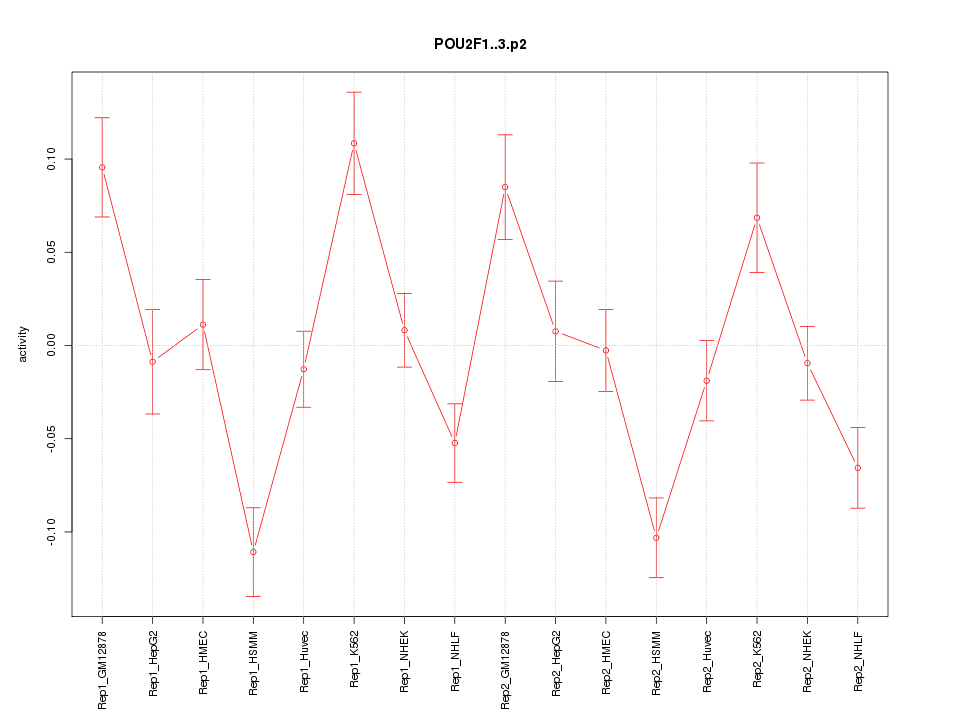
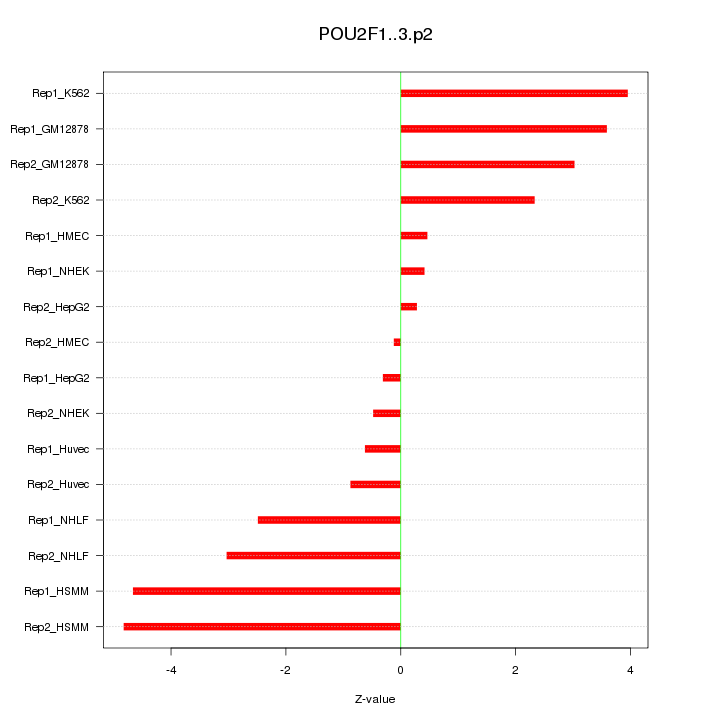
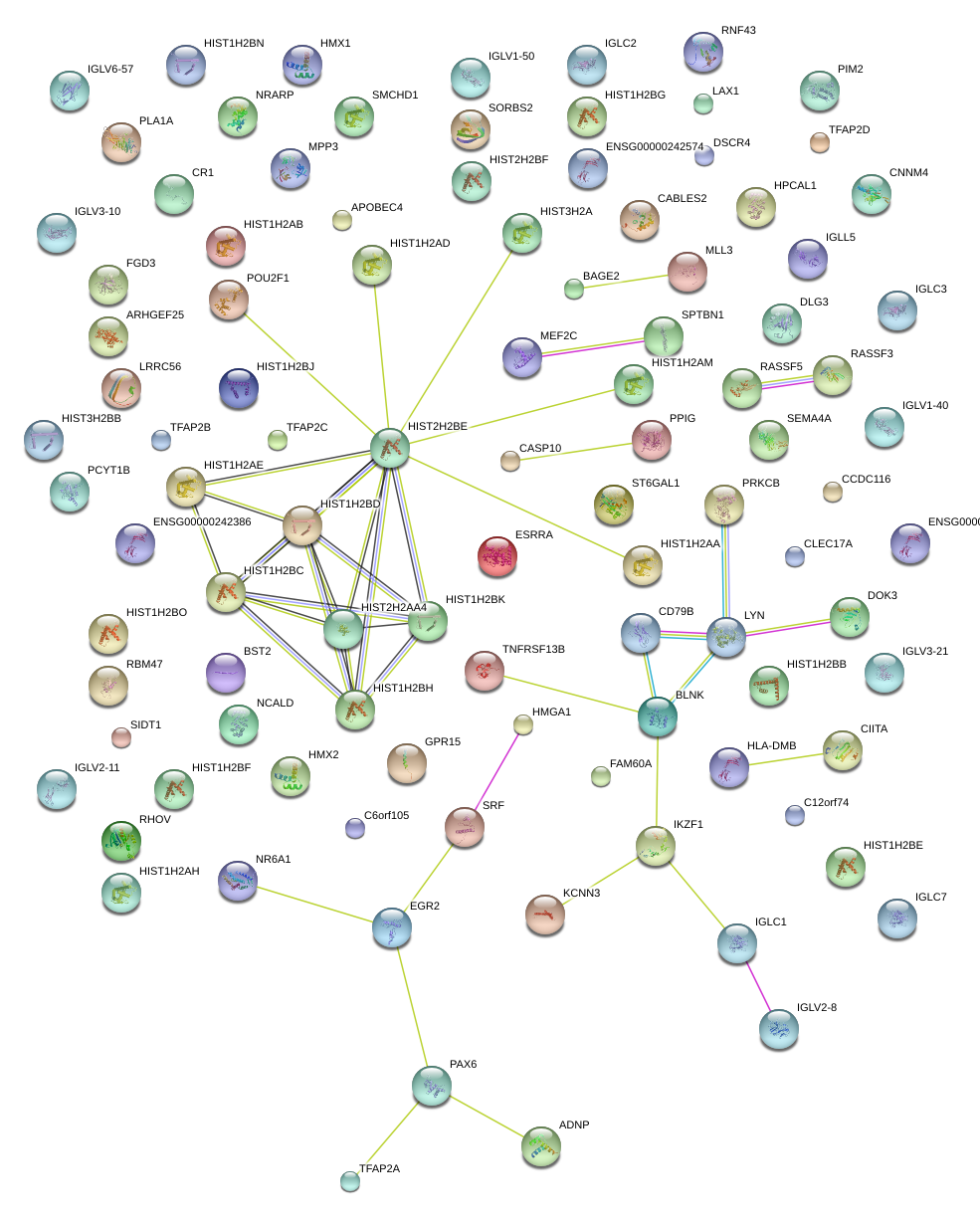
|
chr16_+_23754997
|
7.138
|
|
PRKCB
|
protein kinase C, beta
|
|
chr16_+_23754860
|
6.966
|
|
PRKCB
|
protein kinase C, beta
|
|
chr16_+_23754822
|
6.817
|
|
PRKCB
|
protein kinase C, beta
|
|
chr7_+_50318801
|
6.331
|
|
IKZF1
|
IKAROS family zinc finger 1 (Ikaros)
|
|
chr16_+_23754772
|
6.219
|
NM_002738
NM_212535
|
PRKCB
|
protein kinase C, beta
|
|
chrX_-_48661226
|
4.851
|
NM_006875
|
PIM2
|
pim-2 oncogene
|
|
chr1_+_226712430
|
4.850
|
NM_175055
|
HIST3H2BB
|
histone cluster 3, H2bb
|
|
chr1_-_153098811
|
4.687
|
NM_170782
|
KCNN3
|
potassium intermediate/small conductance calcium-activated channel, subfamily N, member 3
|
|
chr1_-_226712140
|
4.330
|
NM_033445
|
HIST3H2A
|
histone cluster 3, H2a
|
|
chr6_-_33016561
|
4.179
|
NM_002118
|
HLA-DMB
|
major histocompatibility complex, class II, DM beta
|
|
chr1_+_165565006
|
4.021
|
|
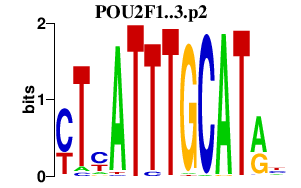
POU2F1
|
POU class 2 homeobox 1
|
|
chr19_-_17377383
|
4.018
|
NM_004335
|
BST2
|
bone marrow stromal cell antigen 2
|
|
chr19_+_17377502
|
4.006
|
|
LOC100507535
|
hypothetical LOC100507535
|
|
chr6_+_26359805
|
3.990
|
NM_003524
|
HIST1H2BH
|
histone cluster 1, H2bh
|
|
chr3_+_99733432
|
3.919
|
NM_005290
|
GPR15
|
G protein-coupled receptor 15
|
|
chr6_-_27968906
|
3.903
|
NM_003514
|
HIST1H2AM
|
histone cluster 1, H2am
|
|
chr6_+_26325124
|
3.898
|
NM_021052
|
HIST1H2AE
|
histone cluster 1, H2ae
|
|
chr1_+_165564904
|
3.793
|
NM_001198783
|
POU2F1
|
POU class 2 homeobox 1
|
|
chr10_-_98021144
|
3.584
|
NM_001114094
NM_013314
|
BLNK
|
B-cell linker
|
|
chr6_+_34312667
|
3.551
|
|
HMGA1
|
high mobility group AT-hook 1
|
|
chr9_-_139316523
|
3.542
|
NM_001004354
|
NRARP
|
NOTCH-regulated ankyrin repeat protein
|
|
chr5_-_88155360
|
3.457
|
NM_001193348
NM_001193349
|
MEF2C
|
myocyte enhancer factor 2C
|
|
chr6_+_34312906
|
3.452
|
|
HMGA1
|
high mobility group AT-hook 1
|
|
chr1_+_204797079
|
3.358
|
NM_182665
|
RASSF5
|
Ras association (RalGDS/AF-6) domain family member 5
|
|
chr15_-_38953628
|
3.308
|
NM_133639
|
RHOV
|
ras homolog gene family, member V
|
|
chr5_-_176868483
|
3.271
|
|
DOK3
|
docking protein 3
|
|
chr6_+_27969181
|
3.261
|
NM_003527
|
HIST1H2BO
|
