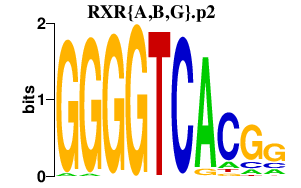
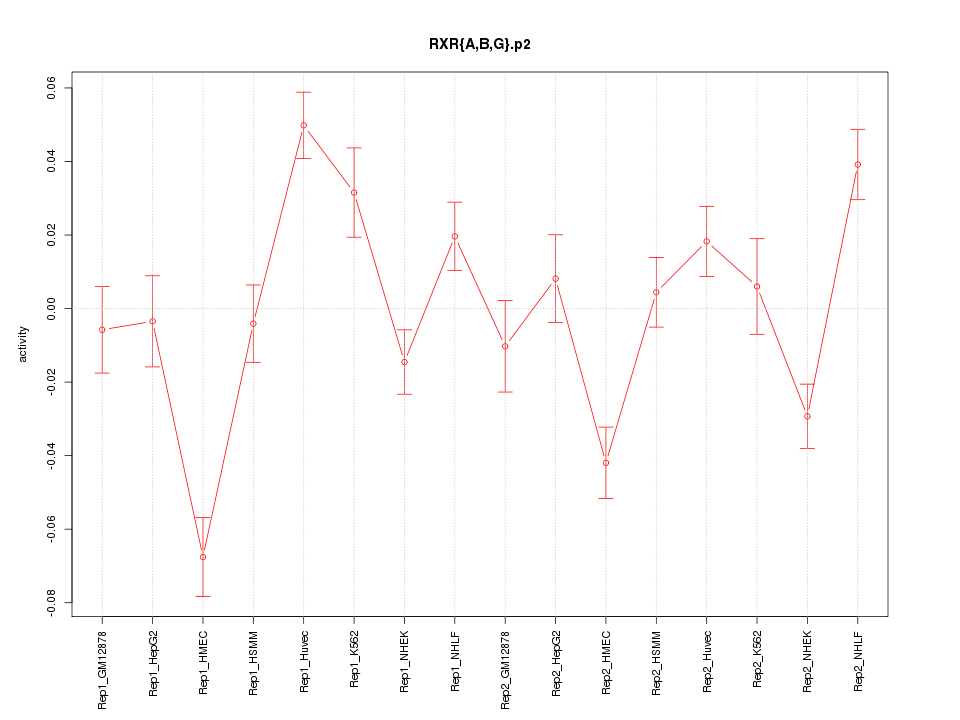
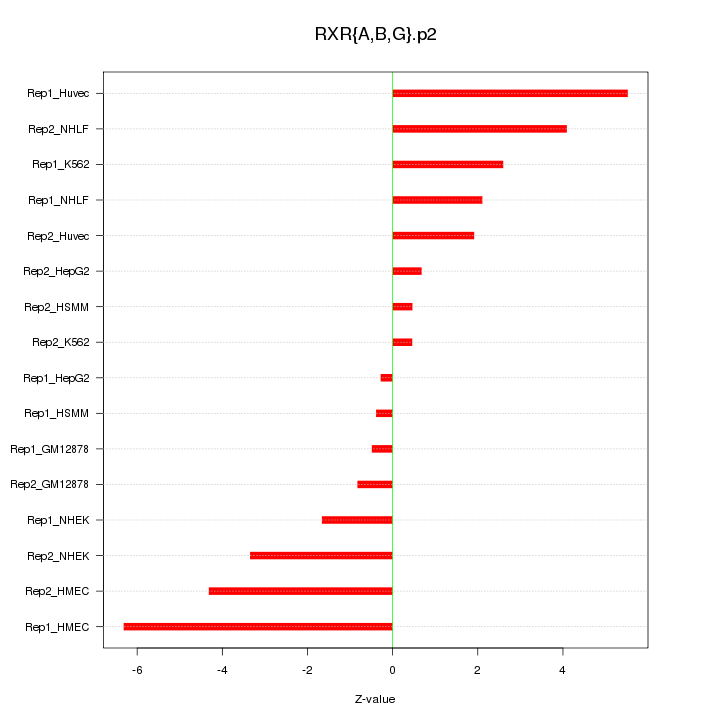
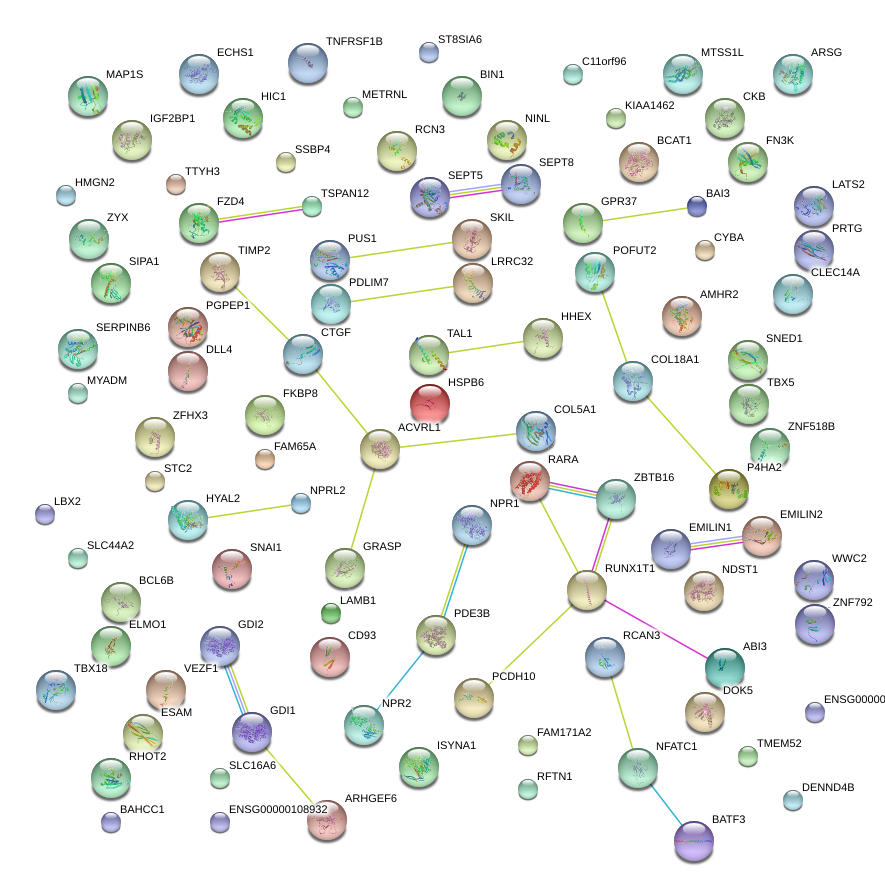
|
chr17_+_44429748
|
6.387
|
NM_001160423
NM_006546
|
IGF2BP1
|
insulin-like growth factor 2 mRNA binding protein 1
|
|
chr19_+_18391423
|
3.316
|
|
|
|
|
chr19_+_18391211
|
3.115
|
NM_001009998
NM_032627
|
SSBP4
|
single stranded DNA binding protein 4
|
|
chr21_-_45532176
|
3.003
|
NM_015227
NM_133635
|
POFUT2
|
protein O-fucosyltransferase 2
|
|
chr22_+_18081968
|
2.676
|
NM_002688
|
SEPT5
|
septin 5
|
|
chr6_-_2916786
|
2.413
|
|
SERPINB6
|
serpin peptidase inhibitor, clade B (ovalbumin), member 6
|
|
chr18_+_75261231
|
2.400
|
NM_172387
NM_172389
|
NFATC1
|
nuclear factor of activated T-cells, cytoplasmic, calcineurin-dependent 1
|
|
chr15_-_53822468
|
2.394
|
NM_173814
|
PRTG
|
protogenin
|
|
chr17_+_35751796
|
2.258
|
NM_001024809
|
RARA
|
retinoic acid receptor, alpha
|
|
chr12_+_50686990
|
2.246
|
NM_181711
|
GRASP
|
GRP1 (general receptor for phosphoinositides 1)-associated scaffold protein
|
|
chr20_+_48032918
|
2.207
|
NM_005985
|
SNAI1
|
snail homolog 1 (Drosophila)
|
|
chr8_-_93184629
|
2.018
|
NM_001198633
NM_175634
|
RUNX1T1
|
runt-related transcription factor 1; translocated to, 1 (cyclin D-related)
|
|
chr21_+_45649719
|
1.976
|
|
|
|
|
chr14_-_103058568
|
1.884
|
|
CKB
|
creatine kinase, brain
|
|
chr3_+_129690735
|
1.880
|
|
|
|
|
chr14_-_37795001
|
1.876
|
|
CLEC14A
|
C-type lectin domain family 14, member A
|
|
chr6_-_132314004
|
1.861
|
NM_001901
|
CTGF
|
connective tissue growth factor
|
|
chr11_+_14621972
|
1.824
|
|
PDE3B
|
phosphodiesterase 3B, cGMP-inhibited
|
|
chr6_-_2916280
|
1.802
|
|
SERPINB6
|
serpin peptidase inhibitor, clade B (ovalbumin), member 6
|
|
chr14_-_37795290
|
1.758
|
NM_175060
|
CLEC14A
|
C-type lectin domain family 14, member A
|
|
chr10_-_135036805
|
1.755
|
NM_004092
|
ECHS1
|
enoyl CoA hydratase, short chain, 1, mitochondrial
|
|
chr17_+_6867090
|
1.741
|
NM_181844
|
BCL6B
|
B-cell CLL/lymphoma 6, member B
|
|
chr1_-_47468029
|
1.698
|
NM_003189
|
TAL1
|
T-cell acute lymphocytic leukemia 1
