Project
GNF SymAtlas + NCI-60 cancer cell lines, comparison of cancers vs non-cancers, human (Su, 2004; Ross, 2000)
Navigation
Downloads
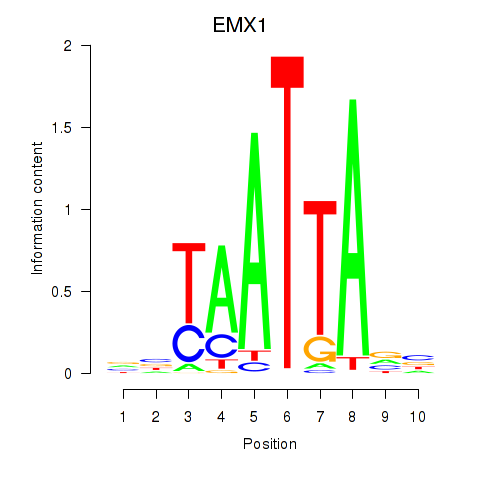
Results for EMX1
Z-value: 0.53
Transcription factors associated with EMX1
| Gene Symbol | Gene ID | Gene Info |
|---|---|---|
|
EMX1
|
ENSG00000135638.9 | empty spiracles homeobox 1 |
Activity-expression correlation:
| Gene | Promoter | Pearson corr. coef. | P-value | Plot |
|---|---|---|---|---|
| EMX1 | hg19_v2_chr2_+_73144604_73144654 | 0.49 | 2.9e-14 | Click! |
Activity profile of EMX1 motif
Sorted Z-values of EMX1 motif
| Promoter | Log-likelihood | Transcript | Gene | Gene Info |
|---|---|---|---|---|
| chr11_-_111794446 | 53.74 |
ENST00000527950.1
|
CRYAB
|
crystallin, alpha B |
| chr11_-_117748138 | 41.30 |
ENST00000527717.1
|
FXYD6
|
FXYD domain containing ion transport regulator 6 |
| chr12_-_91546926 | 41.13 |
ENST00000550758.1
|
DCN
|
decorin |
| chr11_-_117747607 | 34.22 |
ENST00000540359.1
ENST00000539526.1 |
FXYD6
|
FXYD domain containing ion transport regulator 6 |
| chr11_-_117747434 | 33.93 |
ENST00000529335.2
ENST00000530956.1 ENST00000260282.4 |
FXYD6
|
FXYD domain containing ion transport regulator 6 |
| chrX_+_13671225 | 33.37 |
ENST00000545566.1
ENST00000544987.1 ENST00000314720.4 |
TCEANC
|
transcription elongation factor A (SII) N-terminal and central domain containing |
| chr12_-_6233828 | 25.41 |
ENST00000572068.1
ENST00000261405.5 |
VWF
|
von Willebrand factor |
| chrX_-_13835147 | 24.44 |
ENST00000493677.1
ENST00000355135.2 |
GPM6B
|
glycoprotein M6B |
| chr6_+_153552455 | 19.69 |
ENST00000392385.2
|
AL590867.1
|
Uncharacterized protein; cDNA FLJ59044, highly similar to LINE-1 reverse transcriptase homolog |
| chr4_+_175204818 | 16.51 |
ENST00000503780.1
|
CEP44
|
centrosomal protein 44kDa |
| chr4_+_106631966 | 16.27 |
ENST00000360505.5
ENST00000510865.1 ENST00000509336.1 |
GSTCD
|
glutathione S-transferase, C-terminal domain containing |
| chr4_-_89205879 | 15.55 |
ENST00000608933.1
ENST00000315194.4 ENST00000514204.1 |
PPM1K
|
protein phosphatase, Mg2+/Mn2+ dependent, 1K |
| chr10_-_28571015 | 15.39 |
ENST00000375719.3
ENST00000375732.1 |
MPP7
|
membrane protein, palmitoylated 7 (MAGUK p55 subfamily member 7) |
| chr6_+_111580508 | 15.00 |
ENST00000368847.4
|
KIAA1919
|
KIAA1919 |
| chr12_+_53773944 | 14.50 |
ENST00000551969.1
ENST00000327443.4 |
SP1
|
Sp1 transcription factor |
| chr16_+_58283814 | 14.22 |
ENST00000443128.2
ENST00000219299.4 |
CCDC113
|
coiled-coil domain containing 113 |
| chr11_+_92085262 | 13.65 |
ENST00000298047.6
ENST00000409404.2 ENST00000541502.1 |
FAT3
|
FAT atypical cadherin 3 |
| chr19_-_7968427 | 13.54 |
ENST00000539278.1
|
AC010336.1
|
Uncharacterized protein |
| chr11_-_129062093 | 13.24 |
ENST00000310343.9
|
ARHGAP32
|
Rho GTPase activating protein 32 |
| chr3_+_111718036 | 12.67 |
ENST00000455401.2
|
TAGLN3
|
transgelin 3 |
| chr6_+_151042224 | 12.61 |
ENST00000358517.2
|
PLEKHG1
|
pleckstrin homology domain containing, family G (with RhoGef domain) member 1 |
| chr3_+_111717600 | 12.33 |
ENST00000273368.4
|
TAGLN3
|
transgelin 3 |
| chr6_+_151646800 | 11.68 |
ENST00000354675.6
|
AKAP12
|
A kinase (PRKA) anchor protein 12 |
| chr3_+_111717511 | 11.52 |
ENST00000478951.1
ENST00000393917.2 |
TAGLN3
|
transgelin 3 |
| chr1_+_202317815 | 11.28 |
ENST00000608999.1
ENST00000336894.4 ENST00000480184.1 |
PPP1R12B
|
protein phosphatase 1, regulatory subunit 12B |
| chr4_+_169418195 | 10.91 |
ENST00000261509.6
ENST00000335742.7 |
PALLD
|
palladin, cytoskeletal associated protein |
| chr10_-_71169031 | 10.47 |
ENST00000373307.1
|
TACR2
|
tachykinin receptor 2 |
| chr4_-_10686475 | 10.32 |
ENST00000226951.6
|
CLNK
|
cytokine-dependent hematopoietic cell linker |
| chrX_-_73512411 | 10.23 |
ENST00000602576.1
ENST00000429124.1 |
FTX
|
FTX transcript, XIST regulator (non-protein coding) |
| chr5_+_36608422 | 10.16 |
ENST00000381918.3
|
SLC1A3
|
solute carrier family 1 (glial high affinity glutamate transporter), member 3 |
| chr4_+_88896819 | 10.14 |
ENST00000237623.7
ENST00000395080.3 ENST00000508233.1 ENST00000360804.4 |
SPP1
|
secreted phosphoprotein 1 |
| chr10_-_93392811 | 10.02 |
ENST00000238994.5
|
PPP1R3C
|
protein phosphatase 1, regulatory subunit 3C |
| chr3_+_111718173 | 9.79 |
ENST00000494932.1
|
TAGLN3
|
transgelin 3 |
| chr2_+_196313239 | 9.73 |
ENST00000413290.1
|
AC064834.1
|
AC064834.1 |
| chr3_-_123339343 | 9.17 |
ENST00000578202.1
|
MYLK
|
myosin light chain kinase |
| chr14_+_93389425 | 9.12 |
ENST00000216492.5
ENST00000334654.4 |
CHGA
|
chromogranin A (parathyroid secretory protein 1) |
| chr12_-_48164812 | 8.70 |
ENST00000549151.1
ENST00000548919.1 |
RAPGEF3
|
Rap guanine nucleotide exchange factor (GEF) 3 |
| chr5_+_66300446 | 8.56 |
ENST00000261569.7
|
MAST4
|
microtubule associated serine/threonine kinase family member 4 |
| chr1_+_160160283 | 8.52 |
ENST00000368079.3
|
CASQ1
|
calsequestrin 1 (fast-twitch, skeletal muscle) |
| chr6_-_46293378 | 8.52 |
ENST00000330430.6
|
RCAN2
|
regulator of calcineurin 2 |
| chr3_+_130745688 | 8.45 |
ENST00000510769.1
ENST00000429253.2 ENST00000356918.4 ENST00000510688.1 ENST00000511262.1 ENST00000383366.4 |
NEK11
|
NIMA-related kinase 11 |
| chr2_-_217560248 | 7.94 |
ENST00000233813.4
|
IGFBP5
|
insulin-like growth factor binding protein 5 |
| chr5_-_95297534 | 7.88 |
ENST00000513343.1
ENST00000431061.2 |
ELL2
|
elongation factor, RNA polymerase II, 2 |
| chr1_+_160160346 | 7.83 |
ENST00000368078.3
|
CASQ1
|
calsequestrin 1 (fast-twitch, skeletal muscle) |
| chr7_+_130126012 | 7.79 |
ENST00000341441.5
|
MEST
|
mesoderm specific transcript |
| chr11_-_13517565 | 7.72 |
ENST00000282091.1
ENST00000529816.1 |
PTH
|
parathyroid hormone |
| chr3_+_115342349 | 7.71 |
ENST00000393780.3
|
GAP43
|
growth associated protein 43 |
| chr12_-_118796910 | 7.71 |
ENST00000541186.1
ENST00000539872.1 |
TAOK3
|
TAO kinase 3 |
| chr7_+_130126165 | 7.64 |
ENST00000427521.1
ENST00000416162.2 ENST00000378576.4 |
MEST
|
mesoderm specific transcript |
| chr10_-_5046042 | 7.56 |
ENST00000421196.3
ENST00000455190.1 |
AKR1C2
|
aldo-keto reductase family 1, member C2 |
| chr5_-_24645078 | 7.41 |
ENST00000264463.4
|
CDH10
|
cadherin 10, type 2 (T2-cadherin) |
| chr15_+_62853562 | 7.38 |
ENST00000561311.1
|
TLN2
|
talin 2 |
| chr3_-_38071122 | 7.31 |
ENST00000334661.4
|
PLCD1
|
phospholipase C, delta 1 |
| chr4_-_8873531 | 7.26 |
ENST00000400677.3
|
HMX1
|
H6 family homeobox 1 |
| chr12_-_114841703 | 7.26 |
ENST00000526441.1
|
TBX5
|
T-box 5 |
| chr12_+_81110684 | 7.23 |
ENST00000228644.3
|
MYF5
|
myogenic factor 5 |
| chr2_-_152830441 | 7.22 |
ENST00000534999.1
ENST00000397327.2 |
CACNB4
|
calcium channel, voltage-dependent, beta 4 subunit |
| chr4_-_153601136 | 7.12 |
ENST00000504064.1
ENST00000304385.3 |
TMEM154
|
transmembrane protein 154 |
| chrX_+_54947229 | 7.10 |
ENST00000442098.1
ENST00000430420.1 ENST00000453081.1 ENST00000173898.7 ENST00000319167.8 ENST00000375022.4 ENST00000399736.1 ENST00000440072.1 ENST00000420798.2 ENST00000431115.1 ENST00000440759.1 ENST00000375041.2 |
TRO
|
trophinin |
| chr11_-_128894053 | 7.09 |
ENST00000392657.3
|
ARHGAP32
|
Rho GTPase activating protein 32 |
| chr8_+_50824233 | 7.09 |
ENST00000522124.1
|
SNTG1
|
syntrophin, gamma 1 |
| chr1_+_62439037 | 7.00 |
ENST00000545929.1
|
INADL
|
InaD-like (Drosophila) |
| chr6_-_49681235 | 6.98 |
ENST00000339139.4
|
CRISP2
|
cysteine-rich secretory protein 2 |
| chr3_+_35722487 | 6.90 |
ENST00000441454.1
|
ARPP21
|
cAMP-regulated phosphoprotein, 21kDa |
| chr1_+_226013047 | 6.89 |
ENST00000366837.4
|
EPHX1
|
epoxide hydrolase 1, microsomal (xenobiotic) |
| chr4_-_76944621 | 6.88 |
ENST00000306602.1
|
CXCL10
|
chemokine (C-X-C motif) ligand 10 |
| chr12_+_26348246 | 6.82 |
ENST00000422622.2
|
SSPN
|
sarcospan |
| chr3_+_158991025 | 6.72 |
ENST00000337808.6
|
IQCJ-SCHIP1
|
IQCJ-SCHIP1 readthrough |
| chr8_-_17533838 | 6.57 |
ENST00000400046.1
|
MTUS1
|
microtubule associated tumor suppressor 1 |
| chr10_+_7745303 | 6.55 |
ENST00000429820.1
ENST00000379587.4 |
ITIH2
|
inter-alpha-trypsin inhibitor heavy chain 2 |
| chr2_+_171034646 | 6.52 |
ENST00000409044.3
ENST00000408978.4 |
MYO3B
|
myosin IIIB |
| chr4_-_186696425 | 6.45 |
ENST00000430503.1
ENST00000319454.6 ENST00000450341.1 |
SORBS2
|
sorbin and SH3 domain containing 2 |
| chr8_+_98900132 | 6.43 |
ENST00000520016.1
|
MATN2
|
matrilin 2 |
| chr9_+_131062367 | 6.40 |
ENST00000601297.1
|
AL359091.2
|
CDNA: FLJ21673 fis, clone COL09042; HCG2036511; Uncharacterized protein |
| chr8_+_105235572 | 6.38 |
ENST00000523362.1
|
RIMS2
|
regulating synaptic membrane exocytosis 2 |
| chr17_-_74163159 | 6.35 |
ENST00000591615.1
|
RNF157
|
ring finger protein 157 |
| chrX_-_73512177 | 6.33 |
ENST00000603672.1
ENST00000418855.1 |
FTX
|
FTX transcript, XIST regulator (non-protein coding) |
| chr17_-_1532106 | 6.22 |
ENST00000301335.5
ENST00000382147.4 |
SLC43A2
|
solute carrier family 43 (amino acid system L transporter), member 2 |
| chr14_-_70883708 | 6.21 |
ENST00000256366.4
|
SYNJ2BP
|
synaptojanin 2 binding protein |
| chr14_-_23652849 | 6.20 |
ENST00000316902.7
ENST00000469263.1 ENST00000525062.1 ENST00000524758.1 |
SLC7A8
|
solute carrier family 7 (amino acid transporter light chain, L system), member 8 |
| chr3_-_44519131 | 6.16 |
ENST00000425708.2
ENST00000396077.2 |
ZNF445
|
zinc finger protein 445 |
| chr7_-_73038822 | 6.14 |
ENST00000414749.2
ENST00000429400.2 ENST00000434326.1 |
MLXIPL
|
MLX interacting protein-like |
| chr12_+_7013897 | 6.06 |
ENST00000007969.8
ENST00000323702.5 |
LRRC23
|
leucine rich repeat containing 23 |