histone cluster 1, H2bo
|
|
chr22_+_21370394
|
3.190
|
|
IGL@
|
immunoglobulin lambda locus
|
|
chr6_+_34312641
|
3.189
|
|
HMGA1
|
high mobility group AT-hook 1
|
|
chr1_+_205736095
|
3.110
|
NM_000573
NM_000651
|
CR1
|
complement component (3b/4b) receptor 1 (Knops blood group)
|
|
chr17_-_59363352
|
3.098
|
NM_000626
NM_001039933
NM_021602
|
CD79B
|
CD79b molecule, immunoglobulin-associated beta
|
|
chr6_+_34312979
|
3.086
|
|
HMGA1
|
high mobility group AT-hook 1
|
|
chr22_+_20715340
|
2.984
|
|
|
|
|
chr6_+_34312633
|
2.970
|
|
HMGA1
|
high mobility group AT-hook 1
|
|
chr1_+_204797120
|
2.951
|
|
RASSF5
|
Ras association (RalGDS/AF-6) domain family member 5
|
|
chr11_+_63829616
|
2.948
|
NM_004451
|
ESRRA
|
estrogen-related receptor alpha
|
|
chr6_+_34313007
|
2.884
|
|
HMGA1
|
high mobility group AT-hook 1
|
|
chr6_+_34312625
|
2.801
|
NM_145901
NM_145902
|
HMGA1
|
high mobility group AT-hook 1
|
|
chr10_+_124897627
|
2.799
|
NM_005519
|
HMX2
|
H6 family homeobox 2
|
|
chr17_-_53849888
|
2.792
|
|
RNF43
|
ring finger protein 43
|
|
chr6_-_11887265
|
2.752
|
NM_001143948
NM_032744
|
C6orf105
|
chromosome 6 open reading frame 105
|
|
chr8_+_56954946
|
2.711
|
|
LYN
|
v-yes-1 Yamaguchi sarcoma viral related oncogene homolog
|
|
chr6_+_27222839
|
2.709
|
NM_080596
|
HIST1H2AH
|
histone cluster 1, H2ah
|
|
chr8_+_56954899
|
2.676
|
NM_001111097
NM_002350
|
LYN
|
v-yes-1 Yamaguchi sarcoma viral related oncogene homolog
|
|
chr6_+_34312554
|
2.671
|
NM_002131
NM_145899
NM_145903
|
HMGA1
|
high mobility group AT-hook 1
|
|
chrX_-_24575146
|
2.613
|
|
PCYT1B
|
phosphate cytidylyltransferase 1, choline, beta
|
|
chr1_+_202000906
|
2.579
|
NM_001136190
NM_017773
|
LAX1
|
lymphocyte transmembrane adaptor 1
|
|
chr12_+_91620749
|
2.570
|
NM_001037671
NM_001178097
|
C12orf74
|
chromosome 12 open reading frame 74
|
|
chr22_+_20316968
|
2.523
|
NM_152612
|
CCDC116
|
coiled-coil domain containing 116
|
|
chr11_+_527521
|
2.470
|
NM_198075
|
LRRC56
|
leucine rich repeat containing 56
|
|
chr8_+_56954953
|
2.464
|
|
LYN
|
v-yes-1 Yamaguchi sarcoma viral related oncogene homolog
|
|
chr4_-_40212670
|
2.433
|
NM_019027
|
RBM47
|
RNA binding motif protein 47
|
|
chr17_-_53849929
|
2.413
|
NM_017763
|
RNF43
|
ring finger protein 43
|
|
chr6_+_26291936
|
2.321
|
NM_003523
|
HIST1H2BE
