Project
GNF SymAtlas + NCI-60 cancer cell lines, comparison of cancers vs non-cancers, human (Su, 2004; Ross, 2000)
Navigation
Downloads
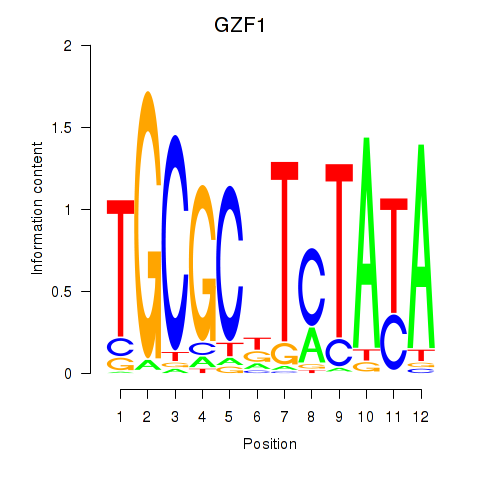
Results for GZF1
Z-value: 0.47
Transcription factors associated with GZF1
| Gene Symbol | Gene ID | Gene Info |
|---|---|---|
|
GZF1
|
ENSG00000125812.11 | GDNF inducible zinc finger protein 1 |
Activity profile of GZF1 motif
Sorted Z-values of GZF1 motif
| Promoter | Log-likelihood | Transcript | Gene | Gene Info |
|---|---|---|---|---|
| chr1_+_26606608 | 14.42 |
ENST00000319041.6
|
SH3BGRL3
|
SH3 domain binding glutamic acid-rich protein like 3 |
| chr5_+_170814803 | 11.23 |
ENST00000521672.1
ENST00000351986.6 ENST00000393820.2 ENST00000523622.1 |
NPM1
|
nucleophosmin (nucleolar phosphoprotein B23, numatrin) |
| chr2_-_150444300 | 10.06 |
ENST00000303319.5
|
MMADHC
|
methylmalonic aciduria (cobalamin deficiency) cblD type, with homocystinuria |
| chr2_-_150444116 | 9.16 |
ENST00000428879.1
ENST00000422782.2 |
MMADHC
|
methylmalonic aciduria (cobalamin deficiency) cblD type, with homocystinuria |
| chr12_-_76478386 | 8.25 |
ENST00000535020.2
|
NAP1L1
|
nucleosome assembly protein 1-like 1 |
| chr12_-_76478417 | 7.16 |
ENST00000552342.1
|
NAP1L1
|
nucleosome assembly protein 1-like 1 |
| chr1_+_32757668 | 6.05 |
ENST00000373548.3
|
HDAC1
|
histone deacetylase 1 |
| chr3_-_79816965 | 5.52 |
ENST00000464233.1
|
ROBO1
|
roundabout, axon guidance receptor, homolog 1 (Drosophila) |
| chr22_-_43355858 | 5.30 |
ENST00000402229.1
ENST00000407585.1 ENST00000453079.1 |
PACSIN2
|
protein kinase C and casein kinase substrate in neurons 2 |
| chr12_-_76478446 | 5.19 |
ENST00000393263.3
ENST00000548044.1 ENST00000547704.1 ENST00000431879.3 ENST00000549596.1 ENST00000550934.1 ENST00000551600.1 ENST00000547479.1 ENST00000547773.1 ENST00000544816.1 ENST00000542344.1 ENST00000548273.1 |
NAP1L1
|
nucleosome assembly protein 1-like 1 |
| chr3_+_30647994 | 5.11 |
ENST00000295754.5
|
TGFBR2
|
transforming growth factor, beta receptor II (70/80kDa) |
| chr12_-_102513843 | 4.80 |
ENST00000551744.2
ENST00000552283.1 |
NUP37
|
nucleoporin 37kDa |
| chr14_-_21979428 | 4.65 |
ENST00000538267.1
ENST00000298717.4 |
METTL3
|
methyltransferase like 3 |
| chr3_+_30648066 | 4.59 |
ENST00000359013.4
|
TGFBR2
|
transforming growth factor, beta receptor II (70/80kDa) |
| chr19_-_14628645 | 4.26 |
ENST00000598235.1
|
DNAJB1
|
DnaJ (Hsp40) homolog, subfamily B, member 1 |
| chr1_+_79115503 | 4.20 |
ENST00000370747.4
ENST00000438486.1 ENST00000545124.1 |
IFI44
|
interferon-induced protein 44 |
| chr1_-_32801825 | 3.60 |
ENST00000329421.7
|
MARCKSL1
|
MARCKS-like 1 |
| chr1_-_226374373 | 3.42 |
ENST00000366812.5
|
ACBD3
|
acyl-CoA binding domain containing 3 |
| chr9_+_96338647 | 3.16 |
ENST00000359246.4