|
|
chr22_+_18085891
|
1.615
|
|
SEPT5
|
septin 5
|
|
chr17_+_6867138
|
1.612
|
|
BCL6B
|
B-cell CLL/lymphoma 6, member B
|
|
chr11_-_124137260
|
1.564
|
NM_138961
|
ESAM
|
endothelial cell adhesion molecule
|
|
chr10_-_17536181
|
1.559
|
NM_001004470
|
ST8SIA6
|
ST8 alpha-N-acetyl-neuraminide alpha-2,8-sialyltransferase 6
|
|
chr22_+_18085990
|
1.530
|
|
SEPT5-GP1BB
SEPT5
|
SEPT5-GP1BB read-through transcript
septin 5
|
|
chr4_+_134292630
|
1.491
|
|
PCDH10
|
protocadherin 10
|
|
chr2_+_27155197
|
1.479
|
|
EMILIN1
|
elastin microfibril interfacer 1
|
|
chr2_+_113709553
|
1.451
|
|
LOC654433
|
hypothetical LOC654433
|
|
chr20_-_23014828
|
1.451
|
NM_012072
|
CD93
|
CD93 molecule
|
|
chr16_-_87244896
|
1.447
|
NM_000101
|
CYBA
|
cytochrome b-245, alpha polypeptide
|
|
chr5_+_149845503
|
1.440
|
|
NDST1
|
N-deacetylase/N-sulfotransferase (heparan glucosaminyl) 1
|
|
chr4_+_134293171
|
1.357
|
|
PCDH10
|
protocadherin 10
|
|
chr7_+_142788521
|
1.351
|
|
ZYX
|
zyxin
|
|
chr3_-_50335113
|
1.345
|
NM_003773
NM_033158
|
HYAL2
|
hyaluronoglucosaminidase 2
|
|
chr16_+_658158
|
1.344
|
|
RHOT2
|
ras homolog gene family, member T2
|
|
chr6_-_85529031
|
1.330
|
|
TBX18
|
T-box 18
|
|
chr11_-_86343737
|
1.315
|
|
FZD4
|
frizzled homolog 4 (Drosophila)
|
|
chr11_-_76059435
|
1.298
|
NM_001128922
|
LRRC32
|
leucine rich repeat containing 32
|
|
chr17_+_63799228
|
1.286
|
|
ARSG
|
arylsulfatase G
|
|
chr17_-_74433048
|
1.285
|
NM_003255
|
TIMP2
|
TIMP metallopeptidase inhibitor 2
|
|
chr21_+_45649429
|
1.276
|
NM_130445
|
COL18A1
|
collagen, type XVIII, alpha 1
|
|
chr5_-_172687798
|
1.259
|
|
STC2
|
stanniocalcin 2
|
|
chr16_+_66129481
|
1.258
|
|
FAM65A
|
family with sequence similarity 65, member A
|
|
chr17_+_78286687
|
1.250
|
NM_022158
|
FN3K
|
fructosamine 3 kinase
|
|
chr17_-_63798851
|
1.249
|
NM_001174166
NM_004694
|
SLC16A6
|
solute carrier family 16, member 6 (monocarboxylic acid transporter 7)
|
|
chr16_-_69277355
|
1.240
|
NM_138383
|
MTSS1L
|
metastasis suppressor 1-like
|
|
chr1_+_12149662
|
1.238
|
|
TNFRSF1B
|
tumor necrosis factor receptor superfamily, member 1B
|
|
chr16_+_658081