|
histone cluster 1, H2be
|
|
chr6_-_27222556
|
2.304
|
NM_080593
|
HIST1H2BK
|
histone cluster 1, H2bk
|
|
chr1_+_154386358
|
2.301
|
NM_001193301
|
SEMA4A
|
sema domain, immunoglobulin domain (Ig), transmembrane domain (TM) and short cytoplasmic domain, (semaphorin) 4A
|
|
chr19_+_14554895
|
2.278
|
NM_207390
|
CLEC17A
|
C-type lectin domain family 17, member A
|
|
chr21_-_10120690
|
2.274
|
NM_001187
NM_182484
NM_181704
NM_182481
NM_182482
|
BAGE
BAGE5
BAGE4
BAGE3
BAGE2
|
B melanoma antigen
B melanoma antigen family, member 5
B melanoma antigen family, member 4
B melanoma antigen family, member 3
B melanoma antigen family, member 2
|
|
chr6_-_26324850
|
2.273
|
NM_003518
|
HIST1H2BG
NCALD
|
histone cluster 1, H2bg
neurocalcin delta
|
|
chr7_+_26404711
|
2.263
|
|
LOC441204
|
hypothetical locus LOC441204
|
|
chr2_+_96790298
|
2.240
|
NM_020184
|
CNNM4
|
cyclin M4
|
|
chr9_+_94766379
|
2.170
|
|
FGD3
|
FYVE, RhoGEF and PH domain containing 3
|
|
chr17_-_16816113
|
2.162
|
NM_012452
|
TNFRSF13B
|
tumor necrosis factor receptor superfamily, member 13B
|
|
chr21_-_38415273
|
2.041
|
NM_005867
|
DSCR4
|
Down syndrome critical region gene 4
|
|
chr6_+_27208763
|
2.027
|
NM_021064
|
HIST1H2AG
|
histone cluster 1, H2ag
|
|
chr2_+_54638957
|
1.997
|
NM_178313
|
SPTBN1
|
spectrin, beta, non-erythrocytic 1
|
|
chr6_-_27208507
|
1.992
|
NM_021058
|
HIST1H2BJ
|
histone cluster 1, H2bj
|
|
chr2_+_201755865
|
1.981
|
NM_001230
NM_032974
NM_032977
|
CASP10
|
caspase 10, apoptosis-related cysteine peptidase
|
|
chr20_-_60415729
|
1.905
|
NM_031215
|
CABLES2
|
Cdk5 and Abl enzyme substrate 2
|
|
chrX_+_69591639
|
1.900
|
NM_001166278
|
DLG3
|
discs, large homolog 3 (Drosophila)
|
|
chr3_+_114733907
|
1.875
|
NM_017699
|
SIDT1
|
SID1 transmembrane family, member 1
|
|
chr22_+_20880159
|
1.850
|
|
IGLV6-57
|
immunoglobulin lambda variable 6-57
|
|
chr6_+_26266311
|
1.842
|
NM_021063
NM_138720
|
HIST1H2BD
|
histone cluster 1, H2bd
|
|
chr12_+_56290229
|
1.838
|
NM_001111270
|
ARHGEF25
|
Rho guanine nucleotide exchange factor (GEF) 25
|
|
chr12_-_31370305
|
1.786
|
|
FAM60A
|
family with sequence similarity 60, member A
|
|
chr6_+_43247179
|
1.784
|
|
SRF
|
serum response factor (c-fos serum response element-binding transcription factor)
|
|
chr6_+_43247168
|
1.777
|
|
SRF
|