| chr17_-_18430160 | 6.03 |
ENST00000392176.3
|
FAM106A
|
family with sequence similarity 106, member A |
| chr10_+_5005598 | 5.98 |
ENST00000442997.1
|
AKR1C1
|
aldo-keto reductase family 1, member C1 |
| chr16_-_29910853 | 5.96 |
ENST00000308713.5
|
SEZ6L2
|
seizure related 6 homolog (mouse)-like 2 |
| chr12_+_7014064 | 5.89 |
ENST00000443597.2
|
LRRC23
|
leucine rich repeat containing 23 |
| chr2_-_214016314 | 5.88 |
ENST00000434687.1
ENST00000374319.4 |
IKZF2
|
IKAROS family zinc finger 2 (Helios) |
| chr4_-_74486347 | 5.77 |
ENST00000342081.3
|
RASSF6
|
Ras association (RalGDS/AF-6) domain family member 6 |
| chr1_-_48937821 | 5.76 |
ENST00000396199.3
|
SPATA6
|
spermatogenesis associated 6 |
| chr5_+_125758813 | 5.76 |
ENST00000285689.3
ENST00000515200.1 |
GRAMD3
|
GRAM domain containing 3 |
| chr19_+_15852203 | 5.76 |
ENST00000305892.1
|
OR10H3
|
olfactory receptor, family 10, subfamily H, member 3 |
| chr9_-_95166884 | 5.67 |
ENST00000375561.5
|
OGN
|
osteoglycin |
| chr1_-_48937838 | 5.66 |
ENST00000371847.3
|
SPATA6
|
spermatogenesis associated 6 |
| chr5_+_174151536 | 5.54 |
ENST00000239243.6
ENST00000507785.1 |
MSX2
|
msh homeobox 2 |
| chr9_-_5304432 | 5.54 |
ENST00000416837.1
ENST00000308420.3 |
RLN2
|
relaxin 2 |
| chr18_+_56530136 | 5.54 |
ENST00000591083.1
|
ZNF532
|
zinc finger protein 532 |
| chr10_-_50970322 | 5.49 |
ENST00000374103.4
|
OGDHL
|
oxoglutarate dehydrogenase-like |
| chr14_+_100485712 | 5.48 |
ENST00000544450.2
|
EVL
|
Enah/Vasp-like |
| chr2_+_113763031 | 5.48 |
ENST00000259211.6
|
IL36A
|
interleukin 36, alpha |
| chr19_-_39368887 | 5.36 |
ENST00000340740.3
ENST00000591812.1 |
RINL
|
Ras and Rab interactor-like |
| chr4_-_74486217 | 5.32 |
ENST00000335049.5
ENST00000307439.5 |
RASSF6
|
Ras association (RalGDS/AF-6) domain family member 6 |
| chr2_-_228582709 | 5.26 |
ENST00000541617.1
ENST00000409456.2 ENST00000409287.1 ENST00000258403.3 |
SLC19A3
|
solute carrier family 19 (thiamine transporter), member 3 |
| chr2_+_113816685 | 5.20 |
ENST00000393200.2
|
IL36RN
|
interleukin 36 receptor antagonist |
| chr8_+_105352050 | 5.19 |
ENST00000297581.2
|
DCSTAMP
|
dendrocyte expressed seven transmembrane protein |
| chr5_+_125758865 | 5.19 |
ENST00000542322.1
ENST00000544396.1 |
GRAMD3
|
GRAM domain containing 3 |
| chrX_-_14047996 | 5.18 |
ENST00000380523.4
ENST00000398355.3 |
GEMIN8
|
gem (nuclear organelle) associated protein 8 |
| chr5_-_95297678 | 5.18 |
ENST00000237853.4
|
ELL2
|
elongation factor, RNA polymerase II, 2 |
| chr9_-_95166841 | 5.16 |
ENST00000262551.4
|
OGN
|
osteoglycin |
| chr4_+_41614909 | 5.06 |
ENST00000509454.1
ENST00000396595.3 ENST00000381753.4 |
LIMCH1
|
LIM and calponin homology domains 1 |
| chr1_-_242162375 | 5.02 |
ENST00000357246.3
|
MAP1LC3C
|
microtubule-associated protein 1 light chain 3 gamma |
| chr18_-_61329118 | 5.02 |
ENST00000332821.8
ENST00000283752.5 |
SERPINB3
|
serpin peptidase inhibitor, clade B (ovalbumin), member 3 |
| chr12_-_112123524 | 4.99 |
ENST00000327551.6
|
BRAP
|
BRCA1 associated protein |
| chr12_+_7014126 | 4.99 |
ENST00000415834.1
ENST00000436789.1 |
LRRC23
|
leucine rich repeat containing 23 |
| chr7_-_73038867 | 4.97 |
ENST00000313375.3
ENST00000354613.1 ENST00000395189.1 ENST00000453275.1 |
MLXIPL
|
MLX interacting protein-like |
| chr12_+_18414446 | 4.95 |
ENST00000433979.1
|
PIK3C2G
|
phosphatidylinositol-4-phosphate 3-kinase, catalytic subunit type 2 gamma |
| chr3_+_158787041 | 4.91 |
ENST00000471575.1
ENST00000476809.1 ENST00000485419.1 |
IQCJ-SCHIP1
|
IQCJ-SCHIP1 readthrough |
| chr4_+_86396265 | 4.90 |
ENST00000395184.1
|
ARHGAP24
|
Rho GTPase activating protein 24 |
| chr1_+_151735431 | 4.88 |
ENST00000321531.5
ENST00000315067.8 |
OAZ3
|
ornithine decarboxylase antizyme 3 |
| chr7_+_142829162 | 4.87 |
ENST00000291009.3
|
PIP
|
prolactin-induced protein |
| chr7_-_83278322 | 4.72 |
ENST00000307792.3
|
SEMA3E
|
sema domain, immunoglobulin domain (Ig), short basic domain, secreted, (semaphorin) 3E |
| chr19_-_58951496 | 4.65 |
ENST00000254166.3
|
ZNF132
|
zinc finger protein 132 |
| chr7_+_43152191 | 4.57 |
ENST00000395891.2
|
HECW1
|
HECT, C2 and WW domain containing E3 ubiquitin protein ligase 1 |
| chr20_+_3052264 | 4.55 |
ENST00000217386.2
|
OXT
|
oxytocin/neurophysin I prepropeptide |
| chr6_+_72926145 | 4.50 |
ENST00000425662.2
ENST00000453976.2 |
RIMS1
|
regulating synaptic membrane exocytosis 1 |
| chr7_-_107642348 | 4.49 |
ENST00000393561.1
|
LAMB1
|
laminin, beta 1 |
| chr10_+_7745232 | 4.48 |
ENST00000358415.4
|
ITIH2
|
inter-alpha-trypsin inhibitor heavy chain 2 |
| chr6_+_39760129 | 4.44 |
ENST00000274867.4
|
DAAM2
|
dishevelled associated activator of morphogenesis 2 |
| chr12_-_11548496 | 4.38 |
ENST00000389362.4
ENST00000565533.1 ENST00000546254.1 |
PRB2
PRB1
|
proline-rich protein BstNI subfamily 2 proline-rich protein BstNI subfamily 1 |
| chr3_-_99594948 | 4.34 |
ENST00000471562.1
ENST00000495625.2 |
FILIP1L
|
filamin A interacting protein 1-like |
| chr18_-_21891460 | 4.33 |
ENST00000357041.4
|
OSBPL1A
|
oxysterol binding protein-like 1A |
| chr4_-_87281196 | 4.33 |
ENST00000359221.3
|
MAPK10
|
mitogen-activated protein kinase 10 |
| chr20_-_23860373 | 4.30 |
ENST00000304710.4
|
CST5
|
cystatin D |
| chr1_+_70876926 | 4.26 |
ENST00000370938.3
ENST00000346806.2 |
CTH
|
cystathionase (cystathionine gamma-lyase) |
| chr17_+_72426891 | 4.25 |
ENST00000392627.1
|
GPRC5C
|
G protein-coupled receptor, family C, group 5, member C |
| chr10_+_32856764 | 4.22 |
ENST00000375030.2
ENST00000375028.3 |
C10orf68
|
Homo sapiens coiled-coil domain containing 7 (CCDC7), transcript variant 5, mRNA. |
| chr1_+_17559776 | 4.22 |
ENST00000537499.1
ENST00000413717.2 ENST00000536552.1 |
PADI1
|
peptidyl arginine deiminase, type I |
| chr8_+_99956662 | 4.22 |
ENST00000523368.1
ENST00000297565.4 ENST00000435298.2 |
OSR2
|
odd-skipped related transciption factor 2 |
| chr3_-_130745571 | 4.21 |
ENST00000514044.1
ENST00000264992.3 |
ASTE1
|
asteroid homolog 1 (Drosophila) |
| chr6_+_72922590 | 4.19 |
ENST00000523963.1
|
RIMS1
|
regulating synaptic membrane exocytosis 1 |
| chr13_+_73632897 | 4.18 |
ENST00000377687.4
|
KLF5
|
Kruppel-like factor 5 (intestinal) |
| chr18_-_53177984 | 4.16 |
ENST00000543082.1
|
TCF4
|
transcription factor 4 |
| chr17_-_39743139 | 4.16 |
ENST00000167586.6
|
KRT14
|
keratin 14 |
| chr16_-_53737722 | 4.13 |
ENST00000569716.1
ENST00000562588.1 ENST00000562230.1 ENST00000379925.3 ENST00000563746.1 ENST00000568653.3 |
RPGRIP1L
|
RPGRIP1-like |
| chr2_+_202047843 | 4.09 |
ENST00000272879.5
ENST00000374650.3 ENST00000346817.5 ENST00000313728.7 ENST00000448480.1 |
CASP10
|
caspase 10, apoptosis-related cysteine peptidase |
| chr3_+_152552685 | 4.09 |
ENST00000305097.3
|
P2RY1
|
purinergic receptor P2Y, G-protein coupled, 1 |
| chr12_+_106751436 | 4.08 |
ENST00000228347.4
|
POLR3B
|
polymerase (RNA) III (DNA directed) polypeptide B |
| chr6_+_72922505 | 4.07 |
ENST00000401910.3
|
RIMS1
|
regulating synaptic membrane exocytosis 1 |
| chr11_-_5255861 | 4.06 |
ENST00000380299.3
|
HBD
|
hemoglobin, delta |
| chr3_-_160823158 | 4.05 |
ENST00000392779.2
ENST00000392780.1 ENST00000494173.1 |
B3GALNT1
|
beta-1,3-N-acetylgalactosaminyltransferase 1 (globoside blood group) |
| chr13_-_28545276 | 4.04 |
ENST00000381020.7
|
CDX2
|
caudal type homeobox 2 |
| chr11_+_24518723 | 4.04 |
ENST00000336930.6
ENST00000529015.1 ENST00000533227.1 |
LUZP2
|
leucine zipper protein 2 |
| chr3_-_99595037 | 4.02 |
ENST00000383694.2
|
FILIP1L
|
filamin A interacting protein 1-like |
| chr11_-_108093329 | 4.00 |
ENST00000278612.8
|
NPAT
|
nuclear protein, ataxia-telangiectasia locus |
| chr5_-_176889381 | 3.97 |
ENST00000393563.4
ENST00000512501.1 |
DBN1
|
drebrin 1 |
| chr22_+_30752606 | 3.95 |
ENST00000399824.2
ENST00000405659.1 ENST00000338306.3 |
CCDC157
|
coiled-coil domain containing 157 |
| chr10_-_50970382 | 3.90 |
ENST00000419399.1
ENST00000432695.1 |
OGDHL
|
oxoglutarate dehydrogenase-like |
| chr1_+_225600404 | 3.89 |
ENST00000366845.2
|
AC092811.1
|
AC092811.1 |
| chr15_-_58571445 | 3.89 |
ENST00000558231.1
|
ALDH1A2
|
aldehyde dehydrogenase 1 family, member A2 |
| chr4_-_89205705 | 3.87 |
ENST00000295908.7
ENST00000510548.2 ENST00000508256.1 |
PPM1K
|
protein phosphatase, Mg2+/Mn2+ dependent, 1K |
| chr7_-_100860851 | 3.85 |
ENST00000223127.3
|
PLOD3
|
procollagen-lysine, 2-oxoglutarate 5-dioxygenase 3 |
| chr6_-_91296602 | 3.84 |
ENST00000369325.3
ENST00000369327.3 |
MAP3K7
|
mitogen-activated protein kinase kinase kinase 7 |
| chr7_-_105029812 | 3.84 |
ENST00000482897.1
|
SRPK2
|
SRSF protein kinase 2 |
| chr2_+_166095898 | 3.81 |
ENST00000424833.1
ENST00000375437.2 ENST00000357398.3 |
SCN2A
|
sodium channel, voltage-gated, type II, alpha subunit |
| chr5_+_31193847 | 3.78 |
ENST00000514738.1
ENST00000265071.2 |
CDH6
|
cadherin 6, type 2, K-cadherin (fetal kidney) |
| chr11_+_65554493 | 3.76 |
ENST00000335987.3
|
OVOL1
|
ovo-like zinc finger 1 |
| chr2_-_179672142 | 3.73 |
ENST00000342992.6
ENST00000360870.5 ENST00000460472.2 ENST00000589042.1 ENST00000591111.1 ENST00000342175.6 ENST00000359218.5 |
TTN
|
titin |
| chr17_-_73663245 | 3.73 |
ENST00000584999.1
ENST00000317905.5 ENST00000420326.2 ENST00000340830.5 |
RECQL5
|
RecQ protein-like 5 |
| chr1_-_95391315 | 3.72 |
ENST00000545882.1
ENST00000415017.1 |
CNN3
|
calponin 3, acidic |
| chr4_+_88754113 | 3.70 |
ENST00000560249.1
ENST00000540395.1 ENST00000511670.1 ENST00000361056.3 |
MEPE
|
matrix extracellular phosphoglycoprotein |
| chr20_+_45947246 | 3.68 |
ENST00000599904.1
|
AL031666.2
|
HCG2018772; Uncharacterized protein; cDNA FLJ31609 fis, clone NT2RI2002852 |