|
1.223
|
NM_138769
|
RHOT2
|
ras homolog gene family, member T2
|
|
chr1_-_152185764
|
1.209
|
NM_014856
|
DENND4B
|
DENN/MADD domain containing 4B
|
|
chr17_+_78630793
|
1.196
|
NM_001004431
|
METRNL
|
meteorin, glial cell differentiation regulator-like
|
|
chr2_+_241586927
|
1.179
|
NM_001080437
|
SNED1
|
sushi, nidogen and EGF-like domains 1
|
|
chr12_-_24946495
|
1.179
|
NM_001178094
|
BCAT1
|
branched chain amino-acid transaminase 1, cytosolic
|
|
chr3_-_16529996
|
1.172
|
|
RFTN1
|
raftlin, lipid raft linker 1
|
|
chr7_+_142788480
|
1.169
|
NM_001010972
NM_003461
|
ZYX
|
zyxin
|
|
chr19_+_17691302
|
1.158
|
NM_018174
|
MAP1S
|
microtubule-associated protein 1S
|
|
chr18_+_2896956
|
1.143
|
|
EMILIN2
|
elastin microfibril interfacer 2
|
|
chr5_-_172688024
|
1.137
|
|
STC2
|
stanniocalcin 2
|
|
chr7_+_2638108
|
1.129
|
NM_025250
|
TTYH3
|
tweety homolog 3 (Drosophila)
|
|
chr17_+_76988134
|
1.128
|
NM_001080519
|
BAHCC1
|
BAH domain and coiled-coil containing 1
|
|
chr9_+_35782405
|
1.120
|
NM_003995
|
NPR2
|
natriuretic peptide receptor B/guanylate cyclase B (atrionatriuretic peptide receptor B)
|
|
chr19_+_10574055
|
1.117
|
NM_001145056
|
SLC44A2
|
solute carrier family 44, member 2
|
|
chr11_+_65162114
|
1.113
|
NM_006747
|
SIPA1
|
signal-induced proliferation-associated 1
|
|
chr19_-_18409788
|
1.108
|
NM_001170939
|
ISYNA1
|
inositol-3-phosphate synthase 1
|
|
chr7_-_107430566
|
1.107
|
NM_002291
|
LAMB1
|
laminin, beta 1
|
|
chr10_+_94439658
|
1.100
|
NM_002729
|
HHEX
|
hematopoietically expressed homeobox
|
|
chr12_+_130979729
|
1.096
|
NM_001002019
NM_025215
|
PUS1
|
pseudouridylate synthase 1
|
|
chr19_+_59064471
|
1.089
|
NM_001020821
NM_001020818
|
MYADM
|
myeloid-associated differentiation marker
|
|
chr9_+_136673534
|
1.085
|
|
COL5A1
|
collagen, type V, alpha 1
|
|
chr10_-_30388439
|
1.084
|
NM_020848
|
KIAA1462
|
KIAA1462
|
|
chr19_+_18312365
|
1.080
|
NM_017712
|
FKBP8
PGPEP1
|
FK506 binding protein 8, 38kDa
pyroglutamyl-peptidase I
|
|
chr19_+_59064795
|
1.077
|
|
MYADM
|
myeloid-associated differentiation marker
|
|
chr12_+_130979696
|
1.067
|
NM_001002020
|
PUS1
|