serum response factor (c-fos serum response element-binding transcription factor)
|
|
chr3_+_188130967
|
1.775
|
NM_173216
NM_173217
|
ST6GAL1
|
ST6 beta-galactosamide alpha-2,6-sialyltranferase 1
|
|
chr16_+_10878539
|
1.761
|
NM_000246
|
CIITA
|
class II, major histocompatibility complex, transactivator
|
|
chr6_-_10523424
|
1.747
|
NM_003220
|
TFAP2A
|
transcription factor AP-2 alpha (activating enhancer binding protein 2 alpha)
|
|
chr4_-_186970366
|
1.726
|
NM_003603
NM_001145670
|
SORBS2
|
sorbin and SH3 domain containing 2
|
|
chr12_-_31370351
|
1.700
|
|
FAM60A
|
family with sequence similarity 60, member A
|
|
chr12_-_31370387
|
1.693
|
NM_001135811
NM_021238
NM_001135812
|
FAM60A
|
family with sequence similarity 60, member A
|
|
chr3_+_120799411
|
1.692
|
NM_015900
|
PLA1A
|
phospholipase A1 member A
|
|
chr6_-_10523203
|
1.659
|
|
TFAP2A
|
transcription factor AP-2 alpha (activating enhancer binding protein 2 alpha)
|
|
chr9_-_126573349
|
1.627
|
NM_001489
NM_033334
|
NR6A1
|
nuclear receptor subfamily 6, group A, member 1
|
|
chr6_-_26141774
|
1.619
|
NM_003513
|
HIST1H2AB
|
histone cluster 1, H2ab
|
|
chr11_-_31789168
|
1.614
|
|
PAX6
|
paired box 6
|
|
chr20_-_48980858
|
1.581
|
NM_015339
NM_181442
|
ADNP
|
activity-dependent neuroprotector homeobox
|
|
chr1_-_181889070
|
1.577
|
NM_203454
|
APOBEC4
|
apolipoprotein B mRNA editing enzyme, catalytic polypeptide-like 4 (putative)
|
|
chrX_-_24575375
|
1.576
|
NM_001163265
NM_004845
|
PCYT1B
|
phosphate cytidylyltransferase 1, choline, beta
|
|
chr6_-_25834733
|
1.557
|
NM_170745
|
HIST1H2AA
|
histone cluster 1, H2aa
|
|
chr10_-_64246129
|
1.534
|
NM_000399
NM_001136177
NM_001136179
|
EGR2
|
early growth response 2
|
|
chr22_+_20880112
|
1.525
|
|
|
|
|
chrX_+_123062143
|
1.522
|
|
STAG2
|
stromal antigen 2
|
|
chr14_-_105763152
|
1.515
|
|
IGHV4-31
IGHA1
IGHD
IGHG1
IGHG3
|
immunoglobulin heavy variable 4-31
immunoglobulin heavy constant alpha 1
immunoglobulin heavy constant delta
immunoglobulin heavy constant gamma 1 (G1m marker)
immunoglobulin heavy constant gamma 3 (G3m marker)
|
|
chr12_+_120548831
|
1.508
|
NM_032790
|
ORAI1
|
ORAI calcium release-activated calcium modulator 1
|
|
chr5_+_68702617
|
1.506
|
NM_133340
NM_133341
|
RAD17
|
RAD17 homolog (S. pombe)
|
|
chr14_-_106064887
|
1.492
|
|
LOC100126583