| chr10_+_18629628 | 3.63 |
ENST00000377329.4
|
CACNB2
|
calcium channel, voltage-dependent, beta 2 subunit |
| chr9_-_5339873 | 3.63 |
ENST00000223862.1
ENST00000223858.4 |
RLN1
|
relaxin 1 |
| chr6_+_29429217 | 3.62 |
ENST00000396792.2
|
OR2H1
|
olfactory receptor, family 2, subfamily H, member 1 |
| chr6_+_28317685 | 3.62 |
ENST00000252211.2
ENST00000341464.5 ENST00000377255.3 |
ZKSCAN3
|
zinc finger with KRAB and SCAN domains 3 |
| chr13_-_38172863 | 3.60 |
ENST00000541481.1
ENST00000379743.4 ENST00000379742.4 ENST00000379749.4 ENST00000541179.1 ENST00000379747.4 |
POSTN
|
periostin, osteoblast specific factor |
| chr2_-_166930131 | 3.59 |
ENST00000303395.4
ENST00000409050.1 ENST00000423058.2 ENST00000375405.3 |
SCN1A
|
sodium channel, voltage-gated, type I, alpha subunit |
| chr16_-_66764119 | 3.59 |
ENST00000569320.1
|
DYNC1LI2
|
dynein, cytoplasmic 1, light intermediate chain 2 |
| chr14_+_32798547 | 3.58 |
ENST00000557354.1
ENST00000557102.1 ENST00000557272.1 |
AKAP6
|
A kinase (PRKA) anchor protein 6 |
| chr11_+_119076745 | 3.55 |
ENST00000264033.4
|
CBL
|
Cbl proto-oncogene, E3 ubiquitin protein ligase |
| chr3_-_160823040 | 3.53 |
ENST00000484127.1
ENST00000492353.1 ENST00000473142.1 ENST00000468268.1 ENST00000460353.1 ENST00000320474.4 ENST00000392781.2 |
B3GALNT1
|
beta-1,3-N-acetylgalactosaminyltransferase 1 (globoside blood group) |
| chr19_-_3600549 | 3.50 |
ENST00000589966.1
|
TBXA2R
|
thromboxane A2 receptor |
| chr6_-_22297730 | 3.48 |
ENST00000306482.1
|
PRL
|
prolactin |
| chr1_-_233431458 | 3.48 |
ENST00000258229.9
ENST00000430153.1 |
PCNXL2
|
pecanex-like 2 (Drosophila) |
| chr4_-_87281224 | 3.47 |
ENST00000395169.3
ENST00000395161.2 |
MAPK10
|
mitogen-activated protein kinase 10 |
| chr7_-_143599207 | 3.47 |
ENST00000355951.2
ENST00000479870.1 ENST00000478172.1 |
FAM115A
|
family with sequence similarity 115, member A |
| chr14_+_68286516 | 3.45 |
ENST00000471583.1
ENST00000487270.1 ENST00000488612.1 |
RAD51B
|
RAD51 paralog B |
| chr5_-_160279207 | 3.42 |
ENST00000327245.5
|
ATP10B
|
ATPase, class V, type 10B |
| chr11_-_102709441 | 3.39 |
ENST00000434103.1
|
MMP3
|
matrix metallopeptidase 3 (stromelysin 1, progelatinase) |
| chr9_-_21202204 | 3.37 |
ENST00000239347.3
|
IFNA7
|
interferon, alpha 7 |
| chr7_-_27169801 | 3.35 |
ENST00000511914.1
|
HOXA4
|
homeobox A4 |
| chr8_+_121137333 | 3.33 |
ENST00000309791.4
ENST00000297848.3 ENST00000247781.3 |
COL14A1
|
collagen, type XIV, alpha 1 |
| chr16_-_53737795 | 3.30 |
ENST00000262135.4
ENST00000564374.1 ENST00000566096.1 |
RPGRIP1L
|
RPGRIP1-like |
| chr21_-_42219065 | 3.29 |
ENST00000400454.1
|
DSCAM
|
Down syndrome cell adhesion molecule |
| chrX_-_51797351 | 3.27 |
ENST00000375644.3
|
RP11-114H20.1
|
RP11-114H20.1 |
| chr14_+_74034310 | 3.23 |
ENST00000538782.1
|
ACOT2
|
acyl-CoA thioesterase 2 |
| chr6_-_100912785 | 3.19 |
ENST00000369208.3
|
SIM1
|
single-minded family bHLH transcription factor 1 |
| chr6_-_91296737 | 3.18 |
ENST00000369332.3
ENST00000369329.3 |
MAP3K7
|
mitogen-activated protein kinase kinase kinase 7 |
| chr11_+_46958248 | 3.16 |
ENST00000536126.1
ENST00000278460.7 ENST00000378618.2 ENST00000395460.2 ENST00000378615.3 ENST00000543718.1 |
C11orf49
|
chromosome 11 open reading frame 49 |
| chr2_-_208031943 | 3.15 |
ENST00000421199.1
ENST00000457962.1 |
KLF7
|
Kruppel-like factor 7 (ubiquitous) |
Network of associatons between targets according to the STRING database.
First level regulatory network of EMX1
Gene Ontology Analysis
Gene overrepresentation in biological_process category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 3.5 | 10.5 | GO:0014057 | positive regulation of acetylcholine secretion, neurotransmission(GO:0014057) |
| 3.4 | 41.1 | GO:1900747 | negative regulation of vascular endothelial growth factor signaling pathway(GO:1900747) |
| 3.4 | 10.1 | GO:2000864 | estrogen secretion(GO:0035937) estradiol secretion(GO:0035938) regulation of estrogen secretion(GO:2000861) regulation of estradiol secretion(GO:2000864) |
| 3.0 | 9.1 | GO:1901899 | dense core granule biogenesis(GO:0061110) positive regulation of relaxation of cardiac muscle(GO:1901899) regulation of dense core granule biogenesis(GO:2000705) |
| 3.0 | 53.7 | GO:0007021 | tubulin complex assembly(GO:0007021) |
| 2.6 | 7.9 | GO:1904204 | regulation of skeletal muscle hypertrophy(GO:1904204) |
| 2.6 | 7.7 | GO:1900158 | positive regulation of osteoclast proliferation(GO:0090290) negative regulation of bone mineralization involved in bone maturation(GO:1900158) |
| 2.5 | 2.5 | GO:0038095 | Fc-epsilon receptor signaling pathway(GO:0038095) |
| 2.4 | 24.4 | GO:0051611 | negative regulation of neurotransmitter uptake(GO:0051581) serotonin uptake(GO:0051610) regulation of serotonin uptake(GO:0051611) negative regulation of serotonin uptake(GO:0051612) |
| 2.4 | 7.3 | GO:0003218 | cardiac left ventricle formation(GO:0003218) |
| 2.4 | 14.5 | GO:1900239 | phenotypic switching(GO:0036166) regulation of phenotypic switching(GO:1900239) |
| 2.2 | 15.4 | GO:0071896 | protein localization to adherens junction(GO:0071896) |
| 2.2 | 8.7 | GO:0030807 | positive regulation of cyclic nucleotide catabolic process(GO:0030807) positive regulation of cAMP catabolic process(GO:0030822) positive regulation of purine nucleotide catabolic process(GO:0033123) |
| 2.0 | 10.2 | GO:0009449 | gamma-aminobutyric acid biosynthetic process(GO:0009449) |
| 1.9 | 13.5 | GO:0009753 | response to jasmonic acid(GO:0009753) cellular response to jasmonic acid stimulus(GO:0071395) |
| 1.8 | 5.5 | GO:0051795 | positive regulation of catagen(GO:0051795) activation of meiosis(GO:0090427) |
| 1.8 | 7.0 | GO:0018352 | protein-pyridoxal-5-phosphate linkage(GO:0018352) |
| 1.8 | 5.3 | GO:0015888 | thiamine transport(GO:0015888) thiamine transmembrane transport(GO:0071934) |
| 1.7 | 8.6 | GO:0060406 | positive regulation of penile erection(GO:0060406) |
| 1.6 | 16.4 | GO:0014894 | response to muscle inactivity involved in regulation of muscle adaptation(GO:0014877) response to denervation involved in regulation of muscle adaptation(GO:0014894) |
| 1.6 | 11.1 | GO:0090324 | negative regulation of oxidative phosphorylation(GO:0090324) |
| 1.5 | 5.9 | GO:0036022 | limb joint morphogenesis(GO:0036022) embryonic skeletal limb joint morphogenesis(GO:0036023) |
| 1.5 | 1.5 | GO:0051919 | positive regulation of fibrinolysis(GO:0051919) |
| 1.4 | 7.2 | GO:0038001 | paracrine signaling(GO:0038001) |
| 1.4 | 105.6 | GO:2000649 | regulation of sodium ion transmembrane transporter activity(GO:2000649) |
| 1.4 | 6.9 | GO:1901509 | regulation of endothelial tube morphogenesis(GO:1901509) |
| 1.4 | 2.7 | GO:0070309 | lens fiber cell morphogenesis(GO:0070309) |
| 1.4 | 19.1 | GO:0098828 | positive regulation of inhibitory postsynaptic potential(GO:0097151) modulation of inhibitory postsynaptic potential(GO:0098828) |
| 1.3 | 5.3 | GO:1902261 | positive regulation of delayed rectifier potassium channel activity(GO:1902261) |
| 1.3 | 3.9 | GO:0000294 | nuclear-transcribed mRNA catabolic process, endonucleolytic cleavage-dependent decay(GO:0000294) |
| 1.3 | 5.2 | GO:0072675 | multinuclear osteoclast differentiation(GO:0072674) osteoclast fusion(GO:0072675) |
| 1.3 | 3.9 | GO:0046947 | hydroxylysine metabolic process(GO:0046946) hydroxylysine biosynthetic process(GO:0046947) |
| 1.2 | 3.7 | GO:0035995 | skeletal muscle myosin thick filament assembly(GO:0030241) detection of muscle stretch(GO:0035995) |
| 1.2 | 3.7 | GO:1990414 | replication-born double-strand break repair via sister chromatid exchange(GO:1990414) |
| 1.2 | 3.6 | GO:1990523 | bone regeneration(GO:1990523) |
| 1.2 | 3.5 | GO:0038193 | thromboxane A2 signaling pathway(GO:0038193) |
| 1.1 | 9.2 | GO:0060414 | aorta smooth muscle tissue morphogenesis(GO:0060414) |
| 1.1 | 4.5 | GO:0021812 | neuronal-glial interaction involved in cerebral cortex radial glia guided migration(GO:0021812) |
| 1.1 | 6.6 | GO:1902514 | regulation of calcium ion transmembrane transport via high voltage-gated calcium channel(GO:1902514) positive regulation of calcium ion transmembrane transport via high voltage-gated calcium channel(GO:1904879) |
| 1.1 | 4.2 | GO:0006114 | glycerol biosynthetic process(GO:0006114) |
| 1.0 | 24.0 | GO:0007597 | blood coagulation, intrinsic pathway(GO:0007597) |
| 1.0 | 5.2 | GO:1902714 | negative regulation of interferon-gamma secretion(GO:1902714) |
| 1.0 | 3.1 | GO:0018153 | isopeptide cross-linking via N6-(L-isoglutamyl)-L-lysine(GO:0018153) isopeptide cross-linking(GO:0018262) |
| 1.0 | 4.0 | GO:0014807 | regulation of somitogenesis(GO:0014807) |
| 1.0 | 4.9 | GO:1902268 | negative regulation of polyamine transmembrane transport(GO:1902268) |
| 1.0 | 7.8 | GO:0007258 | JUN phosphorylation(GO:0007258) |
| 1.0 | 2.9 | GO:0030311 | poly-N-acetyllactosamine metabolic process(GO:0030309) poly-N-acetyllactosamine biosynthetic process(GO:0030311) |
| 1.0 | 2.9 | GO:1903625 | negative regulation of DNA catabolic process(GO:1903625) |
| 1.0 | 2.9 | GO:1901993 | meiotic cell cycle phase transition(GO:0044771) regulation of meiotic cell cycle phase transition(GO:1901993) negative regulation of meiotic cell cycle phase transition(GO:1901994) |
| 0.9 | 5.2 | GO:0000707 | meiotic DNA recombinase assembly(GO:0000707) |
| 0.8 | 19.5 | GO:0003334 | keratinocyte development(GO:0003334) |
| 0.8 | 4.2 | GO:0018101 | protein citrullination(GO:0018101) histone citrullination(GO:0036414) |
| 0.8 | 11.7 | GO:0010739 | positive regulation of protein kinase A signaling(GO:0010739) |
| 0.8 | 7.7 | GO:0016198 | axon choice point recognition(GO:0016198) |