pseudouridylate synthase 1
|
|
chr11_+_43920385
|
1.061
|
NM_001145033
|
C11orf96
|
chromosome 11 open reading frame 96
|
|
chr1_-_1840564
|
1.060
|
NM_178545
|
TMEM52
|
transmembrane protein 52
|
|
chr4_-_10068055
|
1.052
|
NM_053042
|
ZNF518B
|
zinc finger protein 518B
|
|
chr2_-_74583850
|
1.052
|
NM_001009812
|
LBX2
|
ladybird homeobox 2
|
|
chr11_+_113436438
|
1.043
|
NM_001018011
|
ZBTB16
|
zinc finger and BTB domain containing 16
|
|
chr17_-_53420473
|
1.037
|
|
VEZF1
|
vascular endothelial zinc finger 1
|
|
chr7_+_142788569
|
1.033
|
|
ZYX
|
zyxin
|
|
chr19_-_40146792
|
1.020
|
NM_175872
|
ZNF792
|
zinc finger protein 792
|
|
chr7_+_142788566
|
1.018
|
|
ZYX
|
zyxin
|
|
chr12_+_52105131
|
1.011
|
|
AMHR2
|
anti-Mullerian hormone receptor, type II
|
|
chr16_-_71650034
|
1.010
|
NM_001164766
|
ZFHX3
|
zinc finger homeobox 3
|
|
chr17_+_44643067
|
1.005
|
|
ABI3
|
ABI family, member 3
|
|
chr2_+_27154938
|
1.004
|
NM_007046
|
EMILIN1
|
elastin microfibril interfacer 1
|
|
chr19_+_54723044
|
1.004
|
|
RCN3
|
reticulocalbin 3, EF-hand calcium binding domain
|
|
chr21_+_45532377
|
1.001
|
|
LOC642852
|
hypothetical LOC642852
|
|
chr4_+_184257337
|
0.996
|
NM_024949
|
WWC2
|
WW and C2 domain containing 2
|
|
chr11_-_86343859
|
0.993
|
NM_012193
|
FZD4
|
frizzled homolog 4 (Drosophila)
|
|
chr7_+_142788517
|
0.993
|
|
ZYX
|
zyxin
|
|
chr7_+_142788579
|
0.991
|
|
ZYX
|
zyxin
|
|
chr19_-_40146707
|
0.989
|
|
ZNF792
|
zinc finger protein 792
|
|
chr3_+_171558188
|
0.972
|
|
SKIL
|
SKI-like oncogene
|
|
chr17_-_74432800
|
0.968
|
|
TIMP2
|
TIMP metallopeptidase inhibitor 2
|
|
chr12_+_50587468
|
0.957
|
NM_000020
|
ACVRL1
|
activin A receptor type II-like 1
|
|
chr19_-_40938251
|
0.955
|
|
HSPB6
|
heat shock protein, alpha-crystallin-related, B6
|
|
chr5_-_131591380
|
0.953
|
NM_001017974
NM_001142598
NM_001142599
|
P4HA2
|
prolyl 4-hydroxylase, alpha polypeptide II
|
|
chr2_+_74583762
|
0.940
|
|
LOC151534
|
hypothetical LOC151534
|
|
chr7_+_2638156
|
0.939
|
|
TTYH3
|
tweety homolog 3 (Drosophila)
|
|
chr17_-_39796649
|
0.937
|
NM_198475
|
FAM171A2
|
family with sequence similarity 171, member A2
|