IGHV3-48
IGHA1
IGHM
|
hypothetical LOC100126583
immunoglobulin heavy variable 3-48
immunoglobulin heavy constant alpha 1
immunoglobulin heavy constant mu
|
|
chr2_-_157890400
|
1.447
|
NM_020711
|
ERMN
|
ermin, ERM-like protein
|
|
chr9_-_116200569
|
1.430
|
|
AKNA
|
AT-hook transcription factor
|
|
chr6_+_25834983
|
1.427
|
NM_170610
|
HIST1H2BA
|
histone cluster 1, H2ba
|
|
chr19_+_59820413
|
1.419
|
NM_006669
|
LILRB1
|
leukocyte immunoglobulin-like receptor, subfamily B (with TM and ITIM domains), member 1
|
|
chr6_+_20510019
|
1.416
|
NM_001949
|
E2F3
|
E2F transcription factor 3
|
|
chr19_-_18369407
|
1.407
|
NM_145256
|
LRRC25
|
leucine rich repeat containing 25
|
|
chr8_+_19841120
|
1.403
|
|
LPL
|
lipoprotein lipase
|
|
chr12_-_120725210
|
1.398
|
|
LOC338799
|
hypothetical LOC338799
|
|
chr22_+_21495273
|
1.395
|
|
|
|
|
chr16_-_87280380
|
1.389
|
NM_178310
|
SNAI3
|
snail homolog 3 (Drosophila)
|
|
chr22_+_21407066
|
1.378
|
|
|
|
|
chr17_-_38529638
|
1.378
|
NM_007298
|
BRCA1
|
breast cancer 1, early onset
|
|
chr6_-_31668476
|
1.378
|
NM_001145466
NM_001145467
NM_147130
|
NCR3
|
natural cytotoxicity triggering receptor 3
|
|
chr1_-_68735267
|
1.358
|
|
DEPDC1
|
DEP domain containing 1
|
|
chr16_-_73366200
|
1.347
|
NM_024306
|
FA2H
|
fatty acid 2-hydroxylase
|
|
chr14_-_106202262
|
1.347
|
|
IGLJ3
IGHA1
|
immunoglobulin lambda joining 3
immunoglobulin heavy constant alpha 1
|
|
chr19_+_45996881
|
1.336
|
NM_080732
|
EGLN2
|
egl nine homolog 2 (C. elegans)
|
|
chr2_+_201756048
|
1.299
|
|
CASP10
|
caspase 10, apoptosis-related cysteine peptidase
|
|
chr1_-_68735314
|
1.298
|
NM_001114120
NM_017779
|
DEPDC1
|
DEP domain containing 1
|
|
chr22_+_21340757
|
1.298
|
|
|
|
|
chr6_-_10939095
|
1.297
|
NM_005906
|
MAK
|
male germ cell-associated kinase
|
|
chr22_+_21116283
|
1.295
|
|
|
|
|
chr6_+_27890772
|
1.292
|
NM_003521
|
HIST1H2BM
|
histone cluster 1, H2bm
|
|
chr17_-_33179118
|
1.291
|
NM_000458
NM_001165923
|
HNF1B
|
HNF1 homeobox B
|
|
chr17_-_4792542
|
1.285
|
NM_005022
|
PFN1
|
profilin 1
|
|
chr17_+_60563917
|
1.278
|
NM_001081955
NM_001165933
NM_003835
|
RGS9
|
regulator of G-protein signaling 9
|
|
chr1_-_6537139
|
1.269
|
|
NOL9
|
nucleolar protein 9
|
|
chr1_+_200246312
|
1.262
|
NM_001114309
NM_004433
|
ELF3
|