| 0.8 | 3.8 | GO:0035063 | nuclear speck organization(GO:0035063) |
| 0.8 | 3.1 | GO:0086097 | phospholipase C-activating angiotensin-activated signaling pathway(GO:0086097) |
| 0.7 | 7.2 | GO:0048743 | positive regulation of skeletal muscle fiber development(GO:0048743) |
| 0.7 | 3.5 | GO:0090650 | response to oxygen-glucose deprivation(GO:0090649) cellular response to oxygen-glucose deprivation(GO:0090650) |
| 0.7 | 4.2 | GO:0045110 | intermediate filament bundle assembly(GO:0045110) |
| 0.7 | 6.2 | GO:0048312 | intracellular distribution of mitochondria(GO:0048312) |
| 0.7 | 2.0 | GO:0019605 | benzoate metabolic process(GO:0018874) butyrate metabolic process(GO:0019605) |
| 0.7 | 4.0 | GO:0010643 | cell communication by chemical coupling(GO:0010643) |
| 0.7 | 2.6 | GO:1902231 | positive regulation of intrinsic apoptotic signaling pathway in response to DNA damage(GO:1902231) |
| 0.7 | 8.5 | GO:0031573 | intra-S DNA damage checkpoint(GO:0031573) |
| 0.6 | 0.6 | GO:0070668 | regulation of mast cell proliferation(GO:0070666) positive regulation of mast cell proliferation(GO:0070668) |
| 0.6 | 5.7 | GO:0043123 | positive regulation of I-kappaB kinase/NF-kappaB signaling(GO:0043123) |
| 0.6 | 7.4 | GO:0007016 | cytoskeletal anchoring at plasma membrane(GO:0007016) |
| 0.6 | 2.8 | GO:0010513 | positive regulation of phosphatidylinositol biosynthetic process(GO:0010513) |
| 0.6 | 1.7 | GO:2000523 | regulation of T cell costimulation(GO:2000523) positive regulation of T cell costimulation(GO:2000525) |
| 0.5 | 17.5 | GO:0009083 | branched-chain amino acid catabolic process(GO:0009083) |
| 0.5 | 2.7 | GO:1902731 | ovarian cumulus expansion(GO:0001550) fused antrum stage(GO:0048165) negative regulation of chondrocyte proliferation(GO:1902731) |
| 0.5 | 1.9 | GO:0032954 | regulation of cytokinetic process(GO:0032954) regulation of mitotic cytokinetic process(GO:1903436) positive regulation of mitotic cytokinetic process(GO:1903438) positive regulation of mitotic cytokinesis(GO:1903490) |
| 0.5 | 4.2 | GO:0032534 | regulation of microvillus assembly(GO:0032534) |
| 0.5 | 7.4 | GO:0021670 | lateral ventricle development(GO:0021670) |
| 0.5 | 5.5 | GO:1900028 | negative regulation of ruffle assembly(GO:1900028) |
| 0.5 | 3.6 | GO:2000773 | negative regulation of cellular senescence(GO:2000773) |
| 0.4 | 0.4 | GO:0061551 | cerebral cortex tangential migration using cell-axon interactions(GO:0021824) gonadotrophin-releasing hormone neuronal migration to the hypothalamus(GO:0021828) hypothalamic tangential migration using cell-axon interactions(GO:0021856) cranial ganglion development(GO:0061550) trigeminal ganglion development(GO:0061551) facioacoustic ganglion development(GO:1903375) |
| 0.4 | 1.3 | GO:0070781 | arginine biosynthetic process via ornithine(GO:0042450) response to biotin(GO:0070781) |
| 0.4 | 3.9 | GO:0035799 | ureter maturation(GO:0035799) |
| 0.4 | 2.5 | GO:0055059 | asymmetric neuroblast division(GO:0055059) |
| 0.4 | 3.6 | GO:0098870 | neuronal action potential propagation(GO:0019227) action potential propagation(GO:0098870) |
| 0.4 | 2.0 | GO:1904383 | response to sodium phosphate(GO:1904383) |
| 0.4 | 7.0 | GO:0002726 | positive regulation of T cell cytokine production(GO:0002726) |
| 0.4 | 2.3 | GO:1903960 | regulation of plasma membrane long-chain fatty acid transport(GO:0010746) negative regulation of plasma membrane long-chain fatty acid transport(GO:0010748) negative regulation of anion transmembrane transport(GO:1903960) |
| 0.4 | 3.0 | GO:0060665 | regulation of branching involved in salivary gland morphogenesis by mesenchymal-epithelial signaling(GO:0060665) |
| 0.4 | 2.6 | GO:0072513 | positive regulation of secondary heart field cardioblast proliferation(GO:0072513) |
| 0.4 | 1.1 | GO:0036060 | filtration diaphragm assembly(GO:0036058) slit diaphragm assembly(GO:0036060) |
| 0.4 | 7.3 | GO:0060716 | labyrinthine layer blood vessel development(GO:0060716) |
| 0.3 | 9.3 | GO:0044458 | motile cilium assembly(GO:0044458) |
| 0.3 | 7.6 | GO:0009312 | oligosaccharide biosynthetic process(GO:0009312) |
| 0.3 | 2.7 | GO:0034587 | piRNA metabolic process(GO:0034587) |
| 0.3 | 12.8 | GO:0015804 | neutral amino acid transport(GO:0015804) |
| 0.3 | 2.2 | GO:0034350 | regulation of glial cell apoptotic process(GO:0034350) negative regulation of glial cell apoptotic process(GO:0034351) positive regulation of B cell receptor signaling pathway(GO:0050861) |
| 0.3 | 2.2 | GO:0051594 | detection of carbohydrate stimulus(GO:0009730) detection of hexose stimulus(GO:0009732) detection of monosaccharide stimulus(GO:0034287) detection of glucose(GO:0051594) |
| 0.3 | 0.6 | GO:0032489 | regulation of Cdc42 protein signal transduction(GO:0032489) |
| 0.3 | 10.8 | GO:0018146 | keratan sulfate biosynthetic process(GO:0018146) |
| 0.3 | 3.4 | GO:0010727 | negative regulation of hydrogen peroxide metabolic process(GO:0010727) |
| 0.3 | 2.4 | GO:0060020 | Bergmann glial cell differentiation(GO:0060020) |
| 0.3 | 0.9 | GO:0002326 | B cell lineage commitment(GO:0002326) |
| 0.3 | 1.2 | GO:2000253 | positive regulation of feeding behavior(GO:2000253) |
| 0.3 | 2.1 | GO:0019532 | oxalate transport(GO:0019532) |
| 0.3 | 3.8 | GO:0008627 | intrinsic apoptotic signaling pathway in response to osmotic stress(GO:0008627) |
| 0.3 | 2.3 | GO:0090063 | positive regulation of microtubule nucleation(GO:0090063) |
| 0.3 | 2.0 | GO:0021842 | directional guidance of interneurons involved in migration from the subpallium to the cortex(GO:0021840) chemorepulsion involved in interneuron migration from the subpallium to the cortex(GO:0021842) |
| 0.3 | 1.4 | GO:2001023 | regulation of response to drug(GO:2001023) |
| 0.3 | 1.4 | GO:0061107 | seminal vesicle development(GO:0061107) |
| 0.3 | 3.3 | GO:0048842 | positive regulation of axon extension involved in axon guidance(GO:0048842) |
| 0.3 | 4.1 | GO:0015671 | oxygen transport(GO:0015671) |
| 0.3 | 6.5 | GO:0061049 | physiological muscle hypertrophy(GO:0003298) physiological cardiac muscle hypertrophy(GO:0003301) cell growth involved in cardiac muscle cell development(GO:0061049) |
| 0.3 | 2.4 | GO:0000712 | resolution of meiotic recombination intermediates(GO:0000712) |
| 0.3 | 7.8 | GO:0070884 | regulation of calcineurin-NFAT signaling cascade(GO:0070884) |
| 0.3 | 0.5 | GO:0060686 | negative regulation of prostatic bud formation(GO:0060686) |
| 0.2 | 1.4 | GO:0010133 | proline catabolic process to glutamate(GO:0010133) |
| 0.2 | 0.2 | GO:0035992 | tendon cell differentiation(GO:0035990) tendon formation(GO:0035992) |
| 0.2 | 0.7 | GO:0038043 | interleukin-5-mediated signaling pathway(GO:0038043) |
| 0.2 | 4.0 | GO:0043562 | cellular response to nitrogen starvation(GO:0006995) cellular response to nitrogen levels(GO:0043562) |
| 0.2 | 2.8 | GO:0030321 | transepithelial chloride transport(GO:0030321) |
| 0.2 | 11.5 | GO:0030212 | hyaluronan metabolic process(GO:0030212) |
| 0.2 | 0.9 | GO:0014816 | skeletal muscle satellite cell differentiation(GO:0014816) negative regulation of skeletal muscle cell proliferation(GO:0014859) regulation of skeletal muscle tissue growth(GO:0048631) negative regulation of skeletal muscle satellite cell proliferation(GO:1902723) |
| 0.2 | 2.2 | GO:1902260 | negative regulation of delayed rectifier potassium channel activity(GO:1902260) |
| 0.2 | 10.7 | GO:0050911 | detection of chemical stimulus involved in sensory perception of smell(GO:0050911) |
| 0.2 | 1.3 | GO:0051970 | negative regulation of transmission of nerve impulse(GO:0051970) |
| 0.2 | 3.0 | GO:0034372 | very-low-density lipoprotein particle remodeling(GO:0034372) phosphatidylcholine catabolic process(GO:0034638) |
| 0.2 | 4.9 | GO:0070233 | negative regulation of T cell apoptotic process(GO:0070233) |
| 0.2 | 1.0 | GO:0050955 | thermoception(GO:0050955) |
| 0.2 | 32.1 | GO:0007156 | homophilic cell adhesion via plasma membrane adhesion molecules(GO:0007156) |
| 0.2 | 3.7 | GO:0032780 | negative regulation of ATPase activity(GO:0032780) |
| 0.2 | 7.2 | GO:1901385 | regulation of voltage-gated calcium channel activity(GO:1901385) |
| 0.2 | 3.5 | GO:0060397 | JAK-STAT cascade involved in growth hormone signaling pathway(GO:0060397) |
| 0.2 | 1.9 | GO:0009642 | response to light intensity(GO:0009642) |
| 0.2 | 4.9 | GO:0039703 | viral RNA genome replication(GO:0039694) RNA replication(GO:0039703) |
| 0.2 | 3.7 | GO:0030502 | negative regulation of bone mineralization(GO:0030502) |
| 0.2 | 2.0 | GO:0019800 | peptide cross-linking via chondroitin 4-sulfate glycosaminoglycan(GO:0019800) |
| 0.2 | 3.4 | GO:0033141 | positive regulation of peptidyl-serine phosphorylation of STAT protein(GO:0033141) |
| 0.2 | 4.7 | GO:0071526 | semaphorin-plexin signaling pathway(GO:0071526) |
| 0.2 | 4.3 | GO:0034204 | lipid translocation(GO:0034204) phospholipid translocation(GO:0045332) |
| 0.2 | 2.9 | GO:1904380 | endoplasmic reticulum mannose trimming(GO:1904380) |
| 0.2 | 0.8 | GO:0072233 | thick ascending limb development(GO:0072023) metanephric thick ascending limb development(GO:0072233) |
| 0.2 | 0.3 | GO:0033364 | mast cell secretory granule organization(GO:0033364) |
| 0.2 | 1.1 | GO:1901374 | epinephrine transport(GO:0048241) acetate ester transport(GO:1901374) |
| 0.2 | 10.0 | GO:0009250 | glycogen biosynthetic process(GO:0005978) glucan biosynthetic process(GO:0009250) |
| 0.2 | 1.7 | GO:0031284 | positive regulation of guanylate cyclase activity(GO:0031284) |
| 0.1 | 1.6 | GO:0001731 | formation of translation preinitiation complex(GO:0001731) |
| 0.1 | 0.4 | GO:0051697 | protein delipidation(GO:0051697) |