|
chr20_+_52525538
|
0.934
|
NM_018431
|
DOK5
|
docking protein 5
|
|
chr3_+_171558215
|
0.932
|
|
SKIL
|
SKI-like oncogene
|
|
chr12_-_113328271
|
0.925
|
NM_181486
|
TBX5
|
T-box 5
|
|
chr3_+_171558143
|
0.924
|
NM_001145098
NM_005414
|
SKIL
|
SKI-like oncogene
|
|
chr17_+_1905104
|
0.913
|
NM_006497
|
HIC1
|
hypermethylated in cancer 1
|
|
chr15_+_39008822
|
0.909
|
NM_019074
|
DLL4
|
delta-like 4 (Drosophila)
|
|
chr1_+_26671488
|
0.908
|
NM_005517
|
HMGN2
|
high-mobility group nucleosomal binding domain 2
|
|
chr1_-_210939820
|
0.907
|
|
BATF3
|
basic leucine zipper transcription factor, ATF-like 3
|
|
chr7_-_37454933
|
0.902
|
NM_014800
|
ELMO1
|
engulfment and cell motility 1
|
|
chr5_-_176857158
|
0.902
|
|
PDLIM7
|
PDZ and LIM domain 7 (enigma)
|
|
chr20_-_25514137
|
0.899
|
NM_025176
|
NINL
|
ninein-like
|
|
chr5_-_132141368
|
0.899
|
NM_001098813
|
SEPT8
|
septin 8
|
|
chr7_-_120285245
|
0.891
|
|
TSPAN12
|
tetraspanin 12
|
|
chrX_-_135677022
|
0.890
|
|
ARHGEF6
|
Rac/Cdc42 guanine nucleotide exchange factor (GEF) 6
|
|
chr13_-_20533703
|
0.886
|
NM_014572
|
LATS2
|
LATS, large tumor suppressor, homolog 2 (Drosophila)
|
|
chrX_+_153318662
|
0.876
|
|
GDI1
|
GDP dissociation inhibitor 1
|
|
chr6_+_69401660
|
0.875
|
NM_001704
|
BAI3
|
brain-specific angiogenesis inhibitor 3
|
|
chr2_-_127580975
|
0.875
|
NM_004305
NM_139343
NM_139344
NM_139345
NM_139346
NM_139347
NM_139348
NM_139349
NM_139350
NM_139351
|
BIN1
|
bridging integrator 1
|
|
chr7_-_124192265
|
0.871
|
|
GPR37
|
G protein-coupled receptor 37 (endothelin receptor type B-like)
|
|
chr9_-_16860719
|
0.871
|
NM_017637
|
BNC2
|
basonuclin 2
|
|
chr19_+_17691283
|
0.871
|
|
MAP1S
|
microtubule-associated protein 1S
|
|
chr11_+_67563093
|
0.861
|
|
TCIRG1
|
T-cell, immune regulator 1, ATPase, H+ transporting, lysosomal V0 subunit A3
|
|
chr16_+_88315452
|
0.859
|
NM_001113525
|
ZNF276
|
zinc finger protein 276
|
|
chr7_-_150283794
|
0.839
|
NM_172057
|
KCNH2
|
potassium voltage-gated channel, subfamily H (eag-related), member 2
|
|
chr19_-_55046718
|
0.839
|
|
LOC100506033
|
hypothetical LOC100506033
|
|
chr12_+_50104859
|
0.838
|