E74-like factor 3 (ets domain transcription factor, epithelial-specific )
|
|
chr6_-_27890496
|
1.259
|
NM_021066
|
HIST1H2AJ
|
histone cluster 1, H2aj
|
|
chr17_-_8004671
|
1.258
|
|
VAMP2
|
vesicle-associated membrane protein 2 (synaptobrevin 2)
|
|
chr8_+_19841209
|
1.257
|
|
LPL
|
lipoprotein lipase
|
|
chr6_+_27883877
|
1.252
|
NM_003509
|
HIST1H2AI
HIST1H3F
|
histone cluster 1, H2ai
histone cluster 1, H3f
|
|
chr10_-_64246080
|
1.238
|
|
EGR2
|
early growth response 2
|
|
chr10_+_97505398
|
1.236
|
NM_001164178
NM_001164181
NM_001164179
NM_001164182
NM_001164183
NM_001776
|
ENTPD1
|
ectonucleoside triphosphate diphosphohydrolase 1
|
|
chr17_-_33178987
|
1.234
|
|
HNF1B
|
HNF1 homeobox B
|
|
chrX_-_138742050
|
1.233
|
NM_001010986
NM_173694
|
ATP11C
|
ATPase, class VI, type 11C
|
|
chr2_-_113310796
|
1.227
|
NM_000576
|
IL1B
|
interleukin 1, beta
|
|
chr19_-_19142090
|
1.215
|
NM_001145785
|
MEF2B
|
myocyte enhancer factor 2B
|
|
chr21_+_29322100
|
1.202
|
|
USP16
|
ubiquitin specific peptidase 16
|
|
chr1_+_200246370
|
1.197
|
|
ELF3
|
E74-like factor 3 (ets domain transcription factor, epithelial-specific )
|
|
chr7_-_102576570
|
1.190
|
NM_001122838
NM_198990
|
NAPEPLD
|
N-acyl phosphatidylethanolamine phospholipase D
|
|
chr6_+_43246897
|
1.188
|
NM_003131
|
SRF
|
serum response factor (c-fos serum response element-binding transcription factor)
|
|
chr14_-_105862080
|
1.186
|
|
IGHV4-31
IGHA1
IGHD
IGHG1
IGHG2
IGHG3
IGHM
SCFV
|
immunoglobulin heavy variable 4-31
immunoglobulin heavy constant alpha 1
immunoglobulin heavy constant delta
immunoglobulin heavy constant gamma 1 (G1m marker)
immunoglobulin heavy constant gamma 2 (G2m marker)
immunoglobulin heavy constant gamma 3 (G3m marker)
immunoglobulin heavy constant mu
single-chain Fv fragment
|
|
chr19_-_19142053
|
1.178
|
|
MEF2B
|
myocyte enhancer factor 2B
|
|
chr6_+_27323458
|
1.177
|
NM_005865
|
PRSS16
|
protease, serine, 16 (thymus)
|
|
chr17_-_70639382
|
1.176
|
NM_014595
|
NT5C
|
5', 3'-nucleotidase, cytosolic
|
|
chr22_+_21042091
|
1.156
|
|
IGLV1-44
IGL@
|
immunoglobulin lambda variable 1-44
immunoglobulin lambda locus
|
|
chr13_-_45640679
|
1.144
|
|
LCP1
|
lymphocyte cytosolic protein 1 (L-plastin)
|
|
chr2_-_2313894
|
1.137
|
NM_015025
|
MYT1L
|
myelin transcription factor 1-like
|
|
chr5_+_68703036
|
1.123
|
NM_133339
|