| 0.1 | 3.6 | GO:0051642 | centrosome localization(GO:0051642) |
| 0.1 | 2.0 | GO:0003091 | renal water homeostasis(GO:0003091) |
| 0.1 | 0.7 | GO:0048733 | sebaceous gland development(GO:0048733) |
| 0.1 | 1.4 | GO:0034375 | high-density lipoprotein particle remodeling(GO:0034375) |
| 0.1 | 2.0 | GO:0021516 | dorsal spinal cord development(GO:0021516) |
| 0.1 | 6.4 | GO:0008347 | glial cell migration(GO:0008347) |
| 0.1 | 3.2 | GO:0000038 | very long-chain fatty acid metabolic process(GO:0000038) |
| 0.1 | 0.5 | GO:0019240 | citrulline biosynthetic process(GO:0019240) |
| 0.1 | 0.4 | GO:0061113 | pancreas morphogenesis(GO:0061113) |
| 0.1 | 4.3 | GO:0006699 | bile acid biosynthetic process(GO:0006699) |
| 0.1 | 1.6 | GO:0060009 | Sertoli cell development(GO:0060009) |
| 0.1 | 2.4 | GO:0007638 | mechanosensory behavior(GO:0007638) |
| 0.1 | 1.0 | GO:0006883 | cellular sodium ion homeostasis(GO:0006883) |
| 0.1 | 3.3 | GO:0030199 | collagen fibril organization(GO:0030199) |
| 0.1 | 2.9 | GO:0031529 | ruffle organization(GO:0031529) |
| 0.1 | 1.0 | GO:0001867 | complement activation, lectin pathway(GO:0001867) |
| 0.1 | 1.5 | GO:2000353 | positive regulation of endothelial cell apoptotic process(GO:2000353) |
| 0.1 | 12.5 | GO:0007498 | mesoderm development(GO:0007498) |
| 0.1 | 1.9 | GO:0035435 | phosphate ion transmembrane transport(GO:0035435) |
| 0.1 | 1.3 | GO:0046069 | cGMP catabolic process(GO:0046069) |
| 0.1 | 11.2 | GO:0035023 | regulation of Rho protein signal transduction(GO:0035023) |
| 0.1 | 0.5 | GO:0033353 | S-adenosylmethionine cycle(GO:0033353) |
| 0.1 | 11.3 | GO:0006937 | regulation of muscle contraction(GO:0006937) |
| 0.1 | 1.4 | GO:0006706 | steroid catabolic process(GO:0006706) |
| 0.1 | 3.7 | GO:0035418 | protein localization to synapse(GO:0035418) |
| 0.1 | 4.4 | GO:0007368 | determination of left/right symmetry(GO:0007368) |
| 0.1 | 5.0 | GO:0030819 | positive regulation of cAMP biosynthetic process(GO:0030819) |
| 0.1 | 0.6 | GO:0000910 | cytokinesis(GO:0000910) |
| 0.1 | 0.7 | GO:0043010 | camera-type eye development(GO:0043010) |
| 0.1 | 2.8 | GO:0016486 | peptide hormone processing(GO:0016486) |
| 0.1 | 2.1 | GO:0042776 | mitochondrial ATP synthesis coupled proton transport(GO:0042776) |
| 0.1 | 2.6 | GO:0032728 | positive regulation of interferon-beta production(GO:0032728) |
| 0.1 | 0.3 | GO:0051005 | negative regulation of lipoprotein lipase activity(GO:0051005) |
| 0.1 | 2.2 | GO:0060338 | regulation of type I interferon-mediated signaling pathway(GO:0060338) |
| 0.1 | 1.5 | GO:0070584 | mitochondrion morphogenesis(GO:0070584) |
| 0.1 | 1.2 | GO:0034142 | toll-like receptor 4 signaling pathway(GO:0034142) |
| 0.1 | 0.9 | GO:0030277 | maintenance of gastrointestinal epithelium(GO:0030277) |
| 0.1 | 2.9 | GO:0008360 | regulation of cell shape(GO:0008360) |
| 0.1 | 2.9 | GO:0071300 | cellular response to retinoic acid(GO:0071300) |
| 0.1 | 11.9 | GO:0060271 | cilium morphogenesis(GO:0060271) |
| 0.0 | 3.2 | GO:0001657 | ureteric bud development(GO:0001657) mesonephric epithelium development(GO:0072163) mesonephric tubule development(GO:0072164) |
| 0.0 | 1.1 | GO:0086010 | membrane depolarization during action potential(GO:0086010) |
| 0.0 | 0.8 | GO:0036315 | cellular response to sterol(GO:0036315) |
| 0.0 | 1.2 | GO:0016338 | calcium-independent cell-cell adhesion via plasma membrane cell-adhesion molecules(GO:0016338) |
| 0.0 | 0.8 | GO:0090110 | cargo loading into COPII-coated vesicle(GO:0090110) |
| 0.0 | 0.8 | GO:0014059 | dopamine secretion(GO:0014046) regulation of dopamine secretion(GO:0014059) |
| 0.0 | 7.8 | GO:0051056 | regulation of small GTPase mediated signal transduction(GO:0051056) |
| 0.0 | 1.6 | GO:0038128 | ERBB2 signaling pathway(GO:0038128) |
| 0.0 | 0.4 | GO:0006703 | estrogen biosynthetic process(GO:0006703) |
| 0.0 | 0.2 | GO:1901098 | regulation of autophagosome maturation(GO:1901096) positive regulation of autophagosome maturation(GO:1901098) |
| 0.0 | 1.9 | GO:0002763 | positive regulation of myeloid leukocyte differentiation(GO:0002763) |
| 0.0 | 1.0 | GO:0042462 | eye photoreceptor cell development(GO:0042462) |
| 0.0 | 0.5 | GO:0005513 | detection of calcium ion(GO:0005513) |
| 0.0 | 1.3 | GO:0042475 | odontogenesis of dentin-containing tooth(GO:0042475) |
| 0.0 | 0.9 | GO:0007205 | protein kinase C-activating G-protein coupled receptor signaling pathway(GO:0007205) |
| 0.0 | 5.0 | GO:0007265 | Ras protein signal transduction(GO:0007265) |
| 0.0 | 4.6 | GO:0009408 | response to heat(GO:0009408) |
| 0.0 | 0.3 | GO:0001765 | membrane raft assembly(GO:0001765) |
| 0.0 | 2.0 | GO:0042795 | snRNA transcription(GO:0009301) snRNA transcription from RNA polymerase II promoter(GO:0042795) |
| 0.0 | 0.2 | GO:0030574 | collagen catabolic process(GO:0030574) multicellular organism catabolic process(GO:0044243) |
| 0.0 | 2.9 | GO:0007601 | visual perception(GO:0007601) |
| 0.0 | 0.7 | GO:0000387 | spliceosomal snRNP assembly(GO:0000387) |
| 0.0 | 0.7 | GO:0015701 | bicarbonate transport(GO:0015701) |
| 0.0 | 1.4 | GO:0007286 | spermatid development(GO:0007286) |
| 0.0 | 0.2 | GO:0002230 | positive regulation of defense response to virus by host(GO:0002230) |
| 0.0 | 2.4 | GO:0006836 | neurotransmitter transport(GO:0006836) |
| 0.0 | 0.6 | GO:0046847 | filopodium assembly(GO:0046847) regulation of filopodium assembly(GO:0051489) |
| 0.0 | 1.3 | GO:1902476 | chloride transmembrane transport(GO:1902476) |
| 0.0 | 1.5 | GO:0048813 | dendrite morphogenesis(GO:0048813) |
| 0.0 | 0.5 | GO:0014823 | response to activity(GO:0014823) |
| 0.0 | 0.4 | GO:0006099 | tricarboxylic acid cycle(GO:0006099) |
| 0.0 | 0.1 | GO:0002566 | somatic diversification of immune receptors via somatic mutation(GO:0002566) somatic hypermutation of immunoglobulin genes(GO:0016446) |
| 0.0 | 1.0 | GO:0070268 | cornification(GO:0070268) |
Gene overrepresentation in cellular_component category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 3.6 | 14.4 | GO:0097224 | sperm connecting piece(GO:0097224) |
| 3.4 | 53.7 | GO:0097512 | cardiac myofibril(GO:0097512) |
| 2.9 | 41.1 | GO:0098647 | collagen type VI trimer(GO:0005589) collagen beaded filament(GO:0098647) |
| 2.5 | 7.4 | GO:0005879 | axonemal microtubule(GO:0005879) |
| 2.0 | 16.4 | GO:0014802 | terminal cisterna(GO:0014802) |
| 1.5 | 4.5 | GO:0005607 | laminin-2 complex(GO:0005607) |
| 1.3 | 9.4 | GO:0045252 | oxoglutarate dehydrogenase complex(GO:0045252) |
| 1.3 | 15.4 | GO:0097025 | MPP7-DLG1-LIN7 complex(GO:0097025) |
| 1.1 | 7.7 | GO:0032584 | growth cone membrane(GO:0032584) |
| 0.9 | 7.9 | GO:0042567 | insulin-like growth factor ternary complex(GO:0042567) |
| 0.9 | 5.3 | GO:0014701 | junctional sarcoplasmic reticulum membrane(GO:0014701) |
| 0.9 | 5.2 | GO:0033063 | DNA recombinase mediator complex(GO:0033061) Rad51B-Rad51C-Rad51D-XRCC2 complex(GO:0033063) |
| 0.8 | 7.1 | GO:0016013 | syntrophin complex(GO:0016013) |
| 0.8 | 3.9 | GO:0044530 | supraspliceosomal complex(GO:0044530) |
| 0.6 | 3.6 | GO:0002177 | manchette(GO:0002177) |
| 0.6 | 4.7 | GO:0097504 | Gemini of coiled bodies(GO:0097504) |
| 0.5 | 14.2 | GO:0034451 | centriolar satellite(GO:0034451) |
| 0.5 | 2.7 | GO:0071546 | pi-body(GO:0071546) |
| 0.5 | 7.0 | GO:0008385 | IkappaB kinase complex(GO:0008385) |
| 0.5 | 6.6 | GO:1990454 | L-type voltage-gated calcium channel complex(GO:1990454) |
| 0.5 | 8.5 | GO:0001518 | voltage-gated sodium channel complex(GO:0001518) |
| 0.5 | 1.4 | GO:0097182 | protein C inhibitor-TMPRSS7 complex(GO:0036024) protein C inhibitor-TMPRSS11E complex(GO:0036025) protein C inhibitor-PLAT complex(GO:0036026) protein C inhibitor-PLAU complex(GO:0036027) protein C inhibitor-thrombin complex(GO:0036028) protein C inhibitor-KLK3 complex(GO:0036029) protein C inhibitor-plasma kallikrein complex(GO:0036030) serine protease inhibitor complex(GO:0097180) protein C inhibitor-coagulation factor V complex(GO:0097181) protein C inhibitor-coagulation factor Xa complex(GO:0097182) protein C inhibitor-coagulation factor XI complex(GO:0097183) |
| 0.5 | 22.0 | GO:0048786 | presynaptic active zone(GO:0048786) |
| 0.5 | 7.7 | GO:0042583 | chromaffin granule(GO:0042583) |
| 0.4 | 3.3 | GO:0005593 | FACIT collagen trimer(GO:0005593) |
| 0.4 | 3.5 | GO:0016600 | flotillin complex(GO:0016600) |
| 0.4 | 9.7 | GO:0090665 | dystrophin-associated glycoprotein complex(GO:0016010) glycoprotein complex(GO:0090665) |
| 0.3 | 2.4 | GO:0043240 | Fanconi anaemia nuclear complex(GO:0043240) |
| 0.3 | 5.7 | GO:0097342 | ripoptosome(GO:0097342) |
| 0.3 | 2.2 | GO:1990635 | proximal dendrite(GO:1990635) |
| 0.3 | 2.6 | GO:0060077 | inhibitory synapse(GO:0060077) |
| 0.3 | 4.1 | GO:0005833 | hemoglobin complex(GO:0005833) |
| 0.3 | 1.6 | GO:0031466 | Cul5-RING ubiquitin ligase complex(GO:0031466) |
| 0.3 | 1.9 | GO:0070938 | contractile ring(GO:0070938) |
| 0.3 | 3.5 | GO:0031904 | endosome lumen(GO:0031904) |
| 0.3 | 14.0 | GO:0002102 | podosome(GO:0002102) |
| 0.2 | 1.5 | GO:0031232 | extrinsic component of external side of plasma membrane(GO:0031232) |
| 0.2 | 13.1 | GO:0008023 | transcription elongation factor complex(GO:0008023) |
| 0.2 | 1.6 | GO:0097452 | GAIT complex(GO:0097452) |
| 0.2 | 2.7 | GO:1990712 | HFE-transferrin receptor complex(GO:1990712) |
| 0.2 | 6.2 | GO:0031307 | integral component of mitochondrial outer membrane(GO:0031307) |
| 0.2 | 4.9 | GO:0005942 | phosphatidylinositol 3-kinase complex(GO:0005942) |
| 0.2 | 22.6 | GO:0031093 | platelet alpha granule lumen(GO:0031093) |
| 0.2 | 10.7 | GO:0031672 | A band(GO:0031672) |
| 0.2 | 14.0 | GO:0005902 | microvillus(GO:0005902) |