|
PHF2
|
PHD finger protein 2 |
| chrX_+_54835493 | 2.91 |
ENST00000396224.1
|
MAGED2
|
melanoma antigen family D, 2 |
| chr9_+_96338860 | 2.61 |
ENST00000375376.4
|
PHF2
|
PHD finger protein 2 |
| chr16_+_67876180 | 2.58 |
ENST00000303596.1
|
THAP11
|
THAP domain containing 11 |
| chr9_+_131133598 | 2.34 |
ENST00000372853.4
ENST00000452446.1 ENST00000372850.1 ENST00000372847.1 |
URM1
|
ubiquitin related modifier 1 |
| chr1_+_9711781 | 2.29 |
ENST00000536656.1
ENST00000377346.4 |
PIK3CD
|
phosphatidylinositol-4,5-bisphosphate 3-kinase, catalytic subunit delta |
| chr20_+_42086525 | 2.23 |
ENST00000244020.3
|
SRSF6
|
serine/arginine-rich splicing factor 6 |
| chr10_+_89264625 | 2.12 |
ENST00000371996.4
ENST00000371994.4 |
MINPP1
|
multiple inositol-polyphosphate phosphatase 1 |
| chr5_+_92919043 | 1.87 |
ENST00000327111.3
|
NR2F1
|
nuclear receptor subfamily 2, group F, member 1 |
| chr10_+_60028818 | 1.84 |
ENST00000333926.5
|
CISD1
|
CDGSH iron sulfur domain 1 |
| chr19_-_59084647 | 1.72 |
ENST00000594234.1
ENST00000596039.1 |
MZF1
|
myeloid zinc finger 1 |
| chr1_+_41174988 | 1.72 |
ENST00000372652.1
|
NFYC
|
nuclear transcription factor Y, gamma |
| chr17_-_29648761 | 1.68 |
ENST00000247270.3
ENST00000462804.2 |
EVI2A
|
ecotropic viral integration site 2A |
| chr10_-_126849588 | 1.67 |
ENST00000411419.2
|
CTBP2
|
C-terminal binding protein 2 |
| chr19_-_59084922 | 1.48 |
ENST00000215057.2
ENST00000599369.1 |
MZF1
|
myeloid zinc finger 1 |
| chr7_-_26904317 | 1.43 |
ENST00000345317.2
|
SKAP2
|
src kinase associated phosphoprotein 2 |
| chr13_+_98086445 | 1.43 |
ENST00000245304.4
|
RAP2A
|
RAP2A, member of RAS oncogene family |
| chrX_+_102883620 | 1.41 |
ENST00000372626.3
|
TCEAL1
|
transcription elongation factor A (SII)-like 1 |
| chr18_-_53303123 | 1.37 |
ENST00000569357.1
ENST00000565124.1 ENST00000398339.1 |
TCF4
|
transcription factor 4 |
| chr3_+_51705222 | 1.22 |
ENST00000457573.1
ENST00000341333.5 ENST00000412249.1 ENST00000425781.1 ENST00000415259.1 ENST00000395057.1 ENST00000416589.1 |
TEX264
|
testis expressed 264 |
| chr19_-_3786253 | 0.85 |
ENST00000585778.1
|
MATK
|
megakaryocyte-associated tyrosine kinase |
| chr10_-_126849068 | 0.63 |
ENST00000494626.2
ENST00000337195.5 |
CTBP2
|
C-terminal binding protein 2 |
| chr3_-_114035026 | 0.54 |
ENST00000570269.1
|
RP11-553L6.5
|
RP11-553L6.5 |
| chr19_-_3786354 | 0.49 |
ENST00000395040.2
ENST00000310132.6 |
MATK
|
megakaryocyte-associated tyrosine kinase |
| chr14_-_92413353 | 0.45 |
ENST00000556154.1
|
FBLN5
|
fibulin 5 |
| chr1_-_42384343 | 0.39 |
ENST00000372584.1
|
HIVEP3
|
human immunodeficiency virus type I enhancer binding protein 3 |
| chr8_+_23430157 | 0.26 |
ENST00000399967.3
|
FP15737
|
FP15737 |
Network of associatons between targets according to the STRING database.