RAD17
|
RAD17 homolog (S. pombe)
|
|
chr6_+_32515596
|
1.117
|
NM_019111
|
HLA-DRA
|
major histocompatibility complex, class II, DR alpha
|
|
chr6_-_26232109
|
1.114
|
NM_003526
|
HIST1H2BG
HIST1H2BC
|
histone cluster 1, H2bg
histone cluster 1, H2bc
|
|
chr2_-_172458970
|
1.113
|
NM_003705
|
SLC25A12
|
solute carrier family 25 (mitochondrial carrier, Aralar), member 12
|
|
chr2_-_130603264
|
1.113
|
NM_001099771
|
POTEF
|
POTE ankyrin domain family, member F
|
|
chr11_+_74377587
|
1.111
|
NM_006656
|
NEU3
|
sialidase 3 (membrane sialidase)
|
|
chr15_-_38387357
|
1.089
|
|
PLCB2
|
phospholipase C, beta 2
|
|
chr1_-_6537171
|
1.085
|
NM_024654
|
NOL9
|
nucleolar protein 9
|
|
chr6_+_26232351
|
1.082
|
NM_003512
|
HIST1H2AC
|
histone cluster 1, H2ac
|
|
chr12_+_111829191
|
1.081
|
|
OAS1
|
2',5'-oligoadenylate synthetase 1, 40/46kDa
|
|
chr16_-_73365975
|
1.071
|
|
FA2H
|
fatty acid 2-hydroxylase
|
|
chr8_+_19841010
|
1.070
|
|
LPL
|
lipoprotein lipase
|
|
chr19_-_60611136
|
1.069
|
NM_014501
|
UBE2S
|
ubiquitin-conjugating enzyme E2S
|
|
chr7_+_90176647
|
1.065
|
NM_012395
|
CDK14
|
cyclin-dependent kinase 14
|
|
chr8_+_86287161
|
1.060
|
NM_001083589
|
E2F5
|
E2F transcription factor 5, p130-binding
|
|
chr17_+_60563963
|
1.059
|
|
RGS9
|
regulator of G-protein signaling 9
|
|
chr11_-_31789334
|
1.053
|
|
PAX6
|
paired box 6
|
|
chr8_+_19840861
|
1.036
|
NM_000237
|
LPL
|
lipoprotein lipase
|
|
chr17_+_7494964
|
1.033
|
NM_001678
|
ATP1B2
|
ATPase, Na+/K+ transporting, beta 2 polypeptide
|
|
chr11_-_31789360
|
1.032
|
|
PAX6
|
paired box 6
|
|
chr13_-_98757634
|
1.030
|
NM_004951
|
GPR183
|
G protein-coupled receptor 183
|
|
chr14_-_105762742
|
1.026
|
|
|
|
|
chr16_-_87280289
|
1.023
|
|
SNAI3
|
snail homolog 3 (Drosophila)
|
|
chr15_-_38387410
|
1.018
|
NM_004573
|
PLCB2
|
phospholipase C, beta 2
|
|
chr4_-_71751070
|
0.998
|
|
IGJ
|
immunoglobulin J polypeptide, linker protein for immunoglobulin alpha and mu polypeptides
|
|
chr6_+_37245860
|
0.971
|
NM_002648
|
PIM1
|
pim-1 oncogene
|
|
chr12_-_55614311
|
0.967
|
NM_148897
|
SDR9C7
|
short chain dehydrogenase/reductase family 9C, member 7
|
|
chr1_+_40929828
|
0.964
|
NM_001142587
NM_001142588
NM_001142589
NM_014223
|
NFYC
|
nuclear transcription factor Y, gamma
|
|
chr12_+_111829121
|
0.956
|
NM_001032409
NM_002534