| 0.2 | 7.2 | GO:0005891 | voltage-gated calcium channel complex(GO:0005891) |
| 0.2 | 46.4 | GO:0043209 | myelin sheath(GO:0043209) |
| 0.2 | 5.2 | GO:0005921 | gap junction(GO:0005921) |
| 0.2 | 2.3 | GO:0005869 | dynactin complex(GO:0005869) |
| 0.2 | 12.1 | GO:0097610 | cleavage furrow(GO:0032154) cell surface furrow(GO:0097610) |
| 0.2 | 2.1 | GO:0097225 | sperm midpiece(GO:0097225) |
| 0.2 | 17.3 | GO:0017053 | transcriptional repressor complex(GO:0017053) |
| 0.1 | 3.7 | GO:0030867 | rough endoplasmic reticulum membrane(GO:0030867) |
| 0.1 | 2.9 | GO:0044322 | endoplasmic reticulum quality control compartment(GO:0044322) |
| 0.1 | 14.4 | GO:0005796 | Golgi lumen(GO:0005796) |
| 0.1 | 2.1 | GO:0000276 | mitochondrial proton-transporting ATP synthase complex, coupling factor F(o)(GO:0000276) |
| 0.1 | 20.3 | GO:0001650 | fibrillar center(GO:0001650) |
| 0.1 | 3.6 | GO:0005868 | cytoplasmic dynein complex(GO:0005868) |
| 0.1 | 2.6 | GO:0030992 | intraciliary transport particle B(GO:0030992) |
| 0.1 | 2.7 | GO:0030127 | COPII vesicle coat(GO:0030127) |
| 0.1 | 2.6 | GO:0005666 | DNA-directed RNA polymerase III complex(GO:0005666) |
| 0.1 | 4.0 | GO:0000421 | autophagosome membrane(GO:0000421) |
| 0.1 | 2.2 | GO:0005911 | cell-cell junction(GO:0005911) |
| 0.1 | 1.4 | GO:0097440 | apical dendrite(GO:0097440) |
| 0.1 | 2.0 | GO:0030673 | axolemma(GO:0030673) |
| 0.1 | 1.6 | GO:0033276 | transcription factor TFTC complex(GO:0033276) |
| 0.1 | 2.1 | GO:0045178 | basal part of cell(GO:0045178) |
| 0.1 | 22.2 | GO:0005788 | endoplasmic reticulum lumen(GO:0005788) |
| 0.1 | 5.2 | GO:0033116 | endoplasmic reticulum-Golgi intermediate compartment membrane(GO:0033116) |
| 0.1 | 24.3 | GO:0098857 | membrane raft(GO:0045121) membrane microdomain(GO:0098857) |
| 0.1 | 5.1 | GO:0001669 | acrosomal vesicle(GO:0001669) |
| 0.1 | 3.7 | GO:0005913 | cell-cell adherens junction(GO:0005913) |
| 0.1 | 1.8 | GO:0097381 | photoreceptor disc membrane(GO:0097381) |
| 0.1 | 6.5 | GO:0030018 | Z disc(GO:0030018) |
| 0.1 | 3.1 | GO:0001533 | cornified envelope(GO:0001533) |
| 0.1 | 0.2 | GO:0042406 | extrinsic component of endoplasmic reticulum membrane(GO:0042406) |
| 0.1 | 4.1 | GO:0005604 | basement membrane(GO:0005604) |
| 0.1 | 2.4 | GO:0016235 | aggresome(GO:0016235) |
| 0.1 | 8.9 | GO:0001726 | ruffle(GO:0001726) |
| 0.1 | 6.8 | GO:0030496 | midbody(GO:0030496) |
| 0.1 | 92.7 | GO:0005887 | integral component of plasma membrane(GO:0005887) |
| 0.0 | 5.4 | GO:0030027 | lamellipodium(GO:0030027) |
| 0.0 | 5.1 | GO:0016363 | nuclear matrix(GO:0016363) |
| 0.0 | 1.3 | GO:0005791 | rough endoplasmic reticulum(GO:0005791) |
| 0.0 | 1.8 | GO:0005884 | actin filament(GO:0005884) |
| 0.0 | 7.9 | GO:0005578 | proteinaceous extracellular matrix(GO:0005578) |
| 0.0 | 6.9 | GO:0005938 | cell cortex(GO:0005938) |
| 0.0 | 2.4 | GO:0030672 | synaptic vesicle membrane(GO:0030672) exocytic vesicle membrane(GO:0099501) |
| 0.0 | 4.6 | GO:0009897 | external side of plasma membrane(GO:0009897) |
| 0.0 | 1.0 | GO:0031252 | cell leading edge(GO:0031252) |
| 0.0 | 1.1 | GO:0005637 | nuclear inner membrane(GO:0005637) |
| 0.0 | 2.2 | GO:0005741 | mitochondrial outer membrane(GO:0005741) |
| 0.0 | 1.2 | GO:0005923 | bicellular tight junction(GO:0005923) |
| 0.0 | 0.2 | GO:0005832 | chaperonin-containing T-complex(GO:0005832) |
| 0.0 | 9.4 | GO:0005789 | endoplasmic reticulum membrane(GO:0005789) |
| 0.0 | 0.5 | GO:0042645 | nucleoid(GO:0009295) mitochondrial nucleoid(GO:0042645) |
Gene overrepresentation in molecular_function category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 2.5 | 7.6 | GO:0047273 | galactosylgalactosylglucosylceramide beta-D-acetylgalactosaminyltransferase activity(GO:0047273) |
| 2.3 | 13.5 | GO:0047086 | phenanthrene 9,10-monooxygenase activity(GO:0018636) ketosteroid monooxygenase activity(GO:0047086) trans-1,2-dihydrobenzene-1,2-diol dehydrogenase activity(GO:0047115) |
| 2.1 | 10.5 | GO:0004995 | tachykinin receptor activity(GO:0004995) |
| 2.0 | 111.1 | GO:0017080 | sodium channel regulator activity(GO:0017080) |
| 1.9 | 58.9 | GO:0005212 | structural constituent of eye lens(GO:0005212) |
| 1.9 | 9.4 | GO:0004591 | oxoglutarate dehydrogenase (succinyl-transferring) activity(GO:0004591) |
| 1.8 | 9.2 | GO:0004687 | myosin light chain kinase activity(GO:0004687) |
| 1.8 | 5.3 | GO:0015234 | thiamine transmembrane transporter activity(GO:0015234) thiamine uptake transmembrane transporter activity(GO:0015403) |
| 1.5 | 10.2 | GO:0015501 | glutamate:sodium symporter activity(GO:0015501) |
| 1.4 | 4.2 | GO:1990254 | keratin filament binding(GO:1990254) |
| 1.3 | 7.8 | GO:0008597 | calcium-dependent protein serine/threonine phosphatase regulator activity(GO:0008597) |
| 1.3 | 7.8 | GO:0016909 | JUN kinase activity(GO:0004705) SAP kinase activity(GO:0016909) |
| 1.3 | 3.9 | GO:0008475 | procollagen-lysine 5-dioxygenase activity(GO:0008475) procollagen glucosyltransferase activity(GO:0033823) procollagen galactosyltransferase activity(GO:0050211) |
| 1.2 | 3.5 | GO:0004961 | thromboxane receptor activity(GO:0004960) thromboxane A2 receptor activity(GO:0004961) |
| 1.2 | 5.8 | GO:0048248 | CXCR3 chemokine receptor binding(GO:0048248) |
| 1.2 | 3.5 | GO:0005148 | prolactin receptor binding(GO:0005148) |
| 1.2 | 13.9 | GO:0032027 | myosin light chain binding(GO:0032027) |
| 1.1 | 5.7 | GO:0097199 | cysteine-type endopeptidase activity involved in apoptotic signaling pathway(GO:0097199) |
| 1.1 | 17.0 | GO:0008179 | adenylate cyclase binding(GO:0008179) |
| 1.0 | 5.2 | GO:0005152 | interleukin-1 receptor antagonist activity(GO:0005152) |
| 1.0 | 4.1 | GO:0031685 | adenosine receptor binding(GO:0031685) |
| 1.0 | 3.1 | GO:0001596 | angiotensin type I receptor activity(GO:0001596) |
| 1.0 | 32.1 | GO:0019865 | immunoglobulin binding(GO:0019865) |
| 1.0 | 6.9 | GO:0004301 | epoxide hydrolase activity(GO:0004301) |
| 1.0 | 4.9 | GO:0008073 | ornithine decarboxylase inhibitor activity(GO:0008073) |
| 1.0 | 2.9 | GO:0008532 | N-acetyllactosaminide beta-1,3-N-acetylglucosaminyltransferase activity(GO:0008532) |
| 0.8 | 4.2 | GO:0004668 | protein-arginine deiminase activity(GO:0004668) |
| 0.8 | 6.6 | GO:0086056 | voltage-gated calcium channel activity involved in AV node cell action potential(GO:0086056) |
| 0.8 | 2.3 | GO:0098519 | nucleotide phosphatase activity, acting on free nucleotides(GO:0098519) |
| 0.7 | 3.7 | GO:1990430 | extracellular matrix protein binding(GO:1990430) |
| 0.7 | 7.9 | GO:0031995 | insulin-like growth factor II binding(GO:0031995) |
| 0.7 | 4.9 | GO:0035005 | 1-phosphatidylinositol-4-phosphate 3-kinase activity(GO:0035005) |
| 0.7 | 12.4 | GO:0005522 | profilin binding(GO:0005522) |
| 0.7 | 6.8 | GO:0035727 | lysophosphatidic acid binding(GO:0035727) |
| 0.7 | 8.7 | GO:0017034 | Rap guanyl-nucleotide exchange factor activity(GO:0017034) |
| 0.6 | 1.9 | GO:0004613 | phosphoenolpyruvate carboxykinase activity(GO:0004611) phosphoenolpyruvate carboxykinase (GTP) activity(GO:0004613) |
| 0.6 | 3.7 | GO:0009378 | four-way junction helicase activity(GO:0009378) |
| 0.6 | 43.3 | GO:0050840 | extracellular matrix binding(GO:0050840) |
| 0.6 | 13.4 | GO:0051371 | muscle alpha-actinin binding(GO:0051371) |
| 0.6 | 7.2 | GO:0008331 | high voltage-gated calcium channel activity(GO:0008331) |
| 0.5 | 2.6 | GO:0061665 | SUMO ligase activity(GO:0061665) |
| 0.5 | 6.2 | GO:0019534 | toxin transporter activity(GO:0019534) |
| 0.5 | 1.5 | GO:0008502 | melatonin receptor activity(GO:0008502) |
| 0.5 | 1.5 | GO:0005174 | CD40 receptor binding(GO:0005174) |
| 0.5 | 4.5 | GO:0004969 | histamine receptor activity(GO:0004969) |
| 0.5 | 2.8 | GO:0045029 | UDP-activated nucleotide receptor activity(GO:0045029) |
| 0.5 | 5.2 | GO:0000150 | recombinase activity(GO:0000150) |
| 0.5 | 8.5 | GO:1905030 | voltage-gated sodium channel activity(GO:0005248) voltage-gated ion channel activity involved in regulation of postsynaptic membrane potential(GO:1905030) |
| 0.5 | 8.3 | GO:0004993 | G-protein coupled serotonin receptor activity(GO:0004993) |
| 0.4 | 2.2 | GO:0004699 | calcium-independent protein kinase C activity(GO:0004699) |
| 0.4 | 5.2 | GO:0030215 | semaphorin receptor binding(GO:0030215) |
| 0.4 | 7.7 | GO:0048018 | receptor agonist activity(GO:0048018) |
| 0.4 | 7.3 | GO:0032794 | GTPase activating protein binding(GO:0032794) |
| 0.4 | 14.5 | GO:0001103 | RNA polymerase II repressing transcription factor binding(GO:0001103) |
| 0.4 | 1.4 | GO:0004657 | proline dehydrogenase activity(GO:0004657) |
| 0.4 | 3.5 | GO:0036312 | phosphatidylinositol 3-kinase regulatory subunit binding(GO:0036312) |
| 0.4 | 3.9 | GO:0001758 | retinal dehydrogenase activity(GO:0001758) |
| 0.3 | 5.5 | GO:0005149 | interleukin-1 receptor binding(GO:0005149) |
| 0.3 | 2.1 | GO:0019531 | oxalate transmembrane transporter activity(GO:0019531) |
| 0.3 | 1.7 | GO:0008048 | calcium sensitive guanylate cyclase activator activity(GO:0008048) |
| 0.3 | 1.9 | GO:0016812 | hydrolase activity, acting on carbon-nitrogen (but not peptide) bonds, in cyclic amides(GO:0016812) |
| 0.3 | 4.2 | GO:0001011 | transcription factor activity, sequence-specific DNA binding, RNA polymerase recruiting(GO:0001011) transcription factor activity, TFIIB-class binding(GO:0001087) |
| 0.3 | 29.4 | GO:0004722 | protein serine/threonine phosphatase activity(GO:0004722) |
| 0.3 | 4.1 | GO:0005344 | oxygen transporter activity(GO:0005344) |
| 0.3 | 5.1 | GO:0034185 | apolipoprotein binding(GO:0034185) |
| 0.3 | 1.2 | GO:0004346 | glucose-6-phosphatase activity(GO:0004346) sugar-terminal-phosphatase activity(GO:0050309) |