NM_001039960
NM_004858
|
SLC4A8
|
solute carrier family 4, sodium bicarbonate cotransporter, member 8
|
|
chr19_-_40938046
|
0.838
|
|
HSPB6
|
heat shock protein, alpha-crystallin-related, B6
|
|
chr2_-_192767494
|
0.837
|
|
TMEFF2
|
transmembrane protein with EGF-like and two follistatin-like domains 2
|
|
chr9_+_136673464
|
0.836
|
NM_000093
|
COL5A1
|
collagen, type V, alpha 1
|
|
chr11_-_63440891
|
0.834
|
NM_173587
|
RCOR2
|
REST corepressor 2
|
|
chr16_+_30815428
|
0.825
|
NM_001142544
NM_001330
|
CTF1
|
cardiotrophin 1
|
|
chr19_+_6323443
|
0.823
|
NM_032306
|
ALKBH7
|
alkB, alkylation repair homolog 7 (E. coli)
|
|
chr3_-_38666122
|
0.820
|
NM_000335
NM_001099404
NM_001099405
NM_198056
NM_001160160
NM_001160161
|
SCN5A
|
sodium channel, voltage-gated, type V, alpha subunit
|
|
chr10_+_80569464
|
0.820
|
|
ZMIZ1
|
zinc finger, MIZ-type containing 1
|
|
chr19_+_16860807
|
0.814
|
NM_003950
|
F2RL3
|
coagulation factor II (thrombin) receptor-like 3
|
|
chr17_-_53420595
|
0.813
|
NM_007146
|
VEZF1
|
vascular endothelial zinc finger 1
|
|
chr5_-_176857207
|
0.811
|
NM_005451
NM_203352
NM_213636
|
PDLIM7
|
PDZ and LIM domain 7 (enigma)
|
|
chr1_-_210939929
|
0.809
|
NM_018664
|
BATF3
|
basic leucine zipper transcription factor, ATF-like 3
|
|
chr10_+_97794139
|
0.807
|
|
|
|
|
chr17_-_74432671
|
0.805
|
|
TIMP2
|
TIMP metallopeptidase inhibitor 2
|
|
chr22_+_41279811
|
0.801
|
NM_014509
|
SERHL2
|
serine hydrolase-like 2
|
|
chr13_-_20533629
|
0.796
|
|
LATS2
|
LATS, large tumor suppressor, homolog 2 (Drosophila)
|
|
chr19_-_54350457
|
0.796
|
NM_002152
|
HRC
|
histidine rich calcium binding protein
|
|
chr21_+_46342440
|
0.795
|
NM_001849
NM_058174
NM_058175
|
COL6A2
|
collagen, type VI, alpha 2
|
|
chr7_+_128219307
|
0.793
|
NM_022742
|
CCDC136
|
coiled-coil domain containing 136
|
|
chr7_+_142788730
|
0.792
|
|
ZYX
|
zyxin
|
|
chr17_-_75808680
|
0.790
|
NM_000199
|
SGSH
|
N-sulfoglucosamine sulfohydrolase
|
|
chr12_+_105873669
|
0.789
|
NM_152261
|
C12orf23
|
chromosome 12 open reading frame 23
|
|
chr20_-_43972949
|
0.788
|
|
PLTP
|
phospholipid transfer protein
|
|
chr12_+_50104936
|