First level regulatory network of GZF1
Gene Ontology Analysis
Gene overrepresentation in biological_process category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 3.2 | 9.7 | GO:0002661 | B cell tolerance induction(GO:0002514) regulation of B cell tolerance induction(GO:0002661) positive regulation of B cell tolerance induction(GO:0002663) |
| 1.9 | 5.8 | GO:0061188 | negative regulation of chromatin silencing at rDNA(GO:0061188) |
| 1.8 | 5.5 | GO:0021836 | cerebral cortex tangential migration using cell-cell interactions(GO:0021823) postnatal olfactory bulb interneuron migration(GO:0021827) chemorepulsion involved in postnatal olfactory bulb interneuron migration(GO:0021836) regulation of negative chemotaxis(GO:0050923) |
| 1.4 | 11.2 | GO:0060699 | regulation of endoribonuclease activity(GO:0060699) |
| 1.2 | 6.0 | GO:0061198 | fungiform papilla formation(GO:0061198) |
| 1.2 | 4.6 | GO:0006382 | adenosine to inosine editing(GO:0006382) |
| 0.8 | 2.3 | GO:0034227 | tRNA thio-modification(GO:0034227) |
| 0.7 | 19.2 | GO:0009235 | cobalamin metabolic process(GO:0009235) |
| 0.5 | 5.3 | GO:0070836 | caveola assembly(GO:0070836) |
| 0.5 | 2.3 | GO:0060374 | positive regulation of neutrophil apoptotic process(GO:0033031) mast cell differentiation(GO:0060374) |
| 0.4 | 2.9 | GO:0070294 | renal sodium ion absorption(GO:0070294) |
| 0.3 | 4.3 | GO:0070389 | chaperone cofactor-dependent protein refolding(GO:0070389) |
| 0.3 | 2.2 | GO:0060501 | positive regulation of epithelial cell proliferation involved in lung morphogenesis(GO:0060501) |
| 0.3 | 2.3 | GO:0048386 | positive regulation of retinoic acid receptor signaling pathway(GO:0048386) |
| 0.2 | 14.4 | GO:0030834 | regulation of actin filament depolymerization(GO:0030834) |
| 0.1 | 20.6 | GO:0006334 | nucleosome assembly(GO:0006334) |
| 0.1 | 1.4 | GO:0045198 | establishment of epithelial cell apical/basal polarity(GO:0045198) |
| 0.1 | 4.1 | GO:0071431 | tRNA export from nucleus(GO:0006409) tRNA-containing ribonucleoprotein complex export from nucleus(GO:0071431) |
| 0.1 | 1.8 | GO:0043457 | regulation of cellular respiration(GO:0043457) |
| 0.0 | 1.7 | GO:0045540 | regulation of cholesterol biosynthetic process(GO:0045540) |
| 0.0 | 2.1 | GO:0043647 | inositol phosphate metabolic process(GO:0043647) |
| 0.0 | 3.4 | GO:0006694 | steroid biosynthetic process(GO:0006694) |
| 0.0 | 1.3 | GO:0038128 | ERBB2 signaling pathway(GO:0038128) |
| 0.0 | 3.2 | GO:0006367 | transcription initiation from RNA polymerase II promoter(GO:0006367) |
Gene overrepresentation in cellular_component category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 1.5 | 4.6 | GO:0045293 | MIS complex(GO:0036396) mRNA editing complex(GO:0045293) |
| 1.1 | 9.7 | GO:0070022 | transforming growth factor beta receptor homodimeric complex(GO:0070022) |