| 0.3 | 2.7 | GO:0005025 | transforming growth factor beta receptor activity, type I(GO:0005025) |
| 0.3 | 3.2 | GO:0016290 | palmitoyl-CoA hydrolase activity(GO:0016290) |
| 0.3 | 2.2 | GO:0008865 | glucokinase activity(GO:0004340) hexokinase activity(GO:0004396) fructokinase activity(GO:0008865) mannokinase activity(GO:0019158) |
| 0.3 | 1.1 | GO:1901375 | acetylcholine transmembrane transporter activity(GO:0005277) secondary active organic cation transmembrane transporter activity(GO:0008513) acetate ester transmembrane transporter activity(GO:1901375) |
| 0.3 | 6.5 | GO:0008093 | cytoskeletal adaptor activity(GO:0008093) |
| 0.2 | 56.1 | GO:0051015 | actin filament binding(GO:0051015) |
| 0.2 | 2.9 | GO:1904264 | ubiquitin protein ligase activity involved in ERAD pathway(GO:1904264) |
| 0.2 | 3.4 | GO:0005132 | type I interferon receptor binding(GO:0005132) |
| 0.2 | 0.7 | GO:0004914 | interleukin-5 receptor activity(GO:0004914) |
| 0.2 | 0.5 | GO:0004087 | carbamoyl-phosphate synthase (ammonia) activity(GO:0004087) carbamoyl-phosphate synthase (glutamine-hydrolyzing) activity(GO:0004088) |
| 0.2 | 2.5 | GO:0030297 | transmembrane receptor protein tyrosine kinase activator activity(GO:0030297) |
| 0.2 | 8.2 | GO:0005184 | neuropeptide hormone activity(GO:0005184) |
| 0.2 | 2.0 | GO:0004321 | fatty-acyl-CoA synthase activity(GO:0004321) |
| 0.2 | 6.6 | GO:0015175 | neutral amino acid transmembrane transporter activity(GO:0015175) |
| 0.2 | 6.2 | GO:0071889 | 14-3-3 protein binding(GO:0071889) |
| 0.2 | 0.8 | GO:0090554 | phosphatidylcholine-translocating ATPase activity(GO:0090554) |
| 0.2 | 20.4 | GO:0004867 | serine-type endopeptidase inhibitor activity(GO:0004867) |
| 0.2 | 1.9 | GO:0070492 | oligosaccharide binding(GO:0070492) |
| 0.2 | 0.4 | GO:0047006 | 17-alpha,20-alpha-dihydroxypregn-4-en-3-one dehydrogenase activity(GO:0047006) |
| 0.2 | 1.5 | GO:0016846 | carbon-sulfur lyase activity(GO:0016846) |
| 0.2 | 3.6 | GO:0045504 | dynein heavy chain binding(GO:0045504) |
| 0.2 | 1.3 | GO:0048101 | calcium- and calmodulin-regulated 3',5'-cyclic-GMP phosphodiesterase activity(GO:0048101) |
| 0.2 | 4.1 | GO:0001056 | RNA polymerase III activity(GO:0001056) |
| 0.2 | 2.6 | GO:0005229 | intracellular calcium activated chloride channel activity(GO:0005229) |
| 0.2 | 3.5 | GO:0004012 | phospholipid-translocating ATPase activity(GO:0004012) |
| 0.2 | 1.9 | GO:0005436 | sodium:phosphate symporter activity(GO:0005436) |
| 0.2 | 23.4 | GO:0035591 | signaling adaptor activity(GO:0035591) |
| 0.2 | 3.7 | GO:0098641 | cadherin binding involved in cell-cell adhesion(GO:0098641) |
| 0.1 | 1.4 | GO:0008503 | benzodiazepine receptor activity(GO:0008503) |
| 0.1 | 13.3 | GO:0005089 | Rho guanyl-nucleotide exchange factor activity(GO:0005089) |
| 0.1 | 4.3 | GO:0015248 | sterol transporter activity(GO:0015248) |
| 0.1 | 1.3 | GO:0042301 | phosphate ion binding(GO:0042301) |
| 0.1 | 0.8 | GO:0008142 | oxysterol binding(GO:0008142) |
| 0.1 | 1.4 | GO:0004977 | melanocortin receptor activity(GO:0004977) |
| 0.1 | 1.6 | GO:0017162 | aryl hydrocarbon receptor binding(GO:0017162) |
| 0.1 | 11.2 | GO:0005179 | hormone activity(GO:0005179) |
| 0.1 | 5.2 | GO:0004984 | olfactory receptor activity(GO:0004984) |
| 0.1 | 0.5 | GO:0000146 | microfilament motor activity(GO:0000146) |
| 0.1 | 9.7 | GO:0017137 | Rab GTPase binding(GO:0017137) |
| 0.1 | 1.9 | GO:0008574 | ATP-dependent microtubule motor activity, plus-end-directed(GO:0008574) |
| 0.1 | 3.1 | GO:0044183 | protein binding involved in protein folding(GO:0044183) |
| 0.1 | 2.2 | GO:0070402 | NADPH binding(GO:0070402) |
| 0.1 | 1.3 | GO:0042923 | neuropeptide binding(GO:0042923) |
| 0.1 | 0.7 | GO:0004064 | arylesterase activity(GO:0004064) |
| 0.1 | 27.4 | GO:0005096 | GTPase activator activity(GO:0005096) |
| 0.1 | 0.5 | GO:0004013 | adenosylhomocysteinase activity(GO:0004013) trialkylsulfonium hydrolase activity(GO:0016802) |
| 0.1 | 1.2 | GO:0031593 | polyubiquitin binding(GO:0031593) |
| 0.1 | 5.2 | GO:0097110 | scaffold protein binding(GO:0097110) |
| 0.1 | 2.1 | GO:0046933 | proton-transporting ATP synthase activity, rotational mechanism(GO:0046933) |
| 0.1 | 3.0 | GO:0042056 | chemoattractant activity(GO:0042056) |
| 0.1 | 6.8 | GO:0005201 | extracellular matrix structural constituent(GO:0005201) |
| 0.1 | 0.9 | GO:0043325 | phosphatidylinositol-3,4-bisphosphate binding(GO:0043325) |
| 0.1 | 0.2 | GO:0004574 | oligo-1,6-glucosidase activity(GO:0004574) |
| 0.1 | 2.3 | GO:0070840 | dynein complex binding(GO:0070840) |
| 0.1 | 3.6 | GO:0042162 | telomeric DNA binding(GO:0042162) |
| 0.1 | 10.3 | GO:0008083 | growth factor activity(GO:0008083) |
| 0.1 | 18.4 | GO:0003779 | actin binding(GO:0003779) |
| 0.1 | 1.4 | GO:0048487 | beta-tubulin binding(GO:0048487) |
| 0.1 | 5.9 | GO:0004222 | metalloendopeptidase activity(GO:0004222) |
| 0.1 | 0.9 | GO:0004143 | diacylglycerol kinase activity(GO:0004143) |
| 0.1 | 0.9 | GO:0070330 | aromatase activity(GO:0070330) |
| 0.1 | 35.9 | GO:0005509 | calcium ion binding(GO:0005509) |
| 0.1 | 0.3 | GO:0010465 | nerve growth factor receptor activity(GO:0010465) |
| 0.0 | 1.4 | GO:0032934 | cholesterol binding(GO:0015485) sterol binding(GO:0032934) |
| 0.0 | 6.5 | GO:0005516 | calmodulin binding(GO:0005516) |
| 0.0 | 0.5 | GO:0015269 | calcium-activated potassium channel activity(GO:0015269) |
| 0.0 | 1.1 | GO:0019198 | transmembrane receptor protein tyrosine phosphatase activity(GO:0005001) transmembrane receptor protein phosphatase activity(GO:0019198) |
| 0.0 | 1.7 | GO:0005164 | tumor necrosis factor receptor binding(GO:0005164) |
| 0.0 | 0.8 | GO:0042166 | acetylcholine binding(GO:0042166) |
| 0.0 | 5.9 | GO:0001664 | G-protein coupled receptor binding(GO:0001664) |
| 0.0 | 2.0 | GO:0005262 | calcium channel activity(GO:0005262) |
| 0.0 | 3.1 | GO:0060090 | binding, bridging(GO:0060090) |
| 0.0 | 2.5 | GO:0098531 | RNA polymerase II transcription factor activity, ligand-activated sequence-specific DNA binding(GO:0004879) transcription factor activity, direct ligand regulated sequence-specific DNA binding(GO:0098531) |
| 0.0 | 1.4 | GO:0004869 | cysteine-type endopeptidase inhibitor activity(GO:0004869) |
| 0.0 | 2.2 | GO:0008201 | heparin binding(GO:0008201) |
| 0.0 | 0.1 | GO:0030292 | protein tyrosine kinase inhibitor activity(GO:0030292) |
| 0.0 | 1.3 | GO:0004518 | nuclease activity(GO:0004518) |
| 0.0 | 0.2 | GO:0033038 | bitter taste receptor activity(GO:0033038) |
| 0.0 | 2.7 | GO:0061659 | ubiquitin protein ligase activity(GO:0061630) ubiquitin-like protein ligase activity(GO:0061659) |
| 0.0 | 0.2 | GO:0015377 | cation:chloride symporter activity(GO:0015377) |
Gene overrepresentation in C2:CP category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 1.0 | 55.8 | NABA PROTEOGLYCANS | Genes encoding proteoglycans |
| 0.5 | 2.6 | PID SMAD2 3PATHWAY | Regulation of cytoplasmic and nuclear SMAD2/3 signaling |
| 0.3 | 3.1 | PID FRA PATHWAY | Validated transcriptional targets of AP1 family members Fra1 and Fra2 |
| 0.3 | 7.0 | PID TCR JNK PATHWAY | JNK signaling in the CD4+ TCR pathway |
| 0.3 | 8.7 | ST G ALPHA S PATHWAY | G alpha s Pathway |
| 0.3 | 12.7 | ST DIFFERENTIATION PATHWAY IN PC12 CELLS | Differentiation Pathway in PC12 Cells; this is a specific case of PAC1 Receptor Pathway. |
| 0.3 | 12.8 | PID HIF2PATHWAY | HIF-2-alpha transcription factor network |
| 0.3 | 8.6 | SA CASPASE CASCADE | Apoptosis is mediated by caspases, cysteine proteases arranged in a proteolytic cascade. |
| 0.3 | 10.1 | PID INTEGRIN A9B1 PATHWAY | Alpha9 beta1 integrin signaling events |
| 0.2 | 2.4 | PID P38 MK2 PATHWAY | p38 signaling mediated by MAPKAP kinases |
| 0.2 | 14.5 | PID AURORA B PATHWAY | Aurora B signaling |
| 0.2 | 50.7 | NABA ECM GLYCOPROTEINS | Genes encoding structural ECM glycoproteins |
| 0.2 | 10.0 | PID NCADHERIN PATHWAY | N-cadherin signaling events |
| 0.2 | 7.8 | PID UPA UPAR PATHWAY | Urokinase-type plasminogen activator (uPA) and uPAR-mediated signaling |
| 0.2 | 4.4 | SA REG CASCADE OF CYCLIN EXPR | Expression of cyclins regulates progression through the cell cycle by activating cyclin-dependent kinases. |
| 0.2 | 2.3 | SIG IL4RECEPTOR IN B LYPHOCYTES | Genes related to IL4 rceptor signaling in B lymphocytes |
| 0.1 | 3.5 | PID EPHA2 FWD PATHWAY | EPHA2 forward signaling |
| 0.1 | 1.1 | PID KIT PATHWAY | Signaling events mediated by Stem cell factor receptor (c-Kit) |
| 0.1 | 9.3 | PID TRKR PATHWAY | Neurotrophic factor-mediated Trk receptor signaling |
| 0.1 | 9.0 | PID DELTA NP63 PATHWAY | Validated transcriptional targets of deltaNp63 isoforms |
| 0.1 | 4.9 | ST GA13 PATHWAY | G alpha 13 Pathway |
| 0.1 | 5.1 | PID CXCR3 PATHWAY | CXCR3-mediated signaling events |
| 0.1 | 24.0 | NABA ECM REGULATORS | Genes encoding enzymes and their regulators involved in the remodeling of the extracellular matrix |
| 0.1 | 7.4 | ST INTEGRIN SIGNALING PATHWAY | Integrin Signaling Pathway |
| 0.1 | 3.6 | PID ARF6 PATHWAY | Arf6 signaling events |
| 0.1 | 2.9 | PID ECADHERIN STABILIZATION PATHWAY | Stabilization and expansion of the E-cadherin adherens junction |
| 0.1 | 3.3 | NABA COLLAGENS | Genes encoding collagen proteins |
| 0.1 | 1.7 | PID CONE PATHWAY | Visual signal transduction: Cones |
| 0.1 | 24.4 | NABA SECRETED FACTORS | Genes encoding secreted soluble factors |
| 0.1 | 2.7 | PID BMP PATHWAY | BMP receptor signaling |
| 0.1 | 1.5 | PID INTEGRIN2 PATHWAY | Beta2 integrin cell surface interactions |
| 0.1 | 0.7 | PID IL5 PATHWAY | IL5-mediated signaling events |
| 0.1 | 2.5 | PID FCER1 PATHWAY | Fc-epsilon receptor I signaling in mast cells |