0.787
|
|
SLC4A8
|
solute carrier family 4, sodium bicarbonate cotransporter, member 8
|
|
chr12_+_130979994
|
0.786
|
|
PUS1
|
pseudouridylate synthase 1
|
|
chr8_+_69027166
|
0.786
|
|
PREX2
|
phosphatidylinositol-3,4,5-trisphosphate-dependent Rac exchange factor 2
|
|
chr17_+_71097147
|
0.780
|
|
MYO15B
|
myosin XVB pseudogene
|
|
chr7_+_78920846
|
0.780
|
|
LOC100505881
|
hypothetical LOC100505881
|
|
chr19_-_13074669
|
0.778
|
|
LYL1
|
lymphoblastic leukemia derived sequence 1
|
|
chr19_-_55046117
|
0.777
|
|
LOC100506033
|
hypothetical LOC100506033
|
|
chr17_-_6887965
|
0.777
|
NM_153357
|
SLC16A11
|
solute carrier family 16, member 11 (monocarboxylic acid transporter 11)
|
|
chr12_+_12829807
|
0.766
|
NM_030817
|
APOLD1
|
apolipoprotein L domain containing 1
|
|
chr11_+_117906965
|
0.763
|
NM_001144034
NM_001144035
NM_001144036
NM_001144037
NM_001144038
NM_032780
|
TMEM25
|
transmembrane protein 25
|
|
chr10_+_76256305
|
0.759
|
NM_012330
|
MYST4
|
MYST histone acetyltransferase (monocytic leukemia) 4
|
|
chr10_+_17310475
|
0.759
|
|
VIM
|
vimentin
|
|
chr8_+_16929116
|
0.757
|
NM_181723
|
EFHA2
|
EF-hand domain family, member A2
|
|
chr11_-_105398118
|
0.757
|
NM_032424
|
KIAA1826
|
KIAA1826
|
|
chr17_-_45562112
|
0.754
|
NM_174920
|
SAMD14
|
sterile alpha motif domain containing 14
|
|
chr9_-_16860664
|
0.750
|
|
BNC2
|
basonuclin 2
|
|
chr19_-_7244876
|
0.748
|
NM_000208
NM_001079817
|
INSR
|
insulin receptor
|
|
chr17_+_56831951
|
0.741
|
NM_005994
|
TBX2
|
T-box 2
|
|
chr16_-_1369613
|
0.740
|
NM_001193389
NM_023076
|
UNKL
|
unkempt homolog (Drosophila)-like
|
|
chr7_-_81910740
|
0.739
|
|
CACNA2D1
|
calcium channel, voltage-dependent, alpha 2/delta subunit 1
|
|
chr3_-_127558747
|
0.734
|
NM_014079
|
KLF15
|
Kruppel-like factor 15
|
|
chr19_-_13074973
|
0.732
|
NM_005583
|
LYL1
|
lymphoblastic leukemia derived sequence 1
|
|
chr8_+_69027277
|
0.731
|
|
PREX2
|
phosphatidylinositol-3,4,5-trisphosphate-dependent Rac exchange factor 2
|
|
chr17_+_16259573
|
0.731
|
NM_016113
|
TRPV2
|
transient receptor potential cation channel, subfamily V, member 2
|
|
chr11_+_117907083
|
0.729
|
|
