35524060
|
0.501
|
NM_182717
NM_182718
NM_182719
NM_182720
|
CREM
|
cAMP responsive element modulator
|
|
chr8_-_120720200
|
0.499
|
NM_001040092
NM_001130863
NM_006209
|
ENPP2
|
ectonucleotide pyrophosphatase/phosphodiesterase 2
|
|
chr22_-_16080265
|
0.498
|
|
CECR1
|
cat eye syndrome chromosome region, candidate 1
|
|
chr8_-_17599438
|
0.497
|
NM_020749
|
MTUS1
|
microtubule associated tumor suppressor 1
|
|
chr3_+_142587973
|
0.496
|
|
|
|
|
chr11_+_67107852
|
0.493
|
|
GSTP1
|
glutathione S-transferase pi 1
|
|
chr12_+_9871343
|
0.489
|
NM_016523
|
KLRF1
|
killer cell lectin-like receptor subfamily F, member 1
|
|
chr17_-_39094787
|
0.486
|
NM_001040002
|
MEOX1
|
mesenchyme homeobox 1
|
|
chr6_+_26476384
|
0.485
|
NM_001197246
|
BTN3A2
|
butyrophilin, subfamily 3, member A2
|
|
chr2_+_134728299
|
0.483
|
NM_002410
|
MGAT5
|
mannosyl (alpha-1,6-)-glycoprotein beta-1,6-N-acetyl-glucosaminyltransferase
|
|
chr16_-_29797172
|
0.478
|
|
SEZ6L2
|
seizure related 6 homolog (mouse)-like 2
|
|
chr18_+_70471906
|
0.476
|
NM_001146189
NM_001146190
NM_017757
|
ZNF407
|
zinc finger protein 407
|
|
chr19_+_42409205
|
0.476
|
NM_152604
|
ZNF383
|
zinc finger protein 383
|