| 0.1 | 3.3 | PID HNF3A PATHWAY | FOXA1 transcription factor network |
| 0.0 | 2.2 | PID TXA2PATHWAY | Thromboxane A2 receptor signaling |
| 0.0 | 3.0 | PID HIF1 TFPATHWAY | HIF-1-alpha transcription factor network |
| 0.0 | 0.6 | PID NETRIN PATHWAY | Netrin-mediated signaling events |
| 0.0 | 1.0 | PID ERA GENOMIC PATHWAY | Validated nuclear estrogen receptor alpha network |
| 0.0 | 0.6 | PID RXR VDR PATHWAY | RXR and RAR heterodimerization with other nuclear receptor |
| 0.0 | 0.2 | PID EPHRINB REV PATHWAY | Ephrin B reverse signaling |
| 0.0 | 0.7 | PID RHOA REG PATHWAY | Regulation of RhoA activity |
Gene overrepresentation in C2:CP:REACTOME category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 1.5 | 41.1 | REACTOME CS DS DEGRADATION | Genes involved in CS/DS degradation |
| 1.5 | 25.4 | REACTOME PLATELET ADHESION TO EXPOSED COLLAGEN | Genes involved in Platelet Adhesion to exposed collagen |
| 0.7 | 12.8 | REACTOME DOPAMINE NEUROTRANSMITTER RELEASE CYCLE | Genes involved in Dopamine Neurotransmitter Release Cycle |
| 0.7 | 7.0 | REACTOME IRAK2 MEDIATED ACTIVATION OF TAK1 COMPLEX UPON TLR7 8 OR 9 STIMULATION | Genes involved in IRAK2 mediated activation of TAK1 complex upon TLR7/8 or 9 stimulation |
| 0.5 | 0.5 | REACTOME SHC1 EVENTS IN EGFR SIGNALING | Genes involved in SHC1 events in EGFR signaling |
| 0.5 | 7.8 | REACTOME JNK C JUN KINASES PHOSPHORYLATION AND ACTIVATION MEDIATED BY ACTIVATED HUMAN TAK1 | Genes involved in JNK (c-Jun kinases) phosphorylation and activation mediated by activated human TAK1 |
| 0.5 | 15.2 | REACTOME ADHERENS JUNCTIONS INTERACTIONS | Genes involved in Adherens junctions interactions |
| 0.4 | 13.7 | REACTOME KERATAN SULFATE BIOSYNTHESIS | Genes involved in Keratan sulfate biosynthesis |
| 0.4 | 5.7 | REACTOME NFKB ACTIVATION THROUGH FADD RIP1 PATHWAY MEDIATED BY CASPASE 8 AND10 | Genes involved in NF-kB activation through FADD/RIP-1 pathway mediated by caspase-8 and -10 |
| 0.4 | 9.4 | REACTOME REGULATION OF INSULIN LIKE GROWTH FACTOR IGF ACTIVITY BY INSULIN LIKE GROWTH FACTOR BINDING PROTEINS IGFBPS | Genes involved in Regulation of Insulin-like Growth Factor (IGF) Activity by Insulin-like Growth Factor Binding Proteins (IGFBPs) |
| 0.4 | 23.0 | REACTOME AMINO ACID AND OLIGOPEPTIDE SLC TRANSPORTERS | Genes involved in Amino acid and oligopeptide SLC transporters |
| 0.3 | 3.3 | REACTOME DSCAM INTERACTIONS | Genes involved in DSCAM interactions |
| 0.3 | 3.5 | REACTOME PROSTANOID LIGAND RECEPTORS | Genes involved in Prostanoid ligand receptors |
| 0.3 | 3.7 | REACTOME ABACAVIR TRANSPORT AND METABOLISM | Genes involved in Abacavir transport and metabolism |
| 0.3 | 4.2 | REACTOME REGULATION OF KIT SIGNALING | Genes involved in Regulation of KIT signaling |
| 0.2 | 4.9 | REACTOME SYNTHESIS OF PIPS AT THE GOLGI MEMBRANE | Genes involved in Synthesis of PIPs at the Golgi membrane |
| 0.2 | 10.7 | REACTOME MYOGENESIS | Genes involved in Myogenesis |
| 0.2 | 13.8 | REACTOME NCAM1 INTERACTIONS | Genes involved in NCAM1 interactions |
| 0.2 | 2.8 | REACTOME SEROTONIN RECEPTORS | Genes involved in Serotonin receptors |
| 0.2 | 8.2 | REACTOME TIGHT JUNCTION INTERACTIONS | Genes involved in Tight junction interactions |
| 0.2 | 4.7 | REACTOME OTHER SEMAPHORIN INTERACTIONS | Genes involved in Other semaphorin interactions |
| 0.2 | 2.3 | REACTOME SIGNAL ATTENUATION | Genes involved in Signal attenuation |
| 0.2 | 7.3 | REACTOME YAP1 AND WWTR1 TAZ STIMULATED GENE EXPRESSION | Genes involved in YAP1- and WWTR1 (TAZ)-stimulated gene expression |
| 0.2 | 2.8 | REACTOME GLYCOPROTEIN HORMONES | Genes involved in Glycoprotein hormones |
| 0.2 | 9.8 | REACTOME SMAD2 SMAD3 SMAD4 HETEROTRIMER REGULATES TRANSCRIPTION | Genes involved in SMAD2/SMAD3:SMAD4 heterotrimer regulates transcription |
| 0.2 | 10.1 | REACTOME OLFACTORY SIGNALING PATHWAY | Genes involved in Olfactory Signaling Pathway |
| 0.2 | 5.5 | REACTOME REGULATION OF IFNA SIGNALING | Genes involved in Regulation of IFNA signaling |
| 0.2 | 4.1 | REACTOME ADP SIGNALLING THROUGH P2RY1 | Genes involved in ADP signalling through P2Y purinoceptor 1 |
| 0.2 | 4.0 | REACTOME SYNTHESIS SECRETION AND INACTIVATION OF GLP1 | Genes involved in Synthesis, Secretion, and Inactivation of Glucagon-like Peptide-1 (GLP-1) |
| 0.2 | 23.3 | REACTOME INTEGRIN CELL SURFACE INTERACTIONS | Genes involved in Integrin cell surface interactions |
| 0.2 | 2.0 | REACTOME REGULATION OF WATER BALANCE BY RENAL AQUAPORINS | Genes involved in Regulation of Water Balance by Renal Aquaporins |
| 0.2 | 4.1 | REACTOME RNA POL III CHAIN ELONGATION | Genes involved in RNA Polymerase III Chain Elongation |
| 0.2 | 5.3 | REACTOME CASPASE MEDIATED CLEAVAGE OF CYTOSKELETAL PROTEINS | Genes involved in Caspase-mediated cleavage of cytoskeletal proteins |
| 0.2 | 7.2 | REACTOME COLLAGEN FORMATION | Genes involved in Collagen formation |
| 0.2 | 2.0 | REACTOME SHC1 EVENTS IN ERBB4 SIGNALING | Genes involved in SHC1 events in ERBB4 signaling |
| 0.2 | 3.0 | REACTOME CHYLOMICRON MEDIATED LIPID TRANSPORT | Genes involved in Chylomicron-mediated lipid transport |
| 0.2 | 4.5 | REACTOME AMINE LIGAND BINDING RECEPTORS | Genes involved in Amine ligand-binding receptors |
| 0.1 | 3.0 | REACTOME IL 7 SIGNALING | Genes involved in Interleukin-7 signaling |
| 0.1 | 4.9 | REACTOME INTERACTION BETWEEN L1 AND ANKYRINS | Genes involved in Interaction between L1 and Ankyrins |
| 0.1 | 8.9 | REACTOME SMOOTH MUSCLE CONTRACTION | Genes involved in Smooth Muscle Contraction |
| 0.1 | 3.5 | REACTOME PROLACTIN RECEPTOR SIGNALING | Genes involved in Prolactin receptor signaling |
| 0.1 | 5.5 | REACTOME SIGNALING BY ROBO RECEPTOR | Genes involved in Signaling by Robo receptor |
| 0.1 | 1.4 | REACTOME THE ACTIVATION OF ARYLSULFATASES | Genes involved in The activation of arylsulfatases |
| 0.1 | 21.6 | REACTOME G ALPHA Q SIGNALLING EVENTS | Genes involved in G alpha (q) signalling events |
| 0.1 | 14.0 | REACTOME G ALPHA S SIGNALLING EVENTS | Genes involved in G alpha (s) signalling events |
| 0.1 | 2.2 | REACTOME REGULATION OF GENE EXPRESSION IN BETA CELLS | Genes involved in Regulation of gene expression in beta cells |
| 0.1 | 3.4 | REACTOME GASTRIN CREB SIGNALLING PATHWAY VIA PKC AND MAPK | Genes involved in Gastrin-CREB signalling pathway via PKC and MAPK |
| 0.1 | 7.8 | REACTOME INTEGRATION OF ENERGY METABOLISM | Genes involved in Integration of energy metabolism |
| 0.1 | 2.1 | REACTOME FORMATION OF ATP BY CHEMIOSMOTIC COUPLING | Genes involved in Formation of ATP by chemiosmotic coupling |
| 0.1 | 1.0 | REACTOME CREATION OF C4 AND C2 ACTIVATORS | Genes involved in Creation of C4 and C2 activators |
| 0.1 | 3.2 | REACTOME SULFUR AMINO ACID METABOLISM | Genes involved in Sulfur amino acid metabolism |
| 0.1 | 2.2 | REACTOME NUCLEAR SIGNALING BY ERBB4 | Genes involved in Nuclear signaling by ERBB4 |
| 0.1 | 9.9 | REACTOME FACTORS INVOLVED IN MEGAKARYOCYTE DEVELOPMENT AND PLATELET PRODUCTION | Genes involved in Factors involved in megakaryocyte development and platelet production |
| 0.1 | 5.3 | REACTOME METABOLISM OF VITAMINS AND COFACTORS | Genes involved in Metabolism of vitamins and cofactors |
| 0.1 | 1.3 | REACTOME GAP JUNCTION ASSEMBLY | Genes involved in Gap junction assembly |
| 0.1 | 2.7 | REACTOME SIGNALING BY BMP | Genes involved in Signaling by BMP |
| 0.1 | 1.1 | REACTOME APOPTOSIS INDUCED DNA FRAGMENTATION | Genes involved in Apoptosis induced DNA fragmentation |
| 0.1 | 2.2 | REACTOME VOLTAGE GATED POTASSIUM CHANNELS | Genes involved in Voltage gated Potassium channels |
| 0.1 | 6.9 | REACTOME G ALPHA I SIGNALLING EVENTS | Genes involved in G alpha (i) signalling events |
| 0.0 | 0.7 | REACTOME REVERSIBLE HYDRATION OF CARBON DIOXIDE | Genes involved in Reversible Hydration of Carbon Dioxide |
| 0.0 | 1.4 | REACTOME GABA A RECEPTOR ACTIVATION | Genes involved in GABA A receptor activation |
| 0.0 | 0.8 | REACTOME PRESYNAPTIC NICOTINIC ACETYLCHOLINE RECEPTORS | Genes involved in Presynaptic nicotinic acetylcholine receptors |
| 0.0 | 0.6 | REACTOME HDL MEDIATED LIPID TRANSPORT | Genes involved in HDL-mediated lipid transport |
| 0.0 | 2.2 | REACTOME TRANSPORT OF INORGANIC CATIONS ANIONS AND AMINO ACIDS OLIGOPEPTIDES | Genes involved in Transport of inorganic cations/anions and amino acids/oligopeptides |
| 0.0 | 0.7 | REACTOME IL RECEPTOR SHC SIGNALING | Genes involved in Interleukin receptor SHC signaling |
| 0.0 | 0.4 | REACTOME REGULATION OF BETA CELL DEVELOPMENT | Genes involved in Regulation of beta-cell development |
| 0.0 | 0.4 | REACTOME SEMA3A PLEXIN REPULSION SIGNALING BY INHIBITING INTEGRIN ADHESION | Genes involved in SEMA3A-Plexin repulsion signaling by inhibiting Integrin adhesion |
| 0.0 | 1.6 | REACTOME SIGNALING BY ERBB2 | Genes involved in Signaling by ERBB2 |
| 0.0 | 2.3 | REACTOME LOSS OF NLP FROM MITOTIC CENTROSOMES | Genes involved in Loss of Nlp from mitotic centrosomes |
| 0.0 | 1.2 | REACTOME GLUCOSE TRANSPORT | Genes involved in Glucose transport |
| 0.0 | 0.6 | REACTOME NETRIN1 SIGNALING | Genes involved in Netrin-1 signaling |
| 0.0 | 0.5 | REACTOME CGMP EFFECTS | Genes involved in cGMP effects |
| 0.0 | 2.0 | REACTOME PHASE II CONJUGATION | Genes involved in Phase II conjugation |
| 0.0 | 1.1 | REACTOME ION TRANSPORT BY P TYPE ATPASES | Genes involved in Ion transport by P-type ATPases |
| 0.0 | 0.7 | REACTOME NRAGE SIGNALS DEATH THROUGH JNK | Genes involved in NRAGE signals death through JNK |
| 0.0 | 0.9 | REACTOME CYTOCHROME P450 ARRANGED BY SUBSTRATE TYPE | Genes involved in Cytochrome P450 - arranged by substrate type |