| 0.7 | 11.2 | GO:0031616 | spindle pole centrosome(GO:0031616) |
| 0.3 | 4.8 | GO:0031080 | nuclear pore outer ring(GO:0031080) |
| 0.3 | 6.0 | GO:0016580 | Sin3 complex(GO:0016580) |
| 0.2 | 1.7 | GO:0016602 | CCAAT-binding factor complex(GO:0016602) |
| 0.2 | 2.3 | GO:0097470 | ribbon synapse(GO:0097470) |
| 0.1 | 20.6 | GO:0048770 | melanosome(GO:0042470) pigment granule(GO:0048770) |
| 0.1 | 6.7 | GO:0055038 | recycling endosome membrane(GO:0055038) |
| 0.1 | 2.3 | GO:0005942 | phosphatidylinositol 3-kinase complex(GO:0005942) |
| 0.1 | 5.8 | GO:0000777 | condensed chromosome kinetochore(GO:0000777) |
| 0.1 | 2.6 | GO:0045171 | intercellular bridge(GO:0045171) |
| 0.0 | 14.4 | GO:0030027 | lamellipodium(GO:0030027) |
| 0.0 | 4.1 | GO:0031093 | platelet alpha granule lumen(GO:0031093) |
| 0.0 | 21.3 | GO:0005739 | mitochondrion(GO:0005739) |
| 0.0 | 4.7 | GO:0030424 | axon(GO:0030424) |
Gene overrepresentation in molecular_function category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 2.4 | 9.7 | GO:0005026 | transforming growth factor beta receptor activity, type II(GO:0005026) |
| 2.2 | 11.2 | GO:0043023 | ribosomal large subunit binding(GO:0043023) |
| 1.5 | 4.6 | GO:0016422 | mRNA (2'-O-methyladenosine-N6-)-methyltransferase activity(GO:0016422) |
| 1.0 | 14.4 | GO:0030215 | semaphorin receptor binding(GO:0030215) |
| 0.9 | 5.5 | GO:0008046 | axon guidance receptor activity(GO:0008046) |
| 0.5 | 2.1 | GO:0052827 | inositol pentakisphosphate phosphatase activity(GO:0052827) |
| 0.4 | 6.0 | GO:0032041 | histone deacetylase activity (H3-K14 specific)(GO:0031078) NAD-dependent histone deacetylase activity (H3-K14 specific)(GO:0032041) |
| 0.3 | 2.3 | GO:0030267 | hydroxypyruvate reductase activity(GO:0016618) glyoxylate reductase (NADP) activity(GO:0030267) |
| 0.3 | 2.3 | GO:0035005 | 1-phosphatidylinositol-4-phosphate 3-kinase activity(GO:0035005) |
| 0.3 | 5.8 | GO:0032454 | histone demethylase activity (H3-K9 specific)(GO:0032454) |
| 0.2 | 5.3 | GO:0070300 | phosphatidic acid binding(GO:0070300) |
| 0.2 | 1.9 | GO:0044323 | retinoic acid-responsive element binding(GO:0044323) |
| 0.1 | 2.3 | GO:0016783 | sulfurtransferase activity(GO:0016783) |
| 0.1 | 3.4 | GO:0034237 | protein kinase A regulatory subunit binding(GO:0034237) |
| 0.1 | 4.3 | GO:0001671 | ATPase activator activity(GO:0001671) |
| 0.1 | 1.4 | GO:0001087 | transcription factor activity, sequence-specific DNA binding, RNA polymerase recruiting(GO:0001011) transcription factor activity, TFIIB-class binding(GO:0001087) |
| 0.1 | 1.8 | GO:0051537 | 2 iron, 2 sulfur cluster binding(GO:0051537) |
| 0.0 | 2.2 | GO:0036002 | pre-mRNA binding(GO:0036002) |
| 0.0 | 5.8 | GO:0001078 | transcriptional repressor activity, RNA polymerase II core promoter proximal region sequence-specific binding(GO:0001078) |
| 0.0 | 1.4 | GO:0050699 | WW domain binding(GO:0050699) |