NM_016816
|
OAS1
|
2',5'-oligoadenylate synthetase 1, 40/46kDa
|
|
chr19_-_18253413
|
0.924
|
NM_005354
|
JUND
|
jun D proto-oncogene
|
|
chr6_+_10693964
|
0.921
|
NM_145655
|
GCNT2
|
glucosaminyl (N-acetyl) transferase 2, I-branching enzyme (I blood group)
|
|
chr6_+_155579788
|
0.912
|
NM_001010927
|
TIAM2
|
T-cell lymphoma invasion and metastasis 2
|
|
chr1_+_165075304
|
0.911
|
|
POGK
|
pogo transposable element with KRAB domain
|
|
chr22_+_21065134
|
0.910
|
|
LOC100290481
|
immunoglobulin lambda light chain-like
|
|
chrX_+_37524250
|
0.910
|
|
CYBB
|
cytochrome b-245, beta polypeptide
|
|
chr19_-_18253436
|
0.909
|
|
JUND
|
jun D proto-oncogene
|
|
chr17_+_20712337
|
0.907
|
|
|
|
|
chr14_-_106241445
|
0.903
|
|
IGHV4-31
IGHV1-69
IGHA1
IGHG1
IGHG3
IGHM
|
immunoglobulin heavy variable 4-31
immunoglobulin heavy variable 1-69
immunoglobulin heavy constant alpha 1
immunoglobulin heavy constant gamma 1 (G1m marker)
immunoglobulin heavy constant gamma 3 (G3m marker)
immunoglobulin heavy constant mu
|
|
chr3_+_153074118
|
0.901
|
NM_033050
|
SUCNR1
|
succinate receptor 1
|
|
chr14_-_105623358
|
0.899
|
|
IGHA1
IGHD
IGHG1
|
immunoglobulin heavy constant alpha 1
immunoglobulin heavy constant delta
immunoglobulin heavy constant gamma 1 (G1m marker)
|
|
chr11_-_62125826
|
0.895
|
NM_004739
|
MTA2
|
metastasis associated 1 family, member 2
|
|
chr14_+_22422270
|
0.894
|
NM_173527
|
REM2
|
RAS (RAD and GEM)-like GTP binding 2
|
|
chr11_-_31789431
|
0.889
|
NM_000280
NM_001604
|
PAX6
|
paired box 6
|
|
chr1_+_22107137
|
0.883
|
|
RPL21
|
ribosomal protein L21
|
|
chr13_+_51334117
|
0.883
|
NM_031290
|
CCDC70
|
coiled-coil domain containing 70
|
|
chr15_-_56145140
|
0.880
|
NM_003888
NM_170696
|
ALDH1A2
|
aldehyde dehydrogenase 1 family, member A2
|
|
chr10_+_106103503
|
0.880
|
NM_001008723
|
CCDC147
|
coiled-coil domain containing 147
|
|
chr5_-_145875868
|
0.878
|
NM_194251
|
GPR151
|
G protein-coupled receptor 151
|
|
chr16_+_51722285
|
0.870
|
|
CHD9
|
chromodomain helicase DNA binding protein 9
|
|
chr20_+_34019942
|
0.870
|
NM_080834
|
C20orf152
|
chromosome 20 open reading frame 152
|
|
chr17_+_7495140
|
0.866
|
|
ATP1B2
|
ATPase, Na+/K+ transporting, beta 2 polypeptide
|
|
chr14_-_106330781
|
0.866
|
|
IGHV5-78
|
immunoglobulin heavy variable 5-78 pseudogene
|