TMEM25
|
transmembrane protein 25
|
|
chr19_+_55046229
|
0.729
|
|
PTOV1
|
prostate tumor overexpressed 1
|
|
chr9_-_130006309
|
0.727
|
NM_001131015
NM_012127
|
CIZ1
|
CDKN1A interacting zinc finger protein 1
|
|
chr19_+_55046088
|
0.724
|
NM_017432
|
PTOV1
|
prostate tumor overexpressed 1
|
|
chr7_-_81910956
|
0.719
|
NM_000722
|
CACNA2D1
|
calcium channel, voltage-dependent, alpha 2/delta subunit 1
|
|
chr19_-_12929008
|
0.716
|
|
GADD45GIP1
|
growth arrest and DNA-damage-inducible, gamma interacting protein 1
|
|
chr16_-_73857214
|
0.713
|
NM_001170720
NM_001170714
NM_001170718
|
BCAR1
|
breast cancer anti-estrogen resistance 1
|
|
chr17_-_1478293
|
0.713
|
|
SLC43A2
|
solute carrier family 43, member 2
|
|
chr20_+_32755778
|
0.712
|
NM_021202
|
TP53INP2
|
tumor protein p53 inducible nuclear protein 2
|
|
chr17_-_33179118
|
0.707
|
NM_000458
NM_001165923
|
HNF1B
|
HNF1 homeobox B
|
|
chr7_-_126679540
|
0.705
|
NM_001127323
|
GRM8
|
glutamate receptor, metabotropic 8
|
|
chr6_-_2916980
|
0.704
|
|
SERPINB6
|
serpin peptidase inhibitor, clade B (ovalbumin), member 6
|
|
chr5_-_132140786
|
0.704
|
NM_001098811
NM_001098812
NM_015146
|
SEPT8
|
septin 8
|
|
chr9_-_13269562
|
0.702
|
|
MPDZ
|
multiple PDZ domain protein
|
|
chr1_-_234295005
|
0.702
|
|
NID1
|
nidogen 1
|
|
chr10_-_17536351
|
0.701
|
|
ST8SIA6
|
ST8 alpha-N-acetyl-neuraminide alpha-2,8-sialyltransferase 6
|
|
chr6_-_37061902
|
0.700
|
|
MTCH1
|
mitochondrial carrier homolog 1 (C. elegans)
|
|
chr11_-_124175493
|
0.692
|
NM_024631
|
C11orf61
|
chromosome 11 open reading frame 61
|
|
chr19_-_12929033
|
0.691
|
NM_052850
|
GADD45GIP1
|
growth arrest and DNA-damage-inducible, gamma interacting protein 1
|
|
chr7_+_94374862
|
0.690
|
NM_001166160
NM_017650
|
PPP1R9A
|
protein phosphatase 1, regulatory (inhibitor) subunit 9A
|
|
chr14_+_23937766
|
0.687
|
NM_025081
|
NYNRIN
|
NYN domain and retroviral integrase containing
|
|
chr20_+_62158882
|
0.685
|
NM_198723
|
TCEA2
|
transcription elongation factor A (SII), 2
|
|
chr22_-_17891777
|
0.684
|
|
CLDN5
|
claudin 5
|
|
chr17_-_16197433
|
0.678
|
NM_181716
|
CENPV
|
centromere protein V
|