| 0.0 | 1.4 | GO:0019003 | GDP binding(GO:0019003) |
| 0.0 | 1.7 | GO:0000980 | RNA polymerase II distal enhancer sequence-specific DNA binding(GO:0000980) |
| 0.0 | 1.3 | GO:0004715 | non-membrane spanning protein tyrosine kinase activity(GO:0004715) |
| 0.0 | 1.4 | GO:0005070 | SH3/SH2 adaptor activity(GO:0005070) |
| 0.0 | 3.6 | GO:0005516 | calmodulin binding(GO:0005516) |
Gene overrepresentation in C2:CP category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.4 | 6.0 | PID RANBP2 PATHWAY | Sumoylation by RanBP2 regulates transcriptional repression |
| 0.2 | 11.2 | PID BARD1 PATHWAY | BARD1 signaling events |
| 0.2 | 9.7 | PID ALK1 PATHWAY | ALK1 signaling events |
| 0.1 | 2.3 | SA TRKA RECEPTOR | The TrkA receptor binds nerve growth factor to activate MAP kinase pathways and promote cell growth. |
| 0.1 | 5.5 | PID AJDISS 2PATHWAY | Posttranslational regulation of adherens junction stability and dissassembly |
| 0.0 | 1.3 | PID KIT PATHWAY | Signaling events mediated by Stem cell factor receptor (c-Kit) |
| 0.0 | 1.8 | WNT SIGNALING | Genes related to Wnt-mediated signal transduction |
| 0.0 | 1.4 | PID BETA CATENIN NUC PATHWAY | Regulation of nuclear beta catenin signaling and target gene transcription |
| 0.0 | 1.7 | PID MYC REPRESS PATHWAY | Validated targets of C-MYC transcriptional repression |
Gene overrepresentation in C2:CP:REACTOME category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.4 | 9.7 | REACTOME TGF BETA RECEPTOR SIGNALING IN EMT EPITHELIAL TO MESENCHYMAL TRANSITION | Genes involved in TGF-beta receptor signaling in EMT (epithelial to mesenchymal transition) |
| 0.2 | 5.5 | REACTOME ACTIVATION OF RAC | Genes involved in Activation of Rac |
| 0.2 | 6.0 | REACTOME G0 AND EARLY G1 | Genes involved in G0 and Early G1 |
| 0.2 | 11.2 | REACTOME DEPOSITION OF NEW CENPA CONTAINING NUCLEOSOMES AT THE CENTROMERE | Genes involved in Deposition of New CENPA-containing Nucleosomes at the Centromere |
| 0.1 | 4.8 | REACTOME NEP NS2 INTERACTS WITH THE CELLULAR EXPORT MACHINERY | Genes involved in NEP/NS2 Interacts with the Cellular Export Machinery |
| 0.1 | 2.3 | REACTOME REGULATION OF SIGNALING BY CBL | Genes involved in Regulation of signaling by CBL |
| 0.1 | 2.2 | REACTOME TRANSPORT OF MATURE TRANSCRIPT TO CYTOPLASM | Genes involved in Transport of Mature Transcript to Cytoplasm |
| 0.0 | 1.4 | REACTOME SIGNAL REGULATORY PROTEIN SIRP FAMILY INTERACTIONS | Genes involved in Signal regulatory protein (SIRP) family interactions |
| 0.0 | 4.6 | REACTOME PROCESSING OF CAPPED INTRON CONTAINING PRE MRNA | Genes involved in Processing of Capped Intron-Containing Pre-mRNA |
| 0.0 | 1.4 | REACTOME MYOGENESIS | Genes involved in Myogenesis |
| 0.0 | 1.9 | REACTOME NUCLEAR RECEPTOR TRANSCRIPTION PATHWAY | Genes involved in Nuclear Receptor transcription pathway |